Abstract
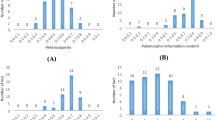
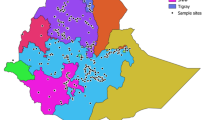
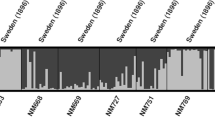
Ethiopia is known for its wide topographically induced variation that favours a large amount of plant genetic diversity, and determination of the genetic structure of the germplasm is a crucial step towards exploiting this variation. Accordingly, a study was initiated to determine population structure patterns based on spatial and temporal factors and thereby establish group accessions depending on genetic similarity. To achieve this goal, 15 simple sequence repeats (SSRs) were used for 199 barley landraces collected from ten administrative regions of Ethiopia. The analysis of the spatial genetic structure was performed on the entire data set and on two groups created based on the year of collection. The results obtained from the model-based Bayesian clustering revealed 16 unstructured groups. Furthermore, the grouping of the accessions based on SSR markers resulted in hierarchical chain-type clusters with most of the accessions in the first cluster. The spatial correlations between the genetic and geographical distances revealed a weak population structure for the entire data set, and a weak temporal population structure was also observed for the barley populations from the recent collection. In general, the results indicated the absence of a distinct and consistent population structure among the regions, which could be presumed because of high gene flow. These findings will facilitate parent selection in order to broaden the genetic base of modern cultivars via breeding efforts and also provide information for the planning of an efficient germplasm collection strategy.





Similar content being viewed by others
References
Abebe TD, Bauer AM, Léon J (2010) Morphological diversity of Ethiopian barleys (Hordeum vulgare L.) in relation to geographic regions and altitudes. Hereditas 147:154–164
Abebe TD, Mathew B, Léon J (2012) Barrier analysis detected genetic discontinuity among Ethiopian barley (Hordeum vulgare L.) landraces due to landscape and human mobility on gene flow. Genet Resour Crop Evol. doi: 10.1007/s10722-012-9834-6
Alemayehu N, Becker H (2002) Genotypic diversity and patterns of variation in a germplasm material of Ethiopian mustard (Brassica carinata A. Braun). Genet Resour Crop Evol 49:573–582
Asfaw Z (2000) The barleys of Ethiopia. In: Brush SB (ed) Genes in the field: on-farm conservation of crop Diversity. IDRC and IPGRI, USA, pp 77–107
Assefa K, Merker A, Tefera H (2003) Multivariate analysis of diversity of tef (Eragrostis tef (Zucc.) Trotter) germplasm from Western and Southern Ethiopia. Hereditas 138:228–236
Ayana A, Bekele E (1999) Multivariate analysis of morphological variation in sorghum (Sorghum bicolor (L.) Moench) germplasm from Ethiopia and Eritrea. Genet Resour Crop Evol 46:273–284
Bishaw Z (2004) Wheat and barley seed system in Ethiopia and Syria. Wageningen University, The Netherlands, PhD Dissertation
Brush SB (1995) In-situ conservation of landraces in center of crop diversity. Crop Sci 35:346–354
Cofi C, Milinkovitch MC, Gibbs JP, Caccone A, Powell JR (2002) Microsatellite analysis of genetic Divergence among populations of giant Galápagos tortoises. Mol Ecol 11:2265–2283
Demissie A, Bjørnstad Å, Kleinhofs A (1998) Restriction fragment length polymorphisms in landrace barleys from Ethiopia in relation to geographic, altitude, and agro-ecological factors. Crop Sci 38:237–243
Dice LR (1945) Measures of the amount of ecologic association between species. Ecology 26:297–302
Doggett H (1988) Sorghum, 2nd edn. Longman, UK
Ennos RA (2001) Inferences about spatial processes in plant populations from the analysis of molecular markers. In: Silvertown J, Antonovics J (eds) Integrating ecology and evolution in spatial context. Blackwell Science, Oxford, pp 45–71
Escudero A, Iriondo JM, Torres ME (2003) Spatial analysis of genetic diversity as a tool for plant conservation. Biol Cons 113:351–365
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Fischbeck G (2003) Diversification through breeding. In: von Bothmer R, van Hintum T, Knüpffer H, Sato K (eds) Diversity in barley (Hordeum vulgare). Elsevier, Amsterdam, pp 29–52
Gao H, Williamson S, Bustamante CD (2007) An MCMC approach for joint inference of population structure and inbreeding rates from multi-locus genotype data. Genetics 176:1635–1651
Hadado TT, Rau D, Bitocchi E, Papa R (2010) Adaptation and diversity along an altitudinal gradient in Ethiopian barley (Hordeum vulgare L.) landraces revealed by molecular analysis. BMC Plant Biol 10:121
Hampton JO, Spencer PBS, Alpers DL, Twigg LE, Woolnough AP, Doust J, Higgs T, Pluske J (2004) Molecular techniques, wildlife management and the importance of genetic population structure and dispersal: a case study with feral pigs. J Appl Ecol 41:735–743
Holcomb J, Tolbert DM, Jain SK (1977) Diversity analysis of genetic resources in rice. Euphytica 26:441–450
Lakis G, Ousmane AM, Sanoussi D, Habibou A, Badamassi M, Lamy F, Jika N, Sidikou R, Adam T, Sarr A, Lexereau A, Robert T (2011) Evolutionary dynamics of cycle length in pearl millet: the role of farmer’s practices and gene flow. Genetica 139:1367–1380
Liu ZW, Biyashev RM, Maroof MAS (1996) Development of simple sequence repeat DNA markers and their integration into a barley linkage map. Theor Appl Genet 93:869–876
McCauley DE (1997) The relative contributions of seed and pollen movement to the local genetic structure of Silene alba. Heredity 88:257–263
Moran PAP (1950) Notes on continuous stocastic phenomena. Biometrika 37:17–23
National Research Council (1996) Lost crops of Africa. Grains, vol 1. National Academy Press, Washington DC
Nevo E (1992) Origin, evolution, population genetics and resources of wild barley, Hordeum spontaneum, in the Fertile Crescent. In: Shewry P(ed) Barley: genetics, molecular biology and biotechnology, CAB International, Wallingford, Oxford, pp 19–43
Ould Med Mahmouda M, Hamzaa S (2009) Genetic diversity in local barley accessions collected from different geographical regions of Tunisia. Plant Genet Resour 7:169–176
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetics software for teaching and research. Mol Ecol Notes 6:288–295
Pecetti L, Damania AB (1996) Geographic variation in tetraploid wheat (Triticum turgidum ssp. turgidum convar. durum) landraces from two provinces in Ethiopia. Genet Resour Crop Evol 43:395–407
PGRC (1996) Ethiopia: country report to the FAO international technical conference on plant genetic resources. Germany, Leipzig
Pillen K, Binder A, Kreuzkam B, Ramsay L, Waugh R, Förster J, Léon J (2000) Mapping new EMB-derived barley microsatellites and their use in differentiating German barley cultivars. Theor Appl Genet 101:652–660
Pritchard JK, Wen W (2003) Documentation for structure software: Version 2. Available: http://pritch.bsd.uchicago.edu
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Ramsay L, Macaulay M, Ivanissevich SD, MacLean K, Cardle L, Fuller J, Edwards KJ, Tuvesson S, Morgante M, Massari A, Maestri E, Marmiroli N, Sjakste T, Ganal M, Powell W, Waugh R (2000) A simple sequence repeat-based linkage map of barley. Genetics 156:1997–2005
Rohlf F (1998) NTSYS-pc: numerical taxonomy and multivariate analysis system. Exeter Publication, New York
Romesburg H (1990) Cluster analysis for researchers. Krieger Publishing Co., Malabar
Rosenberg NA (2004) Distruct: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Rosenberg NA, Mahajan S, Ramachandran S, Zhao C, Pritchard JK, Feldman MW (2005) Clines, clusters, and the effect of study design on the inference of human population structure. PLoS Genet 1:660–671
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley-mendelian inheritance, chromosomal location, and population-dynamics. PNAS 81:8014–8018
Shewayrga H, Sopade AP (2011) Ethnobotany, diverse food uses, claimed health benefits and implications on conservation of barley landraces in North Eastern Ethiopia highlands. J Ethnobiol Ethnomed 7:19
Slatkin M (1987) Gene flow and the geographic structure of natural populations. Science 236:787–792
Sokal RR, Oden NL (1978) Spatial autocorrelation in biology. I. Methodology. Biol J Linn Soc 10:199–228
Spiegelhalter DJ, Best NG, Carlin BP, Linde AVD (2002) Bayesian measures of model complexity and fit. J Roy Stat Soc B 64:583–639
Struss D, Plieske J (1998) The use of microsatellite markers for detection of genetic diversity in barley populations. Theor Appl Genet 97:308–315
Teshome A, Baum BR, Fahrig L, Torrence JK, Arnason TJ, Lambert JD (1997) Sorghum (Sorghum bicolor (L.) Moench) landrace variation and classification in North Shewa and South Welo, Ethiopia. Euphytica 97:255–263
Thiel T, Michalek W, Varshney RK, Graner A (2003) Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). Theor Appl Genet 106:411–422
USDA (2012) http://www.fas.usda.gov/pecad2/highlights/2002/10/ethiopia/baseline/Eth_Crop_Production.htm. Accessed 14 Nov 2012
Vavilov NI (1926) Studies on the origin of cultivated plants. Bull Appl Bot Genet. Plant Breed USSR 16:1–248 (In Russian with English summary)
Vercken E, Fontaine MC, Gladieux P, Hood ME, Jonot O, Giraud T (2010) Glacial refugia in pathogens: European genetic structure of Anther Smut pathogens on Silene latifolia and Silene dioica. PLoS Pathog 6:e1001229
Vernesi C, Crestanello B, Pecchioli E, Tartari D, Carnelli D, Hauffe H, Bertorelle G (2003) The genetic impact of demographic decline and reintroduction in the wild boar (Susscrofa): a microsatellite analysis. Mol Ecol 12:585–595
von Bothmer R, Jacobsen N, Baden C, Jorgensen RB, Linde-Laursen I (1995) An eco-geographical study of the genus Hordeum. Systematic and Ecogeographic Studies on Crop Genepools, 7, 2nd edn. IPGRI, Rome, p 129
Wang Z, Weber JL, Zhang G, Tanksley SD (1994) Survey of plant short tandem DNA repeats. Theor Appl Genet 88:1–6
Zeisset I, Beebee TJC (2001) Determination of bio-geographical range: an application of molecular phylogeography to the European pool frog Rana lessonae. In: Proceedings of the royal society of London. Series B, Biologica Sci 268:933–938
Zohary D, Hopf M (1993) Domestication of plants in the Old World: the origin and spread of cultivated plants in West Asia, Europe and the Nile Valley, 2nd edn. Clarendon Press, Oxford
Acknowledgments
Tiegist Dejene Abebe would like to thank Katholische Akademischer Ausländer-Dienst (KAAD) for financial support of the research and Alexander von Humboldt Foundation for covering the costs incurred during manuscript preparation. The authors would also like to thank the Institute of Biodiversity Conservation (IBC) of Ethiopia for providing the germplasm and the Ethiopian Institute of Agricultural Research and its Holetta and Kulumsa Agricultural Research Centers for hosting the study and providing research facilities during the field evaluation. The authors are thankful to the Faculty of Agriculture of University of Bonn for covering laboratory costs.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Abebe, T.D., Léon, J. Spatial and temporal genetic analyses of Ethiopian barley (Hordeum vulgare L.) landraces reveal the absence of a distinct population structure. Genet Resour Crop Evol 60, 1547–1558 (2013). https://doi.org/10.1007/s10722-012-9941-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-012-9941-4