Abstract
Introgression of desirable traits from wild relatives plays an important role in crop improvement, as wild species have important characters such as high resistance to pests and pathogens. However, use of wild peanut relatives is challenging because almost all wild species are diploid and sexually incompatible with cultivated peanut, which is tetraploid (AABB genome type; 2n = 4x = 40). To overcome the ploidy barrier, we used 2 wild species to make a tetraploid with the same allotetraploid genome composition as cultivated peanut. Crosses were made between 2 diploid wild species, Arachis valida Krapov. and W.C. Greg. (BB genome; 2n = 2x = 20) and Arachis stenosperma Krapov. and W.C. Greg. (AA genome; 2n = 2x = 20). Cuttings from the diploid F1 AB hybrid were treated with colchicine to induce chromosome doubling thus generating an induced allotetraploid. Chromosome counts confirmed polyploidy (AABB genome; 2n = 4x = 40). We named the new allotetraploid ValSten. Plants had well-developed fertile pollen, produced abundant seed and were sexually compatible with cultivated peanut. ValSten exhibits the same high resistance to early and late leaf spot and rust as its diploid parents. Notably, we observed morphological variations, including flower width and branch angles in the earliest generation (S0) of allotetraploids. A SNP array was used to genotype 47 S0 allotetraploids. The great majority of markers showed the additive allelic state from both parents (AABB). However, some loci were AAAA or BBBB, indicating homeologous recombination. ValSten provides a new, vigorous, highly fertile, disease resistant germplasm for peanut research and improvement.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Peanut or groundnut (Arachis hypogaea L., AABB genome type, 2n = 4x = 40) is an important oil and food crop worldwide. Its production is frequently challenged by pests and pathogens including bacteria, fungi, nematodes and viruses that can cause severe yield loss. For example, early and late leaf spot diseases caused by the fungi Passalora arachidicola (syn. Cersospora arachidicola, teleomorph Mycosphaerella arachidis) and Northopassalora personata (syn. Cercosporidium personatum, teleomorph Mycosphaerella berkeleyi) and rust, caused by Puccinia arachidis, can cause over 50% pod yield losses in some areas where fungicides are used rarely or inappropriately (Waliyar 1990; Ouedraogo et al. 1994). In Georgia, the major peanut producing state in USA, more than 35 million dollars was spent on leaf spot control in 2015. Nevertheless, the losses caused by leaf spots alone was about 8.6 million dollars (https://secure.caes.uga.edu/extension). A reduction in reliance on fungicides and other plant protection products is also strategic because, in addition to cost and any real environmental impact, they are the subject of activist campaigns, and consequently very widespread public misconceptions and ever-tightening regulatory restrictions (EPRS 2019; Hill Cks 2012). Additionally, long-term use of fungicides leads to the emergence of resistant strains; in fact, some leaf spot strains already show resistance to benzimidazole fungicides, such as benomyl and thiophanate methyl (Kemerait R, personal communication). Development of disease resistant cultivars is one of the most efficient, environmentally friendly and economical methods to reduce disease losses in peanut (Holbrook and Culbreath 2007; Ouedraogo et al. 1994). Identification of new sources of resistance is vital, as new races or strains of pathogens may emerge and overcome any current resistance (Chen et al. 2017).
The genus Arachis is divided into 9 sections based on morphology, chromosomal characteristics, and cross-compatibility (Krapovickas and Gregory 1994). Whilst cultivated peanut is tetraploid, almost all other species are diploid. The wild species are only found in some regions of South America, are adapted to different growing conditions, and many of them harbor valuable traits, such as disease and pest resistance (Johnson et al. 1977; Leal-Bertioli et al. 2010; Mallikarjuna et al. 2011), which can be used for peanut improvement. Notably, some traits were only found in wild peanuts but absent in the cultivated peanut gene pool. For instance, no cultivated peanut variety shows strong resistance to root-knot nematode (Meloidogyne arenaria) that can cause substantial yield losses in infested fields (Starr et al. 2002). Furthermore, all root-knot nematode resistant peanut cultivars developed thus far harbor the same source of resistance, which was originally transferred from a single wild species, Arachis cardenasii Krapov. and W.C. Greg. (AA, 2n = 2x = 20) (Simpson and Starr 2001). Recently a strong new source of resistance, A. stenosperma Krapov. and W.C. Greg. (AA, 2n = 2x = 20), has also been found to be transferrable into the tetraploid genetic background (Ballén-Taborda et al. 2019).
Despite their great value for peanut improvement and other studies, direct introgression of traits from wild species into cultivated peanut by conventional hybridization is very challenging because interspecific hybrids between diploid wild species and tetraploid cultivated peanut are triploid and sterile. To overcome this ploidy barrier, the production of allotetraploids through genome duplication of the sterile AB diploid hybrids has proven to be a successful strategy for plant breeding programs (reviewed by Stalker 2017). Here we developed a new allotetraploid by inducing whole genome duplication of the hybrid between the two diploid wild species Arachis valida Krapov. and W.C. Greg. (BB genome, 2n = 2x = 20) and A. stenosperma (AA genome, 2n = 2x = 20). We find that the new induced allotetraploid shows strong resistance to early and late leaf spots and rust. We also observed phenotypic variation despite relatively high genomic stability in the neoallotetraploid individuals. The new synthetic allotetraploid provides a valuable genetic resource for breaking the ploidy barrier and speeding up the introgression of desirable traits from wild species into cultivated peanut.
Materials and methods
Plant material
Seeds of A. valida (accession PI 468,154 = GK 30,011) and A. stenosperma (accession PI666100 = V10309) were obtained from USDA seed bank. Seeds of the commercial peanut cultivars, including Tifrunner, TifNV-High O/L, GA06G, GA09B, GA12Y and NC3033 were obtained from the breeders.
Production of induced allotetraploids
Interspecific hybridization
Seeds of the 2 diploid wild species were treated overnight with 0.5% of Monterey Florel brand growth regulator to break dormancy and then maintained in a growth chamber for 2 weeks. Seedlings were transplanted into pots in greenhouse at the University of Georgia, GA, USA. Arachis species are autogamous, therefore, to avoid self-pollination and maximize cross efficiency, flower buds of A. valida (female parent) were emasculated after 4 PM of the day prior to pollination using fresh pollen of A. stenosperma (male parent), next morning, before 10 AM.
Identification of hybrid plants
Hybrid plants were identified by their aggregated pollen masses and low pollen fertility. Pollen fertility was determined by counting stained (fertile) and unstained (sterile) pollen grains as described by Fishman and Willis (2001). Flowers from different genotypes were taken between 9 and 11 AM and stored at 4 °C for subsequent analysis. To detect the pollen viability, anthers were squashed with tweezers and pollen grains detached from anthers were stained with 2% acetocarmine solution (Tokyo Chemical Industry Co., Ltd. Tokyo, Japan) for 30 s. The numbers of stained and non-stained pollen grains were determined using the Olympus BX 51 microscope. For each individual hybrid plant, 3 slides were prepared from three different anthers and at least 240 pollen grains were observed for each slide.
Induction of whole genome duplication
Chromosome doubling was performed essentially as previously described (Leal-Bertioli et al. 2015a). Briefly, approximately 20 cm sections of lateral branches (cuttings) of confirmed diploid hybrids were collected and immersed in 0.2% (50 mM) colchicine (Sigma-Aldrich, MO) for 18 h. Cuttings were then rinsed 3 times with for 15 min and transplanted into pots filled with sand and kept in the greenhouse. The survival rate was recorded after 30 days, with only the cuttings with new shoots counted and maintained in the same pots until pegs emerged and developed into mature pods. All pods harvested from the treated branches were considered as containing putative induced allotetraploid seed, used for cytological and phenotypic analysis.
Morphological variations of induced allotetraploid plants
Three characteristics were used to evaluate morphological variations in the induced allotetraploids (A. valida x A. stenosperma)4x: flower width, branch angle and pod weight. Flower width was determined using a 0–200 mm Digital caliper (Ted Pella. INC, Redding, CA). At least 8 flowers were measured for each individual and the data was collected on more than 3 different days. The angles of branches were measured for each individual, when the first pair of branches were fully developed, and the second pair of branches had emerged. The pod and seed weights were determined by weighing 5 pods or seeds, and analysis of 3 replicates (15 pods or seeds) were conducted for each sample. To observe variations in architecture/plant growth of ValSten, 24 S1 plants were observed in the field. Seed were germinated and grown for 4 weeks before transplanting to the UGA Southeast Georgia Research and Education Center experimental field in Midville, GA.
Preparation of chromosome spreads and fluorescence in situ hybridization (FISH)
Seeds of the putative induced allotetraploid were germinated on moist filter papers, in Petri dishes and maintained at 25 °C for 5 days. Plantlets were then grown in the greenhouse, for 4 weeks. After that, root tips (5–10 mm long) were isolated from 5 different individuals and the apical meristem was treated with 2 mM 8-hydroxyquinoline for 3 h, at 25 °C. Subsequently, the samples were incubated in a fixative solution containing absolute ethanol: glacial acetic acid (3:1, v/v), at 4 °C for 12 h and then treated with proteolytic enzymes (2% cellulase and 20% pectinase in sodium citrate buffer, pH7.0), at 37 °C for 2 h. Meristematic cells of each root were spread on drop of 45% acetic acid on slides and chromosome spreads were obtained using mechanical pressure onto the coverslip (Schwarzacher and Heslop-Harrison 2000). Slides containing well spread metaphases were selected using the phase contrast mode in the AxiosKop microscope (Zeiss, Oberkochen, Germany).
To observe and count chromosomes, selected slides were mounted with the Vectashild media (Vector Laboratories, Burlingame, California-USA), added to 4′, 6-diamidino-2-phenylindole (DAPI) fluorophore solution (2 µg/ml McIlvaine buffer pH 7.0) and visualized in the epifluorescence Zeiss AxioPhot Microscope (Zeiss, Oberkochen, Germany).
Ribosomal DNA sequences coding for the 5S (rDNA 5S) of Lotus japonicus (Pedrosa et al. 2002) and 18S-5.8S-25S sequences (rDNA 45S) of Arabidopsis thaliana (Wanzenböck et al. 1997) were used to obtain the DNA probes for Flourescent In Situ Hybridization (FISH). DNA sequences were labelled with, either digoxigenin-11-dUTP or Cy3-dUTP, by Nick Translation using the Nick Translation Mix kit from Roche (Basel, Switzerland). FISH was performed according to the established methods (Schwarzacher and Heslop-Harrison 2000). Chromosomes were observed in the epifluorescence microscope and the images captured using the Axiocam MRc digital camera, with the Axiovision Rel. 4.8 software. Images were treated using the Adobe Photoshop CS software.
Evaluation of disease resistance
Six different events originated from the same cross (A. valida x A. stenosperma)4× were evaluated for resistance to rust (Puccinia arachidis), early and late leaf spot (Passalora arachidicola syn. Cercospora arachidicola) and Northopassalora personata syn. Cercosporidium personatum). The method of detached leaf assays was used, as described in Leal-Bertioli et al. (2015b). In short, spores were collected from infested peanut plants in Tifton, GA. Rust inoculum was collected from the cultivars Tifguard, Tifrunner and NC3033 and LLS and ELS were collected from a population C1799 (cross between Tifrunner and NC3033). Leaves were inoculated with 105 spores/ml of 0.005% Tween 20 and incubated at 20–24 °C (ELS and LLS) or 26–28 °C (rust) and photoperiod of 10 h light and 14 h dark. Each experiment was conducted twice.
DNA extraction and genotyping
Genomic DNA was extracted from young leaves using the DNeasy Plant Mini Kit from QIAGEN (Hilden, Germany). DNA concentrations were measured using Qbit Fluorometric Quantification (Waltham, MA), and ~ 50 ng of DNA of each sample was used for genotyping.
In order to observe structural genomic variation in different events of polyploidization, a total of 47 synthetic allotetraploids (A. valida × A. stenosperma)4x individuals here obtained were genotypsed. Genotyping was conducted with the version 02 of the Affymetrix microarray (Thermo Fisher Scientific, Waltham, MA) containing 47,836 single nucleotides polymorphisms (Korani et al. 2019). SNP calling was performed with Axiom Analysis Suite software (Version1.1) provided by Affymetrix. Genotyping data was manually curated. For genomic analyses, only SNPs that had the following characteristics were used: (1) SNPs that showed polymorphic alleles between the diploid wild parents (2) SNPs that showed consistent calling between replicates. SNPs that were monomorphic and/or inconsistent between the replicates were discarded.
Results
Production of induced allotetraploids
In 2015, between July 2 and August 7, 185 crosses were made between A. valida (female) and A. stenosperma (male). No reciprocal crosses were attempted as flowers of AA genome species tend not to support artificial pollination (not published). Thirty-one putative hybrid pods were harvested on october 28, 2015. Eleven seedlings prematurely germinated in the same pot before harvest, a characteristic sign of hybridity. Ten out of the 11 seedlings exhibited yellow flower, a dominant character present in the male parent, therefore, this phenotypic marker confirmed hybridity (Online Resource 1A). Anthers of F1 hybrids were thin and shriveled, compared the diploid parents (Online Resource 1B). Pollen grains of both A. valida and A. stenosperma were mostly stained, indicating high levels of viability. However, pollen from the hybrid seedlings were smaller and lacked staining, indicating high infertility (Online Resource 1C).
Eighty cuttings from diploid hybrids were treated with 0.2% colchicine solution to induce chromosome doubling (summer of 2016). Fifty-three cuttings survived and developed new branches and flowers. Some flowers developed into budding ovaries (pegs), which penetrated the sand and developed into pods (Fig. 1). A total of 47 pods and 44 seedlings (totaling 91 individuals) were obtained from the colchicine-treated cuttings. The pods and seedlings produced from the treated cuttings imply successful whole chromosome duplication in the diploid cuttings. We refer to these pods and seedlings as ValSten hereafter. Untreated diploid hybrids did not produce pods even after two years being maintained in the vegetative state.
The process of generation of an induced allotetraploid peanut via the tetraploid route. Branches (cuttings) are collected from the diploid F1 hybrid plant (AB genome) a, treated with colchicine solution and maintained in pots filled with wet sand b; surviving cuttings can grow and develop new branches, leaves and flowers c, pegs d and pods e. All the process, from cutting treatment to pod harvest, takes over seven months
Morphological variations of the induced allotetraploid
We investigated the morphological characters of the diploid wild parents and 48 synthetic tetraploid plants at S0, which is the generation immediately after chromosome doubling. The ValSten had a prominent main stem, which is similar to A. valida, and different from A. stenosperma (Fig. 2a). All 48 ValSten produced yellow flowers, which were as same as that of the male parent, A. stenosperma and the diploid hybrid. Figure 2b, Online Resource 1A). We also investigated the viability of mature pollen grains in the diploid parents and ValSten allotetraploids, with all of them showing high number of stained pollen, indicating high fertility (Fig. 2c, Table 1). ValSten plants produced pods similar in shape and size as A. valida (Figs. 2d, 3e).
As all the induced allotetraploid were derived from a same F1 hybrid combination and could have the same expected genome composition, and therefore, similar phenotypes. However, the 48 induced allotetraploids plants from different events of tetraploidization varied. For instance, the average flower width of A. valida and A. stenosperma was 16.32 ± 1.28 and 15.54 ± 0.9 mm, respectively. Flower width of the 48 tetraploids varied from 14.1 to 20.9 mm; most of them (83%, 40/48) produced larger flowers than TifNV-High O/L (17.21 ± 0.86 mm) (Fig. 3a). Most flowers produced by the 48 ValSten individuals showed regular development and morphology: they consisted of a large banner and 2 lateral wings (Fig. 3b). However, some flowers with morphological irregularities were occasionally observed, including double or anomalously developed layers of banner (Fig. 3c) or with chimeric color patterns (white/yellow) (Fig. 3d). The angle between basal branches is an important trait of peanut, since it defines plant architecture and is one of the traits that distinguish the market types (Kayam et al 2017). The first pair of branches of both A. valida and A. stenosperma are spread and nearly parallel to the soil (Figs. 4a, b). Most of the induced allotetraploids had more upright branches with angles lower than the parents and of TifNV-High O/L, with the smallest angle corresponding to 40° (Fig. 4c–e, Online Resource 2). Pods and seeds of six events of ValSten were also compared. All were significantly heavier (t test, p < 0.003) than that of A. stenosperma. Three were generally similar to those of A. valida, with three ValSten-2, ValSten-3 and ValSten-9 producing heavier pods and seeds than that of A. valida (t test, p < 0.01) although, as expected significantly lighter (T test, p < 0.007) than those of TifNV-High O/L (Online Resource 3). Dramatic phenotypic differences were also observed among the S1 ValSten plants in the field. Some plants exhibited more spreading growth habits (Fig. 5a–c), while others were more compact (Fig. 5d–g). Large variations were also observed in plant sizes (Fig. 5a–k).
Flowers from Valsten plants a histogram showing the range of flower width of different ValSten plants: 14.3 to 20.9 mm. Arrows show flower width averages of A. stenosperma (Sten, 15.5 mm), A. valida (Val, 16.3) and A. hypogaea TifNV-H/O (TifNV, 17.2); b a regular yellow flower, with two wings and a single banner; c two abnormal flowers with unusual shape and 2 layers of banners; d a flower with mosaic white and yellow patterns
Cytogenetic analyses
We examined the chromosome numbers from root apical meristem cells of ValSten. Forty mitotic chromosomes, including 38 metacentric and 2 submetacentric chromosomes were observed (Fig. 6a), which are equal to the sum of its diploid parents, A. stenosperma (AA, 2n = 2x = 20) and A. valida (BB, 2n = 2x = 20) (Robledo and Seijo 2010; Robledo et al. 2009). Ten pairs of chromosomes contained DAPI+ bands on their centromeric regions, which correspond to the A subgenome chromosomes from A. stenosperma. The other ten pairs lacked detectable DAPI+ bands, indicating that they were derived from A. valida. It is worth to note that cyt-B3 of ValSten presented distal DAPI+ bands (Fig. 6a), despite being a metacentric chromosome in A. valida (Robledo and Seijo 2010) (Because most correspondences between cytogenetic and sequenced chromosome assignments are unknown we will distinguish cytogenetic numbering by using the prefix “cyt- “). The small pair of chromosomes cyt-A9 and the submetacentric satellite (SAT) chromosomes, the cyt-A10, were also observed (Fig. 6a).
The number and localization of the 5S rDNA and 45S rDNA loci can be used as additive characters to distinguish different chromosomes in the diploid species. The 5S rDNA loci were found on the proximal regions of chromosomes cyt-A3 and cyt-B3 of ValSten (Fig. 6b), consistent with the corresponding chromosomes in the diploid progenitors (Robledo and Seijo 2010; Robledo et al. 2009). Seven 45S rDNA loci were identified in ValSten (Fig. 6b), including the strong signals at the proximal regions of cyt-A2, cyt-A7, cyt-B4 and cyt-B7 chromosomes. The cyt-B10 chromosome contained one 45S rDNA locus on the proximal and another on the terminal region of the arm. However, the 45S rDNA locus on cyt-A10 was situated near the secondary constriction of the SAT chromosome (Fig. 6b), forming a thread-like region between the SAT and the long arm of the chromosome. This thread-like region is typically observed whenever DNA is under transcription, a characteristic of nucleolus organizer regions (NORs). Following earlier description by Robledo et al. (2009) for the corresponding chromosomes of A. stenosperma, here we also conclude that cyt-A10 has a NOR.
Cytogenetic analysis of ValSten. DAPI bands (bright white) on centromere regions of the chromosomes of subgenome A (red arrows) and subgenome B (green arrows) a. FISH using DNAr 5S and 45S probes and sites detected in green and red (arrows), respectively b. A10: short arm and proximal segment of the long arm (A10*) and the satellite (A10°). Barra: 5 μm
Disease evaluation
Resistance to early leaf spot (ELS), late leaf spot (LLS) and rust in ValSten and eight peanut cultivars was evaluated. Susceptibility to leaf spot was determined based on the number of lesions (incidence) and percentage of leaf area affected by the symptoms of the disease (DLA, severity). Five control peanut cultivars were common to all experiments. These controls were all infected by these pathogens, showing varying numbers of lesions. All 5 cultivars were highly susceptible to rust (Online Resource 4). No ELS, LLS or rust lesions were detected on the leaves of ValSten (Online Resource 4, Fig. 7a). All other cultivars, including Bailey, that presents partial resistance to ELS, showed lesions (Figs. 7b, c).
Genome-wide SNP analysis of genome stability of synthetic allotetraploids
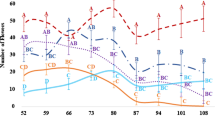
Since phenotypic variations were detected among the induced allotetraploid individuals, we hypothesized that there may have been genomic changes during polyploidization. To test this, DNAs of A. stenosperma (AA genome), A. valida (BB genome) and 47 induced allotetraploids (AABB genome) were genotyped with the 48 K Affymetrix chip (Korani et al. 2019). After filtering out all loci which or detected no polymorphisms between of A. stenosperma and A. valida, 1,312 loci were kept for genome-wide comparison. All ValSten presented SNPs from both parents on most loci (99.36%), having the expected genome configuration: AABB. This is consistent with the cytogenetic analysis (Fig. 6). A small number of loci deviated from this configuration, they were either AAAA (169, 0.27%) or BBBB (222, 0.36%), indicating some type of homeologous recombination or gene conversion, and a slight bias towards conversion to A. valida alleles BBBB (Fig. 8, Online Resource 5). These loci were present all chromosomes and tended to occur in short stretches of adjacent markers, representing short chromosome segments. Chromosome A06/B06 (1.47%), A03/B03 (0.98%) and A02/B02 (0.81%) had the highest level of homeologous recombination (Figs. 8a, b, Online Resource 5). Estimated genetic similarities between different ValSten plants ranged between 0.86 and 0.96 (Online Resource 6), these estimates being certainly inflated by noise. Estimated genetic similarities between the virtually genetically identical sibling seeds from the same accessions ranged between 0.91 and 0.97.
Discussion
A new source of allelic variation for peanut improvement
Modern plant breeding has made great progress in improving the yield and nutritional qualities of crops and continues making enormous impacts on food security worldwide. However, a key vulnerability is relative lack of genetic variation among commercial cultivars of important species (Fu 2015). This provides a fundamental limitation to the genetic gains that can be made, and also increases the risk of crop disease epidemics (Keneni et al. 2012). Wild crop relatives play pivotal roles in crop improvement and sustainable agriculture. They represent a large pool of genetic diversity, a valuable resource for improving yield, and adaptation to changing disease pressures, and other traits (Dempewolf et al. 2017). Thus far, many desirable traits from wild crop relatives have been successfully used for crop improvement, including improved yield traits, drought tolerance, disease resistance and nutritional value in legumes (reviewed by Zhang et al. 2019. In the case of peanut, introgression of new alleles from wild species is hindered by the ploidy differences. In this work, from the wild species A. valida and A. stenosperma, we created a new induced allotetraploid (ValSten) that is sexually compatible with cultivated peanut. This neo allotetraploid is being used to transfer wild alleles from the wild species into cultivated peanut, by crossing and backcrossing. Notably, ValSten is very vigorous, produces large biomass and is prolific in seed production. The male parent A. stenosperma V10309 is highly resistant to root-knot nematode (Proite et al. 2008) and this resistance is genetically mapped and transferrable to tetraploid peanut (Ballén-Taborda et al. 2019). Furthermore, ValSten showed complete resistance to three peanut major diseases: early and late leaf spot and rust.
The creation of ValSten (and others being developed in our research program) are intended for use as a source of valuable alleles in breeding. They have the advantage of being directly sexually compatible with cultivated peanut, and of having identified DNA markers linked to a subset of their alleles. These alleles are valuable once incorporated into the genetic background of cultivated peanut, but as outlined by Tanksley and Nelson (1996) and Tanksley and McCouch (1997), their value is mostly 'masked' whilst in a wild genetic background. In peanut, a clear example of that is the increase in seed size and biomass by introgressing alleles from the neo allotetraploid IpaDur1 that has small seed and low biomass (Faye et al. 2016; Leal-Bertioli et al. 2018; Suassuna et al. 2015). This indicates that ValSten has the potential to contribute useful alleles besides the ones that confer resistance to fungal pathogens.
Cytological characterization of the synthetic allotetraploid
In this study, the chromosomes of the allotetraploid ValSten were characterized by the visualization of heterochromatic regions of DNA rich in A-T using DAPI staining, and in situ localization of 5S and 45S rDNA loci. Karyotypes were compared to the progenitor species and other allotetraploids. Our data confirmed that ValSten has the expected 40 chromosomes and karyotype almost identical to the sum of its diploid parents. The majority of chromosomes were metacentric, except for 2 pairs of chromosomes that are submetacentric. The metaphase chromosomes from 2 different genomes, A and B, could be distinguished by their staining with DAPI. Chromosomes of the subgenome A of ValSten (A. stenosperma) having bands on the centromeric and pericentromeric regions (as is well established for the A genome species; Seijo et al. 2004; Robledo et al. 2009; Nascimento et al. 2018). Notably, however, although no DAPI band is detected in the chromosomes of the diploid B genome species, A. valida, A. ipaensis and A. magna (Robledo and Seijo 2010), we found distal DAPI+ bands on cyt-B3 of ValSten. Ribosomal DNA (rDNA) locations and number in ValSten were consistent with the sum of the diploid parents. The ValSten putative Nuclear Organizing Region (NOR) shows A genome nucleolar dominance, being present only on cyt-A10, and not cyt-B10.
These results can be compared to allotetraploids derived from A. ipaënsis and A. duranensis, the spontaneous allotetraploids A. hypogaea and A. monticola (Seijo et al. 2004; Robledo and Seijo 2010; Robledo et al. 2009), and induced allotetraploids, IpaDur1 (Nascimento et al. 2018). At least in the initial generations studied here, ValSten seems more similar in behavior to the more karyotypically stable spontaneous allotetraploids A. hypogaea and A. monticola, than the induced IpaDur1. All of these allotetraploids show the sum of DAPI bands essentially being the sum of their parental diploid species. All show A genome nucleolar dominance. However, in the IpaDur1 the rDNAs are unexpected, they are not additive of the parental species.
Phenotypic and genomic variations in synthetic allotetraploids
Polyploidy plays a critical role in the evolution and diversification of flowering plants it is thought to increase biological complexity, generate evolutionary novelties and improve adaption to changing environmental stresses (Van de Peer et al. 2009). It is now known that virtually all angiosperm lineages have undergone polyploidy at some stage in their evolutionary pasts. Indeed, most lineages have undergone multiple rounds of polyploidy. Species that have undergone recent whole genome duplications are easily identifiable as polyploid. Species whose most recent whole genome duplication is more ancient have undergone evolutionary processes, such as gene loss, that return them to a “diploid” state (Soltis et al. 2015). Cultivated peanut is an allotetraploid that arose very recently, less than 10,000 years ago. The interspecific hybridization between the diploid ancestors, A. duranensis and A. ipaensis was enabled by the human movement of A. ipaensis into the range of A. duranensis by proto-agriculturalists in prehistory (Bertioli et al. 2016). Although multiple diploid wild species were cultivated before the polyploid A. hypogaea, it was the polyploid species that became fully domesticated—compelling evidence that there was some “polyploid advantage” for domestication (Bertioli et al. 2019). By the observation of new induced allotetraploids, like ValSten, it is possible to begin to glimpse at what this advantage may have been. Here we observed that ValSten is exceptionally vigorous. It has larger flowers (Online Resource 2), larger leaves and more biomass (data not shown). However, the sizes of pods and seeds were only slightly larger than that of the diploid parents (Fig. 2d, e, Online Resource 3). This result indicates that artificial selection, more than the immediate effect of polyploidy has been the main cause of larger seed and pod sizes in cultivated peanut, as previously found by the observation of other induced allotetraploids (Leal-Bertioli et al., 2017).
The production of Arachis synthetic allotetraploids is a time-consuming task, starting with the generation of diploid hybrids. Once produced, they must be allowed to grow sufficiently to produce cuttings for treatment with colchicine. Survival rate of these cuttings is generally very low as is the recovery of fertility and the production of tetraploid seeds. In the case of ValSten, the recovery of fertility and production of seeds was the highest we have ever observed during more than ten years of our work producing induced allotetraploids (Leal-Bertioli et al. 2015a, 2017 and unpublished data). More than 90 S0 ValSten plants were obtained, allowing us to compare many different events at early generations. The naïve expectation, for plants of pure allotetraploid genetic behavior would be that all plants would be the same phenotypically. This was certainly not the case: extensive morphological differences were found among the S0 and S1 tetraploids, including flower width and color, angle between side branches, pod weight, plant architecture among others (These morphological changes are similar to those observed in smaller number in other allotetraploid plants; unpublished). Nevertheless, in spite of this morphological variability, as was consistent with karyotypic characterization, genotyping revealed relative genome stability. Most SNP loci had the expected AABB composition and that there was little variation between different ValSten genotypes. Only 0.64% of markers had unexpected tetrasomic/nullisomic composition (AAAA or BBBB).
An alternative approach for wild species conservation
Increasing anthropic action has destroyed the natural habitats of many crop wild relatives (Schoen and Brown 2001) many are now are endangered (Humphreys et al. 2019). Indeed, the location of the collection of the A. stenosperma (V10309; 16°28′14.54"S 54°39′19.30"W) used to make ValSten has been overtaken by urban sprawl, and the population is now extinct in the wild. Seed banks are of profound importance for species conservation and maintenance of genetic diversity (Schoen and Brown 2001; Hay and Probert 2013). However, the climates and photoperiods of the banks are often different from the native habitats. Thus, some wild species, do not grow well and/or produce only few or no seeds in the places where the seedbanks are located.
sThe genus Arachis is native to South America (Krapovickas and Gregory, 1994), the USDA seedbank holds around 500 accessions of wild peanut species. However, some accessions such as Grif 14,535 (A. simpsonii) can develop flowers, but don’t produce seeds in greenhouse, therefore they are maintained and exchanged via vegetative tissue. Arachis stenosperma V10309 (PI666100), grows vigorously in Brazil but it grows weakly in the USA, even in greenhouse conditions and produces only about 20 seeds per plant. In contrast, the induced allotetraploid ValSten grows well and often produces over 100 seeds per plant. We propose that the allotetraploids derived from wild relatives of peanut can serve as an additional option for the maintenance and distribution of genetic diversity. Induced allotetraploids often produce abundant seed, harbor essentially the genetic information from both parents, and critically, are directly compatible with peanut, thus being directly usable in breeding programs seeking to increase genetic diversity and introgress desirable traits from wild species into the peanut crop.
Data availability
All data is in the Online Resources. The genotypes are being release as germplasm and will be deposited at the USDA-PGRCU (Plant Genetic Resources Conservation Unit at the United States Department of Agriculture).
References
Ballén-Taborda C, Chu Y, Ozias-Akins P, Timper P, Holbrook CC, Jackson SA, Bertioli DJ, Leal-Bertioli SCM (2019) A new source of root-knot nematode resistance from Arachis stenosperma incorporated into allotetraploid peanut (Arachis hypogaea). Sci Rep 9:17702. https://doi.org/10.1038/s41598-019-54183-1
Bertioli DJ, Cannon SB, Froenicke L, Huang G, Farmer AD, Cannon EK, Liu X, Gao D, Clevenger J, Dash S, Ren L, Moretzsohn MC, Shirasawa K, Huang W, Vidigal B, Abernathy B, Chu Y, Niederhuth CE, Umale P, Araújo AC, Kozik A, Kim KD, Burow MD, Varshney RK, Wang X, Zhang X, Barkley N, Guimarães PM, Isobe S, Guo B, Liao B, Stalker HT, Schmitz RJ, Scheffler BE, Leal-Bertioli SC, Xun X, Jackson SA, Michelmore R, Ozias-Akins P (2016) The genome sequences of Arachis duranensis and Arachis ipaensis, the diploid ancestors of cultivated peanut. Nat Genet 48:438–446. https://doi.org/10.1038/ng.3517
Bertioli DJ, Jenkins J, Clevenger J, Dudchenko O, Gao D, Seijo G, Leal-Bertioli SCM, Ren L, Farmer AD, Pandey MK, Samoluk SS, Abernathy B, Agarwal G, Ballén-Taborda C, Cameron C, Campbell J, Chavarro C, Chitikineni A, Chu Y, Dash S, El Baidouri M, Guo B, Huang W, Kim KD, Korani W, Lanciano S, Lui CG, Mirouze M, Moretzsohn MC, Pham M, Shin JH, Shirasawa K, Sinharoy S, Sreedasyam A, Weeks NT, Zhang X, Zheng Z, Sun Z, Froenicke L, Aiden EL, Michelmore R, Varshney RK, Holbrook CC, Cannon EKS, Scheffler BE, Grimwood J, Ozias-Akins P, Cannon SB, Jackson SA, Schmutz J (2019) The genome sequence of segmental allotetraploid peanut Arachis hypogaea. Nat Genet 51(5):877–884. https://doi.org/10.1038/s41588-019-0405-z
Chen J, Upadhyaya NM, Ortiz D, Sperschneider J, Li F et al (2017) Loss of AvrSr50 by somatic exchange in stem rust leads to virulence for Sr50 resistance in wheat. Science 358:1607–1610
Dempewolf H, Baute G, Anderson J, Kilian B, Smith C, Guarino L (2017) Past and future use of wild relatives in crop breeding. Crop Sci 57:1070–1108
EPRS | European Parliamentary Research Service. Farming without plant protection products, Can we grown without using herbicides, fungicides and insecticides (2019) In-depth analysis, Panel for the Future of Science and Technology. https://www.europarl.europa.eu/RegData/etudes/IDAN/2019/634416/EPRS_IDA(2019)634416_EN.pdf
Faye I, Hodo-Abalo T, Ndoye O, Fonceka D (2016) Dossier technique d’homologation de nouvelles variétés d’arachide. In Institut Sénegalais de Rescherches Agricoles (ISRA) [ed.], Senegal, 34. ISRA, Dakar, Senegal
Fishman L, Willis JH (2001) Evidence for Dobzhansky-Muller incompatibilities contributing to the sterility of hybrids between Mimulus guttatus and M. nasutus. Evolution 55:1932–1942
Fu YB (2015) Understanding crop genetic diversity under modern plant breeding. Theor Appl Genet 128(11):2131–2142. https://doi.org/10.1007/s00122-015-2585-y
Hay FR, Probert RJ (2013) Advances in seed conservation of wild plant species: a review of recent research. Conserv Physiol 1(1):030
Hillocks RJ (2012) Farming with fewer pesticides: EU pesticide review and resulting challenges for UK agriculture. Crop Prot 31:85–93
Holbrook CC, Culbreath AK (2007) Registration of ‘Tifrunner’ Peanut. J Plant Reg 1:124
Humphreys AM, Govaerts R, Ficinski SZ, Nic Lughadha E, Vorontsova MS (2019) Global dataset shows geography and life form predict modern plant extinction and rediscovery. Nat Ecol Evol 3:1043–1047. https://doi.org/10.1038/s41559-019-0906-2
Johnson DR, Wynne JC, Campbell WV (1977) Resistance of wild species of Arachis to the two spotted spider mite, Tetranychus urticae. Peanut Sci 4:9–11
Kayam G, Brand Y, Faigenboim-Doron A, Patil A, Hedvat I, Hovav R (2017) Fine-mapping the branching habit trait in cultivated peanut by combining bulked segregant analysis and high-throughput sequencing. Front Plant Sci 8:467
Keneni G, Bekele E, Imtiaz M, Dagne K (2012) Genetic vulnerability of modern crop cultivars: causes, mechanism and remedies. Int J Plant Res 2:69–79
Korani W, Clevenger JP, Chu Y, Ozias-Akins P (2019) Machine learning as an effective method for identifying true Single Nucleotide Polymorphisms in polyploid plants. Plant Genome 12. https://doi.org/10.3835/plantgenome2018.05.0023
Krapovickas A, Gregory WC (1994) Taxonomía del género Arachis (Leguminosae). Bonplandia 8:1–186
Leal-Bertioli SCM, de Farias MP, Silva PIT, Guimarães PM, Brasileiro ACM, Bertioli DJ, Guerra de Araújo AC (2010) Ultrastructure of the initial interaction of Puccinia arachidis and Cercosporidium personatum with leaves of Arachis hypogaea and Arachis stenosperma. J Phytopathol 158:792–796
Leal-Bertioli SCM, Cavalcante U, Gouveia EG, Ballen-Taborda C, Shirasawa K, Guimaraes PM, Jackson SA, Bertioli DJ, Moretzsohn MC (2015a) Identification of QTLs for rust resistance in the peanut wild species Arachis magna and the development of KASP markers for Marker Assisted Selection. Genes/Genomes/Genet 5(7):1403–1413. https://doi.org/10.1534/g3.115.018796
Leal-Bertioli SCM, Santos SP, Dantas KM, Inglis PW, Nielen S, Araujo ACG, Silva JP, Cavalcante U, Guimarães PM, Brasileiro ACM, Carrasquilla-Garcia N, Penmetsa RV, Cook D, Moretzsohn MC, Bertioli DJ (2015b) Arachis batizocoi: a study of its relationship to cultivated peanut (A. hypogaea) and its potential for introgression of wild genes into the peanut crop using induced allotetraploids. Ann Bot 115:237–249. https://doi.org/10.1093/aob/mcu237
Leal-Bertioli SCM, Moretzsohn MC, Santos SP, Brasileiro ACM, Guimarães PM, Bertioli DJ, Araujo ACG (2017) Phenotypic effects of allotetraploidization of wild Arachis and their implications for peanut domestication. Am J Bot 104:379–388. https://doi.org/10.3732/ajb.1600402
Leal-Bertioli SCM, Godoy IJ, Santos JF, Doyle JJ, Guimarães PM, Abernathy BL, Jackson SA, Moretzsohn MC, Bertioli DJ (2018) Segmental allopolyploidy in action: Increasing diversity through polyploid hybridization and homoeologous recombination. Am J Bot 105:1–14
Mallikarjuna N, Senthilvel S, Hoisington D (2011) Development of new sources of tetraploid Arachis to broaden the genetic base of cultivated groundnut (Arachis hypogaea L.). Genet Resour Crop Evol 58:889–907
Nascimento EFDM, Santos BV, Marques LO, Guimarães PM, Brasileiro AC, Leal-Bertioli SC, Bertioli DJ, Araujo AC (2018) The genome structure of Arachis hypogaea (Linnaeus, 1753) and an induced Arachis allotetraploid revealed by molecular cytogenetics. Comp cytogenet 12:111–140
Ouedraogo M, Smith OD, Simpson CE (1994) Smith DH (1994) Early and late leaf spot resistance and agronomic performance of nineteen interspecific derived peanut lines. Peanut Science 21:99–104
Pedrosa A, Sandal N, Stougaard J, Schweizer D, Bachmair A (2002) Chromosomal map of the model legume Lotus japonicus. Genet 161:1661–1672
Proite P, Carneiro R, Falcão R, Gomes A, Leal-Bertioli S, GuimarãesBertioli PD (2008) Post-infection development and histopathology of Meloidogyne arenaria race 1 on Arachis spp. Plant Pathol on line. https://doi.org/10.1111/j.1365-3059.2008.01861.x
Robledo G, Seijo G (2010) Species relationships among the wild B genome of Arachis species (section Arachis) based on FISH mapping of rDNA loci and heterochromatin detection: A new proposal for genome arrangement. Theor Appl Genet 121:1033–1046. https://doi.org/10.1007/s00122-010-1369-7
Robledo G, Lavia GI, Seijo G (2009) Species relations among wild Arachis species with the A genome as revealed by FISH mapping of rDNA loci and heterochromatin detection. Theor Appl Genet 118:1295–1307. https://doi.org/10.1007/s00122-009-0981-x
Schwarzacher T, Heslop-Harrison J (2000) Practical in situ hybridization. BIOS Scientific Publishers Ltd., Oxford
Schoen DJ, Brown AHD (2001) The Conservation of wild plant Species in seed banks: attention to both taxonomic coverage and population biology will improve the role of seed banks as conservation tools. Bioscience 51:960–966
Seijo JG, Lavia GI, Fernández A, Krapovickas A, Ducasse D, Moscone EA (2004) Physical mapping of the 5S and 18S–25S rRNA genes by FISH as evidence that Arachis duranensis and A. ipaënsis are the wild diploid progenitors of A. hypogaea (Leguminosae). Am J Bot 91:1294–1303. https://doi.org/10.3732/ajb.91.9.1294
Simpson CE, Starr JL (2001) Registration of COAN peanut. Crop Sci 41:918
Soltis PS, Marchant DB, Van de Peer Y, Soltis DE (2015) Polyploidy and genome evolution in plants. Curr Opin Genet Dev 35:119–125
Stalker HT (2017) Utilizing wild species for peanut improvement. Crop Sci 57:1102–1120
Starr JL, Morgan ER, Simpson CE (2002) Management of the peanut root-knot nematode, Meloidogyne arenaria, with host resistance. Plant Health Progr. https://doi.org/10.1094/PHP-2002-1121-01-HM
Suassuna TMF, Suassuna ND, Moretzsohn MC, Leal-Bertioli SCM, Bertioli DJ, Medeiros EP (2015) Yield, market quality, and leaf spots partial resistance of interspecific peanut progenies. Crop Breed Applied Biotechnol 15:175–180
Tanksley SD, McCouch SR (1997) Seed banks and molecular maps: unlocking genetic potential from the wild. Science 277:1063–1066
Tanksley SD, Nelson JC (1996) Advanced backcross QTL analysis: a method for the simultaneous discovery and transfer of valuable QTLs from unadapted germplasm into elite breeding lines. Theor Appl Genet 92:191–203
Van de Peer Y, Maere S, Meyer A (2009) The evolutionary significance of ancient genome duplications. Nat Rev Genet 10:725–732
Waliyar F (1990) Evaluation of yield losses due to groundnut leaf diseases in West Africa. In: Nduguru BJ, Waliyar F, Ntare BR (eds) Summary Proceedings of the Second ICRISAT Regional Groundnut Meeting for West Africa. Niger, Niamey, pp 32–33
Wanzenböck EM, Schöfer C, Schweizer D, Bachmair A (1997) Ribosomal transcription units integrated via T-DNA transformation associate with the nucleolus and do not require upstream repeat sequences for activity in Arabidopsis thaliana. Plant J 11:1007–1016. https://doi.org/10.1046/j.1365-313X.1997.11051007.x
Zhang H, Yasmin F, Song BH (2019) Neglected treasures in the wild-legume wild relatives in food security and human health. Curr Opin Plant Biol 49:17–26
Acknowledgements
The authors appreciate Jenny Leveret, Carlos Ruiz and Rida Osman for their technical assistance. This work was supported by Agriculture and Food Research Initiative Competitive Grant no. 2018-67013-28139 co-funded by the USDA National Institute of Food and Agriculture and the National Peanut Board, grants from National Science Foundation (grant # MCB-1543922), Georgia Peanut Commission, Peanut Research Foundation and Mars Wrigley.
Funding
This work was supported by Agriculture and Food Research Initiative Competitive Grant no. 2018–67013-28139 co-funded by the USDA National Institute of Food and Agriculture and the National Peanut Board, grants from National Science Foundation (grant # MCB-1543922), Georgia Peanut Commission, Peanut Research Foundation and Mars Wrigley.
Author information
Authors and Affiliations
Contributions
DG, SLB, DJB and SJ conceived and designed the study. DG and HX made cross. DG generated synthetic allotetraploids. EN and AA conducted cytogenetic analysis. SB, CC, DG and DJB performed SNP array analysis. SB conducted in vitro disease evaluation. DG and SLB drafted the manuscript. All authors were involved in manuscript revision and approved the submitted version.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no competing interests.
Human or Animal rights
This research does not involve human participants or animals.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary information.
10722_2020_1076_MOESM1_ESM.pptx
Online Resource 1. Comparison of flowers (A), anthers (B) and carmin acetic stained pollens (C) of two diploid wild parents A. valida and A. stenosperma and their F1 hybrid. (PPTX 290 kb)
10722_2020_1076_MOESM2_ESM.pptx
Online Resource 2. Angle between basal branches of 46 S0 ValsSten individuals (blue), A. stenosperma (red), A. valida (yellow) and A. hypogaea TifNV-H/O (green). (PPTX 47kb)
10722_2020_1076_MOESM3_ESM.pptx
Online Resource 3. Seed weight of six S0 ValsSten individuals (blue), A. stenosperma (red), A. valida (yellow) and A. hypogaea TifNV-H/O (green). Note that seeds of three individuals (ValSten2, 3 and 9) are significantly heavier than those of either parent. (PPTX 46 kb)
10722_2020_1076_MOESM4_ESM.xlsx
Online Resource 4. Data of in vitro bioassays of early and late leaf spot, and rust on the induced allotetraploid ValSten and seven different peanut cultivars. (XLSX 13kb)
10722_2020_1076_MOESM5_ESM.xlsx
Online Resource 5. Filtered genotyping data of 47 S0 ValsSten individuals and the diploid parents A. stenosperma V10309 (in duplicate) and A. valida PI468154 (in triplicate) obtained using the version 02 of the Affymetrix microarray (Thermo Fisher Scientific, Waltham, MA). (XLSX 271kb)
10722_2020_1076_MOESM6_ESM.xlsx
Online Resource 6. Similarity matrix constructed with 1,312 loci from SNP genotyping data (Online Resource 5). Similarity between 47 different S0 ValSten individuals ranged between 86 and 96%.(XLSX 31kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gao, D., Araujo, A.C.G., Nascimento, E.F.M.B. et al. ValSten: a new wild species derived allotetraploid for increasing genetic diversity of the peanut crop (Arachis hypogaea L.). Genet Resour Crop Evol 68, 1471–1485 (2021). https://doi.org/10.1007/s10722-020-01076-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-020-01076-2