Abstract
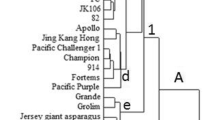
Destruction in wild resource and decline in genetic diversity of Scutellaria baicalensis are becoming a big problem urgently needed be dealt in China. To help efficiently utilize and conserve this medicinal plant resources, ISSR analysis was employed to reveal genetic diversity of 107 S. baicalensis accessions consisting of 82 wild and 25 cultivated germplasm resources from 6 populations in two main producing provinces in China. Only eighteen ISSR primers that resulted in accuracy and credibility, high polymorphism, well stability PCR products were selected. The maximal and minimal bands resulted from an ISSR primer are 35 and 18 respectively with a total of 480 bands were resolved in the agarose gels with an average of two alleles per locus. As indicated by the index of genetic diversity (Na = 1.96, Ne = 1.39, He = 0.25, I = 0.39), there were richness of genetic diversity in the collected samples and that the S. baicalensis germplasm had a large representativeness as primary collection. Cluster analysis by UPGMA and STRUCTURE placed the 107-germplasm resources into three linkage clusters. The samples from all the collection locations were clustered together, but the populations from one province were not clustered together. Compared to the RS and SW strategies, the LDSS proved to be more representative for core collection construct of S. baicalensis and the best sampling ratio was 10.3 % of the total germplasms. Finally, we found that the mini core collection by LDSS was composed of 11 accessions with high genetic diversity (Na = 1.82, Ne = 1.40, He = 0.25, I = 0.39). To verify the reliability of sampling strategy, the t test was used to evaluate the representativeness between core collections (constructed by LDSS), primary and reserve collection. Our results showed that no significant difference was found in the two important genetic parameters He and I (P > 0.05), thus suggesting that the LDSS will provide a rational sampling strategy for constructing a core collection of S. baicalensis germplasm resources in the future.




Similar content being viewed by others
References
Albrecht E, Zhang DP, Saftner RA, Stommel JR (2012) Genetic diversity and population structure of Capsicum baccatum genetic resources. Genet Resour Crop Evol 59:517–538
Bai CK, Wen MM, Yu F, Li GS (2010) Methods on construction of core germplasm collection of S. baicalensis by ISSR marker. J Chin Med Mater 33:1689–1694
Baranger A, Aubert G, Arnau G, Laine AL, Deniot G, Potier J, Weinachter C, Lejeune HI, Lallemand J, Burstin J (2004) Genetic diversity within Pisum sativum using protein and PCR-based markers. Theor Appl Genet 108:1309–1321
Brown AHD (1989) Core collection: a practical approach to genetic resources management. Genome 31:818–824
Brown AHD, Grace JP, Speer SS (1987) Designation of a core collection of perennial Glycine. Soybean Genet Newsl 14:59–70
Chabane JCK, Valkoun J (2004) Characterizations of genetic diversity in ICARDA core collection of cultivated barley (Hordeum vulgare L.). Czech J Genet Plant Breed 40:134–136
Chang WH, Chen CH, Lu FJ (2002) Different effects of baicalein, baicalin and wogonin on mitochondrial function, glutathione content and cell cycle progression in human hepatoma cell lines. Planta Med 68:128–132
Chiang Su New Medical College (ed) (1977) Dictionary of Chinese crude drugs. Shanghai Scientific Techonological Publishers, Shanghai, p 783
Cui L, Lu JX, Lin HB, Lin JQ, Lin JQ (2009) Advances in research on S. baicalensis resources and production status in China. Lishizhen Med Materia Medica Res 20:2279–2280
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure from small quantities of fresh leaf tissues. Phytochem Bull 19:11–15
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Esayas A, Endashaw B, Tomas B (2005) Inter-simple sequence repeat (ISSR) variation in forest coffee trees (Coffea arabica L.) populations from Ethiopia. Genetica 124:213–221
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Flora of China editorial committee (1977) Flora Reipublicae Popularis Sinicae (FRPS), vol 65. Science press, Beijing, pp 195–198
Frank OH, Rotem D (1993) Random sampling from databases: a survey. Stat Comput 5:25–42
Frankel OH (1984) Genetic perspectives of germplasm conservation. In: Arber WK, Llimensee K, Peacock WJ, Starlinger P (eds) Genetic manipulation: impact on man and society. Cambridge University Press, Cambridge, pp 161–170
Frankel OH, Brown AHD (1984) Plant genetic resources today: a critical appraisal. In: Hoden HW, Williams JT (eds) Crop genetic resources: conservation and evaluation. George Allen and Urwin, London, pp 249–257
Golkar P, Arzani A, Rezaei AM (2011) Genetic variation in safflower (Carthamus tinctorius L.) for seed quality-related traits and inter-simple sequence repeat (ISSR) markers. Int J Mol Sci 12:2664–2677
Hanelt P (2001) Scutellarioideae (2027–2028). In: Hanelt P, Institute of Plant Genetics and Crop Plant Research (eds) Mansfeld’s encyclopedia of agricultural and horticultural crops. Springer, Berlin
Himejia M, Ohtsukia T, Fukazawaa H, Tanakaa M, Yazakia S, Uia S, Nishiob K, Yamamotoc H, Tasakad K, Mimuraa A (2007) Difference of growth-inhibitory effect of Scutellaria baicalensis producing flavonoid wogonin among human cancer cells and normal diploid cell. Cancer Lett 245:269–274
Hintum TL (1995) Core collections of plant genetic resources. Wiley, Chichester, pp 23–34
Holbrook CC, Anderson WF (1995) Evaluation of a core collection to identify resistance to late leaf spot in peanut. Crop Sci 35:1700–1702
Hu J, Xu HM, Zhu J (2000a) A method of constructing core collection reserving special germplasm materials. J Biomath 16:348–352
Hu J, Zhu J, Xu HM (2000b) Methods of constructing core collections by stepwise clustering with three sampling strategies based on the genotypic values of crops. Theor Appl Genet 101:264–268
Jansen J, van Hintum T (2007) Genetic distance sampling: a novel sampling method for obtaining core collections using genetic distances with an application to cultivated lettuce. Theor Appl Genet 114:421–428
Lee DH, Kim C, Zhang L, Lee YJ (2008) Role of p53, PUMA, and Bax in wogonin-induced apoptosis in human cancer cells. Biochem Pharmacol 75:2020–2033
Nagaoka T, Ogihara Y (1997) Applicability of inter-simple sequence repeat polymorphism in wheat for use as DNA markers in comparison to RFLP and RAPD markers. Theor Appl Genet 93:133–139
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci 70:3321–3323
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rohlf FJ (1998) NTSYS-PC, Numerical taxonomy and multivariate analysis system, version 2.02a. Exeter Publications, Setauket
Rosenberg NA, Prichard JK, Weber JL, Cann HM, Kidd KK, Zhivotovsky LA, Feldman MW (2002) Genetic structure of human populations. Science 298:2381–2385
Sekiya K, Okuda H (1982) Selective inhibition of platelet lipoxygenase by baicalein. Biochem Biophys Res Commun 105:1090–1095
Shannon CE, Weaver W (1949) The mathematical theory of communication. University of Illinois Press, Urbana pp 117
Shao AJ, Li X, Huang LQ, Lin SF, Chen J (2006) RAPD analysis of S. baicalensis from different germplasms. China J Chin Materia Medica 31:452–455
Tang W, Eisenbrand G (1992) Chinese drugs of plant origin: chemistry, pharmacology, and use in traditional and modern medicine. Springer, Heidelberg, pp 919–929
The State Pharmacopoeia Commission of People’s Republic of China (2010) Pharmacopoeia of People’s Republic of China. China medical science press, Beijing, pp 282–283
Tu YY (2011) The discovery of artemisinin (qinghaosu) and gifts from Chinese medicine. Nat Med 17:1217–1220
Wang KJ, Li XH (2012) Genetic diversity and geographical peculiarity of Tibetan wild soybean (Glycine soja). Genet Resour Crop Evol 59:479–490
Wang JC, Hu J, Xu HM, Zhang S (2007) A strategy on constructing core collections by least distance stepwise sampling. Theor Appl Genet 115:1–8
Wen MM, Li GS, Zhang LJ, Zheng P, Bai CK (2012) Analysis and evaluation on genetic diversity of Scutellaria baicalensis G. by ISSR markers. Bull Bot Res 32:32–37
Yang Q, Luo LW, Luo GC, Wang WQ, Zang H (2008) AFLP analysis on genetic diversity of morphological variation types in Scutellaria baicalensis Georgi. J Guangdong Coll Pharm 24:103–105
Yeh FC, Yang RC, Boyle TBJ, Ye ZH, Mao JX (1997) POPGENE. The User-friendly Shareware for Population Genetic Analysis, Molecular Biology and Biotechnology Center, University of Alberta, Canada
Yonezawa K, Nomura T, Morishima H (1995) Sampling strategies for use in stratified germplasm collections. In: Hodgkin T, Brown AHD, Hintum TJL, van Morales EAV (eds) Core collections of plant genetic resources. Wiley, Chichester, pp 35–53
Yu J, Chen J, Xiao XY, Zhu XH, Yang SL, Cheng HZ (2005) Study on yield and quality of Scutellaria baicalensis from different habitats. China J Chin Materia Medica 30:491–494
Zewdie Y, Tong NK, Bosland P (2004) Establishing a core collection of Capsicum using a cluster analysis with enlightened selection of accessions. Genet Resour Crop Evol 51:147–151
Zhang GY, Wang XF, Liu SJ, Ma ZY (2004) Cluster analysis and sampling methods for core collection construction in glandless cotton. Cotton Sci 16:8–12
Zietkiewicz E, Rafalski A, Labuda D (1994) Genome finger printing by simple sequence repeat (ISSR) anchored polymerase chain reaction amplification. Genomics 20:176–183
Acknowledgments
This work was partially founded by Natural Science Foundation of China (NSFC no. 31100241) and Shaanxi key scientific and technological project (2011K16-0205). Thanks to Dr. Guang Wu (College of Life Science, Shaanxi Normal University) and Aaron Follansbee (College of Botany, University of Wisconsin-Madison) for valuable advices and language improvement.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bai, C., Wen, M., Zhang, L. et al. Genetic diversity and sampling strategy of Scutellaria baicalensis germplasm resources based on ISSR. Genet Resour Crop Evol 60, 1673–1685 (2013). https://doi.org/10.1007/s10722-012-9949-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-012-9949-9