Abstract
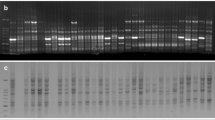
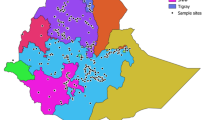
Naked barley, due to its favorable attributes such as high feed value, good human nutrition and easy processing, increasingly attracts people’s attention. Qinghai–Tibet Plateau is very rich in naked barley resources and has a long growing history. Genetic diversity of these cultivated naked barley is, however, poorly documented. This study analyzed the genetic diversity of monomeric prolamins (protein fraction corresponding to wheat gliadins) using the Acid -PAGE technique in eighty-six cultivated naked barley cultivars from Qinghai–Tibet Plateau. Extensive diversity was observed. A total of 43 different bands and 76 distinct patterns were found. Jaccard’s coefficient of similarity was calculated, and the accessions were divided into five main groups by cluster analysis using UPGMA. Differentiation among the populations from different collecting regions was investigated, based on the polymorphism of monomeric prolamins. The genetic diversity within the accessions from Tibet was slightly higher with an average diversity index of 0.29 than those from Sichuan. This study supports Tibetan Plateau is the diversity center of the cultivated naked barley and suggests that cultivated naked barley from Qinghai–Tibet Plateau is an important pool of variability that could be used in cereal breeding.



Similar content being viewed by others
References
Atanassov P, Borries C, Zaharieva M, Monneveux P (2001) Hordein polymorphism and variation of agromorphological traits in a collection of naked barley. Genet Resour Crop Evol 48:353–360
Atienza SG, Giménez MJ, Martín A, Martín LM (2000) Variability in monomeric prolamins in Hordeum chilense. Theor Appl Genet 101:970–976
Badr A, Müller K, Schäfer-Pregl R, El Rabey H, Effgen S, Ibrahim HH, Pozzi C, Rohde W, Salamini F (2000) On the origin and domestication history of barley (Hordeum vulgare). Mol Biol Evol 17:499–510
Berglung PT, Holm ET, Fastnaught CE (1993) Hulless barley: alternative uses. Barley Newslett 36:130–131
Bhatty RS (1986) The potential of hull-less barley, a review. Cereal Chem 63:97–103
Ciaffi M, Dominici L, Lafiandra D (1997) Gliadin polymorphism in wild and cultivated einkorn wheats. Theor Appl Genet 94:68–74
Dakir EH, Ruiz ML, García P, Marcelino PV (2002) Genetic variability evaluation in a Moroccan collection of barley, Hordeum vulgare L., by means of storage proteins and RAPDs. Genet Resour Crop Evol 49:619–631
Doll H, Brown AHD (1979) Hordein variation in wild (Hordeum spontaneum) and cultivated (H. vulgare) barley. Can J Genet Cytol 21:391–404
Draper SR (1987) ISTA Variety Committee report of the working group for biochemical tests for cultivar identification 1983–1986. Seed Sci Technol 15:431–434
Francis CY, Yang RC, Tim B (1999) POPGENE. Microsoft Window-based Freeware for Population Genetic Analysis. Version 1.31
Heisel SE, Peterson DM, Jones BL (1986) Identification of US barley cultivars by SDS-PAGE of hordeins. Cereal Chem 63:500–505
Jaradat AA (1991) Grain protein variability among populations of wild barley (Hordeum spontaneum C. Koch) from Jordan. Theor Appl Genet 83:164–168
Luan YF, He Y (2004) Tendency and counter measures on breeds improvement of Tibet highland barley. Barley Sci 12:1–4
Liu CT, Wesenberg DM, Hunt CW, Branen AL, Robertson LD, Burrup DE, Dempster KL, Haggerty RJ (1996) Hulless barley: a new look for barley in Idaho. Resources for Idaho. http://info.ag.uidaho.edu/
Ma DQ (2000) Genetic resources of Tibetan barley in China. Agriculture Press, Beijing, pp 216–254
Ma DQ (2002) Genetic resources of Tibetan barley in China. Agriculture Press, Beijing, p 37
Metakovsky EV, Branlard G (1998) Genetic diversity of French common wheat germplasm based on gliadin alleles. Theor Appl Genet 96:209–218
Moralejo M, Romagosa I, Salcedo G, Sanchez-Monge R, Molina-Cano JL (1994) On the origin of Spanish two-rowed barleys. Theor Appl Genet 87:829–836
Newman RK, Ore KC, Abbott J, Newman CW (1998) Fiber enrichment of baked products with a barley milling fraction. Cereal Foods World 43:23–25
Nevo E, Beiles A, Storch N (1983) Microgeographic edaphic differentiation in hordein polymorphism of wild barley. Theor Appl Genet 64:123–132
Nieto-Taladriz MT, Branlard G, Dardevet M (1994) Polymorphism of omega-gliadins in durum wheat as revealed by the two-step APAGE/SDS-PAGE technique. Theor Appl Genet 87:1001–1005
Nimazhaxi (1998) Hulless barley and food safety in the region of plateau. Tibetan Agric Technol 20:20–25
Oscarsson M, Andersson R, Salomonsson AC, Aman P (1996) Chemical composition of barley samples focusing on dietary fibre components. J Cereal Sci 24:161–170
Pomortsev AA (2001) Hordein polymorphism in Ethiopian barley. Genetika 37:1371–1382
Radovic D, Vapa L (1996) Hordein composition of Yugoslav barley cultivars. Cereal Res Com 24: 3:332–337
Rohlf FJ (1998) NTSYS-pc. Numerical taxonomy and multivariate analysis system, version 2.0, Exeter Software, New York
Roininen J, Nissila E, Puolimatka M, Pulli S (1992) Identification of barley cultivars using SDS-PAGE. Agric Sci Finl 1:73–82
Romanova YA, Gubareva NK, Konarev AV, Mitrofanova OP, Lyapunova OA, Anfilova NA, Strel’chenko PP (2001) Analysis of gliadin polymorphism in a Triticum spelta collection. Russian J Genet 37:1054–1060
Shewry PR, Pratt HM, Miflin BJ (1978) Varietal identification of single seeds of barley by analysis of hordein polypeptides. J Sci Food Agric 29:587–596
Shannon CE, Weaver W (1949) The mathematical theory of communication. University of Illinois Press, Urbana
Shewry PR, Finch RA, Parmar S, Franklin J, Miflin BJ (1983) Chromosomal location of Hor-3, a new locus governing storage proteins in barley. Heredity 50:179–189
Sneath PHA, Sokai RR (1973) Numerical taxonomy. The priciples and practices of numerical classification. W. H. Freeman, San Francisco
Sun LJ, Lu W, Zhang J, Zhang WX (1999) Investigation of barley germplasm in China. Genet Resour Crop Evol 46:361–369
Taketa S, Kikuchi S, Awayama T, Yamamoto S, Ichii M, Kawasaki S (2004) Monophyletic origin of naked barley inferred from molecular analyses of a marker closely linked to the naked caryopsis gene (nud). Theor Appl Genet 108:1236–1242
Vapa L, Radovic D (1998) Genetics and molecular biology of barley hordeins. Cereal Res Com 26:31–38
Vyhnánek T, Bednář J (2003) Detection of the varietal purity in sample of harvested wheat and triticale grains by prolamin marker. Plant Soil Environ 49:95–98
Williams MDHM, Peña RJ, Mujeeb-Kazi A (1993) Seed protein and isozyme variations in Triticum tauschii (Aegilops squarrosa). Theor Appl Genet 87:257–263
Yang RW, Wei XH, Zhou YH, Zhen YL (2004) Genetic polymorphism of gliadin in leymus. Acta Botanica Yunnanica 26:103–110
Yin YQ, Ma DQ, Ding Y (2003) Analysis of genetic diversity of hordein in wild close relatives of barley from Tibet. Theor Appl Genet 107:837–842
Zemede AF (1989) Variation in hordein polypeptide pattern within Ethiopian barley, Hordeum vulgare L. (Poaceae). Hereditas 110:185–191
Zhan XY, Yu ZL, Huang PZ (1991) Research of hordein polymorphism in barley from China. Acta Genet Sin 18:252–262
Zhang QF, Yang GP, Dai XK, Sun JZ (1994) A comparative analysis of genetic polymorphism in wild and cultivated barley from Tibet using isozyme and ribosomal DNA markers. Genome 37:631–638
Acknowledgements
This research was supported by grant No. KSCX2-SW-304 from the Chinese Academy of Sciences and No. 2006FY110700 from the Department of Science and Technology of China.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pan, Z.F., Deng, G.B., Zhai, X.G. et al. Genetic diversity of Acid-PAGE monomeric prolamins in cultivated hulless barley (Hordeum vulgare L.) from Qinghai–Tibet plateau in China. Genet Resour Crop Evol 54, 1691–1699 (2007). https://doi.org/10.1007/s10722-006-9177-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-006-9177-2