Abstract
This study investigates the potential role of Glycosyltransferases (GTs) in the glycosylation process and their association with malignant tumors. Specifically, the study focuses on PARP14, a member of GTs, and its potential as a target for tumors in the diagnosis and treatment of cervical cancer. To gather data, the study used somatic mutation data, gene expression data and clinical information from TCGA-CESE dataset as well as tissue samples from cervical cancer patients. Further verification was conducted through RT-qPCR and immunohistochemistry staining on cervical cancer tissues to confirm the expression of PARP14. The study utilized Kaplan-Meier for survival analysis of cervical cancer patient and found significant mutational abnormalities in GTs. The high frequency mutated gene was identified as PARP14. RT-qPCR revealed significantly higher mRNA expression of PARP14 compared to precancerous tissue. Using IHC combined with Kaplan-Meier,patients in the PARP14 high expression group had a better prognosis than the low expression group. The study identified PARP14 as a frequently mutated gene in cervical cancer and proposed its potential role in diagnosis and treatment.
Similar content being viewed by others
Introduction
Glycosyltransferases (GTs) are a large family of enzymes that play an important role in the process of glycosylation. [1]. GTs are present in the endoplasmic reticulum (ER) and Golgi apparatus. Currently, more than 200 genes encoding GTs have been identified and classified into different families such as fucosyltransferase, sialyltransferase and N-acetylglucosaminyltransferase [1, 2]. Previous studies have suggested that the abnormal expression of GT may be associated with various diseases, especially related with the proliferation, invasion, metastasis, epithelial-mesenchymal transition (EMT) and chemoresistance in malignant tumor [3,4,5,6,7]. Therefore, GTs have been suggested as crucial biomarkers for the timely detection and prediction of prognosis in various types of cancer. [4, 5, 7].
Cervical cancer is one of the malignant tumors of the reproductive system that seriously threatens women’s health. Its incidence and mortality rate are ranked in the 5th and 4th place in the female population, and the trend is still increasing [8, 9]. Recent studies have shown that the glycosylation varients are closely related to the proliferation, metastasis and drug resistance of cervical cancer [10, 11] and certain GTs have been associated with diagnosis and prognosis [12]. These fingdings suggest that GTs may play a significant role in cervical cancer and could potentially become new targets for diagnosis and treatment.
Poly (ADP-ribose) polymerase member 14 (PARP14), also known as ARTD8, BAL2, or CoaST6, belongs to the group of glycosyltransferases (GTs). In the past years, PARP14 was found to be associated with STAT6 activation, B-cell differentiation, and the JNK1/JNK2 signaling pathway. There has been growing interest in PARP14 as a potential target in allergic inflammation and tumors [13].PARP14’s role in inflammatory diseases including atopic dermatitis (AD), emphysema and asthma [14,15,16].PARP14 also plays an important role in tumors. PARP14 high expression in human primary pancreatic cancer (PCa) is associated with poor prognosis, and PARP14 promotes the proliferation and gemcitabine chemoresistance of pancreatic cancer cells through activation of NF-κ pathway [17]. In hepatocellular carcinoma (HCC) cells, PARP14 plays a role in promoting cell survival. One way it does this is by inhibiting the activity of JNK1, which is a pro-survival protein that functions as a serine/threonine kinase. This inhibition leads to a suppression of the nuclear function of downstream PKM2, which ultimately promotes the survival of tumor cells [18].
This paper analyzed 244 GTs that were previously identified [19] and examined the mutation and expression levels of GTs in both cervical cancer patients from the Cancer Genome Atlas (TCGA) database and clinical patients. The study identified several GTs that were linked to either positive or negative prognosis. In addition, the paper delves into the role of PARP14in the prognosis of cervical cancer.
Materials and methods
TCGA dataset: We obtained a dataset from The Cancer Genome Atlas (TCGA, https://portal.gdc.cancer.gov) consisting of 178 tumor samples with somatic mutation data, 3 normal samples and 306 tumor samples with gene expression data and clinical information. The dataset was selected from TCGA-CESE. This study adheres to the publication guidelines provided by TCGA (http://cancergenome.nih.gov/publications/ publicationguidelines).
Clinical tissue specimens: The use of human tumor specimen was approved by the local Ethics Committee and informed consents were obtained from all patients. The clinical cohort contained data from 10 cervical cancer patients from the Obstetrics and Gynecology Hospital of Fudan University. We collected the clinical characteristic of the 10 patients of cervical cancer, with a mean age of: 51.8 years (Table 1). Of the 10 patients, 8 had squamous cell carcinoma and 2 had adenocarcinoma. Additionally, there were 5 patients in stage I, 2 in stage II and 3 in stage III (Table 1). We performed whole genome sequencing (WGS) on the samples to obtain data related to their somatic mutations. Additionally, we established a cohort of 35 cervical cancer samples and a cohort of 4 cervical cancer tissues which were matched with para-cancer tissues using immunohistochemistry staining and RT-qPCR, respectively. All clinical specimens were untreated with preoperative chemotherapy or radiotherapy.
Quantitative Real-Time PCR Analysis: The mRNA of cervical cancer tissue was isolated and reverse transcripted to cDNAs by the MolPure® Cell/Tissue Total RNA Kit (19221ES50, Yeasen, Shanghai, China) and Hifair® II 1st Strand cDNA Synthesis Kit (gDNA digester plus) (11123ES10, Yeasen, Shanghai, China). Then the cDNAs was used for quantitative PCR analysis using Hieff® qPCR SYBR Green Master Mix(Low Rox Plus) (11202ES03, Yeasen, Shanghai, China).The RNA concentration and quality were determined by NanoDrop 2000 (Applied Biosystems, USA)and the RT-qPCR analysis was performed on QuantStudio5 Real-Time PCR (Applied Biosystems, USA). The PCR primers were as follows: GAPDH forward, 5’-GGAGCGAGATCCCTCCAAAAT-3’ and reverse, 5’-GGCTGTTGTCATACTTCTCATGG-3’; PRAP14 forward, 5’ -TGTTAGTGGAGAACATAAGTGGC-3’ and reverse, 5’ -TGAATGGTGCTTGGTACAATCAT-3’. GAPDH was used as the internal control. Data were analyzed using the 2-ΔΔCt method.
Immunohistochemistry (IHC) Staining: The samples from the tissue were embedded in paraffin. Tissue sections (3 μm) were prepared for IHC analyses of PARP14. In brief, the sections were de-paraffinized and, dehydrated, and were then subjected to antigen retrieval. Endogenous peroxidase was eliminated by the use of 3% H2O2.The slides were then incubated with anti-PARP14(1:100, Rabbit, Affinity, DF14173) overnight at 4℃. The samples were incubated with a second antibody (Mouse anti-rabbit IgG, Recordbio, RC0080RM) at room temperature, developed with DAB, counterstained with hematoxylin for 3 min, and photographed under a microscope. All images were acquired and processed in TIFF format, and the integrated option densities (IODs) of in each image were counted and measured using ImageJ (1.53t, Java 1.8.0_322) and IHC-Toolbox plugin [20].
Gene set enrichment analysis (GSEA):
Gene set enrichment analysis (GSEA) is a commenly used computational method for analyzing genome-wide expression matrices [21]. In this study, we utilized GSEA to assess related pathways and molecular mechanisms in cervical cancer patients by setting GTs gene mutation levels as population phenotypes. The nominal P-value and normalized enrichment score (NES) were used to sort the pathways enriched in each phenotype. Gene sets with a nominal P value of less than 0.05 were considered statistically significant.
Statistics Analysis: The R software (4.0.2, http://www.R-project.org, RRID:SCR_001905) was used to derive all statistical analyses. Somatic mutation data for 178cervical cancer samples from TCGA-CESE cohort and 10 samples from the clinical cohort were extracted by “Maftools” plugin. Overall survival (OS) was predicted using Kaplan-Meier survival plots. GraphPad Prism V8.0.1 (GraphPad Software, Inc., La Jolla, CA, USA, RRID:SCR_002865) software was used to determine statistical differences by using the Welch’s t test or one-way analysis of variance (ANOVA). p < 0.05 was considered to be statistically significant.
Results
Glycosyltransferase (GT) genes Associated with Mutation in TCGA cohorts
We analyzed the association between the mutation of 244 GTs and cervical cancer in TCGA cohorts. Somatic mutation profiles from 178 cervical cancer patients were downloaded and analyzed using “Maftools” plugin. The mutations were classified into different categories with missense mutations accounting for the largest proportion based on protein structure (Fig. 1A). Here we presented the top 30 mutated genes in cervical cancer, ranked by percentage, the top 10 including PARP14, ALG13, UGT2B15, ART4, C1GALNT1C1, GALNT9, MGAT4C, OGT, TNKS2, UGGT2 (Fig. 1B). In addition, single nucleotide polymorphisms (SNPs) occurred more frequently than insertions or deletions (Fig. 1C), with G > A being the most common single nucleotide variant (SNV) in cervical cancer (Fig. 1D) according to gene structure classification.
Summary of the GT mutation information from TCGA (A) Mutation types were classified by protein structure categories, with missense mutations accounting for the largest proportion. (B) The top 30 mutated GT genes in cervical cancer. (C, D) Mutation types were classified by gene structure categories, SNP occurring more frequency than insertion or deletion, and G > A being the most common mutation type in SNV
Enrichment analysis of GTs based on TCGA cohort
To investigate the potential mechanism of GT effect on cervical cancer, we utilized Gene Set Enrichment Analysis (GSEA) to examine the pathways and biological processes associated with GTs. We divided the 178 cervical cancer samples from the TCGA database into mutation and non-mutation groups based on the mutation status of 244 GTs. Out of these patients, 104 were classified as the GTs mutant-type group while 90 were classified as the GTs wild-type group. As shown, we obtained significant enrichment in cell cycle (NES = 1.87, P = 8e-7), HPV infection (NES = 2.03, P = 1e-7) and T cell (NES = 1.76, P = 0.03) pathways (Fig. 2A,B,D). In HPV infection analysis, we further found that HPV31 infection was higher in mutant-type than wild-type (Fig. 2C). In our investigation of T cell-related pathways, we observed an increase in T.helper cell and M1 macrophage in mutant-type(Fig. 2E), and further analysis revealed that the increase in SNV neoantigen may be the reason for the high T cell content(Fig. 2F). In addition, the ratio of silent and non-silent mutations was significantly elevated in the mutant-type (Fig. 2G). Finally, we identified the gene sets with the most significant differences in the enrichment of GSEA genes, including histone modifications, doxorubicin resistance, early T lymphocyte, metastasis, cell cycle, HPV positive tumors and G2M cell cycle(Fig. 2H).
Enrichment analysis of GTs in TCGA cohort (A) GSEA Enrichment plot of Cell cycle in mutation group. (B) GSEA Enrichment plot of HPV infection in mutation group. (C) Comparison of HPV31 burden between GTs mutant-type and wild-type cervical cancer patients in TCGA cohorts. (D) GSEA Enrichment plot of T cell in mutation group. (E) Comparison of T-helper cells follicular and M1 macrophages between GTs mutant-type and wild-type cervical cancer patients in TCGA cohorts. (F) SNV-induced neoantigen was significantly higher in the mutant group, which may be the reason for the increase in T cells. (G) Both silent and non-silent mutations were significantly increased in mutant-type compared to wild-type. (H) Name of the gene set with the most significant enrichment differences in the GEEA gene set
Glycosyltransferase (GT) genes Associated with mutation in clinical cohorts
We conducted whole genome sequencing of 10 cervical cancer tissues, and obtained somatic mutation data for analysis. These mutations were classified into different categories, with the largest proportion of intron variants based on protein structure (Fig. 3A). We identified the top 30 mutated genes, sorted by percentage, and the top 10 included MGAT4C, ST8SIA1, GLT1D1, B4GALNT3, GALNT13, GALNTL6, GXYLT1, ST6GALNAC3, GTDC1, XYLT (Fig. 3B). In addition, SNP occurred more frequently than insertions or deletions (Fig. 3C), with G > A being the most common SNV in cervical cancer (Fig. 3D). To compare with the mutations in the TCGA cohort, we screened for GT genes with missense mutations. A total of 12 GT genes in the clinical cohort had missense mutations, including PARP14 (Table 2).
Somatic mutation patterns of GTs identified by whole-genome sequencing (WGS) in 10 patients of cervical cancer. (A) Mutation types were classified by protein structure categories, with intron variant accounting for the largest proportion. (B) The top 30 mutated GT genes in cervical cancer. (C, D) Mutation types were classified by gene structure categories, SNP occurring more frequency than insertion or deletion, and G > A being the most common mutation type in SNV
Somatic mutation of GTs from patients with cervical cancer in TCGA and clinical cohort
Furthermore, we summarized the mutation categories of the most frequently mutated GTs in TCGA datasets and tissues (Fig. 4A, B). Notably, PARP14 had a high frequency of missense mutations in both cohorts, despite not having a higher mutation frequency in the tissue cohort. This arouse our interest in investigating the role of PARP14 in cervical cancer, leading to the following related studies.
PARP14 expression in cervical cancer tissues and correlation analysis
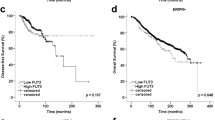
According to TCGA datasets, there was an up-regulation of PARP14 in cervical cancer when compared to normal tissues (Fig. 5A). And it was also validated that the mRNA expression of PARP14 in tumor tissues was significantly increased when compared to the matched para-cancer tissues (Fig. 5B). To investigate the association between glycosyltransferase mutations and cervical cancer prognosis, we further conducted a survival analysis of 244 GTs in the TCGA cohort. Our analysis revealed that the majority of the 244 GTs had either a protective or hazardous effect on the prognosis of cervical cancer (Online Resource 1, Online Resource 2). We listed the 25 GT genes in which mutations had the greatest protective or hazardous effect on the prognosis of cervical cancer, with mutations in the PARP14 gene exhibiting a protective effect against cervical cancer (Fig. 5C). At the same time, we performed immunohistochemical staining of cancer tissues from a clinical cohort of 35 cervical cancer patients to analyze the qualitative and quantitative expression levels of PARP14. Clinical characteristics and prognostic data were collected. The median follow-up time was 23 months. We divided them into PARP14 high expression group (N = 17) and PARP14 low expression group (N = 18) (Fig. 5D) based on quantitative immunohistochemical analysis. The study utilized the Kaplan-Meier method to assess the prognosis of two groups, and the results were statistically different (p = 0.036), with patients in the high PARP14 expression group having a better prognosis compared to those in the low PARP14 expression group (Fig. 5E). In addition, we also conducted a differential analysis based on various factors such as pathological type, clinicopathological stage, menopause and body mass index (BMI). However, there was no significant difference in PARP14 expression among any of these factors(Table 3).
PARP14 expression in cervical cancer tissues and correlation analysis. (A) Volcano plot of mRNA expression of GTs in tumor and normal cervical tissues from TCGA dataset. In the volcano map, red: genes up-regulated in tumor group; blue: genes down-regulated in tumor groups; gray: no differentially expressed genes in the tumor group; green: PARP14. (B) PARP14 expression in tumor and normal cervical tissues. (C) GT genes which had the greatest protective or hazardous effect on the prognosis of cervical cancer from TCGA cohort. (D) Immunohistochemical staining of PARP14 protein was performed on 35 cervical cancer tissues, which were divided into low expression group (N = 18) and high expression group (N = 17) according to expression level. (E) Kaplan-Meier survival curves for overall survival of cervical cancer patients from two groups
Discussion
Glycosyltransferases are a large group of enzymes which play a crucial role in glycosylation. Over 200 GTs have been identified and their abnormal expression has been closely related to tumors. In colorectal cancer, the expression of GTs has been found to contribute significantly to tumor cell proliferation, survival, stem-like cell properties induction, epithelial-mesenchymal transition (EMT), metastasis and resistance to chemotherapy and radiotherapy [4]. Jahanshah Ashkani et al. analyzed that the expression of 210 GT genes from 1893 cancer patient samples in The Cancer Genome Atlas (TCGA) microarray data. Their findings showed that these genes were effective in classifying six types of cancers, including breast, ovarian, glioblastoma, kidney, colon and lung. Furthermore, these GT genes were able to classify subgroups of breast cancer [22]. Yousra Mohamed Abd-El-Halim et al. proposed a gene signature funded based on 19 GT genes that can effectively stratify the prognosis of patients with pancreatic ductal adenocarcinoma (PDAC) [23]. These studies indicates that the expression of GTs has an impact on the fate of tumors.
Our study summarized the mutation pattern of GTs in cervical cancer tissues based on TCGA database and whole genome sequencing (WGS). According to the analysis of TCGA database, it revealed that missense mutations were the most common protein structure mutation types in GTs, with the top 30 mutation frequencies all having missense mutations. Meanwhile, the number of SNPs was also more than other gene structure mutation types. Missense variants were the most common consequence of SNPs in humans [24]. We concerned that the effect of missense mutations on glycosyltransferase activity was arguably more difficult to judge than in the case of mutations with other consequences (such as total gene deletions or point nonsense mutation) [25]. Among the top 30 GTs in terms of mutation frequency, a de-novo missense mutation in C1GALT1C1: c.266 C > T, p. (T89I) caused a novel X-linked form of atypical hemolytic-uremic syndrome (aHUS) [26]. Another study by Marilena De Mariano et al. revealed that a GALNT14 mutation (c.802 C > T) had been associated to the development of neuroblastoma (NB) and GALNT14 might act as a novel gene potentially involved in NB predisposition [27].
Recent studies confirmed the correlation between GTs and cervical cancer. UDP-glucose ceramide glycosyltransferase (UGCG) contributed to proliferation and glycolysis of cervical cancer cells by regulating the PI3K/AKT pathway [10]. Additionally, Lixia Zhou et al. demonstrated an increase in the mRNA expression and protein level of GALNT2 in cervical high-grade intraepithelial neoplasia and tumor tissues compared to normal cervix tissues. GALNT2 was associated with worse overall survival and could be utilized as a prognostic biomarker for cervical cancer [12]. The findings of the current study confirmed that there was a correlation between the expression of GTs and the prognosis of cervical cancer. Out of the 244 GTs analyzed, almost all of them were found to have either a protective or detrimental effect on the prognosis of cervical cancer.
PARP14, a member of the glycosyltransferase family, has been shown to be closely associated with the development of a number of tumors, particularly in pancreatic cancer, hepatocellular carcinoma, and hematologic malignancy [17, 18, 28,29,30]. In diffuse large B-cell lymphoma (DLBCL), PARP14 regulated the IL-4-STAT6 signaling pathway, enhanced the expression of several downstream genes, and enabled DLBCL tumor cell survival [31]. In this study, we found that PARP14 showed a trend of high expression in cervical cancer tissues compared to para cancerous tissues, and the high expression of PARP14 was closely related to the prognosis of patients, suggesting that patients with high PARP14 expression have a better prognosis than those with low expression. Although our case sample was limited and our study was restricted to a descriptive study, the result still led us to focus on the role of PARP14 in cervical cancer. GSEA enrichment analysis identified cell cycle, HPV infection, and T cell-associated immunity as the probable mechanistic pathways. It had been demonstrated that cyclin D1 was over-expressed in many tumors. PARP14 deletion led to a decrease of cyclin D1, resulting in G1 cell-cycle arrest and reduced proliferation [32]. This confirmed the involvement of PARP14 in regulating cell cycle progression. However, the role of this mechanism in cervical cancer was unclear. Among many allergic diseases, PARP14 interacted with STAT6 to enhance IL-4-induced gene expression in T cells, thereby promoting Th2 differentiation [33, 34]. Research on the mechanism of PARP14 and HPV infection has not known yet. Regrettably, our study has not yet established a correlation between mutation and expression of PARP14 in cervical cancer. This would hold significant importance for future studies. The potential mechanism of PARP14 in cervical cancer will be explored in our further studies.
Conclusions
In this study, we analyzed the high-frequency mutated GT gene PARP14 from both TCGA database and clinical case samples. We found that PARP14 was significantly overexpressed at both the molecular and protein levels in cervical cancer tissues, and that higher expression of PARP14 caused better prognosis. Our findings suggested a potential role of PARP14 in the diagnosis and treatment of cervical cancer.
Data Availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Varki, A., Cummings, R.D., Esko, J.D., et al. (eds.): Essentials of Glycobiology. 4th ed. (2022)
Taniguchi, N., Kizuka, Y.: Glycans and cancer: Role of N-glycans in cancer biomarker, progression and metastasis, and therapeutics. Adv. Cancer Res. 126, 11–51 (2015). https://doi.org/10.1016/bs.acr.2014.11.001
Ohtsubo, K., Marth, J.D.: Glycosylation in cellular mechanisms of health and disease. Cell. Sep. 8(5), 855–867 (2006). https://doi.org/10.1016/j.cell.2006.08.019
Fernandez-Ponce, C., Geribaldi-Doldan, N., Sanchez-Gomar, I., et al.: The role of Glycosyltransferases in Colorectal Cancer. Int. J. Mol. Sci. May. 30(11) (2021). https://doi.org/10.3390/ijms22115822
Fernandez, L.P., Sanchez-Martinez, R., Vargas, T., et al.: The role of glycosyltransferase enzyme GCNT3 in colon and ovarian cancer prognosis and chemoresistance. Sci. Rep. May. 31(1), 8485 (2018). https://doi.org/10.1038/s41598-018-26468-4
Li, N., Xu, H., Fan, K., et al.: Altered beta1,6-GlcNAc branched N-glycans impair TGF-beta-mediated epithelial-to-mesenchymal transition through smad signalling pathway in human lung cancer. J. Cell. Mol. Med. Oct. 18(10), 1975–1991 (2014). https://doi.org/10.1111/jcmm.12331
Shan, M., Yang, D., Dou, H., Zhang, L.: Fucosylation in cancer biology and its clinical applications. Prog Mol. Biol. Transl Sci. 162, 93–119 (2019). https://doi.org/10.1016/bs.pmbts.2019.01.002
Sung, H., Ferlay, J., Siegel, R.L., et al.: Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and Mortality Worldwide for 36 cancers in 185 countries. CA Cancer J Clin. May. 71(3), 209–249 (2021). https://doi.org/10.3322/caac.21660
Bray, F., Ferlay, J., Soerjomataram, I., Siegel, R.L., Torre, L.A., Jemal, A.: Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. Nov. 68(6), 394–424 (2018). https://doi.org/10.3322/caac.21492
Zhang, F., Zhang, H.: UDP-Glucose Ceramide Glycosyltransferase contributes to the proliferation and glycolysis of Cervical Cancer cells by regulating the PI3K/AKT pathway. Ann. Clin. Lab. Sci. Sep. 51(5), 663–669 (2021)
You, X., Wang, Y., Meng, J., et al.: Exosomal miR663b exposed to TGFbeta1 promotes cervical cancer metastasis and epithelialmesenchymal transition by targeting MGAT3. Oncol. Rep. Apr. 45(4) (2021). https://doi.org/10.3892/or.2021.7963
Zhou, L., Wu, H., Bai, X., Min, S., Zhang, J., Li, C.: O-Glycosylating enzyme GALNT2 predicts worse prognosis in Cervical Cancer. Pathol. Oncol. Res. 28, 1610554 (2022). https://doi.org/10.3389/pore.2022.1610554
Tauber, A.L., Levonis, S.M., Schweiker, S.S.: Recent developments in PARP14 research. Future Med. Chem. Sep. 12(18), 1657–1667 (2020). https://doi.org/10.4155/fmc-2020-0166
Krishnamurthy, P., Da-Silva-Arnold, S., Turner, M.J., Travers, J.B., Kaplan, M.H.: Poly-ADP ribose polymerase-14 limits severity of allergic skin disease. Immunol. Nov. 152(3), 451–461 (2017). https://doi.org/10.1111/imm.12782
Zaffini, R., Gotte, G., Menegazzi, M.: Asthma and poly(ADP-ribose) polymerase inhibition: A new therapeutic approach. Drug Des. Devel Ther. 12, 281–293 (2018). https://doi.org/10.2147/DDDT.S150846
Hu, W.P., Zeng, Y.Y., Zuo, Y.H., Zhang, J.: Identification of novel candidate genes involved in the progression of emphysema by bioinformatic methods. Int. J. Chron. Obstruct Pulmon Dis. 13, 3733–3747 (2018). https://doi.org/10.2147/COPD.S183100
Yao, N., Chen, Q., Shi, W., Tang, L., Fu, Y.: PARP14 promotes the proliferation and gemcitabine chemoresistance of pancreatic cancer cells through activation of NF-kappaB pathway. Mol. Carcinog. Jul. 58(7), 1291–1302 (2019). https://doi.org/10.1002/mc.23011
Iansante, V., Choy, P.M., Fung, S.W., et al.: PARP14 promotes the Warburg effect in hepatocellular carcinoma by inhibiting JNK1-dependent PKM2 phosphorylation and activation. Nat. Commun. Aug. 10, 6:7882 (2015). https://doi.org/10.1038/ncomms8882
Joud, M., Moller, M., Olsson, M.L.: Identification of human glycosyltransferase genes expressed in erythroid cells predicts potential carbohydrate blood group loci. Sci. Rep. Apr. 16(1), 6040 (2018). https://doi.org/10.1038/s41598-018-24445-5
Shu, J., Dolman, G.E., Duan, J., Qiu, G., Ilyas, M.: Statistical colour models: An automated digital image analysis method for quantification of histological biomarkers. Biomed. Eng. Online Apr. 27, 15:46 (2016). https://doi.org/10.1186/s12938-016-0161-6
Subramanian, A., Tamayo, P., Mootha, V.K., et al.: Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. U S A Oct. 25(43), 15545–15550 (2005). https://doi.org/10.1073/pnas.0506580102
Ashkani, J., Naidoo, K.J.: Glycosyltransferase gene expression profiles classify Cancer types and propose prognostic subtypes. Sci. Rep. May. 20, 6:26451 (2016). https://doi.org/10.1038/srep26451
Mohamed Abd-El-Halim, Y., El Kaoutari, A., Silvy, F., et al.: A glycosyltransferase gene signature to detect pancreatic ductal adenocarcinoma patients with poor prognosis. EBioMedicine Sep. 71, 103541 (2021). https://doi.org/10.1016/j.ebiom.2021.103541
Lek, M., Karczewski, K.J., Minikel, E.V., et al.: Analysis of protein-coding genetic variation in 60,706 humans. Nat. Aug. 18(7616), 285–291 (2016). https://doi.org/10.1038/nature19057
Cao, X., Durairaj, P., Yang, F., Bureik, M.: A comprehensive overview of common polymorphic variants that cause missense mutations in human CYPs and UGTs. Biomed. Pharmacother Mar. 111, 983–992 (2019). https://doi.org/10.1016/j.biopha.2019.01.024
Hadar, N., Schreiber, R., Eskin-Schwartz, M., et al.: X-linked C1GALT1C1 mutation causes atypical hemolytic uremic syndrome. Eur. J. Hum. Genet. Jan. 4 (2023). https://doi.org/10.1038/s41431-022-01278-5
De Mariano, M., Gallesio, R., Chierici, M., et al.: Identification of GALNT14 as a novel neuroblastoma predisposition gene. Oncotarget Sep. 22(28), 26335–26346 (2015). https://doi.org/10.18632/oncotarget.4501
Camicia, R., Bachmann, S.B., Winkler, H.C., et al.: BAL1/ARTD9 represses the anti-proliferative and pro-apoptotic IFNgamma-STAT1-IRF1-p53 axis in diffuse large B-cell lymphoma. J. Cell. Sci. May. 1(Pt 9), 1969–1980 (2013). https://doi.org/10.1242/jcs.118174
Goenka, S., Cho, S.H., Boothby, M.: Collaborator of Stat6 (CoaSt6)-associated poly(ADP-ribose) polymerase activity modulates Stat6-dependent gene transcription. J. Biol. Chem. Jun. 29(26), 18732–18739 (2007). https://doi.org/10.1074/jbc.M611283200
Barbarulo, A., Iansante, V., Chaidos, A., et al.: Poly(ADP-ribose) polymerase family member 14 (PARP14) is a novel effector of the JNK2-dependent pro-survival signal in multiple myeloma. Oncogene Sep. 5. 32(36), 4231–4242 (2013). https://doi.org/10.1038/onc.2012.448
Cho, S.H., Goenka, S., Henttinen, T., et al.: PARP-14, a member of the B aggressive lymphoma family, transduces survival signals in primary B cells. Blood Mar. 12(11), 2416–2425 (2009). https://doi.org/10.1182/blood-2008-03-144121
O’Connor, M.J., Thakar, T., Nicolae, C.M., Moldovan, G.L.: PARP14 regulates cyclin D1 expression to promote cell-cycle progression. Oncogene. Jul. 40(30), 4872–4883 (2021). https://doi.org/10.1038/s41388-021-01881-8
Mehrotra, P., Hollenbeck, A., Riley, J.P., et al.: Poly (ADP-ribose) polymerase 14 and its enzyme activity regulates T(H)2 differentiation and allergic airway disease. J Allergy Clin Immunol. ;131(2):521 – 31 e1-12. (2013). https://doi.org/10.1016/j.jaci.2012.06.015
Mehrotra, P., Riley, J.P., Patel, R., Li, F., Voss, L., Goenka, S.: PARP-14 functions as a transcriptional switch for Stat6-dependent gene activation. J. Biol. Chem. Jan. 21(3), 1767–1776 (2011). https://doi.org/10.1074/jbc.M110.157768
Acknowledgements
In addition to the listed authors, we acknowledge the following individuals and teams for their assistance with this study: NHC Key Laboratory of Glycoconjugates Research, Department of Biochemistry and Molecular Biology, School of Basic Medical Sciences, Fudan University; Leading Talents Project of Huangpu District of Shanghai District Committee and Government.
Funding
This research was funded by Shanghai Shenkang Hospital Development Center’s Shenkang Promotion of Clinical Skills and Clinical Innovation in Municipal Hospitals Three Year Action Plan (2020–2023) Major Clinical Research Project (Grant No. SHDC2020CR1048B), the General Program of National Natural Science Foundation of China (Grant No.82271654), General program of Natural Science Foundation of Shanghai (Grant No. 19ZR1407000), Pilot construction project of high level universities in Shanghai (Grant No.DGF501017-06), “ZaiDing-Le” Foundation from Beijing Kanghua Foundation for the development of Traditional Chinese and Western Medicine (Grant No. KH-2020-LJJ-008).
Author information
Authors and Affiliations
Contributions
Conceptualization, X.W. and Y.R.; methodology, H.W., S.L. and X.T.; software, H.W. and W.Y.; validation, H.F., L.Q. and S.Z.; formal analysis, Y.Z.; investigation, Y.Y.; resources, M.L.; data curation, S.L.; writing—original draft preparation, H.W.; writing—review and editing, X.W., Y.R. and H.J.; visualization, Y.Z.; supervision, H.J.; project administration, X.W. and H.J.; funding facquisition, H.J. and X.W. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Ethics approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. The study was approved by Ethics Committee of Obstetrics & Gynecology Hospital of Fudan University (protocol code 2021 − 210, Nov.29, 2021 and protocol code 2022-51, Mar. 8, 2022).
Additional information
Hui Wang and Shen Luo contributed equally to this work.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, H., Luo, S., Wu, X. et al. Exploration of glycosyltransferases mutation status in cervical cancer reveals PARP14 as a potential prognostic marker. Glycoconj J 40, 513–522 (2023). https://doi.org/10.1007/s10719-023-10134-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10719-023-10134-7