Abstract
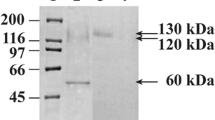
A gene for processing α-glucosidase I from a filamentous fungus, Aspergillus brasiliensis (formerly called Aspergillus niger) ATCC 9642 was cloned and fused to a glutathione S-transferase tag. The active construct with the highest production level was a truncation mutant deleting the first 16 residues of the hydrophobic N-terminal domain. This fusion enzyme hydrolyzed pyridylaminated (PA-) oligosaccharides Glc3Man9GlcNAc2-PA and Glc3Man4-PA and the products were identified as Glc2Man9GlcNAc2-PA and Glc2Man4-PA, respectively. Saturation curves were obtained for both Glc3Man9GlcNAc2-PA and Glc3Man4-PA, and the K m values for both substrates were estimated in the micromolar range. When 1 μM Glc3Man4-PA was used as a substrate, the inhibitors kojibiose and 1-deoxynojirimycin had similar effects on the enzyme; at 20 μM concentration, both inhibitors reduced activity by 50%.






Similar content being viewed by others
Abbreviations
- GST:
-
glutathione S-transferase
- PA:
-
pyridylaminated
- AbPGI:
-
Aspergillus brasiliensis processing α-glucosidase I
- IC50 :
-
concentration for 50% inhibition
References
Dhanawansa, R., Faridmoayer, A., van der Merwe, G., Li, Y.X., Scaman, C.H.: Overexpression, purification, and partial characterization of Saccharomyces cerevisiae processing alpha glucosidase I. Glycobiology 12, 229–234 (2002)
Herscovics, A.: Importance of glycosidases in mammalian glycoprotein biosynthesis. Biochim. Biophys. Acta 1473, 96–107 (1999)
Moremen, K.W., Trimble, R.B., Herscovics, A.: Glycosidases of the asparagine-linked oligosaccharide processing pathway. Glycobiology 4, 113–125 (1994)
Cantarel, B.L., Coutinho, P.M., Rancurel, C., Bernard, T., Lombard, V., Henrissat, B.: The Carbohydrate-Active EnZymes database (CAZy): an expert resource for Glycogenomics. Nucleic Acids Res. 37, Database issue, D233-D238 (2009)
Spiro, R.G.: Role of N-linked polymannose oligosaccharides in targeting glycoproteins for endoplasmic reticulum-associated degradation. Cell Mol. Life Sci. 61, 1025–1041 (2004)
Wang, T., Hebert, D.N.: EDEM an ER quality control receptor. Nat. Struct. Biol. 10, 319–321 (2003)
Herscovics, A.: Processing glycosidases of Saccharomyces cerevisiae. Biochim. Biophys. Acta 1426, 275–285 (1999)
Barker, M.K., Wilkinson, B.L., Faridmoayer, A., Scaman, C.H., Fairbanks, A.J., Rose, D.R.: Production and crystallization of processing α-glucosidase I: Pichia pastoris expression and a two-step purification toward structural determination. Protein Expr. Purif. 79, 96–101 (2011)
Frade-Pérez, M.D., Hernández-Cervantes, A., Flores-Carreón, A., Mora-Montes, H.M.: Biochemical characterization of Candida albicans α-glucosidase I heterologously expressed in Escherichia coli. Antonie Van Leeuwenhoek 98, 291–298 (2010)
Varga, J., Kocsubé, S., Tóth, B., Frisvad, J.C., Perrone, G., Susca, A., Meijer, M., Samson, R.A.: Aspergillus brasiliensis sp. nov., a biseriate black Aspergillus species with world-wide distribution. Int. J. Syst. Evol. Microbiol 57, 1925–1932 (2007)
Mizuno, M., Koide, A., Yamamura, A., Akeboshi, H., Yoshida, H., Kamitori, S., Sakano, Y., Nishikawa, A., Tonozuka, T.: Crystal structure of Aspergillus niger isopullulanase, a member of glycoside hydrolase family 49. J. Mol. Biol. 376, 210–220 (2008)
Aoki, H., Yopi, Sakano, Y.: Molecular cloning and heterologous expression of the isopullulanase gene from Aspergillus niger A.T.C.C. 9642. Biochem. J. 323, 757–764 (1997)
Zhang, L., Zhou, H., Ouyang, H., Li, Y., Jin, C.: Afcwh41 is required for cell wall synthesis, conidiation, and polarity in Aspergillus fumigatus. FEMS Microbiol. Lett. 289, 155–165 (2008)
Galagan, J.E., Calvo, S.E., Cuomo, C., et al.: Sequencing of Aspergillus nidulans and comparative analysis with A. fumigatus and A. oryzae. Nature 438, 1105–1115 (2005)
Wood, V., Gwilliam, R., Rajandream, M.A., et al.: The genome sequence of Schizosaccharomyces pombe. Nature 415, 871–880 (2002)
Jiang, B., Sheraton, J., Ram, A.F., Dijkgraaf, G.J., Klis, F.M., Bussey, H.: CWH41 encodes a novel endoplasmic reticulum membrane N-glycoprotein involved in β1,6-glucan assembly. J. Bacteriol. 178, 1162–1171 (1996)
Matsuo, I., Kashiwagi, T., Totani, K., Ito, Y.: First chemical synthesis of triglucosylated tetradecasaccharide (Glc3Man9GlcNAc2), a common precursor of asparagine-linked oligosaccharides. Tetrahedron Lett. 46, 4197–4200 (2005)
Fujimoto, I., Menon, K.K., Otake, Y., Tanaka, F., Wada, H., Takahashi, H., Tsuji, S., Natsuka, S., Nakakita, S., Hase, S., Ikenaka, K.: Systematic analysis of N-linked sugar chains from whole tissue employing partial automation. Anal. Biochem. 267, 336–343 (1999)
Yanagida, K., Natsuka, S., Hase, S.: A pyridylamination method aimed at automatic oligosaccharide analysis of N-linked sugar chains. Anal. Biochem. 274, 229–234 (1999)
Matsuda, K., Kurakata, Y., Miyazaki, T., Matsuo, I., Ito, Y., Nishikawa, A., Tonozuka, T.: Heterologous expression, purification, and characterization of an α-mannosidase belonging to glycoside hydrolase family 99 of Shewanella amazonensis. Biosci. Biotechnol. Biochem. 75, 797–799 (2011)
Mount, S.M.: A catalogue of splice junction sequences. Nucleic Acids Res. 10, 459–472 (1982)
Pel, H.J., de Winde, J.H., Archer, D.B., et al.: Genome sequencing and analysis of the versatile cell factory Aspergillus niger CBS 513.88. Nat. Biotechnol 25, 221–231 (2007)
Faridmoayer, A., Scaman, C.H.: Truncations and functional carboxylic acid residues of yeast processing α-glucosidase I. Glycoconj. J. 24, 429–437 (2007)
Basehoar, A.D., Zanton, S.J., Pugh, B.F.: Identification and distinct regulation of yeast TATA box-containing genes. Cell 116, 699–709 (2004)
Khan, F.A., Varma, G.M., Vijay, I.K.: Genomic organization and promoter activity of glucosidase I gene. Glycobiology 9, 797–806 (1999)
Makrides, S.C.: Strategies for achieving high-level expression of genes in Escherichia coli. Microbiol. Rev. 60, 512–538 (1996)
Schweden, J., Borgmann, C., Legler, G., Bause, E.: Characterization of calf liver glucosidase I and its inhibition by basic sugar analogs. Arch. Biochem. Biophys. 248, 335–340 (1986)
Bause, E., Erkens, R., Schweden, J., Jaenicke, L.: Purification and characterization of trimming glucosidase I from Saccharomyces cerevisiae. FEBS Lett. 206, 208–212 (1986)
Bause, E., Schweden, J., Gross, A., Orthen, B.: Purification and characterization of trimming glucosidase I from pig liver. Eur. J. Biochem. 183, 661–669 (1989)
Neverova, I., Scaman, C.H., Srivastava, O.P., Szweda, R., Vijay, I.K., Palcic, M.M.: A spectrophotometric assay for glucosidase I. Anal. Biochem. 222, 190–195 (1994)
Kalz-Fuller, B., Bieberich, E., Bause, E.: Cloning and expression of glucosidase I from human hippocampus. Eur. J. Biochem. 231, 344–351 (1995)
Zheng, Y.C., Elbein, A.D.: Purification to homogeneity and properties of plant glucosidase I. Arch. Biochem. Biophys. 355, 26–34 (1998)
Ugalde, R.A., Staneloni, R.J., Leloir, L.F.: Microsomal glucosidases of rat liver. Partial purification and inhibition by disaccharides. Eur. J. Biochem. 113, 97–103 (1980)
Kurakata, Y., Uechi, A., Yoshida, H., Kamitori, S., Sakano, Y., Nishikawa, A., Tonozuka, T.: Structural insights into the substrate specificity and function of Escherichia coli K12 YgjK, a glucosidase belonging to the glycoside hydrolase family 63. J. Mol. Biol. 381, 116–128 (2008)
Gibson, R.P., Gloster, T.M., Roberts, S., Warren, R.A.J., Storch de Gracia, I., García, Á., Chiara, J.L., Davies, G.J.: Molecular basis for trehalase inhibition revealed by the structure of trehalase in complex with potent inhibitors. Angew. Chem. Int. Ed 46, 4115–4119 (2007)
Aleshin, A., Golubev, A., Firsov, L.M., Honzatko, R.B.: Crystal structure of glucoamylase from Aspergillus awamori var. X100 to 2.2-Å resolution. J. Biol. Chem 267, 19291–19298 (1992)
Mizuno, M., Tonozuka, T., Suzuki, S., Uotsu-Tomita, R., Kamitori, S., Nishikawa, A., Sakano, Y.: Structural insights into substrate specificity and function of glucodextranase. J. Biol. Chem. 279, 10575–10583 (2004)
Jorge, C.D., Sampaio, M.M., Hreggvidsson, G.Ó., Kristjánson, J.K., Santos, H.: A highly thermostable trehalase from the thermophilic bacterium Rhodothermus marinus. Extremophiles 11, 115–122 (2007)
Lee, J.H., Saito, S., Mori, H., Nishimoto, M., Okuyama, M., Kim, D., Wongchawalit, J., Kimura, A., Chiba, S.: Molecular cloning of cDNA for trehalase from the European honeybee, Apis mellifera L., and its heterologous expression in Pichia pastoris. Biosci. Biotechnol. Biochem 71, 2256–2265 (2007)
de Almeida, F.M., Bonini, B.M., Beton, D., Jorge, J.A., Terenzi, H.F., da Silva, A.M.: Heterologous expression in Escherichia coli of Neurospora crassa neutral trehalase as an active enzyme. Protein Expr. Purif. 65, 185–189 (2009)
Frandsen, T.P., Christensen, T., Stoffer, B., Lehmbeck, J., Dupont, C., Honzatko, R.B., Svensson, B.: Mutational analysis of the roles in catalysis and substrate recognition of arginines 54 and 305, aspartic acid 309, and tryptophan 317 located at subsites 1 and 2 in glucoamylase from Aspergillus niger. Biochemistry 34, 10162–10169 (1995)
Ichikawa, K., Tonozuka, T., Uotsu-Tomita, R., Akeboshi, H., Nishikawa, A., Sakano, Y.: Purification, characterization, and substrate affinities of Thermoactinomyces vulgaris R-47 maltooligosaccharide-metabolizing enzyme homologous to glucoamylases. Biosci. Biotechnol. Biochem. 68, 413–420 (2004)
Mertens, J.A., Braker, J.D., Jordan, D.B.: Catalytic properties of two Rhizopus oryzae 99–880 glucoamylase enzymes without starch binding domains expressed in Pichia pastoris. Appl. Biochem. Biotechnol. 162, 2197–2213 (2010)
Nitta, Y., Mizushima, M., Hiromi, K., Ono, S.: Influence of molecular structure of substrates and analogues on Taka-amylase A catalyzed hydrolyses. I. Effect of chain length of linear substrates. J. Biochem. 69, 567–576 (1971)
Yokota, T., Tonozuka, T., Kamitori, S., Sakano, Y.: The deletion of amino-terminal domain in Thermoactinomyces vulgaris R-47 α-amylases: effects of domain N on activity, specificity, stability and dimerization. Biosci. Biotechnol. Biochem. 65, 401–408 (2001)
Yoshigi, N., Okada, Y., Sahara, H., Koshino, S.: Expression in Escherichia coli of cDNA encoding barley β-amylase and properties of recombinant β-amylase. Biosci. Biotechnol. Biochem. 58, 1080–1086 (1994)
Kato, N., Suyama, S., Shirokane, M., Kato, M., Kobayashi, T., Tsukagoshi, N.: Novel α-glucosidase from Aspergillus nidulans with strong transglycosylation activity. Appl. Environ. Microbiol. 68, 1250–1256 (2002)
Sim, L., Quezada-Calvillo, R., Sterchi, E.E., Nichols, B.L., Rose, D.R.: Human intestinal maltase-glucoamylase: crystal structure of the N-terminal catalytic subunit and basis of inhibition and substrate specificity. J. Mol. Biol. 375, 782–792 (2008)
Kitamura, M., Okuyama, M., Tanzawa, F., Mori, H., Kitago, Y., Watanabe, N., Kimura, A., Tanaka, I., Yao, M.: Structural and functional analysis of a glycoside hydrolase family 97 enzyme from Bacteroides thetaiotaomicron. J. Biol. Chem. 283, 36328–36337 (2008)
Acknowledgements
We would like to thank Tomoko Fujii for her technical assistance. This study was supported by a grant-in-aid for Scientific Research (20570103 and 23570132) from the Ministry of Education, Culture, Sports, Science, and Technology of Japan. We thank Hayashibara Biochemical Laboratories Inc. for providing kojibiose and nigerose.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Miyazaki, T., Matsumoto, Y., Matsuda, K. et al. Heterologous expression and characterization of processing α-glucosidase I from Aspergillus brasiliensis ATCC 9642. Glycoconj J 28, 563–571 (2011). https://doi.org/10.1007/s10719-011-9356-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10719-011-9356-z