Abstract
If we predict the host range of new or mutant influenza virus in advance, we are able to measure against pandemic human influenza immediately after the new virus emerges somewhere. Influenza viral hemagglutinin(HA)–sialoside receptor interaction is a target event for in silico chemical prediction studies about the virus host range determination. We theoretically studied avian and human influenza A virus HA H3 subtype complexed with avian or human type receptor Neu5Acα(2-3 or 2-6)Gal analogues by ab initio fragment molecular orbital (FMO) method at the second order Møller–Plesset (MP2)/6–31G level, which can evaluate correctly not only electrostatic interactions but also lipophilic interactions based on van der Waals dispersion force. Avian H3 bound to avian α2-3 11.4 kcal/mol stronger than to human α2-6 in the model complexes with taking account of intermolecular lipophilic interaction. A substitution at the position 226 between Gln(avian) and Leu(human) on influenza H3 HA1 has altered its virus host range between avian and human. In the ab initio FMO studies, binding energy of avian Gln226Leu H3–human α2-6 was quite similar to that in the human H3–human α2-6 complex with amino acid sequence differences at nine positions in the models. This similarity indicates that avian Gln226Leu H3 virus can infect human with the same level as human H3 virus. Opposite mutation Leu226Gln in the human H3 gave the moderate binding energies to avian α2-3 with similarity to avian H3–α2-3 complex that supported our previous virus-sialoside binding assay. Ab initio FMO studies have revealed the relationship between influenza H3 virus host range and H3–α(2-3 or 2-6) receptors binding. Our theoretical approach may predict the infectious level of new viruses and point out some unknown dangerous mutation positions on HA in advance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Avian H5N1 virus has a high potential for pandemic influenza. The original H5N1 avian viruses can bind to avian type receptors on human lower respiratory tract [1, 2], however, this infection mechanism does not cause pandemic human influenza. We should always take precaution against when single or double point mutations occur in H5N1 viral hemagglutinin and its higher binding affinity to human type receptor [3–6].
If we predict the host range of new or mutant influenza virus in advance, we are able to measure against pandemic human influenza immediately after the new virus emerges somewhere. Influenza virus host range is mainly determined by HA binding affinity to host cell surface receptors sialo-glycolipids and -glycoproteins terminating in sialic acid α2-3 or α2-6 galactose [7–10], therefore the HA–sialoside interaction is a target event for in silico chemical prediction studies about the virus host range determination. Quantum chemical calculations will provide various chemical foresights about HA–sialosaccharide interaction without treatments of unknown dangerous influenza virus mutants. We are attempting to construct HA–sialoside complex models in order to explain the HA binding specificity using H3 subtype system [11–12]. Our current interest is focused on avian virus HA–Neu5Acα(2-3/6)Gal bindings and comparison between avian HA mutant–human α2-6 interaction and the original human HA–human α2-6 interaction.
Avian influenza A virus H3 subtype has Gln226 as one of the amino acids on sialoside binding site HA1 that binds to avian type receptor N-acetylneuraminic acid (Neu5Ac) α2-3 galactose (Gal) oligosaccharide stronger than to human type receptor Neu5Acα(2-6)Gal [13]. We remark that avian H3 weakly and certainly recognize human receptor α2-6 [14] on TLC-virus binding assay using synthetic sialylparaglobosides or corresponding B30 derivatives [14–18] that is supported by X-ray crystallographic structure of avian H3–human α2-6 complex [19]. Thus, it is significant to analyze the relationship between avian H3 binding affinity and the corresponding avian H3–Neu5Acα(2-3 or 2-6)Gal complexes. Human H3 virus has Leu226 instead of Gln that strongly recognizes human receptor α2-6 sialosaccharide [13, 20–24]. In the human H3 subtype, point mutations on the sialoside binding site at Tyr98Phe, His183Phe, and Leu194Ala decrease their hemagglutination to human erythrocytes [25]. Human Ser193Ile H3 agglutinates α2-6 sialoside expressed erythrocytes stronger than the original human H3 [26]. In particular, a substitution from Leu226 to Gln at the position 226 in human H3 HA1 changes its binding specificity from human α2-6 to avian α2-3 [16, 27–29]. Weis et al. have reported X-ray crystal structure of human Leu226Gln H3 complexed with avian type receptor α2-3 sialyllactose [20].
For the purpose of theoretical clarification about the H3–sialoside binding properties, we plausibly investigate the binding affinities of H3–Neu5Acα(2-3 or 2-6)Gal complexes by ab initio molecular orbital studies. It is worth while researching the energy profile on all steps in the HA–sialoside interaction composed of several events: encounter the sialoside binding site with sialoside receptor, leaving needless water solvent in the interaction process, thermodynamical relaxation on HA–sialoside complex, and multivalent effect on HA–sialoside interaction. However, we just urgently request a simple and essential approach with a reasonable cost. Ab initio fragment molecular orbital (FMO) calculations [30–39] to evaluate the binding energies between HA and Neu5Acα(2-3/6)Gal receptors are one of the promising strategies.
Ab initio FMO method has been applied in large biochemical systems for quantum chemical analysis to molecular interaction [40–45]. In this method, HA–sialoside complex is divided into fragments, and MO calculations are carried out on each fragment and fragment pairs. The total energy on the HA–sialoside complex is obtained as a summation of the fragment energies and interfragment interaction energies (IFIEs) [30–32, 34, 46, 47].
Our previous ab initio FMO studies estimated the binding energies between influenza H3 and Neu5Acα(2-3/6)Gal receptors at the FMO-Hartree-Fock(HF)/STO–3G level, and analyzed IFIEs between Neu5Ac-Gal receptor and amino acid residues on the sialoside binding site to confirm qualitatively avian H3–avian α2-3 binding affinity based on electrostatic interactions [11]. This approach would be valid qualitatively, but the FMO-HF/STO–3G method cannot evaluate significantly weak interactions such as intermolecular lipophilic stabilization which play important roles in biochemical systems. Taking account of electron correlation effects is necessary for more quantitative treatment of these stabilizations based on van der Waals dispersion force. Recently, the FMO method has been extended to correlated calculations, which can be performed at the MP2 method [36, 48]. Including electron correlation effect, we applied the FMO-MP2 method to HA–sialoside model complexes and analyzed the hydrophilic and lipophilic interactions between Neu5Ac-Gal and amino acid residues on the sialoside binding site HA1 using IFIEs.
2 Methods
Energy minimized structures of avian H3 complexed with Neu5Acα(2-3 or 2-6)Gal disaccharide analogues were prepared from the corresponding X-ray crystallographic structures [19] by molecular mechanics (MM) energy calculation with the CFF force field [49] using Discovery Studio 1.5.1 program as shown in our previous works[11, 12]. We mutated in silico avian H3 Gln226 to Leu in the avian H3–human α2-6 disaccharide complex, changed α2-6 bond dihedral angle to human Leu226 H3 type orientation referring the crystal structure of Neu5Acα(2-6)Galβ(1-4)GlcNAcβ(1-3)Galβ(1-4)Glc: LSTc complexed with human H3 [50], and optimized its geometry by MM calculation to make the energy minimum avian Gln226Leu H3–human α2-6 complex. Optimum human H3–human α2-6 complex was given in silico by modification of crystallographic structure of human H3–Neu5Acα(2-3)Galβ(1-4)Glc complex [51]. We replaced the α2-3 trisaccharide receptor with Neu5Acα(2-6)Gal disaccharide based on the common Neu5Ac residue and changed its α2-6 bond dihedral angle referring LSTc conformation complexed with human H3 [50] that was similar manner to the previous study [12]. Human Leu226Gln H3–avian α2-3 complex was prepared by in silico point mutation from Leu226 to Gln in the human H3–avian α2-3 complex [51], changed α2-3 bond dihedral angle to avian Gln226 H3 type orientation [19] referring the crystal structure of human Leu226Gln–avian α2-3 complex [20] followed by geometry optimization. MM calculations were carried out in vacuo with structural determined water molecules in the crystal structures.
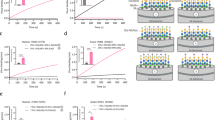
We cut out the H3–sialoside models (HA: N96–P99, G129–Y161, G181–V196, and N216–I232, 70 amino acid residues) from the optimum structures for ab initio FMO calculations (Fig. 1A). This approach covers the amino acid residues on the sialoside binding site especially position 226, besides whose range is outside of substitutions far away from the binding site. Peptide terminals in the models were treated as NH3 and COO similar manner to the previous study [11]. Amino acid sequences of avian and human H3 differ at ten positions with same secondary structure as shown in Fig. 1B. The differences at positions 137, 145, 226, and 228 are located on the direct interaction site to Neu5Ac-Gal disaccharides. Since there are few electron density of water molecules around the sialoside binding site–Neu5Acα(2-3/6)Gal analogue complex in the H3 crystal structures, we computed the model complexes by the FMO method without water molecules.
Interaction site of avian influenza A virus HA H3 subtype complexed with avian type receptor Neu5Acα(2-3)Gal analogue for ab initio FMO studies. A Ribbon model; avian H3 sialoside binding site; N96–P99, G129–Y161, G181–V196, and N216–I232. CPK model; Neu5Acα(2-3)Gal disaccharide, B sequence alignments of avian/human H3 in the model complexes. yellow; non-matching residues, green; weak matching residues, light blue; strong matching residues, red; helix, blue; sheet
Single point energies of the model complexes were computed at the FMO-HF level with STO–3G [11], 6–31G, and 6–31G(d) basis sets, and the correlated FMO-MP2/6–31G level of theory. The later method evaluates interfragment van der Waals type stabilization in the protein–ligand complex. Stabilizations of CH–π interaction between tryptophan and carbohydrate in the β-galactosidase-substrates or products complex were calculated at the MP2/6–31+G(d) level as 2.4–5.2 kcal/mol [52]. Stabilizing interaction energy of the fucose-benzene complex was estimated 3.0 kcal/mol at the MP2/6–31G(d,p) level of theory [53].
The 70 amino acids on the H3-sialoside complexes were divided into one amino acid residue as a single fragment using automatic fragmentation program in the ABINIT-MP package, and Neu5Ac-Gal receptors were also treated as a single fragment. The receptors and sialoside binding sites were charged to −1 and +1. We calculated single point energies of the complexes (E complex), Neu5Ac-Gal (E receptor), and the binding sites (E H3) to estimate binding energies (ΔE) between the receptor and H3 by the following expression; ΔE = (E receptor + E H3) − E complex. Ab initio FMO calculations were performed by using ABINIT-MP program [30–39].
3 Results and discussion
Figures 2, 3, 4, and 5 show the direct interactions between Neu5Ac-Gal and sialoside binding site on H3 HA1. Binding energies ΔE and the selected IFIEs of Neu5Acα(2-3 or 2-6)Gal with amino acid residues at the FMO-HF/STO–3G and FMO-MP2/6–31G levels are summarized in Tables 1, 2, 3, 4, and 5.
Intermolecular interactions of Neu5Acα(2-3/6)Gal with amino acid residues on the sialoside binding site in avian H3. A–C; avian H3–avian α2-3 complex. D–F; avian H3–human α2-6 complex. The red and black dotted lines represent hydrogen bonds and long range interactions whose distances are given in angstrom
ΔEs are estimated with electron correlation at the FMO-MP2/6–31G level (Table 1). In the avian H3 complexes, MP2/6–31G ΔEs are 43.5 and 40.3 kcal/mol larger than the corresponding HF/STO–3G energies to give ΔE α2-3–ΔE α2-6 11.4 kcal/mol (entries 1, 3). Extension from minimal basis sets to valence double zeta basis sets at the HF level affords the differences ΔE α2-3–ΔE α2-6 13.1 kcal/mol in the avian H3–α(2-3/6) complexes.
As shown in Tables 2 and 3, various intermolecular lipophilic interactions benefit forming H3–Neu5Acα(2-3/6)Gal complexes at the FMO-MP2/6–31G level. Neu5-NHCOCH 3 makes CH–π interaction network with Trp153 and Ile230 to afford MP2/6–31G IFIEs 8.5 and 8.6 kcal/mol in avian H3–α(2-3/6) complexes, 4.0 kcal/mol in human H3–human α2-6 complex (Figs. 3B, E, and 5B, Tables 2 and 3; entry 5), while the corresponding HF/STO–3G energies do not evaluate the CH–π interaction stabilization. Gly134 and Thr155 directly interact with Neu 5-NHCOCH 3 , but do not contribute to the HA–α(2-3/6) binding (Tables 2 and 3; entries 6, 12). Hydrophobic association between Leu194 and Neu7-CH, 9-CH2 has bonding IFIEs 2.4–3.7 kcal/mol at the MP2/6–31G level (Figs. 3C, F, and 5C, Tables 2 and 3; entry 13). Intermolecular hydrophilic interactions also advantage the forming HA–sialoside complex. In the avian H3 complexes, MP2/6–31G IFIEs are 5.4–9.1 kcal/mol larger than the corresponding HF/STO–3G energies (Table 2; entries 1, 2, 4, 8, 10). Especially, Gln226 on avian H3 interacts with avian α2-3 whose MP2/6–31G IFIE is estimated 15.1 kcal/mol larger than its HF/STO–3G energy (Fig. 3A, Table 2; entry 3). In contrast, IFIE between Ser137 and Neu 1-CO1O1’ has 33.9 kcal/mol at the MP2/6–31G which is 2.6 kcal/mol smaller than the HF/STO–3G energy (Fig. 3B, Table 2; entry 9). Gly135 clearly forms an intermolecular hydrogen bond with Neu 5-NHCOCH3, but whose IFIE are quite smaller than the others (Tables 2 and 3; entry 7).
In the following section, we discuss molecular interaction between Neu5Acα(2-3/6)Gal and amino acid residues on the sialoside binding site in each complex with ΔEs and IFIEs at the FMO-MP2/6–31G level.
3.1 Avian H3–Neu5Acα(2-3 or 2-6)Gal complexes
Avian H3 binds to avian receptor Neu5Acα(2-3)Gal 11.4 kcal/mol stronger than to human receptor Neu5Acα(2-6)Gal with taking account of intermolecular lipophilic stabilization at the FMO-MP2/6–31G level (Table 1; entries 1, 3). In the avian H3–avian α2-3 complex, side chain NH2CO on Gln226 forms intermolecular hydrogen bond network with 8-OH, 1-COO on Neu5Ac, Gal 4-OH, and weakly with glycoside oxygen O3 to give IFIE 23.9 kcal/mol, which is 6.5 kcal/mol larger than the corresponding IFIE in avian H3–human α2-6 complex (Fig. 3A, D, Table 2; entry 3). Besides, hydrogen bond formations of Neu 1-COO with Ser136, Ser137 and Asn145 strongly stabilize the avian H3–avian α2-3 complex by IFIEs 27.1, 33.9, and 13.2 kcal/mol (Fig. 3B, E, Table 2; entries 8–10). The later interaction consists of intermolecular hydrogen bonds between Asn145 side chain CONH2 and O1’O1C1 on Neu5Acα(2-3)Gal with 2.78 Å. Gly228 on influenza A virus H3 subtype correlates with the shift of virus host range between avian and human. In our study, Gly228 interacts with Neu5Acα(2-3/6)Gal disaccharides by IFIEs 4.7 and 4.3 kcal/mol (Fig. 3C, F, Table 2; entry 11).
Comparison between ΔEs and sums of 13 IFIEs, the formers are about 20 kcal/mol larger than the 13 IFIEs sums (Table 1; entries 1, 3, Table 2; entry 14). IFIEs sum difference of 9.3 kcal/mol mainly causes the binding energy difference (ΔE α2-3–ΔE α2-6) of 11.4 kcal/mol. In the X-ray crystal structure of avian H3 complexed with α(2-3 or 2-6)-pentasaccharide receptors, the sialoside binding site reliably recognizes the non-reducing terminal Neu5Ac-Gal [19] that approximately allows our theoretical approach.
3.2 Human H3–Human Neu5Acα(2-6)Gal complex
Binding energy ΔE in the human H3–human Neu5Acα(2-6)Gal disaccharide was estimated 154.3 kcal/mol at the FMO-MP2/6–31G level (Table 1; entry 5). Human H3 complex has an intramolecular hydrogen bond at Neu 8-OH∙∙∙O1C1O1’ instead of intermolecular Gln226∙∙∙Neu5Ac interaction observed in the avian H3 complex (Fig. 5A), thus ΔE in the human H3 complex is smaller by 15 kcal/mol than the corresponding ΔE in the avian Gln226 H3 complex (Table 1; entries 3, 5). Neu 1-COO interacts with Ser136 by 29.3 kcal/mol, Asn137 by 45.5 kcal/mol, and Ser145 by 2.6 kcal/mol (Table 3; entries 8–10) whose summation is larger by 7.3 kcal/mol than that in the avian H3–human α2-6 complex (Table 2; entries 8–10). Intermolecular hydrogen bond network between His183, Tyr98 and Neu 9,8-OH stabilizes the human H3–human α2-6 complex with IFIEs of 8.9 and 16.2 kcal/mol (Fig. 5C, Table 3; entries 1, 2), therefore human Tyr98Phe or His183Phe H3 viruses lose this stabilization by hydrogen bonds decomposition to decrease severely their binding affinities [25]. Tyr98 forms a hydrogen bond with His183 by IFIE of 10.5 kcal/mol, and CH–π interaction with Leu226 by 1.2 kcal/mol.
Inter- and intra-molecular lipophilic association is significant for the human H3–human α2-6 binding. Leu226 interacts with Gal 6-CH2 on human α2-6 by lipophilic IFIE of 6.1 kcal/mol (Fig. 5A, Table 3; entry 3) that is supported by a lipophilic network beyond Leu226 consisted Tyr98, Pro99, Ala138, (CH2)3 on Arg220 and Arg229, Ile230, and Trp153. Hydrophobic site on Neu 7-CH 9-CH2 associates with lipophilic Leu194 by IFIE of 2.4 kcal/mol (Fig. 5C, Table 3; entry 13) supported by hydrophobic groups around Leu194 such as Thr155, Tyr195, (CH2)2 on Glu190, and CH2 on Ser193. Since human Leu194Ala H3 virus does not agglutinate human erythrocyte [25], the lipophilic association around Leu194 is significant for the human H3–human α2-6 binding affinity. A substitution at Ser193 to Ile in human H3 HA1 increases the hydrophobic capacity around Leu194 to give higher binding affinity to human α2-6 [26]. The two lipophilic networks around Leu226∙∙∙Gal 6-CH2 and Leu194∙∙∙Neu 7-CH 9-CH2 are connected by a hydrogen bond formation of indole NH Trp153 with OH Tyr195 (Fig. 5C). However, both substitutions Trp153Phe or Tyr195Phe partially decrease its binding affinity [8, 25].
3.3 Avian Gln226Leu H3–Human Neu5Acα(2-6)Gal and human Leu226Gln H3–Avian Neu5Acα(2-3)Gal complexes
Figures 6 and 7 show the interaction of Neu5Ac-Gal with avian Gln226Leu/human Leu226Gln H3s. Avian Gln226Leu H3 moderately binds to human α2-6 with ΔE 157.6 kcal/mol at the FMO-MP2/6–31G level (Table 1; entry 4). This binding energy is quite similar to ΔE α2-6 154.3 kcal/mol in the human H3 complex with amino acid difference at nine positions (Fig. 1B), therefore avian Gln226Leu H3 virus can infect human with the same level as human H3 virus. In the avian Gln226Leu H3–human α2-6 complex, Leu226 associates with lipophilic Gal 6-CH2 whose IFIE is smaller by 1.3 kcal/mol than that of Leu226∙∙∙Gal 6-CH2 interaction in the human H3 complex (Tables 3 and 4; entry 3). Besides Neu 8-OH makes an intramolecular hydrogen bond with O1C1O1’-Neu similar manner to that in the human Leu226 H3 complex (Figs. 5A and 6).
Human Leu226Gln H3 interacts with avian α2-3 by ΔE α2-3 of 179.7 kcal/mol that is almost the same value as ΔE α2-3 of 180.4 kcal/mol in the avian H3 complex at the FMO-MP2/6–31G level (Table 1; entries 1, 2). In the human Leu226Gln H3–avian α2-3 complex, Gln226 makes intermolecular hydrogen bond network with 8-OH, 1-CO1O1’ on Neu5Ac and Gal 4-OH to give IFIE 31.1 kcal/mol (Fig. 7, Table 5; entry 3). We previously reported that A/Udorn Leu226Gln (human Leu226Gln H3N2) bound to avian α2-3 receptor [14]. Our ab initio FMO studies have confirmed that human Leu226Gln H3 moderately interacts with avian α2-3 analogue at the correlated FMO-MP2/6–31G level.
4 Conclusion
We theoretically studied influenza A virus hemagglutinin H3 subtype complexed with avian/human type receptor Neu5Acα(2-3 or 2-6)Gal analogues by ab initio FMO method at the correlated MP2/6–31G level. Avian H3 bound to avian α2-3 11.4 kcal/mol stronger than to human α2-6 in the model complexes with taking account of intermolecular lipophilic interaction. Single point substitution at the position 226 on H3 subtype sialoside binding site HA1 changes its binding affinity between avian α2-3 and human α2-6. Our ab initio FMO studies showed that the binding energy of avian Gln226Leu H3 with human α2-6 was similar value to that in the human H3–human α2-6 complex at the FMO-MP2/6–31G level with amino acid differences at nine positions in our models. Thus avian Gln226Leu H3 virus can infect human with the same level as human H3 virus. Opposite mutation Leu226Gln in the human H3 gave the moderate binding energy to avian α2-3 that supported our previous virus-sialoside binding assay.
As shown in a schematic summary (Fig. 8), ab initio FMO studies revealed the relationship between influenza H3 virus host range and H3–α(2-3 or 2-6) receptors binding. Our theoretical approach will predict the infectious level of new viruses and point out some unknown dangerous mutation positions on HA in advance against human pandemic influenza.
Binding energy profile at the FMO-MP2/6–31G level on the relationship between influenza H3 virus host range and H3–α(2-3 or 2-6) receptors binding. ΔEs are shown in Table 1
References
Shinya, K., Ebina, M., Yamada, S., Ono, M., Kasai, N., Kawaoka, Y.: Avian flu: influenza virus receptors in the human airway. Nature 440, 435–436 (2006)
van Riel, D., Munster, V.J., de Wit, E., Rimmelzwaan, G.F., Fouchier, R.A.M., Osterhaus, A.D.M.E., Kuiken, T.: H5N1 virus attachment to lower respiratory tract. Science 312, 399 (2006)
Yamada, S., Suzuki, Y., Suzuki, T., Le, M.Q., Nidom, C.A., Sakai-Tagawa, Y., Muramoto, Y., Ito, M., Kiso, M., Horimoto, T., Shinya, K., Sawada, T., Kiso, M., Usui, T., Murata, T., Lin, Y., Hay, A., Haire, L.F., Stevens, D.J., Russell, R.J., Gamblin, S.J., Skehel, J.J., Kawaoka, Y.: Haemagglutinin mutations responsible for the binding of H5N1 influenza A viruses to human-type receptors. Nature 444, 378–382 (2006)
Stevens, J., Blixt, O., Tumpey, T.M., Taubenberger, J.K., Paulson, J.C., Wilson, I.A.: Structure and receptor specificity of the hemagglutinin from an H5N1 influenza virus. Science 312, 404–410 (2006)
Yang, Z-Y., Wei, C-J., Kong, W-P., Wu, L., Xu, L., Smith, D.F., Nabel, G.J.: Mutants with altered receptor binding specificity immunization by avian H5 influenza hemagglutinin. Science 317, 825–828 (2007)
Auewarakul, P., Suptawiwat, O., Kongchanagul, A., Sangma, C., Suzuki, Y., Ungchusak, K., Louisirirotchanakul, S., Lerdsamran, H., Pooruk, P., Thitithanyanont, A., Pittayawonganon, C., Guo, C-T., Hiramatsu, H., Jampangern, W., Chunsutthiwat, S., Puthavathana, P.: An avian influenza H5N1 virus that binds to a human-type receptor. J. Virol. 81, 9950–9955 (2007)
Suzuki, Y.: Sialobiology of influenza: molecular mechanism of host range variation of influenza viruses. Biol. Pharm. Bull. 28, 399–408 (2005)
Skehel, J.J., Wiley, D.C.: Receptor binding and membrane fusion in virus entry: the influenza hemagglutinin. Ann. Rev. Biochem. 69, 531–569 (2000)
Horimoto, T., Kawaoka, Y.: Influenza: lessons from past pandemics, warnings from current incidents. Nat. Rev. Microbiol. 3, 591–600 (2005)
Matrosovich, M.N., Klenk, H.D., Kawaoka, Y.: Receptor specificity, host-range, and pathogenicity of influenza viruses. In: Kawaoka, Y. (ed.) Influenza Virology Current Topics, pp. 95–137. Caister Academic Press, Norfolk (2006)
Sawada, T., Hashimoto, T., Nakano, H., Suzuki, T., Ishida, H., Kiso, M.: Why does avian influenza A virus hemagglutinin bind to avian receptor stronger than to human receptor? Ab initio fragment molecular orbital studies. Biochem. Biophys. Res. Commun. 351, 40–43 (2006)
Sawada, T., Hashimoto, T., Nakano, H., Suzuki, T., Suzuki, Y., Kawaoka, Y., Ishida, H., Kiso, M.: Influenza viral hemagglutinin complicated shape is advantageous to its binding affinity for sialosaccharide receptor. Biochem. Biophys. Res. Commun. 355, 6–9 (2007)
Ito, T., Suzuki, Y., Mitnaul, L., Vines, A., Kida, H., Kawaoka, Y.: Receptor specificity of influenza A viruses correlates with the agglutination of erythrocytes from different animal species. Virology 227, 493–499 (1997)
Ito, T., Suzuki, Y., Suzuki, T., Takada, A., Horimoto, T., Wells, K., Kida, H., Otsuki, K., Kiso, M., Ishida, H., Kawaoka, Y.: Recognition of N-glycolylneuraminic acid linked to galactose by the a2,3 linkage is associated with intestinal replication of influenza A virus in ducks. J. Virol. 74, 9300–9305 (2000)
Suzuki, Y., Nakano, T., Ito, T., Watanabe, N., Toda, Y., Guiyun, X., Suzuki, T., Kobayashi, T., Kimura, Y., Yamada, A., Sugawara, K., Nishimura, H., Kitame, F., Nakamura, K., Deya, E., Kiso, M., Hasegawa, A.: Structural determination of gangliosides that bind to influenza A, B, and C viruses by an improved binding assay: strain-specific receptor epitopes in sialo-sugar chains. Virology 189, 121–131 (1992)
Ito, T., Suzuki, Y., Takada, A., Kawamoto, A., Otsuki, K., Masuda, H., Yamada, M., Suzuki, T., Kida, H., Kawaoka, Y.: Differences in sialic acid–galactose linkages in the chicken egg amnion and allantois influence human influenza virus receptor specificity and variant selection. J. Virol. 71, 3357–3362 (1997)
Masuda, H., Suzuki, T., Sugiyama, Y., Horiike, G., Murakami, K., Miyamoto, D., Hidari, K.I.-P.J., Ito, T., Kida, H., Kiso, M., Fukunaga, K., Ohuchi, M., Toyoda, T., Ishihama, A., Kawaoka, Y., Suzuki, Y.: Substitution of amino acid residue in influenza A virus hemagglutinin affects recognition of sialyl-oligosaccharides containing N-glycolylneuraminic acid. FEBS Lett. 464, 71–74 (1999)
Shinya, K., Hatta, M., Yamada, S., Takada, A., Watanabe, S., Halfmann, P., Horimoto, T., Neumann, G., Hyun Kim, J., Lim, W., Guan, Y., Peiris, M., Kiso, M., Suzuki, T., Suzuki, Y., Kawaoka, Y.: Characterization of a human H5N1 influenza A virus isolated in 2003. J. Virol. 79, 9926–9932 (2005)
Ha, Y., Stevens, D.J., Skehel, J.J., Wiley, D.C.: X-ray structure of the hemagglutinin of a potential H3 avian progenitor of the 1968 Hong Kong pandemic influenza virus. Virology 309, 209–218 (2003)
Weis, W., Brown, J.H., Cusack, S., Paulson, J.C., Skehel, J.J., Wiley, D.C.: Structure of the influenza virus haemagglutinin complexed with its receptor, sialic acid. Nature 333, 426–431 (1988)
Connor, R.J., Kawaoka, Y., Webster, R.G., Paulson, J.C.: Receptor specificity in human, avian, and equine H2 and H3 influenza virus isolates. Virology 205, 17–23 (1994)
Gambaryan, A.S., Piskarev, V.E., Yamskov, I.A., Sakharov, A.M., Tuzikov, A.B., Bovin, N.V., Nifant’ev, N.E., Matrosovich, M.N.: Human influenza virus recognition of sialyloligosaccharides. FEBS Lett. 366, 57–60 (1995)
Gambaryan, A.S., Tuzikov, A.B., Piskarev, V.E., Yamnikova, S.S., Lvov, D.K., Robertson, J.S., Bovin, N.V., Matrosovich, M.N.: Specification of receptor-binding phenotypes of influenza virus isolates from different hosts using synthetic sialylglycopolymers: nonegg-adapted human H1 and H3 influenza A and influenza B viruses share a common high binding affinity for 6’-sialyl(N-acetyllactosamine). Virology 232, 345–350 (1997)
Nobusawa, E., Ishihara, H., Morishita, T., Sato, K., Nakajima, K.: Change in receptor-binding specificity of recent human influenza A viruses (H3N2): a single amino acid change in hemagglutinin altered its recognition of sialyloligosaccharides. Virology 278, 587–596 (2000)
Martin, J., Wharton, S.A., Lin, Y.P., Takemoto, D.K., Skehel, J.J., Wiley, D.C., Steinhauer, D.A.: Studies of the binding properties of influenza hemagglutinin receptor-site mutants. Virology 241, 101–111 (1998)
Daniels, R.S., Jeffries, S., Yates, P., Schild, G.C., Rogers, G.N., Paulson, J.C., Wharton, S.A., Douglas, A.R., Skehel, J.J., Wiley, D.C.: The receptor-binding and membrane-fusion properties of influenza virus variants selected using anti-hemagglutinin monoclonal antibodies. EMBO J. 6, 1459–1465 (1987)
Rogers, G.N., Paulson, J.C., Daniels, R.S., Skehel, J.J., Wilson, I.A., Wiley, D.C.: Single amino acid substitutions in influenza haemagglutinin change receptor binding specificity. Nature 304, 76–78 (1983)
Naeve, C.W., Hinshaw, V.S., Webster, R.G.: Mutations in the hemagglutinin receptor-binding site can change the biological properties of an influenza virus. J. Virol. 51, 567–569 (1984)
Rogers, G.N., Daniels, R.S., Skehel, J.J., Wiley, D.C., Wang, X.F., Higs, H.H., Paulson, J.C.: Host-mediated selection of influenza virus receptor variants. Sialic acid-alpha 2, 6 gal-specific clones of A/duck/Ukraine/1/63 revert to sialic acid-alpha 2,3Gal-specific wild type in ovo. J. Biol. Chem. 260, 7362–7367 (1985)
Kitaura, K., Sawai, T., Asada, T., Nakano, T., Uebayasi, M.: Pair interaction molecular orbital method: an approximate computational method for molecular interactions. Chem. Phys. Lett. 312, 319–324 (1999)
Kitaura, K., Ikeo, E., Asada, T., Nakano, T., Uebayasi, M.: Fragment molecular orbital method: an approximate computational method for large molecules. Chem. Phys. Lett. 313, 701–706 (1999)
Nakano, T., Kaminuma, T., Sato, T., Akiyama, Y., Uebayasi, M., Kitaura, K.: Fragment molecular orbital method: application to polypeptides. Chem. Phys. Lett. 318, 614–618 (2000)
Kitaura, K., Sugiki, S., Nakano, T., Komeiji, Y., Uebayasi, M.: Fragment molecular orbital method: analytical energy gradients. Chem. Phys. Lett. 336, 163–170 (2001)
Nakano, T., Kaminuma, T., Sato, T., Fukuzawa, K., Akiyama, Y., Uebayasi, M., Kitaura, K.: Fragment molecular orbital method: use of approximate electrostatic potential. Chem. Phys. Lett. 351, 475–480 (2002)
Mochizuki, Y., Nakano, T., Koikegami, S., Tanimori, S., Abe, Y., Nagashima, U., Kitaura, K.: A parallelized integral-direct second-order Møller–Plesset perturbation theory method with a fragment molecular orbital scheme. Theor. Chem. Acc. 112, 442–452 (2004)
Mochizuki, Y., Koikegami, S., Nakano, T., Amari, S., Kitaura, K.: Large scale MP2 calculations with fragment molecular orbital scheme. Chem. Phys. Lett. 396, 473–479 (2004)
Mochizuki, Y., Koikegami, S., Amari, S., Segawa, K., Kitaura, K., Nakano, T.: Configuration interaction singles method with multilayer fragment molecular orbital scheme. Chem. Phys. Lett. 406, 283–288 (2005)
Mochizuki, Y.: A size-extensive modification of super-CI for orbital relaxation. Chem. Phys. Lett. 410, 165–171 (2005)
Mochizuki, Y., Fukuzawa, K., Kato, A., Tanaka, S., Kitaura, K., Nakano, T.: A configuration analysis for fragment interaction. Chem. Phys. Lett. 410, 247–253 (2005)
Fedorov, D.G., Kitaura, K.: Extending the power of quantum chemistry to large systems with the fragment molecular orbital method. J. Phys. Chem. A 111, 6904–6914 (2007)
Fukuzawa, K., Kitaura, K., Uebayasi, M., Nakata, K., Kaminuma, T., Nakano, T.: Ab initio quantum mechanical study of the binding energies of human estrogen receptor with its ligands: an application of fragment molecular orbital method. J. Comput. Chem. 26, 1–10 (2005)
Fukuzawa, K., Komeiji, Y., Mochizuki, Y., Kato, A., Nakano, T., Tanaka, S.: Intra- and intermolecular interactions between cyclic-AMP receptor protein and DNA: ab initio fragment molecular orbital study. J. Comput. Chem. 27, 948–960 (2006)
Fukuzawa, K., Mochizuki, Y., Shigenori, T., Kitaura, K., Nakano, T.: Molecular interactions between estrogen receptor and its ligand studied by the ab initio fragment molecular orbital method. J. Phys. Chem. B 110, 16102–16110 (2006)
Yamamoto, K., Abe, D., Yoshimoto, N., Choi, M., Yamagishi, K., Tokiwa, H., Shimizu, M., Makishima, M., Yamada, S.: Vitamin D receptor: ligand recognition and allosteric network. J. Med. Chem. 49, 1313–1324 (2006)
Yamagishi, K., Yamamoto, K., Yamada, S., Tokiwa, H.: Functions of key residues in the ligand-binding pocket of vitamin D receptor: fragment molecular orbital-interfragment interaction energy analysis. Chem. Phys. Lett. 420, 465–468 (2006)
Nakano, T., Mochizuki, Y., Amari, S., Kobayashi, M., Fukuzawa, K., Tanaka, S.: Applications of the fragment molecular orbital (FMO) method to nano-bio field. J. Comput. Chem. Jpn. 6, 173–184 (2007)
Fukuzawa, K., Nakano, T., Kato, A., Mochizuki, Y., Tanaka, S.: Applications of the fragment molecular orbital method for bio-macromolecules. J. Comput. Chem. Jpn. 6, 185–197 (2007)
Fedorov, D.G., Kitaura, K.: Second order Moller–Plesset perturbation theory based upon the fragment molecular orbital method (FMO). J. Chem. Phys., 121, 2483–2490 (2004)
Maple, J.R., Hwang, M.-J., Jalkanen, K.J., Stockfisch, T.P., Hagler, A.T.: Derivation of class II force fields: quantum force field for amides, peptides, and related compounds. J. Comput. Chem. 19, 430–458 (1998)
Eisen, M.B., Sabesan, S., Skehel, J.J., Wiely, D.C.: Binding of the influenza A virus to cell-surface receptors: structures of five hemagglutinin-sialyloligosaccharide complexes determined by x-ray crystallography. Virology 232, 19–31 (1997)
Sauter, N.K., Hanson, J.E., Glick, G.D., Brown, J.H., Crowther, R.L., Park, S.J., Skehel, J.J., Wiley, D.C.: Binding of influenza virus hemagglutinin to analogs of its cell-surface receptor, sialic acid: analysis by proton nuclear magnetic resonance spectroscopy and x-ray crystallography. Biochemistry 31, 9609–9621 (1992)
Spiwok, V., Lipovová, P., Skálová, T., Buchtelová, E., Hašekb, J., Královaá, B.: Role of CH/π interactions in substrate binding by Escherichia coli β-galactosidase. Carbohydr. Res. 339, 2275–2280 (2004)
Fernández-Alonso, M.D.C., Cañada, F.J., Jiménez-Barbero, J., Gabriel Cuevas, G.: Molecular recognition of saccharides by proteins. insights on the origin of the carbohydrate–aromatic interactions. J. Am. Chem. Soc. 127, 7379–7386 (2005)
Acknowledgment
This work was supported in part by CREST, Japan Science and Technology Agency (JST), and a Grant-in-Aid (No. 17101007 to M. Kiso) for Scientific Research from Japan Society for the Promotion of Science.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Sawada, T., Hashimoto, T., Tokiwa, H. et al. Ab initio fragment molecular orbital studies of influenza virus hemagglutinin–sialosaccharide complexes toward chemical clarification about the virus host range determination. Glycoconj J 25, 805–815 (2008). https://doi.org/10.1007/s10719-008-9141-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10719-008-9141-9