Abstract
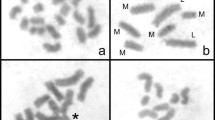
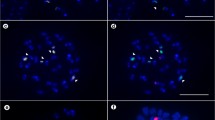
Studies of rDNA location in holocentric chromosomes of the Cyperaceae are scarce, but a few reports have indicated the occurrence of multiple 45S rDNA sites at terminal positions, and in the decondensed state of these regions in prometaphase/metaphase. To extend our knowledge of the number 45S and 5S rDNA sites and distribution in holocentric chromosomes of the Cyperaceae, 23 Brazilian species of Eleocharis were studied. FISH showed 45S rDNA signals always located in terminal regions, which varied from two (E. bonariensis with 2n = 20) to ten (E. flavescens with 2n = 10 and E. laeviglumis with 2n = 60). 5S rDNA showed less variation, with 16 species exhibiting two sites and 7 species four sites, preferentially at terminal positions, except for four species (E. subarticulata, E. flavescens, E. sellowiana and E. geniculata) that showed interstitial sites. The results are discussed in order to understand the predominance of terminal rDNA sites, the mechanisms involved in the interstitial positioning of 5S rDNA sites in some species, and the events of amplification and dispersion of 45S rDNA terminal sites.



Similar content being viewed by others
References
Berjano R, Roa F, Talavera S, Guerra M (2009) Cytotaxonomy of diploid and polyploid Aristolochia (Aristolochiaceae) species based on the distribution of CMA/DAPI bands and 5S and 45S rDNA sites. Plant Syst Evol 280:219–227. doi:10.1007/s00606-009-0184-6
Da Silva CRM, González-Elizondo MS, Vanzela ALL (2005) Reduction of chromosome number in Eleocharis subarticulata (Cyperaceae) by multiple translocation. Bot J Linn Soc 149:457–464. doi:10.1111/j.1095-8339.2005.00449.x
Da Silva CRM, González-Elizondo MS, Rego LNAA, Torezan JMD, Vanzela ALL (2008a) Cytogenetical and cytotaxonomical analysis of some Brazilian species of Eleocharis. Aust J Bot 56:82–90. doi:10.1071/BT070170067-1924/08/010082
Da Silva CRM, González-Elizondo MS, Vanzela ALL (2008b) Chromosome reduction in Eleocharis maculosa (Cyperaceae). Cytogenetic and Genome Res 122:175–180. doi:10.1159/000163096
Da Silva CRM, Trevisan R, González-Elizondo MS, Ferreira JM, Vanzela ALL (2010) Karyotypic diversification and its contribution to the taxonomy of Eleocharis (Cyperaceae) from Brazil. Aust J Bot 58:1–12. doi:10.1071/BT091850067-1924/10/010001
De Melo NF, Guerra MS (2003) Variability of the 5S and 45S rDNA sites in Passiflora L. species with distinct base chromosome numbers. Ann Bot 92:309–316. doi:10.1093/aob/mcg138
Furuta T, Kondo K (1999) Sites of 18S–5.8S–26S rDNA sequences in diffused-centromeric chromosomes of Drosera falconerii. Chromosome Sci 3:69–73. doi:00A0408029
Gerlach WL, Bedbrook JR (1979) Cloning and characterization of ribosomal RNA genes from wheat and barley. Nucleic Acids Res 7:1869–1885. doi:10.1093/nar/7.7.1869
González-Elizondo MS, Peterson PM (1997) A classification of and key to the supraspecific taxa in Eleocharis (Cyperaceae). Taxon 46:433–449. doi:10.2307/1224386
Govaerts R, Simpson DA, Bruhl JJ, Egorova T, Goetghebeur P, Wilson K (2007) World checklist of Cyperaceae. Kew Publishing, Sedges. Kew
Guerra MS (2000) Patterns of heterochromatin distribution in plant chromosomes. Genet Mol Biol 23:1029–1041. doi:10.1590/S1415-47572000000400049
Guerra MS, García M (2004) Heterochromatin and rDNA sites distribution in the holocentric chromosomes of Cuscuta approximata Bab(Convolvulaceae). Genome 47:134–140. doi:10.1139/G03-098
Håkansson A (1958) Holocentric chromosomes in Eleocharis. Hereditas 44:531–540
Hasterok R, Maluszynska J (2000) Cytogenetic markers of Brassica napus L. chromosome. J Appl Genet 41:1–9
Hipp AL, Rothrock PE, Roalson EH (2009) The evolution of chromosome arrangements in Carex (Cyperaceae). Bot Rev 75:96–109. doi:10.1007/s12229-008-9022-8
Hoshi Y (1995) Chromosome studies in Drosera, the Droseraceae in connection with possible origin of the basic chromosome number of x = 10 well-differentiated in the North-hemisphere. Dissertation, Hiroshima University
Kwon J, Kim B (2009) Localization of 5S and 25S rRNA genes on somatic and meiotic chromosomes in Capsicum species of chili pepper. Mol Cells 27:205–209. doi:10.1007/s10059-009-0025-z
Leitch IJ, Bennett MD (1997) Polyploidy in angiosperms. Trend in Plant Science 2:470–476. doi:10.1016/S1360-1385(97)01154-0
Lim K, Wennekes J, Jong JH, Jacobsen E, Tuyl JM (2001) Karyotype analysis of Lilium longiflorum and Lilium rubellum by chromosome banding and fluorescence in situ hybridization. Genome 44:911–918. doi:10.1139/gen-44-5-911
Löve A, Löve D, Raymond M (1957) Cytotaxonomy of Carex section Cappilares. Can J Bot 35:715–761
Moscone EA, Klein F, Lambrou M, Fuchs J, Schweizer D (1999) Quantitative karyotyping and dual-color FISH mapping of 5S and 18S–25S rDNA probes in the cultivated Phaseolus species (Leguminosae). Genome 42:1224–1233. doi:10.1139/gen-42-6-1224
Pedersen C, Linde-Laursen I (1994) Chromosomal locations of four minor rDNA loci and a marker microsatellite sequence in barley. Chromosome Res 2:65–71. doi:10.1007/BF01539456
Raskina O, Barber JC, Nevo E, Belyayev A (2008) Repetitive DNA and chromosomal rearrangements: speciation-related events in plant genomes. Cytogenet Genome Res 120:351–357. doi:10.1159/000121084
Rego LNAA, Da Silva CRM, Torezan JMD, Gaeta ML, Vanzela ALL (2009) Cytotaxonomical study in Brazilian species of Solanum, Lycianthes and Vassobia (Solanaceae). Plant Syst Evol 279:93–102. doi:10.1007/s00606-009-0149-9
Roalson EH (2008) A synopsis of chromosome number variation in the Cyperaceae. Bot Rev 74:209–393. doi:10.1007/s12229-008-9011-y
Schubert I, Wobus U (1985) In situ hybridization confirms jumping nucleolus organizing regions in Allium. Chromosoma 92:143–148. doi:10.1007/BF00328466
Schweizer D, Loidl J (1987) A model for heterochromatin dispersion and the evolution of C-banded patterns. Chromosom today 9:61–74
Vanzela ALL, Cuadrado A, Jouve N, Luceño M, Guerra M (1998) Multiple locations of the rDNA sites in holocentric chromosomes of Rhynchospora (Cyperaceae). Chromosome Res 6:345–349. doi:10.1023/A:1009279912631
Vanzela ALL, Luceño M, Guerra M (2000) Karyotype evolution and cytotaxonomy in Brazilian species of Rhynchospora Vahl (Cyperaceae). Bot J Linn Soc 134:557–566. doi:10.1006/bojl.2000.0352
Vanzela ALL, Cuadrado A, Guerra M (2003) Localization of 45S rDNA and telomeric sites on holocentric chromosomes of Rhynchospora tenuis Link (Cyperaceae). Genet Mol Biol 26:199–201. doi:10.1590/S1415-47572003000200014
Yano O, Katsuyama T, Tsubota H, Hoshino T (2004) Molecular phylogeny of Japanese Eleocharis (Cyperaceae) based on ITS sequence data, and chromosomal evolution. J Plant Res 117:409–419. doi:10.1007/s10265-004-0173-3
Acknowledgments
The authors would like to thank the Brazilian agencies Fundação Araucária and CNPq for financial support. Dr. A. Leyva helped with English editing of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
da Silva, C.R.M., Quintas, C.C. & Vanzela, A.L.L. Distribution of 45S and 5S rDNA sites in 23 species of Eleocharis (Cyperaceae). Genetica 138, 951–957 (2010). https://doi.org/10.1007/s10709-010-9477-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-010-9477-5