Abstract
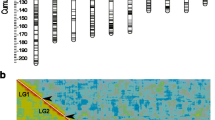
Recombination plays an important role in species adaptation since it acts as an evolutionary force that can influence genome pattern organization. However, recombination can be detrimental in some situations, causing the breakdown of some adaptive gene combinations such as coadapted gene complexes. Genetic and cytological chromosome maps allow recombination throughout the genome to be analyzed. In this study we compare the recombination rate of two types of homokaryotypic lines of D. subobscura (OST and O 3+4 ) using a set of at least 13 microsatellite loci. The genetic maps obtained present similar lengths: 184 and 196 cM for OST and O 3+4 chromosomes, respectively. For most pairs of markers analyzed, a sample size of about 150 individuals appeared sufficient to obtain appropriate recombination values, with the exception of markers located in the same cytological band. Recombination rates seemed to be fairly uniform along the O chromosome, but some regional differences were observed. Several recombination hot and coldspots were detected, and their numbers were different in the homokaryotypic line types (OST and O 3+4 ). This variability could be attributed to differences between the genetic content of the two arrangements or to differences between the lines.




Similar content being viewed by others
References
Araúz PA, Mestres F, Pegueroles C, Arenas C, Tzannidakis G, Krimbas CB, Serra L (2009) Tracking the origin of the American colonization by Drosophila subobscura: genetic comparison between Eastern and Western Mediterranean populations. J Zool Syst Evol Res 47:25–34
Ashburner M (1989) Drosophila, a laboratory handbook. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Betrán E, Rozas J, Navarro A, Barbadilla A (1997) The estimation of the number and the length distribution of gene conversion tracts from population DNA sequence data. Genetics 146:89–99
Carpenter ATC (1988) Thoughts on recombination nodules, meiotic recombination, and chiasmata. In: Kucherlapati R, Smith GR (eds) Genetic recombination. American Society of Microbiology, Washington, pp 529–548
Cirulli ET, Kliman RM, Noor MAF (2007) Fine-scale crossover rate heterogeneity in Drosophila pseudoobscura. J Mol Evol 64:129–135
Clark AG, Eisen MB, Smith DR, Bergman CM, Oliver B et al (2007) Evolution of genes and genomes on the Drosophila phylogeny. Nature 450:203–218
Dobzhansky Th (1950) Genetics of natural populations. XIX. Origin of heterosis through natural selection in populations of Drosophila pseudoobscura. Genetics 35:288–302
Felsenstein J, Yokoyama S (1974) The evolutionary advantage of recombination. Genetics 78:737–756
Ford CF, Hamerton JL (1956) The chromosomes of man. Nature 178:1020–1023
Gloor GB, Preston CR, Johnson-Schlitz DM, Nassif NA, Phillis RW, Benz WK, Robertson HM, Engels WR (1993) Type I repressors of P element mobility. Genetics 135:81–95
Hamblin MT, Aquadro CF (1999) DNA sequence variation and the recombinational landscape in Drosophila pseudoobscura: a study of the second chromosome. Genetics 153:859–869
Kolkman JA, Stemmer WPC (2001) Directed evolution of proteins by exon shuffling. Nat Biotechnol 19:423–428
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Krimbas CB (1993) Drosophila subobscura: biology, genetics and inversion polymorphism. Verlag Dr. Kovac, Hamburg
Krimbas CB, Loukas M (1980) The inversion polymorphism of Drosophila subobscura. Evol Biol 12:163–234
Kulathinal RJ, Bennett SM, Fitzpatrick CL, Noor MAF (2008) Fine-scale mapping of recombination rate in Drosophila refines its correlation to diversity and divergence. Proc Natl Acad Sci USA 105:10051–10056
Kunze-Mühl E, Müller E (1958) Weitere Untersuchungen über die chromosomale Struktur und die natürlichen Strukturtypen von Drosophila subobscura Coll. Chromosoma 9:559–570
Lander ES, Green P, Abrahamsom J, Barlow A, Daly MJ, Lincoln SE, Newburg L (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Lindsley DL, Zimm GG (1992) The genome of Drosophila melanogaster. Academic Press, San Diego
Loukas M, Krimbas CB, Mavragani-Tsipidou P, Kastritsis CD (1979) Genetics of Drosophila subobscura populations. VIII. Allozyme loci and their chromosome maps. J Heredity 70:17–26
Lynch M, Walsh B (1998) Genetics and analysis of quantitative traits. Sinauer Associates Inc., Sunderland
Mestres F, Pegueroles G, Prevosti A, Serra L (1990) Colonization of America by Drosophila subobscura: lethal genes and the problem of the O5 inversion. Evolution 44:1823–1836
Mestres F, Sanz J, Serra L (1998) Chromosomal structure and recombination between inversions in Drosophila subobscura. Hereditas 128:105–113
Munté A, Rozas J, Aguadé M, Segarra C (2005) Chromosomal inversion polymorphism leads to extensive genetic structure: a multilocus survey in Drosophila subobscura. Genetics 169:1573–1581
Nicklas RB (1974) Chromosome segregation mechanisms. Genetics 78:205–213
Ortiz-Barrientos D, Chang AS, Noor MAF (2006) A recombinational portrait of the Drosophila pseudoobscura genome. Genet Res 87:23–31
Otto SP, Barton NH (1997) The evolution of recombination: removing the limits to natural selection. Genetics 147:879–906
Pascual M, Schug MD, Aquadro CF (2000) High density of long dinucleotide microsatellites in Drosophila subobscura. Mol Biol Evol 17:1259–1267
Pascual M, Aquadro C, Soto V, Serra L (2001) Microsatellite variation in colonizing and palearctic populations of Drosophila subobscura. Mol Biol Evol 18:731–740
Patthy L (1999) Genome evolution and exon-shuffling—a review. Gene 238:103–114
Rozen S, Skaletsky HJ (2000) Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics methods and protocols: methods in molecular biology. Humana Press, Totowa, pp 365–386
Ruiz-Martín H (2006) Estudi de les associacions entre al.lels de loci microsatèl.lits i inversions cromosòmiques en poblacions europees i colonitzadores de D. subobscura. Diploma d’Estudis Avançats. Dept. Genètica, Universitat de Barcelona, Spain
Santos J, Serra L, Solé E, Pascual M (2010) FISH mapping of microsatellite loci from Drosophila subobscura and its comparison to related species. Chrom Res 18:213–226
Schultz J, Redfield H (1951) Interchromosomal effects on crossing-over in Drosophila. Cold Spring Harb Symp Quant Biol 16:175–197
Singh ND, Aquadro CF, Clark AG (2009) Estimation of fine-scale recombination intensity variation in the white-echinus interval of D. melanogaster. J Mol Evol 69:42–53
Staten R, Schully SD, Noor MAF (2004) A microsatellite linkage map of Drosophila mojavensis. BMC Genetics 5:12
True JR, Mercer JM, Laurie CC (1996) Differences in crossover frequency and distribution among three sibling species of Drosophila. Genetics 142:507–523
Acknowledgments
We would like to thank Prof. Joan Balanyà (Dept. of Genetics, Universitat de Barcelona) for the analyses of the polytene chromosomes, Prof. Concepció Arenas (Dept. of Statistics Universitat de Barcelona) for her statistical advice and Mr. Robin Rycroft (Language Advisory Service, Universitat de Barcelona) for the English grammar corrections. Furthermore, we would also like to thank Prof. Charles F. Aquadro, Prof. Martha Hamblin (Cornell University), Prof. Luís Serra and Prof. Enrique Blanco (Universitat de Barcelona) for their valuable comments on the manuscript. This work was supported by a pre-doctoral fellowship to CP (2009FIC-00096) from the Generalitat de Catalunya (Spain). The research was funded by projects CGL2006-13423-C02-02 from the Ministerio de Ciencia y Tecnología (MCYT, Spain) and SGR2009-636 from the Generalitat de Catalunya (Spain).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pegueroles, C., Araúz, P.A., Pascual, M. et al. A recombination survey using microsatellites: the O chromosome of Drosophila subobscura . Genetica 138, 795–804 (2010). https://doi.org/10.1007/s10709-010-9461-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-010-9461-0