Abstract
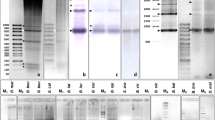
Major satellites of species in the genus Pimelia comprise large portions of their genomes and belong to seven major satellite families which all originate from a common ancestral sequence. Here we present the results of comprehensive screening of 26 Pimelia species belonging to three distinct geographic groups (Ibero-Balearic, African and Canary Islands) for the presence of different Pimelia satellite families in their genomes. Dot-blot hybridization experiments suggest that together with one dominant, highly abundant satellite family, other families are also present in genomes of the majority of examined Pimelia species, but as low-copy number repeats. The estimated abundance of these underrepresented repeats is about 4,000 copies per haploid genome. Signals of highly abundant satellite family from P. scabrosa (PSCA) in examined congeneric species, obtained after PCR amplification and Southern hybridization under high stringency conditions, corroborate sequence preservation of low-copy representatives of satellite families. PRINS localized low-copy repeats within the pericentromeric regions of all chromosomes. These results point to the existence of an extensive library of repetitive DNAs that was already present in the genome of the common ancestor of extant Pimelia taxa, and shifts the period of diversification of Pimelia satellites far in the history of this genus.
Similar content being viewed by others
References
Bruvo B, Pons J, Ugarković Đ, Juan C, Petitpierre E, Plohl M (2003) Evolution of low-copy number and major satellite DNA sequences in two Pimelia species-groups (Coleoptera). Gene 312:85–94
Cesari M, Luchetti A, Passamonti M, Scali V, Mantovani B (2003) Polymerase chain reaction amplification of the Bag320 satellite family reveals ancestral library and past gene conversion events in Bacillus rossius (Insecta, Phasmatodea). Gene 312:289–295
Charlesworth B, Sniegowski P, Stephan W (1994) The evolutionary dynamics of repetitive DNA in eukaryotes. Nature 371:215–220
Dover GA (1986) Molecular drive in multigene families: how biological novelties arise, spread and are assimilated. Trends Genet 168:159–165
Fry K, Salser W (1977) Nucleotide sequences of HS-α satellite DNA from kangaroo rat Dipodomys ordii and characterization of similar sequences in other rodents. Cell 12:1069–1084
Juan C, Gosalvez J, Mezzanotte R, Petitpierre E (1991) Cytological and biochemical characterization of the in situ endonuclease digestion of fixed Tenebrio molitor chromosomes. Chromosoma 100:432–438
Kwieton E (1977) Esquisse phylogenetique du genre Pimelia F. Acta Ent Mus Nat Pragae 39:559–589
Landais I, Chavigny P, Castagnone C, Pizzol J, Abad P, Vanlerberghe-Massuti F (2000) Characterization of a highly conserved satellite DNA from the parasitic wasp Trichogramma brassicae. Gene 255:65–73
Meštrović N, Plohl M, Mravinac B, Ugarković Đ (1998) Evolution of satellite DNAs from the genus Palorus – experimental evidence for the “library” hypothesis. Mol Biol Evol 15:1062–1068
Mravinac B, Plohl M, Meštrović N, Ugarković Đ (2002) Sequence of PRAT satellite DNA “frozen” in some coleopteran species. J Mol Evol 54:774–783
Mravinac B, Plohl M, Ugarkovic Đ (2005) Preservation and high sequence conservation of satellite DNAs suggest functional constraints. J Mol Evol 61:542–550
Pinkel D, Gray J, Trask B, Van Den Engh G, Fuscoe J, Van Dekken H (1986) Cytogenetic analysis by in situ hybridization with fluorescently labeled nucleic acid probes. Cold Spring Harb Symp Quant Biol 51:151–157
Plohl M, Bruvo B, Meštrović N, Mravinac B, Petrović V, Durajlija-Žinić S, Ugarković Đ (2004) Satellite DNA sequences in centromeric heterochromatin. Period Biol 106:95–102
Pons J (1999) Evolucion del DNA satellite en el genero Pimelia. PhD Thesis, University of Balearic Islands
Pons J, Bruvo B, Juan C, Petitpierre E, Plohl M, Ugarković Đ (1997) Conservation of satellite DNA in species of the genus Pimelia (Tenebrionidae, Coleoptera). Gene 205:183–190
Pons J, Petitpierre E, Juan C (2002a) Evolutionary dynamics of satellite DNA family PIM357 in species of the genus Pimelia. Mol Biol Evol 19:1329–1340
Pons J, Petitpierre E, Juan C (2002b) Higher-order organization and compartmentalization of satellite DNA PIM357 in species of the coleopteran genus Pimelia. Chromosome Res 10:597–606
Pons J, Bruvo B, Petitpierre E, Plohl M, Ugarković Đ, Juan C (2004) Complex structural features of satellite DNA sequences in the genus Pimelia (Coleoptera, Tenebrionidae): random differential amplification from a common “satellite DNA library”. Heredity 92:418–427
Steiner JJ, Poklemba CJ, Fjellstrom RG, Elliott LF (1995) A rapid one-tube genomic DNA extraction process for PCR and RAPD analyses. Nucleic Acids Res 23:2569–2570
Ugarković Đ, Plohl M (2002) Variation in satellite DNA profiles – causes and effects. EMBO J 21:5955–5959
Ugarković Đ, Petitpierre E, Juan C, Plohl M (1995) Satellite DNAs in tenebrionid species – structure, organization and evolution. Croat Chem Acta 68:627–638
Acknowledgements
Dr. Joan Pons kindly provided the samples of the species used in this study. We thank anonymous referees for critical reading and constructive suggestions that substantially improved the manuscript. This work was supported by the Research Fund of the Republic of Croatia (projects P0098074 and P0098075).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bruvo-Mađarić, B., Plohl, M. & Ugarković, D. Wide distribution of related satellite DNA families within the genus Pimelia (Tenebrionidae). Genetica 130, 35–42 (2007). https://doi.org/10.1007/s10709-006-0017-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-006-0017-2