Abstract
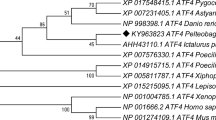
The 14-3-3 proteins are a family of widely expressed acidic proteins, which are involved in the regulation of many biological processes of animals. However, no research regarding 14-3-3 has been described in sturgeon to date, one of the most primitive Actinopterygii species. Here, we identified the first 14-3-3 gene from Siberian sturgeon (Acipenser baeri), named Ab14-3-3β/α (GenBank Accession No. KY094076.1). The cDNA of Ab14-3-3β/α is 1212 bp in length, containing a 5′-untranslated region (UTR) of 82 bp, a 3′UTR of 392 bp, and an open reading frame (ORF) of 738 bp, encoding a polypeptide of 245 amino acids which contains a 14-3-3 homologs domain (PF00244). Phylogenetic analysis showed that the 14-3-3 gene product from Acipenser baeri is a counterpart of vertebrate 14-3-3β/α. The deduced Ab14-3-3β/α protein shares high identities of 46.5–95.5% with the homologs of other species. Ab14-3-3β/α mRNA was constitutively expressed in all examined tissues, with high expression levels in the blood and gill. Furthermore, the expression level of Ab14-3-3β/α mRNA increased significantly in the gill at 1 h under acute salinity shock by transfer of Siberian sturgeons from fresh water (FW) to 15 ppt. In fish subjected to a high temperature (31 °C), Ab14-3-3β/α showed a significant upregulation in the liver at 3 h compared with the control group (24 °C). A 4.85-fold increase of Ab14-3-3β/α expression in the spleen of Siberian sturgeon was observed at 24 h following Aeromonas hydrophila challenge. Collectively, these results indicated that Ab14-3-3β/α might play a certain role in sturgeon in response to some environmental stresses and bacterial challenge.









Similar content being viewed by others
References
Aghazadeh Y, Papadopoulos V (2016) The role of the 14-3-3 protein family in health, disease, and drug development. Drug Discov Today 21:278–287. https://doi.org/10.1016/j.drudis.2015.09.012
Aitken A (2006) 14-3-3 proteins: a historic overview. Semin Cancer Biol 16:162–172. https://doi.org/10.1016/j.semcancer.2006.03.005
Aitken A (2011) Post-translational modification of 14-3-3 isoforms and regulation of cellular function. Semin Cell Dev Biol 22:673–680. https://doi.org/10.1016/j.semcdb.2011.08.003
Andrea K, Devulapalli C, Dietmar K (2003) Teleost Fh14-3-3a protein protects Xenopus oocytes from hyperosmolality. J Exp Zool A Comp Exp Biol 299:103–109. https://doi.org/10.1002/jez.a.10294
Baldin V (2000) 14-3-3 proteins and growth control. Prog Cell Cycle Res 4:49–60. https://doi.org/10.1007/978-1-4615-4253-7_5
Bemis WE, Findeis EK, Grande L (1997) An overview of Acipenseriformes. Environ Biol Fish 48:25–71. https://doi.org/10.1023/a:1007370213924
Besser J, Bagowski CP, Salasvidal E, van Hemert MJ, Bussmann J, Spaink HP (2007) Expression analysis of the family of 14-3-3 proteins in zebrafish development. Gene Expr Patterns 7:511–520. https://doi.org/10.1016/j.modgep.2006.10.007
Campanella JJ, Bitincka L, Smalley J (2003) MatGAT: an application that generates similarity/identity matrices using protein or DNA sequences. BMC Bioinformatics 4:29. https://doi.org/10.1186/1471-2105-4-29
Cheng CH, Guo ZX, Luo SW, Wang AL (2018) Effects of high temperature on biochemical parameters, oxidative stress, DNA damage and apoptosis of pufferfish ( Takifugu obscurus ). Ecotoxicol Environ Saf 150:190–198. https://doi.org/10.1016/j.ecoenv.2017.12.045
Darling DL, Yingling J, Wynshaw-Boris A (2005) Role of 14-3-3 proteins in eukaryotic signaling and development. Curr Top Dev Biol 68:281–315. https://doi.org/10.1016/S0070-2153(05)68010-6
Dougherty MK, Morrison DK (2004) Unlocking the code of 14–3-3. J Cell Sci 117:1875–1884. https://doi.org/10.1242/jcs.01171
Dytham CM (1999) Choosing and using statistics: a biologist’s guide. J Ecol 87:734–735. https://doi.org/10.1046/j.1365-2745.1999.00393-4.x
Finnie C, Borch J, Collinge DB (1999) 14–3-3 proteins: eukaryotic regulatory proteins with many functions. Plant Mol Biol 40:545–554. https://doi.org/10.1023/a:1006211014713
Ford JC, Alkhodairy F, Fotou E, Sheldrick KS, Griffiths DJ, Carr AM (1994) 14-3-3 protein homologs required for the DNA damage checkpoint in fission yeast. Science 265:533–535. https://doi.org/10.1126/science.8036497
Funami K, Matsumoto M, Obuse C, Seya T (2016) 14-3-3-zeta participates in TLR3-mediated TICAM-1 signal-platform formation. Mol Immunol 73:60–68. https://doi.org/10.1016/j.molimm.2016.03.010
Gómezsuárez M et al (2016) 14-3-3 proteins regulate Akt Thr308 phosphorylation in intestinal epithelial cells. Cell Death Differ 23:1060–1072. https://doi.org/10.1038/cdd.2015.163
Graidist P, Phusantisampan T, Wanlem S, Wanna W (2010) Detection of PinX1 and 14-3-3 in the shrimp (Litopenaeus vannamei) and study on gene expressions during viral infection and environmental stresses. Songklanakarin J Sci Technol 32:571–579
Kaeodee M, Pongsomboon S, Tassanakajon A (2011) Expression analysis and response of Penaeus monodon 14-3-3 genes to salinity stress. Comp Biochem Physiol B Biochem Mol Biol 159:244–251. https://doi.org/10.1016/j.cbpb.2011.05.004
Kültz D, Chakravarty D, Adilakshmi T (2001) A novel 14-3-3 gene is osmoregulated in gill epithelium of the euryhaline teleost Fundulus heteroclitus. J Exp Biol 204:2975–2985
Lassalle G, Crouzet P, Gessner J, Rochard E (2010) Global warming impacts and conservation responses for the critically endangered European Atlantic sturgeon. Biol Conserv 143:2441–2452. https://doi.org/10.1016/j.biocon.2010.06.008
Liu D, Bienkowska J, Petosa C, Collier RJ, Fu H, Liddington R (1995) Crystal structure of the zeta isoform of the 14-3-3 protein. Nature 376:191–194. https://doi.org/10.1038/376191a0
Liu N, Wang XW, Sun JJ, Wang L, Zhang HW, Zhao XF, Wang JX (2016) Akirin interacts with Bap60 and 14-3-3 proteins to regulate the expression of antimicrobial peptides in the kuruma shrimp (Marsupenaeus japonicus). Dev Comp Immunol 55:80–89. https://doi.org/10.1016/j.dci.2015.10.015
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Lu Q, Wu S, Zhen H, Deng H, Song Q, Ma K, Cao Z, Pang Q, Zhao B (2017) 14-3-3 α and 14-3-3 ζ contribute to immune responses in planarian Dugesia japonica. Gene 615:25–34. https://doi.org/10.1016/j.gene.2017.03.017
Macdonald N, Welburn JPI, Noble MEM, Nguyen A, Yaffe MB, Clynes D, Moggs JG, Orphanides G, Thomson S, Edmunds JW, Clayton AL, Endicott JA, Mahadevan LC (2005) Molecular basis for the recognition of phosphorylated and phosphoacetylated histone h3 by 14-3-3. Mol Cell 20:199–211. https://doi.org/10.1016/j.molcel.2005.08.032
Masters SC, Fu H (2001) 14-3-3 proteins mediate an essential anti-apoptotic signal. J Biol Chem 276:45193–45200. https://doi.org/10.1074/jbc.M105971200
Mercedes PR (2012) 14-3-3 proteins are regulators of autophagy. Cells 1:754–773. https://doi.org/10.3390/cells1040754
Meyer A, Zardoya R (2003) Recent advances in the (molecular) phylogeny of vertebrates. Annu Rev Ecol Evol Syst 34:311–338. https://doi.org/10.1146/annurev.ecolsys.34.011802.132351
Mhawech P (2005) 14-3-3 proteins—an update. Cell Res 15:228–236. https://doi.org/10.1038/sj.cr.7290291
Moore BW, Perez VJ (1967) Specific acidic proteins of the nervous system. In: Carlson FD (ed) Physiological and biochemical aspects of Neverous integration. Prentice–Hall, Englewood Cliffs, NJ, pp 343–359
Rosner M, Hengstschlager M (2006) 14-3-3 proteins are involved in the regulation of mammalian cell proliferation. Amino Acids 30:105–109. https://doi.org/10.1007/s00726-005-0240-7
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Shandala T, Woodcock JM, Ng Y, Biggs L, Skoulakis EMC, Brooks DA, Lopez AF (2011) Drosophila 14-3-3ε has a crucial role in anti-microbial peptide secretion and innate immunity. J Cell Sci 124:2165–2174. https://doi.org/10.1242/jcs.080598
Shu M, Zhang L, Xu B, Hu H, Guo X (2012) The full length cDNA cloning and expression profile of 14-3-3 gene from the mud crab( Scylla paramamosain). J Fish China 36:1193–1200. https://doi.org/10.3724/SP.J.1231.2012.27996
Steinacker P, Aitken A, Otto M (2011) 14-3-3 proteins in neurodegeneration. Semin Cell Dev Biol 22:696–704. https://doi.org/10.1016/j.semcdb.2011.08.005
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680. https://doi.org/10.1093/nar/22.22.4673
Tzivion G, Avruch J (2002) 14-3-3 proteins: active cofactors in cellular regulation by serine/threonine phosphorylation. J Biol Chem 277:3061–3064. https://doi.org/10.1074/jbc.R100059200
Wang W, Shakes DC (1996) Molecular evolution of the 14-3-3 protein family. J Mol Evol 43:384–398. https://doi.org/10.1007/BF02339012
Wang D, Lu T, Liu H (2009) Characterization and genetic diversity of the sturgeon Acipenser schrenskii Ig heavy chain. Immunobiology 214:359–366. https://doi.org/10.1016/j.imbio.2008.10.006
Wang QF, Shen WL, Hou CC, Liu C, Wu XF, Zhu JQ (2017) Physiological responses and changes in gene expression in the large yellow croaker Larimichthys crocea following exposure to hypoxia. Chemosphere 169:418–427. https://doi.org/10.1016/j.chemosphere.2016.11.099
Wanna W, Rexroad CE, Yao J (2010) Identification of a functional splice variant of 14-3-3E1 in rainbow trout. Mar Biotechnol 12:70–80. https://doi.org/10.1007/s10126-009-9201-6
Xiao B, Smerdon SJ, Jones DH, Dodson GG, Soneji Y, Aitken A, Gamblin SJ (1995) Structure of a 14-3-3 protein and implications for coordination of multiple signalling pathways. Nature 376:188–191. https://doi.org/10.1038/376188a0
Zhang X et al (2014) Molecular cloning and expression of 14-3-3ζ in Haliotis diversicolor under stresses. J Fish China 38:492–502. https://doi.org/10.3724/SP.J.1231.2014.49122
Zhu H, Song R, Wang X, Hu H, Zhang Z (2017) Peritoneal bacterial infection repressed the expression of IL17D in Siberia sturgeon a chondrostean fish in the early immune response. Fish Shellfish Immunol 64:39–48. https://doi.org/10.1016/j.fsi.2017.03.011
Funding
This study was funded by Beijing Municipal Excellent Talents Foundation (2017000020060G122), Foundation of Beijing Municipal Science and Technology Project (Z161100004516003), Natural Science Foundation of China (31602137), and Beijing Sturgeon and Trout Innovation Team (BAIC08-2019).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The institutional Animal Care and Use Committee at the Beijing Fisheries Research Institute approved this study. The experimental procedures were performed with strict accordance with the standards of the Chinese Council on Animal Care.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 322 kb)
Rights and permissions
About this article
Cite this article
Wang, X., Ma, G. & Zhu, H. Regulation of 14-3-3β/α gene expression in response to salinity, thermal, and bacterial stresses in Siberian sturgeon (Acipenser baeri). Fish Physiol Biochem 46, 519–531 (2020). https://doi.org/10.1007/s10695-019-00702-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10695-019-00702-w