Abstract
Pathogenic mutations in CYLD can be identified in patients affected with Brooke-Spiegler syndrome, (Familial) Cylindromatosis or multiple familial trichoepithelioma. To date, only technologies which are able to identify small point mutations in CYLD, such as sequence and WAVE analysis, were used. Here we describe the identification of a larger rearrangement identified by Quantitative PCR analysis of CYLD, indicating that a combination of these technologies is necessary when searching for pathogenic mutations in CYLD.
Similar content being viewed by others
Introduction
Brooke-Spiegler syndrome (BSS; OMIM #60541) is an autosomal dominant disorder characterised by cylindromas, trichoepitheliomas and spiradenomas, especially in the head and neck region. Due to the clinical overlap with (familial) Cylindromatosis (FC; OMIM #132700) and multiple Familial Trichoepithelioma (MFT; OMIM #601606), these disorders are considered to represent a single disease entity [1–3].
Linkage analysis was used to define the familial cylindromatosis region, resulting in the identification of CYLD [4–6]. Germline mutations in CYLD have been identified in BSS, FC and MFT patients, in agreement with the clinical findings indicating that these disorders have a common genetic basis [6–11]. CYLD is encoded by 20 exons of which exon 4 contains the ATG start codon [6]. The CYLD protein is a deubiquitinating enzyme regulating cell signalling via several pathways, including the NF-κB and JNK pathways [12–14].
Recently, a nice overview was published by Blake and Toro where they described 51 distinct germline mutations in CYLD in 73 families with BSS, FC and MFT [7]. All types of mutations were identified: frameshift (41%), nonsense (35%), missense (14%) and putative splice site (10%). No large rearrangements were identified, probably because all techniques used were not suitable for identifying this type of mutation. Loss of heterozygosity is shown in cylindromas and trichoepitheliomas, indicating that CYLD acts as a tumour suppressor gene [6, 8, 15–17]. This observation is in agreement with the function of the CYLD protein in regulating several pathways in cell signalling. In most tumour suppressor genes, e.g. BRCA1, NF1, TSC2, large rearrangements are identified as pathogenic germline mutations [18–20]. Therefore we hypothesized that larger rearrangements in CYLD could be identified in patients with BSS, FC and MFT.
Here we describe the identification of a large rearrangement in CYLD in a patient with FC using Quantitative (Q)-PCR, indicating that a quantitative test should be performed on DNA samples of patients if no mutation was identified by techniques which can only identify small sequence changes, such as sequence or WAVE analysis.
Materials and methods
Patient samples
In our diagnostic setting we received 13 samples from patients with Brooke-Spiegler syndrome (BSS, n = 2), (Familial) Cylindromatosis [(F)C; n = 7] or Trichoepithelioma (T; n = 2). Of the remaining two patients the indication for testing CYLD was not provided by the applicant (Table 1). Clinical details were obtained only from the patient described in this paper.
Mutation analysis
Extraction of DNA from peripheral blood cells was performed according to standard techniques. Mutation analysis of the coding exons 4–20 and exon/intron boundaries of CYLD was performed by sequence analysis (primers available on request) using an automated sequencer (ABI 3730XL, Applied Biosystems, Foster City, CA, USA). Data were analysed using SeqScape software (version 2.6; Applied Biosystems). If a sequence change could not be classified as a pathogenic mutation, the sequence change was analysed using Alamut software (Mutation Interpretation Software; version 1.5 May 2009; Interactive Biosoftware, Rouen, France).
In case no pathogenic mutation or an unclassified variant was identified, Quantitative (Q)-PCR analysis was performed to search for large rearrangements.
Quantitative-PCR, Long-range-PCR and sequence analysis of breakpoints
Real-time Q-PCR was performed using Fam-labelled Taqman assays. Oligonucleotides were designed for all (non) coding exons. If the CG content was too high (exon 1) or too low (exons 14–17), a Taqman assay was designed using oligonucleotides in the promoter region (instead of exon 1) and in the intronic regions between exons 14 and 17 as close as possible to the exons (Table 2). Oligonucleotides were designed with Primer Express 2.0.0 (Applied Biosystems). Primer specificity was checked by performing BLAST analysis. Taqman probes were synthesised with a melting temperature (Tm) 8–10°C higher than the primers by incorporating Locked Nucleic Acid (LNA) monomers in the probe. Tm values for the LNA probes were calculated using the Exiqon website (http://lna-tm.com/). The LNA-based Taqman assays were manufactured by Eurogentec (Maastricht, The Netherlands). Since LNA probes show a high thermal stability and are resistant to exo- and endonuclease activity [21], we prefer the use of these probes in the Taqman assay.
Gene dosage alterations were detected on an ABI7500 Real time PCR system (Applied Biosystems) by performing a relative quantification run. Real time PCR reactions were performed in a total volume of 25 μl, containing 20 ng DNA, 1 × qPCR mastermix Plus–low ROX (Eurogentec: RT-QP2x-03-WOULR), 1 × RNAse P (endogenous control; Applied Biosystems), 30 μM forward and reverse primers and 10 μM probe. PCR conditions were as follows: an initial 2 min incubation at 50°C, followed by 95°C for 10 min and then 40 cycles of 95°C for 15 s and 60°C for 1 min. All samples were analysed in triplicate and compared to a normal control sample.
LR-PCR was performed with the Expand Long Template PCR System (Roche Applied Science, Indianapolis, IN, USA). The product obtained by LR-PCR was sequenced using an automated sequencer (ABI 3730XL). Nomenclature of the deletion was according to the recommendations of the Human Genome Variation Society, using reference sequence NM_015247_2. After characterisation of the breakpoints in patient 37999, a deletion-specific PCR analysis was designed. Three primers (Table 2) were used: a common primer, a primer specific for the deletion and a primer specific for the wildtype allele. The primers were designed in such a way that the PCR product from the allele carrying the deletion was shorter than the product from the wildtype allele. The PCR conditions were: an initial 2 min incubation at 94°C, followed by 30 cycles of 94°C for 60 s, 60°C for 30 s and 72°C for 90 s and completed by 72°C for 10 min.
Results
Mutation analysis
Using sequence analysis of all coding exons and exon intron boundaries of CYLD, a pathogenic mutation was identified in 6 out of 13 patients of our diagnostic cohort (Table 1). Of these mutations, c.1112C>A (p.Ser371X) and c.2272C>T (p.Arg758X) were previously described. The c.2272C>T (p.Arg758X) mutation was found in two families with different haplotypes, indicating that this mutation is a recurrent mutation in CYLD [6]. In three patients, an unclassified variant was identified: c.2146C>A (p.Gln716K), c.2350+5G>A and c.2662_2664delTTT. One of these variants, c.2350+5G>A was previously described, but no formal proof that this sequence change is a pathogenic mutation was provided [6]. Three splice site prediction programs of Alamut (SpliceSiteFinder-like, MaxEntScan and NNSPLICE) showed that it is very likely that the sequence change c.2350+5G>A will have an effect on RNA splicing (data not shown). The fourth program, GeneSplicer, did not recognise the wildtype donor site and therefore no conclusion could be drawn from this program. Since we did not receive relevant family members to analyse the presence/absence of c.2350+5G>A, or a new sample of the index patient to perform RNA analysis, we were not able to prove that c.2350+5G>A is a pathogenic splice site mutation. No DNA of relevant relatives of the patients with the unclassified variants c.2146C>A (p.Gln716K) and c.2662_2664delTTT was received to obtain further information with respect to the pathogenic character of these sequence changes.
No pathogenic mutation or unclassified variant was identified in four patients.
Identification and characterisation of the large rearrangement
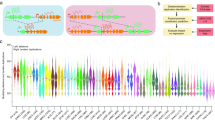
To identify larger rearrangements, a quantitative technique must be applied. We developed Q-PCR analyses for all exons/introns of CYLD and DNA of all patients without a pathogenic mutation, including the patients carrying an unclassified variant, was analysed. Q-PCR analysis of DNA of patient 37999 showed a pattern in agreement with a deletion of exon 20 (Fig. 1). No abnormal patterns were identified in DNA of the other 6 patients, indicating that no large rearrangements are present in these patients (data of the other exons: not shown).
Q-PCR result of exon 20 of patients with no pathogenic mutation or with an unclassified variant after sequence analysis of CYLD. Lane 1 no DNA; lane 2 negative control used as calibrator; lane 3 patient 28597; lane 4 patient 30945; lane 5 patient 32565; lane 6 patient 35680; lane 7 patient 37999; lane 8 patient 41440 and lane 9 patient 11468. Bars represent Relative Quantification (RQ) calculated by 2− ΔΔCT. On top of the bars, the standard error of the mean RQ value is displayed. Inside the bars the calculated RQ value is given of a triplicate measurement
To characterise the breakpoints, LR-PCR was performed and the PCR product was sequenced. The deletion started 60 nucleotides after exon 19 and extended 3340 nucleotides after the translation stopcodon of CYLD. In total 5362 nucleotides were deleted (c.2686+60_*3340del5362; Fig. 2a and 2b). No (direct) repeat structures were present in the vicinity of the breakpoint which could explain the nature of the deletion. A deletion-specific PCR was developed, showing not only the wildtype allele of 913 basepairs, but also the mutant specific fragment of 462 basepairs (Fig. 2c). This analysis confirmed the presence of the pathogenic mutation in two independent DNA samples of the patient and can be used for further family analysis.
Characterization of the breakpoint (c.2686+60_*3340del5362) of patient 37999. a Result of sequence analysis of the long-range PCR product. The arrow indicates the last nucleotide of intron 19 that is still present. b Schematic overview of the deletion present in patient 37999. Exon 19 and the coding part of exon 20 of CYLD are represented by open boxes and the noncoding part of exon 20 by a black box. The intergenic region is given by dot with broken line c Agarose gel electrophoresis of the deletion-specific PCR product. Lane 1 and 2, two independent DNA samples of patient 37999; lane 3, negative control DNA and lane 4, no DNA. The wildtype fragment is 913 bp in length and the mutant fragment 462 bp
Clinical details of patient 37999
The patient, a 47-year old female, was referred because of the presence of multiple lenticular, smooth papules on the forehead and the scalp (Fig. 3). The lesions were reported to be present for 10–15 years. Similar lesions were confirmed to be present in her sister, and were reported by the patient to be present in her father, grandfather and her son. No other clinical features were present.
Discussion
Our diagnostic cohort of patients with a pathogenic mutation in CYLD comprised the clinical phenotypes resembling Brooke-Spiegler syndrome (BSS), (Familial) Cylindromatosis (FC) and (Multiple Familial) Trichoepithelioma (MFT). One BSS patient was heterozygous for the unclassified variant c.2350+5G>A. This sequence change was previously described as being a pathogenic mutation, but there was no formal proof for this conclusion [6]. In silico analysis showed that the sequence change c.2350+5G>A might have an effect on RNA splicing and therefore might be a pathogenic mutation. No pathogenic mutation was identified in the other BSS patient. Of the seven (F)C patients, five carried a pathogenic mutation, whereas the two other patients were carrier of an unclassified variant. In one of two MFT patients a pathogenic mutation was identified. In one of the two patients without information with respect to the indication for testing, a pathogenic mutation could be identified. Using a combination of sequence and Q-PCR analyses, 7 pathogenic and 3 unclassified variants in 13 patients (77%) were identified, comparable with the overall detection rate of 83% previously described [7]. All missense mutations described to date were located within the USP (Ubiquitin Specific Protease) domain (amino acids 583–956) [7, 22]. Therefore it is very likely that the unclassified variants c.2146C>A (p.Gln716K) and c.2662_2664delTTT (p.Phe888del) identified in our cohort, were also pathogenic mutations. In that case, a pathogenic mutation was identified in all our (F)C patients. This was comparable with the findings of Saggar et al., who identified pathogenic mutations in 100% (3/3) and 44% (4/9) of their FC and MFT patients, respectively, [9]. In agreement with all previously identified sequence change in CYLD [7], the localisation of the abnormalities described in this paper were in exons 9–20. In patient 37999 a large genomic rearrangement was identified using a quantitative analysis. This deletion (c.2686+60_*3340del5362) encompassed almost complete intron 19 and extended in the 3′ UTR of CYLD. If the allele carrying the deletion will lead to a stable mRNA, the mutant RNA will encode a protein lacking amino acids 896–956, which are part of the USP domain (aa 583–956). The mutant CYLD protein will lack a part of its functional domain and this explains the clinical phenotype of the patient. The frequency of this type of rearrangments in CYLD in our cohort is about 10% and is comparable with the frequencies observed in other genetic disorders such as Neurofibromatosis type 1 and Tuberous sclerosis complex [18, 19].
Since there is intra- en interfamilial variability in the clinical expression, it will be very hard to get a genotype-phenotype correlation. Our patient with a large rearrangement in CYLD was not severely affected. This might be due to the fact that the large deletion only encompassed CYLD and no additional gene(s). It had been suggested that there is a higher incidence of BSS, (F)C or MFT in females [23], most likely as a result of reduced penetrance in males [24]. In our cohort, 12 out of 13 patients were females (Table 1), in agreement with this hypothesis.
To our knowledge, this is the first large germline deletion identified in CYLD, indicating that screening for this type of mutations in a diagnostic setting is recommended in patients with Brooke-Spiegler syndrome, (familial) Cylindromatosis or multiple Familial Trichoepithelioma.
References
Welch JP, Wells RS, Kerr CB (1968) Ancell-Spiegler cylindromas (turban tumours), Brooke-Fordyce Trichoepitheliomas: evidence for a single genetic entity. J Med Genet 5(1):29–35
Young AL, Kellermayer R, Szigeti R, Teszas A, Azmi S, Celebi JT (2006) CYLD mutations underlie Brooke-Spiegler, familial cylindromatosis, multiple familial trichoepithelioma syndromes. Clin Genet 70(3):246–249
Oranje AP, Halley D, den Hollander JC et al (2008) Multiple familial trichoepithelioma, familial cylindroma: one cause!. J Eur Acad Dermatol Venereol 22(11):1395–1396
Biggs PJ, Wooster R, Ford D et al (1995) Familial cylindromatosis (turban tumour syndrome) gene localised to chromosome 16q12–q13: evidence for its role as a tumour suppressor gene. Nat Genet 11(4):441–443
Takahashi M, Rapley E, Biggs PJ et al (2000) Linkage and LOH studies in 19 cylindromatosis families show no evidence of genetic heterogeneity, refine the CYLD locus on chromosome 16q12–q13. Hum Genet 106(1):58–65
Bignell GR, Warren W, Seal S et al (2000) Identification of the familial cylindromatosis tumour-suppressor gene. Nat Genet 25(2):160–165
Blake PW, Toro JR (2009) Update of cylindromatosis gene (CYLD) mutations in Brooke-Spiegler syndrome: novel insights into the role of deubiquitination in cell signaling. Hum Mutat 30(7):1025–1036
Kazakov DV, Thoma-Uszynski S, Vanecek T, Kacerovska D, Grossmann P, Michal M (2009) A case of Brooke-Spiegler syndrome with a novel germline deep intronic mutation in the CYLD gene leading to intronic exonization, diverse somatic mutations, unusual histology. Am J Dermatopathol 31(7):664–673
Saggar S, Chernoff KA, Lodha S et al (2008) CYLD mutations in familial skin appendage tumours. J Med Genet 45(5):298–302
Zhang XJ, Liang YH, He PP et al (2004) Identification of the cylindromatosis tumor-suppressor gene responsible for multiple familial trichoepithelioma. J Invest Dermatol 122(3):658–664
Zhang G, Huang Y, Yan K et al (2006) Diverse phenotype of Brooke-Spiegler syndrome associated with a nonsense mutation in the CYLD tumor suppressor gene. Exp Dermatol 15(12):966–970
Brummelkamp TR, Nijman SM, Dirac AM, Bernards R (2003) Loss of the cylindromatosis tumour suppressor inhibits apoptosis by activating NF-kappaB. Nature 424(6950):797–801
Trompouki E, Hatzivassiliou E, Tsichritzis T, Farmer H, Ashworth A, Mosialos G (2003) CYLD is a deubiquitinating enzyme that negatively regulates NF-kappaB activation by TNFR family members. Nature 424(6950):793–796
Xue L, Igaki T, Kuranaga E, Kanda H, Miura M, Xu T (2007) Tumor suppressor CYLD regulates JNK-induced cell death in Drosophila. Dev Cell 13(3):446–454
Biggs PJ, Chapman P, Lakhani SR, Burn J, Stratton MR (1996) The cylindromatosis gene (cyld1) on chromosome 16q may be the only tumour suppressor gene involved in the development of cylindromas. Oncogene 12(6):1375–1377
Thomson SA, Rasmussen SA, Zhang J, Wallace MR (1999) A new hereditary cylindromatosis family associated with CYLD1 on chromosome 16. Hum Genet 105(1–2):171–173
Leonard N, Chaggar R, Jones C, Takahashi M, Nikitopoulou A, Lakhani SR (2001) Loss of heterozygosity at cylindromatosis gene locus, CYLD, in sporadic skin adnexal tumours. J Clin Pathol 54(9):689–692
De Luca A, Bottillo I, Dasdia MC et al (2007) Deletions of NF1 gene, exons detected by multiplex ligation-dependent probe amplification. J Med Genet 44(12):800–808
Kozlowski P, Roberts P, Dabora S et al (2007) Identification of 54 large deletions/duplications in TSC1, TSC2 using MLPA, genotype-phenotype correlations. Hum Genet 121(3–4):389–400
van den Ouweland AM, Dinjens WN, Dorssers LC et al (2009) Deletion of exons 1a–2 of BRCA1: a rather frequent pathogenic abnormality. Genet Test Mol Biomarkers 13(3):399–406
Ugozzoli LA, Latorra D, Puckett R, Arar K, Hamby K (2004) Real-time genotyping with oligonucleotide probes containing locked nucleic acids. Anal Biochem 324(1):143–152
Komander D, Lord CJ, Scheel H et al (2008) The structure of the CYLD USP domain explains its specificity for Lys63-linked polyubiquitin and reveals a B box module. Mol cell 29(4):451–464
van Balkom ID, Hennekam RC (1994) Dermal eccrine cylindromatosis. J Med Genet 31(4):321–324
Anderson DE, Howell JB (1976) Epitheliomaadenoides cysticum: genetic update. Br J Dermatol 95(3):225–232
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
van den Ouweland, A.M.W., Elfferich, P., Lamping, R. et al. Identification of a large rearrangement in CYLD as a cause of familial cylindromatosis. Familial Cancer 10, 127–132 (2011). https://doi.org/10.1007/s10689-010-9393-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10689-010-9393-y