Abstract
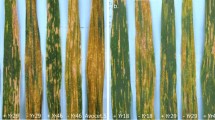
Crop improvement by means of traditional or molecular breeding is a key strategy to accomplish the European Green Deal target of reducing pesticides by 50% by 2030. Regarding viticulture, this is exacerbated by the massive use of chemicals to control pathogen infections. Black rot is an emergent disease caused by the ascomycete Phyllosticta ampelicida, and its destructiveness is alarming vine growers. Implementing and improving effective phenotyping strategies are fundamental preliminary steps to breed disease resistant varieties and this work suggests good practices adopted for this purpose. Primarily, the pedigree of black rot resistance donors was reconstructed based on the collection of phenotypic historical data, highlighting unexplored sources of black rot resistance. Strains used for artificial infections were isolated, genetically characterized and mixed to avoid race-specific resistance selection. A new inoculation protocol based on the use of leaf mature lesions was developed. Ex vivo inoculation on detached leaves was effective for the evaluation of conidia germination and hyphal growth, but not for disease progression. Finally, the pedigree was used for the identification of 23 genotypes to be tested. Two breeding selections (NY39 and NY24) resulted symptomless in all assessments and a third one (F25P52) also showed very high resistance, although with a greater variability. Other two genotypes (F12P19 and ‘Charvir’) fell within the medium resistance category, making them good candidates in a regime of well-timed preventive treatments. In conclusion, this work was effective to a comprehensive parental line characterization and preparatory towards grapevine breeding programs for black rot resistance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Crop improvement has always been an intrinsic activity in the history of agriculture. Through natural selection, traditional breeding or with the most recent molecular techniques, the main targets have always been augmented productivity, abiotic stress resilience and disease resistance. Susceptibility to biotic stresses is an urgent problem that afflicts all cultures and causes significant economic losses in terms of both yield and pecuniary investments in plant protection strategies, the latter also having an additive cost in terms of human health and environmental impact. For this reason, the European Green Deal, the ambitious policy initiative of the European Union to achieve carbon neutrality by 2050, allocated 30% of the budget for research and innovation activities in the fields of agriculture, forestry, and rural areas towards organic sector related issues such as genetic biodiversity with the goal of 50% pesticides reduction by 2030 (European Commission 2020). With this perspective, the exploitation of resistance traits is a promising complementary strategy to control plant disease. Unfortunately, these traits are more commonly controlled by a complex interplay of different biochemical mechanisms. Consequently, the resistance behavior is expressed in a quantitative manner as well as strongly affected by the interaction with the environment. The identification of the genomic region (locus) associated with a phenotypic trait is called quantitative trait locus (QTL) analysis, and feasibility of this kind of studies depends on a fundamental prerequisite: the high quality of both genetic and phenotypic data. While the advances in molecular biology and sequencing technology enable today a deep characterization of the genome, adding multiple layers of complexity, the bottleneck is now represented by the development of optimized and standardized approaches for the acquisition of robust phenotypic data, made difficult by the high dimensionality of the phenome, namely the full set of phenotypes of an individual (Houle et al. 2010).
While for herbaceous plants as well as for well-known diseases the technological improvements are more advanced (high throughput) at the expense of low resolution, the applied research for woody plants or emergent pathogens is more focused on low-throughput and high-resolution approaches. The grapevine is one of the most important fruit crops cultivated worldwide, and many agricultural communities depend on its production. However, the Eurasian species Vitis vinifera L. is both the most widespread and the most sensitive to several fungal diseases. Besides the more common mildews – powdery (PM) and downy (DM), caused respectively by Erysiphe necator and Plasmopara viticola – newly emerging diseases are threatening viticulture, as in the case of black rot (BR). The causal agent of BR is the ascomycete Phyllosticta ampelicida (Engelm.) Van der Aa (syn. Guignardia bidwellii, Rossman et al. 2015) and was introduced in Europe already in 1885 from the USA (Ramsdell and Milholland 1988). This fungus was indirectly controlled by the massive treatments to protect the vineyards from mildews imported from the USA some years earlier (Pirrello et al. 2019). Toward the end of the last century, the increasing interest for a sustainable viticulture encouraged many winegrowers to reduce fungicide treatments and to introduce mildew-tolerant varieties, known as “PIWI”, a German abbreviation for pilzwiderstandsfähig/ fungus-resistant (https://piwi-international.de/en/). Worldwide the hectarage of PIWI varieties of the main wine-growing countries accounts for 0.6% in France and Italy, 2.5% in Germany, 1.8% in the USA, about 10% in Romania and Hungary, almost 20% in Moldova, Ukraine and Russia and up to 67% in Brazil (Sartori 2016). However, most of those mildew-tolerant varieties are susceptible to BR. This situation, together with the advent of more favorable condition of higher temperature in humid continental areas, fostered a new spread of the disease through France and Switzerland (Jermini and Gessler 1996), Romania, Hungary, Bulgaria, Austria, Italy, and Spain (Roznik 2018), and in Germany where severe outbreaks of BR have occurred in the Moselle valley and other wine-growing regions since 2002 (Harms et al. 2005). Because of this situation, the breeders are now facing the challenge of introducing BR resistance in their programs to produce new resistant varieties, possibly combining it with mildews resistance (R)-loci.
Phyllosticta ampelicida is a non-obligate fungus that can be grown on a cultural medium and produces spores after approximately two weeks. After inoculation by water suspension spray on the leaves, the hemi-biotrophic pathogen penetrates the cuticle and starts to grow subcutaneously creating a 2D-hyphal net and acquiring nutrients from the host tissue indirectly (Ullrich et al. 2009). After eight to ten days, it switches to necrotrophy, and immature lesions appear on leaves as discolored spots. With the disease progression, lesions dry up, turn brown with darker-reddish borders, and then develop black dots growths, that are the fruiting bodies (pycnidia) containing the new asexual spores (conidia) (Pirrello et al. 2019). Unfortunately, when population studies or germplasm screening are done, the production of spores from cultured mycelium is exceptionally space and time consuming, since one plate per plant is needed.
The aim of this work was to optimize propagation and inoculation protocols for future high-throughput screening, as well as efficiently evaluate the BR resistance of a subset of parental lines and selections of the breeding program for biotic stresses at Edmund Mach Foundation (FEM, Italy).
Materials and methods
Genetic material and pedigree reconstruction
A panel of 23 genotypes were selected within the plant material available at the FEM breeding program as putative BR resistance donors and grown in the greenhouse as potted grafted plants on Kober 5BB rootstock (Table S1). The plant material, that already carried mildew resistance traits, was divided into two classes: (i) parental lines, comprehending commercially available varieties and two wild genotypes from FEM collection, and (ii) breeding selections of FEM grapevine breeding program for biotic stresses. The genotypes were selected based on their relationship with BR resistant donors, after a deep study of available online resources (Table S2) – scientific literature and reports – about BR resistance evaluation and pedigree information, the last taken principally from Vitis International Variety Catalog (VIVC, Maul and Töpfer 2022). To focus the dataset on BR resistance, the pedigree of all susceptible V. vinifera varieties was excluded and the prefix ‘VV_’ was added, except for two Georgian V. vinifera accessions with documented BR resistance (Töpfer and Maul 2017). Resistance level records were converted (Table S3) to the rating proposed to be included in the OIV descriptors (OIV 2009) by Rex et al. (2014), based on the commonly recognized five-step ordinal scale: (1) very low, (3) low, (5) medium, (7) high, (9) very high resistant. The overall median for each genotype was then calculated, together with the minimum, the maximum, the standard deviation, and also the median for each organ (bunch or leaf) and for the assesments made in USA or Europe regions. All the collected information was then employed for the pedigree reconstruction of the historical origin of the BR resistance trait and visualized through the software Helium (Shaw et al. 2014). Making use of the ‘size based on pedigree contribution’ tool implemented in Helium settings, the species that are more recurrent within the BR resistance gene pool were highlighted. The software Pedimap (Voorrips et al. 2012) was then used to extract the subpopulation of multiple genotypes through the tool ‘select relatives’, to have a snapshot of specific subgroups.
Pathogen isolation and characterization
Black rot outbreaks were documented in two experimental vineyards (Telve and Tenna) of the Valsugana valley in the Trentino region (Italy), where the high humidity caused by the nearby Caldonazzo lake favored BR infection, respectively on two grapevine varieties, ‘Prior’ and ‘Civit6’. Infected leaves and bunches were collected at the end of the season 2018, but P. ampelicida isolation was effective only from mummified berries (mummies), because on leaves many other fungi were concomitantly colonizing the tissue. The procedure consists of the sterilization of small fragments of BR lesions (ca. 5×5 mm) with 1% NaOH for one minute, then rinsed three times in sterile water for five minutes and, after air drying, placed on ½ PDA (potato dextrose agar) Petri dishes (Ø 92 mm) with chloramphenicol (0.1 g/l). Monoconidic cultures of the samples from the two regions (named TN1 and TN2) together with three isolates (18.1, 80.88 and 10.133) collected in Germany and provided by Julius Kühn Institute (JKI)–Institute for Grapevine Breeding Geilweilerhof (Siebeldingen, Germany) were transferred and grown on new ½ PDA plates covered with sterilized micropore cellophane film (Disco Cell) to facilitate mycelium collection with a spatula in 2 ml Eppendorf tubes. The mycelium was stored at -80 °C, lyophilized until completely dried and ground with the addition of a steel bead using the TissueLyzer (QIAGEN, Hilden, Germany). Genomic DNA was extracted with Nucleospin Plant II kit (Macherey-Nagel, Oensingen, Switzerland). Internal transcribed spacer 1 (ITS1) region of the two samples isolated in Italy was amplified with ITS5 and ITS4 primers (White et al. 1990), sequenced and blasted (GenBank KF851314.1, KF851310.1, KF851306.1, KF851304.1, KF851296.1, KF851295.1, KF851294.1, KF851293.1, KF851292.1, KF851291.1, KF015265.1) to confirm the isolation of pure cultures of P. ampelicida.
The five cultures were genetically characterized following the guidelines published by Narduzzi-Wicht et al. (2014) based on 11 microsatellite markers. The protocol was slightly modified, with regards to the usage of different fluorochromes 5’ labels (Table S4). Dual colony assays were carried to support genetic data, as suggested by Roznik (2019): briefly, strains TN1, TN2 and 18.1 were plated together to evaluate if they can grow without forming any barrier or if the genetic diversity inhibits the growth at the interface.
Inoculum propagation
The maintenance of P. ampelicida strains was carried on organic oatmeal agar (OMA) medium according to Rex et al. (2014), since there was evidence that this fungus grows faster on this substrate compared to ½ PDA (Hausmann L. personal communication). The protocol was followed with slight modification i.e., the medium was supplemented with 0.1 g/l of chloramphenicol, and no blacklight was used. Briefly, the studied strains were incubated in a growing chamber under continuous light at 24 °C and refreshed every 14 days by transferring a distal piece of mycelium (ca. 5×5 mm) on a fresh plate. After 14 days of growth, the mycelium was collected to produce inoculum for the resistance tests. Ten ml of sterile distilled water was added to the plate and the lightly hydrophobic mycelium was gently rubbed with a spatula to facilitate direct contact with the water. The condition of 100% humidity for 10 min leads to the release of the spores. The suspension was filtered, the concentration was defined using a BRAND® counting chamber BLAUBRAND® Thoma pattern (Merck, Sigma-Aldrich, Burlington, MA, United States) and then the concentration adjusted to 104 conidia/ml. This primary inoculum derived from the mycelium grown on plates was then propagated in planta as follows. In a greenhouse chamber, the inoculum was sprayed until saturation of the adaxial leaf face (dripping) of leaves of young growing shoots of potted grafted cuttings of the susceptible V. vinifera variety ‘Teroldego’, previously treated to prevent PM infection (CIDELY, Syngenta, Basel, Switzerland), and insects (EPIK SL, Sipcam Italia S.p.A., Pero, Italy) and thrip infestation (Neemilk, Serbios, Badia Polesine, Italy). Inoculated plants were kept overnight at 100% relative humidity (RH) through the live-controlled humidifier of the chamber avoiding direct UV irradiation, and then maintained at 50% RH at 24 °C, with an artificial light cycle of 12 hours. Approximately 14 days post inoculation (dpi) the leaves started presenting the typical mature brown BR lesions. Pycnidia development was induced with 24 hours 100% RH treatment. During the fall, the development of the disease is slower, taking approximately a week longer for lesion onset.
To optimize artificial infection for large-scale experiments, a new protocol was developed to avoid the necessity to produce one plate per plant, by using fresh symptomatic leaves (14 dpi) directly as inoculum source by soaking them in distilled water, in continuous shaking with a stirring bar. To estimate the conidia release curve over time and therefore determine the optimal leaf soaking time, simple linear regression ‘lm()’ function from Rstudio environment (RStudio Team, 2022) – which use as test static the t value from a two-sided t test – was applied to the dataset of three independent experiments, which were conducted as follow: after placing leaf tissue with mature lesions in 15 ml of distilled water, the conidia concentration was measured as previously described at 5, 10, 15, 30, 60 and 120 min. Since the concentrations of conidia strictly depend on the number of mature pycnidia present on the infected leaves used for the experiment, the data were normalized by subtracting the mean of the experiment at each time point. After dipping the leaves, the suspension was filtered, adjusted to a concentration of 104 conidia/ml, and readily used for its reiterative propagation in planta on V. vinifera or for inoculation experiments. Since the project was oriented to breeding purposes, the strains producing active inoculum were combined into a mix for resistance assessment experiments, to avoid the detection of race-specific responses.
In vivo inoculation experiments and macroscopic inspection
For each genotype, three biological replicates (potted plants) per experiment were tested. Ten inoculations were carried out during 2020 and 2021 to obtain three evaluations per each genotype. For the artificial infection, the same protocol as described for inoculum propagation in planta was followed. Resistance evaluation was performed through visual inspection of the first five inoculated leaves from apex per each biological replicate, following Rex et al. (2014). To ensure the homogeneity of the data within and among experiments, the V. vinifera susceptible variety ‘Teroldego’ was included as a reference in each inoculation experiment for data normalization.
Ex vivo inoculation assays and microscopic symptom evaluation
Sterile Linfaboxes (Micropoli, Cesano Boscone, Italy) were prepared with 0.1% water agar medium (0.1 g/l chloramphenicol) covered with sterilized filter paper. The third to fifth growing leaves of the genotypes of interest (‘Teroldego’ = S, ‘Merzling’ = MR, ‘Seyval’ = HR) were collected with the full petiole and sterilized in 1% NaOH for 1 min, then rinse for three times in sterile distilled water for five minutes and left on sterile filter paper under a vertical flux biological hood until dried. Leaves were subsequently placed in the boxes adaxial face-up by cutting the filter paper to introduce the petiole in the medium. After preparing nine leaves per genotype (3×3 time points), 4 to 9 drops of 10 µl suspension (104 conidia/ml) depending on the leaf size were placed on one half of the leaves in proximity of the main veins; on the second half the same number of sterile distilled water drops were placed as mock inoculation. The boxes were left overnight in the dark, then the drops dried with sterile filter paper and samples incubated in a growing chamber with continuous light at 24 °C for 14 days. The inoculated spots were made traceable by a mark on the lid of the plates. Microscopic evaluation was carried out at 1, 3 and 7 dpi on three inoculated fragments per leaf after clearing with KOH for one hour at 62 °C, rinsed with sterile distilled water three times for five minutes, stained with aniline blue (0.1% in glycine buffer pH 9.4) (Vanacker et al. 2000) and viewed under bright field and fluorescent light (argon laser excitation at 405 nm) through a confocal microscope (Leica DM 6000 CS) equipped with a CCD camera (DFC310 FX, Leica).
Results
The pedigree reconstruction reveals a large pool of black rot resistance donors
The review of the overall looked-up resources allowed to find 20 BR resistance reports (Table S2), for a total of 498 scores of 263 entities, 246 varieties (V. vinifera or hybrids) and 18 species screened, of which 148 and 14 respectively resulted resistant in at least one survey (Table S3). Some of those (78 varieties and 13 species) have been tested more than once, to a maximum of nine times. All the collected data regarding the 148 resistant varieties are summarized in Fig. 1, and all species in Fig. 2, listed based on median resistance. The dataset has then been integrated with the 23 genotypes phenotyped in this work. Finally, including their relatives, a list of 586 entries (Table S5 and S6) was used for the graphical depiction of the overall pedigree of BR resistance donors (Fig. S1).
Graphical representation of historical phenotypic data of black rot resistance evaluation of Vitis species accessions (whole plant) collected in this work. Genotypes (y-axes) are sorted and colored by the median (x-axes), represented by a mark (×), from very low resistance (1, violet) to very high (9, yellow)
Graphical representation of historical black rot resistance phenotypic data collected in this work of grapevine varieties evaluated at least once as resistant (maximum ≥ 5, bunch and leaf). Genotypes (y-axes) are sorted by the median (x-axes), represented with a mark (×), from very low resistance (1, violet) to very high (9, yellow)
Three works that screened different wild species accessions were found and included in this study (Soursac 1908 and Viala 1889, reviewed by Galet 1977; and Hausmann et al. 2017), along with a fourth one describing the screening of pure V. rotundifolia varieties (Jabco et al. 1985). Those works suggested that very high (median = 9) level of resistance is present in different species: V. mustangensis and V. vulpina. Sources of high resistance (7 ≤ median < 9) are present within V. acerifolia, V. aestivalis, V. aestivalis var. aestivalis (syn. lincecumii), V. cinerea, V. cinerea var. helleri (syn. berlandieri), V. monticola, V. riparia, V. rupestris and V. rotundifolia and V. shuttleworthii, while medium (5 ≤ median < 7) resistance has been detected in V. amurensis and V. palmata (syn. rubra) (Soursac 1908) (Fig. 1). Conversely, V. arizonica, V. californica and V. labrusca have been described to be susceptible. Even if V. vinifera is generally consider highly susceptible (Fig. 2), two Georgian varieties, ‘Muradouli’ and ‘Ojaleshi’, have been described to be medium resistant (Roznik et al. 2017). Upon pedigree contribution analysis of the genotypes considered in this study, the species that resulted most often as BR resistance donor was V. rotundifolia, represented by nine resistant cultivars evaluated by Jabco et al. (1985). To evaluate FEM breeding material, five seedlings originating from this species were included in the screening test: F13P86, NY24, NY30, NY39 and NY42; in particular, they have VRH3082-1-42 as a grandparent, namely a V. rotundifolia-derived back cross line called by antonomasia BC4 worldwide. The second most prominent species was V. labrusca, whose accessions evaluated to date have been found to be susceptible (Soursac 1908). However, by inspecting its subpopulation (Fig. S2) nine varieties shown from 5 to 9 resistance level, and the three varieties ‘Carman’, Fredonia’ and ‘Triumph’ (Fig. 3), showed only V. labrusca as wild ancestor, without any unknown (‘UNK’) or open pollinated (‘OP’) parent reported in the pedigree. To consider the possibility of a V. labrusca-derived resistance trait, two FEM wild parental lines ‘Coia 9’ and ‘Corella 2’ – carrying evident labrusca-like phenotypic characteristics – were tested for BR resistance.
Pedigree of black rot resistant V. labrusca-derived hybrids ‘Triumph’, ‘Fredonia’ and ‘Carman’. Color code from very low resistance (1, violet) to very high (9, yellow). Graphical representation by Helium (Shaw et al. 2014)
Focusing on varieties data (Fig. 2, Table S3), cases of strong discrepancy between evaluations were inspected. For instance, ‘Bianca’ is a complex hybrid widely exploited in Hungary, where it is considered very susceptible to BR (Roznik et al. 2017). On the contrary, its evaluation with P. ampelicida strains collected in Germany (Hausmann et al. 2017) classified this variety as medium resistant, and the same difference was detected also for its parental donor ‘Villard blanc’ (Table S3). A lower resistance have also been detected in North American assessments compared to Europe for ‘Catawba’, ‘Clinton’, ‘Concord’, ‘Jacquez’, ‘Ironclad’, ‘Missouri Riesling’ and ‘Triumph’, ‘De Chaunac’, ‘Chancellor’, ‘Fredonia’ and ‘Seyval blanc’ (Fig. 4a). Moreover, a consistent lower resistance of bunches compared to leaves evaluations was highlighted by Roznik et al. (2017) and Töpfer and Maul (2017) (Table S3), and confirmed in the overall dataset (Fig. 4b). The only exception was given by ‘Felicia’, which nevertheless showed high resistance of all organs. In other cases, a significant difference among USA states or years of evaluations was found, as for ‘La Crosse’, medium resistant in Minnesota (MGGA 1991, 2016) but susceptible in Nebraska (Read and Gamet 2022), or ‘La Crescent’, medium resistant in Nebraska (Read and Gamet year 2022) and Minnesota (MGGA 2016) but recently found to be susceptible in Minnesota (MGGA 2018) (Table S3).
Within the varieties, considering the comprehensive dataset, the most frequent genotype determined by pedigree contribution was ‘Seyval blanc’ with 24 resistant descendants documented (Fig. 5), while it was ‘Merzling’ for the current studied subset (Fig. S3). The high contribution of ‘Merzling’ to the pedigree of the studied panel was determined by its extensive use in FEM breeding, giving the possibility to evaluate the inheritance of the BR resistant trait coming from its parent, ‘Seyval blanc’, a French hybrid well-known for its BR resistance as confirmed by ten evaluations in previous reports (Table S3). The remaining genotypes studied in this work were therefore chosen based on kinship degree with ‘Seyval blanc’.
Progeny of black rot resistant French hybrids ‘Seyval blanc’. Color code from very low resistance (1, violet) to very high (9, yellow). Graphical representation by Helium (Shaw et al. 2014)
Strains isolated in the Valsugana valley are genetically identical
The samples isolated in Italy were confirmed as P. ampelicida by the ITS analysis. Following the molecular characterization of the five studied cultures, the most polymorphic SSRs resulted to be GBMS10, 08 and 11 with three alleles, and GBMS06 and 07 with two. All samples were monomorphic at GBMS01 to 04 and at GBMS09 SSR loci. Upon three assays, no amplification occurred with GBMS05 primers, suggesting a common monomorphic null allele. The three strains provided by JKI resulted to be different among each other and compared to the samples isolated in Trentino (genotype B, C, D). Instead, TN1 and TN2 isolates resulted to be identical (genotype A) (Table 1). Dual colony assays did not show any inhibition between the isolates, in contrast with the outcomes by Roznik (2019). Furthermore, no statistical difference in performance was detected among TN1, TN2, and 18.1 strains. By contrast, it was not possible to produce the primary inoculum from strains 80.88 and 10.133, probably due to loss of virulence caused by long term propagation on plates without re-isolation from infected material.
Based on the estimated standard error predicted by the linear model, conidia concentration changed significantly from 5 to ten minutes, reaching a plateau thereafter (Fig. S4). Therefore, the optimal leaf soaking time was settled at ten minutes, consistently with the timing guidelines provided for inoculum production from plates (Hoffman et al. 2002, Wicht et al. 2012, Rex et al. 2014).
Both mildew and black rot resistances are combined into the FEM breeding
The foliar artificial inoculations led to the identification of 12 resistant genotypes (mean ≥ 5), which were grouped based on the median end the minimum resistance level (Fig. 6). Three genotypes resulted symptomless in all assessments (mean and minimum = 9). One was ‘Seyval blanc’, the most exploited resistance donor as highlighted by its pedigree contribution (Figure S1), which already carry two R-loci against PM and two haplotypes of an R-locus against DM. The other two genotypes were NY24 and NY39, two seedlings of the population Eger 99-11.01 × NY 95.0308.02 that were selected in the FEM breeding program for their high resistance to both DM and PM. ‘Calardis musqué’ and ‘Calardis blanc’, deriving from ‘Seyval blanc’ and carrying the same mentioned R-loci, showed a median resistance of 9 and a minimum of 7. The same for F25P52, a breeding line derived from the variety ‘Bianca’ and descendant of ‘Villard blanc’, originally selected for its resistance to DM. ‘Felicia’, ‘Merzling’ and its offspring ‘Solaris’, cultivars known for their resistance to DM, also exhibited very high resistance to BR (median = 9), but with an increased variability of the response (minimum = 5). Only ‘Villard blanc’ showed a median resistance of 7, but unlike the previous group it never showed minimum resistance lower than 7, and it carries an R-locus for DM. Finally, two genotypes exhibited medium resistance (mean = 5). One of them, the breeding line F12P19, was characterized by a resistance distribution that tended to 9 (minimum = 5). ‘Charvir’ instead, an already patented cultivar due to its resistance to both DM and PM, showed a variability from 3 to 7. All the other genotypes of the studied panel revealed from low to very low resistance (median < 5).
Black rot resistance evaluation on parental lines and breeding selections. Resistance performance of 24 genotypes was assessed by artificial leaf infection under controlled conditions of grafted potted plants, by means of ten inoculations, three evaluation per genotype (three biological replicates). The resistance scale goes from very low (1) to very high (9). Genotypes (yaxes) are sorted by the median (x-axes), represented by a dot (•)
Veins and trichomes are hotspots for conidia germination and hyphal growth
The objective of this experiment was to follow disease progression in genotypes with different degrees of resistance through microscopic assessment of spore germination (1 dpi) and mycelium growth (3, 7 and 14 dpi) on detached leaves. However, no evidence of different germination or hyphal production was detectable between the genotypes, and the progression of disease did not occur, which also happens on leaf discs (Hausmann L. personal communication). Nonetheless, general considerations were made about shared features across the different genotypes. First, at 1 dpi germinated conidia were detectable through the formation of the melanized appressoria that preferentially adhered near the veins (asterisks, Fig. 7A and B). Second, increased concentration of conidia (1 dpi) and hyphal growth was observed around the trichomes that appeared to trap the spores (Fig. 7C and D).
Microscopic visualization of spore germination and hyphal growth on leaf tissue from the susceptible variety ‘Teroldego’. Disease progression assessment through bright field (A and C) and fluorescence (B and D) confocal microscopy at 1 (A and B, 50 μm) and 4 dpi (C and D, 200 μm). Appressoria were clearly detectable at 1 dpi (asterisks) for the characteristic dark brown color due to melanization, and different stages of germinating tube development were visible (triangles). Un-germinated conidia keep a translucent aspect (cross mark), while after germination they become visible just through fluorescence microscopy (squares). Conidia were preferentially trapped by trichomes, where the hyphal growth was concentrated (arrows). Leica DM 6000 CS
Discussion
Gathered phenotypic and pedigree information to support decision-making process in breeding
Although phenotypic and pedigree data are often publicly available, their use and association are not straightforward since they are dispersed and often recorded without following official standard references and ontologies. Therefore, the value of that information strictly depends on their level of correspondence to FAIR (findability, accessibility, interoperability, and reusability) principles (Wilkinson et al. 2016). In fact, breeders’ choices are commonly based on their know-how, but with the lack of a systematic screening of available resources. For grapevine, VIVC is a very comprehensive database of pedigree information, also specifying variety prime names and synonyms, not a trivial issue in viticulture (Maul et al. 2022). When available, universal marker profiles are also included, to support varietal and parentage identification. However, despite the efforts of VIVC, phenotypic data are scattered in a multitude of publications and reports. In this work all available BR resistance data were collected (Table S2 and S3) to give an overview of the available trait donors (Figure S1) and the instruments for the exploration of their pedigree through the viewer Helium (Shaw et al. 2014) (Table S5 and S6). In fact, digitalization and visualization through dedicated software provide a powerful tool for the association and interoperability of pedigree and phenotypic information to support decision-making process in breeding. A comprehensive view of the trait of interest allows to highlight interconnections and inheritance patterns otherwise difficult to recognize within complex datasets. For this purpose, the graphic representation and navigation of the BR resistance pedigree reconstructed in this work will also be freely available through the user-friendly web application PERSEUS (Jurado-Ruiz et al. 2022).
The availability of a comprehensive database of genetic and phenotypic data is the ultimate challenge. Effective examples are given for instance by the effort of the Rice Research and Development institute of Sri Lanka (Rathnayake et al. 2020) that organized the pedigree, genetic and phenotypic information of its breeding program to support accurate breeding decisions. The same approach was followed by the Crop Genetics Department of John Innes Centre for the historical datasets of national variety performance trials (NVPT) of winter wheat grown in the United Kingdom between 2002 and 2017 (Shorinola et al. 2022), and many other application are given regarding both herbaceous (i.e., white yam, Norman et al. 2020; maize, White et al. 2020; strawberry, Pincot et al. 2021; red clover, Egan et al. 2019, 2021; soybean, Lee et al. 2015; wheat, Fradgley et al. 2019) and woody (i.e., grapevine, Van Heerden et al. 2018; pear, Montanari et al. 2020; peach, Fresnedo-Ramírez et al. 2016) crops. In this regard, FEM is actively working on the creation of a database for FAIR management and integration of its data relating to the grapevine (FEMVitisDB, Vezzulli et al. 2022).
Exploration of black rot resistance resources for pyramiding strategy
The review of the available information on BR resistance revealed a broad spectrum of Vitis species that have genotypes that could be used as BR resistance donors. Pedigree reconstruction gives an important overview of both founders and interesting hybrids to be used in grapevine breeding. Thus, assuming that BR resistant genotypes with unlinked pedigrees will carry different types of resistance mechanisms, there is a concrete possibility to combine them with a pyramiding strategy for durable protection against P. ampelicida (Pilet-Nayel et al. 2017). Of course, the validation of the historical pedigrees by means of molecular markers is a fundamental step to support this hypothesis. Pyramiding different resistance loci against the same pathogen is an efficient strategy to increase resistance durability. Recently it has been estimated that four pyramided R-genes would be theoretically impossible to overcome for a pathogen population of cereal PM, since the occurrence of a quadruple mutant capable of eluding four different resistance mechanisms is extremely unlikely (Dry et al. 2019). The importance of the identification of different sources of resistance is even exacerbated for woody perennial plants species. This is because the plants remain in the vineyard for dozens of years, while the fast life cycle of pathogens facilitates the selection of new strains able to overcome a single specific resistance mechanism. The utilization of donors that have undergone selection for fruit quality is also an efficient strategy, far more desirable compared to the use of wild accessions or first-generation hybrids that often carry undesirable traits (Roznik et al. 2017). In this regard, it is of interest that also some Georgian V. vinifera demonstrated BR resistance evidence (Töpfer and Maul 2017). This result is supported by the discovery of PM (Hoffmann et al. 2008; Possamai et al. 2021) and DM (Toffolatti et al. 2018) R-loci in V. vinifera varieties of the same region, with the one identified in the variety ‘Kishmish vatkana’ (Hoffmann et al. 2008) being already widely used in grapevine breeding programs (Katula-Debreceni et al. 2010; Agurto et al. 2017; Fresnedo-Ramírez et al. 2017; Zini et al. 2019). In addition, it is known that there are V. vinifera with lower susceptibility to BR (Soursac 1908; Jabco et al. 1985), and it was already noted that varieties with large juicy grains are more susceptible (Galet 1977). As for botrytis (Possamai et al. 2021), it can be presumed that there are specific physiological traits that influence the degree of resistance. This hypothesis is also supported by evidence of an effect of the V. vinifera background on DM and PM resistant seedlings with the same R-loci assessment (Bettinelli et al. 2021). We also found cases of acquisition of increased BR resistance in crosses between a hybrid with a V. vinifera variety, i.e., ‘Calardis musqué’ and ‘Carman’. Regarding this peculiar occurrence, some hypotheses on the possible causes can be made, such as the segregation of an antagonistic or susceptibility trait in the resistant parent, or epistatic effects determined by the V. vinifera background (Bettinelli et al. 2021). Nonetheless, different susceptibility degrees to the strain used for the screening must also be considered. Another interesting case is given by V. labrusca, mostly described as susceptible (Fig. 1). Nevertheless, some V. labrusca crosses with V. vinifera i.e., ‘Catawba’, have been described as resistant to BR (Fig. 3) (Hausmann et al. 2017). If their pedigree is confirmed by means of molecular markers, it will open a new perspective for breeders, especially in the field of grape juice varieties. Finally, concerning the possible inheritance of the resistant trait from V. rotundifolia for BC4 derived genotypes, this is extremely unlikely (Kozma personal communication). In fact, so far, no BR resistance has been described within the 99–1 V. rotundifolia hybrids (Fig. S1) extensively used at the breeding institute of Pécs University (Hungary), where BR is heavily present and studied (Roznik et al. 2017).
Interesting varieties that show a high difference between organ evaluations were found and specific trends were highlighted. Primarily, it was evident a divergent resistance of bunches (lower) compared to leaves (higher). The issue of unpaired susceptibility between fruits and leaves had already been raised by Arnaud in 1931 (as reported by Galet 1977) and confirmed by the results of Töpfer and Maul (2017) and Roznik et al. (2017) (Table S3). In general, analyzing the overall collected data (Table S3), within the 48 varieties with both leaf and bunch evaluations, there were only ten cases of higher fruit resistance. Of those, just three of them, ‘Brianna’, ‘Vidal blanc’ and ‘Verdelet’, actually showed resistant bunches and susceptible leaves, all the others were resistant at both organs. However, it must be noticed that leaf assessments were evaluated with high-pressure greenhouse artificial infection experiments, while bunch response was derived from field observations of natural infection, certainly influenced by environmental conditions and seasonality. Nevertheless these outcomes, supported by the cited previous works, suggest the importance to focus also on bunch resistance i.e., for QTL detection and identification of associated molecular markers, as demonstrated in our recent work (Bettinelli et al. 2023). Moreover, numerous cases of increased resistance were also detected in experiments carried out in Europe compared to North America. This may be due to the occurrence in the USA of strains that overcame specific resistance mechanisms due to the selective pressure caused by long-standing coexistence of the resistant varieties with the pathogen, while these strains have not yet been imported or evolved in the EU. Therefore, some specific R-loci remain still effective vs. European P. ampelicida population, and this is what we presume for the possible V. labrusca derived resistance trait carried by ‘Catawba’, ‘Clinton’, ‘Concord’ and ‘Triumph’, and maybe ‘Ironclad’ and ‘Missouri Riesling’ (V. labrusca × V. riparia). The same hypothesis can be done about the lower resistance displayed by ‘Bianca’ and ‘Villard blanc’ in Hungary. In fact, the extensive use of resistant hybrids in that region and the consequent reduction in treatments may have led to the selection of strains that overcome ‘Villard blanc’ derived resistance mechanism, and apparently those strains are not present in Germany and, as suggested by the current study, also in Trentino region. Similarly, strains diversity has also to be considered regarding differences within years of assessment or USA regions. Those evidences suggested that particular attention should be given to the topic of strain genetic variability, selection and diffusion, and therefore genetic characterization when resistance assessments are made. Finally, where the discrepancy was very large and with no apparent justification, the identity of the evaluated genetic material was questioned, since cases of misnomers have already been reported (Maul and Töpfer 2015). This was the case of the questionable susceptibility of ‘Seyval blanc’ in Nebraska (Read and Gamet 2022), or the resistance of ‘Chardonnay blanc’ in Kentucky (Ward and Kaiser 2012).
Good practices for pathogen maintenance and inoculation
Genetic characterization of pathogen strains is a fundamental step for the harmonization of phenotypic data, as expressed by the International Union for the Protection of New Varieties of Plants (UPOV) in the European project “Harmorescoll” for the harmonization of reference collections of isolates for resistance DUS-testing (Distinctness, Uniformity and Stability). In modern breeding programs, this should be a mandatory step to take trace of the strain that have been used for selection, and a mix of strains should be recommended to avoid the selection of race-specific resistance mechanisms. Furthermore, during the process of breeding for biotic stresses, it is also very important to propagate the pathogen just on susceptible (V. vinifera) plants, avoiding selecting for new mutation overcoming the genetic barriers that are being introgressed.
The optimization of propagation and inoculation protocols is also crucial for an efficient phenotyping process when a relevant number of seedlings need to be screened. Being independent from plate propagation made the workflow much more rapid and efficient. Actually, despite the positive aspect of the possibility to grow P. ampelicida on cultural medium, the principle of ‘one plate-one plant’ generally adopted for artificial infection is not easily feasible because of the need of a lot of space in growing chambers, while a single infected leaf produces enough inoculum for many plants. Moreover, the routinary inoculation of V. vinifera permitted continuous feedback on infection dynamics during different seasons. In fact, according to our experience – even the controlled conditions of the greenhouse – the development of the disease was influenced by seasonal factors such as the cycle of natural light and the different rate of growth of the plants, suggesting spring and the first half of summer as the best period for BR inoculation. This introduced an additional level of complexity in the phenotyping procedures in terms of synchronization of pathogen and host life cycles. Lastly, this new approach permits to maintain a highly active and virulent inoculum, while there is evidence of a loss of these important characteristics producing the inoculum from the fungus propagated on the cultural medium.
Deployment of parental selection into disease resistance breeding programs
Although ex vivo assay experiments were useful for visualizing the early stages of conidia adhesion and hypha growth, assessment of disease progression was probably not feasible because the pathogen did not perceive adequate signals on detached leaves. According to in vivo assessments on leaves, ‘Seyval blanc’ was a symptom-free BR resistant genotype. It originated from the cross between ‘Seibel 5656’ and ‘Rayon d’or’ (Fig. S5), the latest being also the BR resistance donor of the fruit symptomless variety ‘Csillam’ (Kiss et al. 2017). Furthermore, ‘Seibel 5656’, brother of ‘Subereux’, is most probably the BR resistance donor of ‘Villard blanc’, as its mother ‘Seibel 6468’ was documented susceptible by Soursac (1908). Accordingly, excluding ‘Seibel 5656’ from the donors, we can conclude that both the resistance trait carried by ‘Seyval blanc’ and ‘Villard blanc’ originated from V. rupestris and V. aestivalis var. lincecumii as wild ancestors, respectively (Fig. 4). NY39 and NY24 were also found to be very high resistant to BR, and both share a complex genealogical tree combining V. aestivalis, V. cinerea, V. labrusca, V. rotundifolia, V. rupestris and V. riparia. ‘Merzling’ and ‘Solaris’ are respectively one and two steps forward in the pedigree (Fig. S3) in respect to ‘Seyval blanc’. Even if they still have a high degree of resistance, their response is characterized by an increased variability (Fig. 6). The decrease of resistance toward ‘Seyval blanc’ pedigree is evident also inspecting the different scores of the other ‘Merzling’ offspring (Fig. S3). This could be indirect evidence that the resistance of ‘Seyval blanc’ is based on several QTL, some of them not inherited by the ‘Merzling’ offspring. The breeding selection F25P52 resulted also very resistant and since it is descendant of ‘Villard blanc’, thus again suggested for the segregation of negative epistatic traits through ‘Villard blanc’ pedigree, as for ‘Seyval blanc’. Finally, the genotypes that showed medium resistance in the high pressure of greenhouse experiments, as ‘Charvir’ and the breeding selection F12P19, are also promising candidates for the fight against BR under a well-timed organic farming management.
Conclusion
This work outlined good practices for black rot resistance evaluation for breeding, developing a new propagation and inoculation protocol based on fresh infected leaves. Moreover, it gave a comprehensive overview of black rot resistance resources to the grapevine community, and it allowed the spotlight on a subset of genotypes to be evaluated to identify new BR resistance donors within FEM parental lines and breeding selections that were readily introduced in the breeding pipeline. Those genotypes in fact have been already crossed both with V. vinifera varieties or with hybrids for the introgression of the trait into DM and PM resistance genotypes, for the generation of disease resistance super-donors.
Data Availability
The data collected in the current study are partially available in the VIVC (Vitis International Variety Catalogue) repository (https://www.vivc.de/). There is the intention to upgrade this web resource with the missing data. Moreover, the reconstructed pedigree together with the black rot resistance trait information will be uploaded to the visualization webtool PERSEUS (Jurado-Ruiz et al. 2022).
References
Agurto M, Schlechter RO, Armijo G et al (2017) RUN1 and REN1 pyramiding in grapevine (Vitis vinifera cv. crimson seedless) displays an improved defense response leading to enhanced resistance to powdery mildew (Erysiphe necator). Front Plant Sci 8:25. https://doi.org/10.3389/fpls.2017.00758
Bettinelli P, Camponogara Tomazetti T, Zulini L et al (2021) Forward marker-assisted selection for mildew resistance in grapevine: an optimized applied process. In: 21st general congress Eucarpia. Rotterdam, the Netherlands. https://hdl.handle.net/11572/377380
Bettinelli P, Nicolini D, Costantini L et al (2023) Towards marker-assisted breeding for black rot bunch resistance: identification of a major QTL in the Grapevine Cultivar ‘Merzling. Int J Mol Sci 24:3568. https://doi.org/10.3390/ijms24043568
Dry IB, Riaz S, Fuchs M et al (2019) Scion breeding for resistance to biotic stresses. In: Cantu D, Walker MA (eds) The grape genome. Springer International Publishing, Cham, Switzerland, pp 319–347. https://doi.org/10.1007/978-3-030-18601-2_15
Egan LM, Hofmann RW, Ghamkhar K, Hoyos-Villegas V (2019) Identification of founding accessions and patterns of relatedness and inbreeding derived from historical Pedigree Data in a Red Clover Germplasm Collection in New Zealand. Crop Sci 59:2100–2108. https://doi.org/10.2135/cropsci2019.01.0045
Egan LM, Hofmann RW, Ghamkhar K, Hoyos-Villegas V (2021) Prospects for Trifolium Improvement through Germplasm Characterisation and Pre-breeding in New Zealand and Beyond. Front Plant Sci 12:1056. https://doi.org/10.3389/FPLS.2021.653191/BIBTEX
European C (2020) EUR-Lex – 52020DC0381. In: Eur. Comm. https://eur-lex.europa.eu/legal-content/EN/TXT/?uri=CELEX:52020DC0381. Accessed 5 Aug 2022
Fradgley N, Gardner KA, Cockram J et al (2019) A large-scale pedigree resource of wheat reveals evidence for adaptation and selection by breeders. PLOS Biol 17:e3000071. https://doi.org/10.1371/journal.pbio.3000071
Fresnedo-Ramírez J, Frett TJ, Sandefur PJ et al (2016) QTL mapping and breeding value estimation through pedigree-based analysis of fruit size and weight in four diverse peach breeding programs. Tree Genet Genomes 12:25. https://doi.org/10.1007/s11295-016-0985-z
Fresnedo-Ramírez J, Yang S, Sun Q et al (2017) An integrative AmpSeq platform for highly multiplexed marker-assisted pyramiding of grapevine powdery mildew resistance loci. Mol Breed 37:1–16. https://doi.org/10.1007/S11032-017-0739-0/TABLES/4
Galet P (1977) Black rot. In: Les maladies et les parasites de la vigne. Le Paysan du Midi, Montpellier, France, pp 223–260, Tome 1
Harms M, Holz G, Hoffmann C et al (2005) Occurrence of Guignardia bidwellii, the causal fungus of Black Rot on grapevine, in the vine growing areas of Rhineland-Palatinate, Germany M Harms. In: BCPC Symposium. pp 127–132
Hausmann L, Rex F, Töpfer R (2017) Evaluation and genetic analysis of grapevine black rot resistances. Acta Hortic 1188:285–290. https://doi.org/10.17660/ActaHortic.2017.1188.37
Hoffman LE, Wilcox WF (2002) Utilizing epidemiological investigations to optimize management of grape black rot. Phytopathology 92:676–680. https://doi.org/10.1094/PHYTO.2002.92.6.676
Hoffmann S, Di Gaspero G, Kovács L et al (2008) Resistance to Erysiphe necator in the grapevine ‘Kishmish vatkana’ is controlled by a single locus through restriction of hyphal growth. Theor Appl Genet 116:427–438. https://doi.org/10.1007/s00122-007-0680-4
Houle D, Govindaraju DR, Omholt S (2010) Phenomics: the next challenge. Nat Rev Genet 11:855–866. https://doi.org/10.1038/nrg2897
Jabco JP, Nesbitt WB, Werner DJ (1985) Resistance of various classes of grapes to the bunch and muscadine grape forms of Black Rot. J Am Soc Hortic Sci 110:762–765
Jermini M, Gessler C (1996) Epidemiology and control of grape black rot in Southern Switzerland. Plant Dis 80:322. https://doi.org/10.1094/PD-80-0322
Jurado-Ruiz F, Pradas N, Arús P, Aranzana MJ (2022) Molecular-based pedigree reconstruction of peach cultivars. Acta Hortic 1352:133–140. https://doi.org/10.17660/ActaHortic.2022.1352.18
Katula-Debreceni D, Lencsés AK, Szőke A et al (2010) Marker-assisted selection for two dominant powdery mildew resistance genes introgressed into a hybrid grape population. Sci Hortic (Amsterdam) 126:448–453. https://doi.org/10.1016/J.SCIENTA.2010.08.012
Kiss E, Tóth-Lencsés K, Szke A et al (2017) Origin of Csillam, a promising source for black rot resistance. Vitis - J Grapevine Res 56:53–54. https://doi.org/10.5073/vitis.2017.56.53-54
Lee C, Choi M-S, Kim H-T et al (2015) Soybean [Glycine max (L.) Merrill]: importance as a crop and Pedigree Reconstruction of Korean Varieties. Plant Breed Biotechnol 3:179–196. https://doi.org/10.9787/PBB.2015.3.3.179
Maul E, Töpfer R (2015) Vitis International Variety Catalogue (VIVC): a cultivar database referenced by genetic profiles and morphology. BIO Web Conf 5:01009. https://doi.org/10.1051/bioconf/20150501009
Maul et al (2022) Vitis International Variety Catalogue. https://www.vivc.de
MGGA (1991) Growing grapes in Minnesota, Fifth edit. Minnesota Grape Growers Association
MGGA (2016) Growing grapes in Minnesota, Tenth edit. Minnesota Grape Growers Association
MGGA (2018) Cold Hardy Grape Varieties, indigenous to Minnesota.mngrapes.org/page/varieties
Montanari S, Postman J, Bassil NV, Neale DB (2020) Reconstruction of the largest pedigree network for pear cultivars and evaluation of the genetic diversity of the USDA-ARS national pyrus collection. G3 Genes Genomes Genet 10:3285–3297. https://doi.org/10.1534/g3.120.401327
Narduzzi-Wicht B, Jermini M, Gessler C, Broggini GAL (2014) Microsatellite markers for population studies of the ascomycete Phyllosticta ampelicida, the pathogen causing grape black rot. Phytopathol Mediterr 53:470–479. https://doi.org/10.14601/phytopathol_mediterr-14481
Norman PE, Paterne AA, Danquah A et al (2020) Paternity assignment in white guinea Yam (Dioscorea Rotundata) half-sib progenies from polycross mating design using SNP markers. Plants 9:527. https://doi.org/10.3390/plants9040527
OIV (2009) 2nd edition of the OIV Descriptor. list for grape varieties and Vitis species
Pilet-Nayel ML, Moury B, Caffier V et al (2017) Quantitative resistance to plant pathogens in pyramiding strategies for durable crop protection. Front Plant Sci 8:1838. https://doi.org/10.3389/FPLS.2017.01838/BIBTEX
Pincot DDA, Ledda M, Feldmann MJ et al (2021) Social network analysis of the genealogy of strawberry: retracing the wild roots of heirloom and modern cultivars. G3 Genes Genomes Genet 11. https://doi.org/10.1093/g3journal/jkab015
Pirrello C, Mizzotti C, Tomazetti TC, Colombo M et al (2019) Emergent ascomycetes in viticulture: an interdisciplinary overview. Front Plant Sci 10:1–30. https://doi.org/10.3389/fpls.2019.01394
Possamai T, Wiedemann-Merdinoglu S, Merdinoglu D et al (2021) Construction of a high-density genetic map and detection of a major QTL of resistance to powdery mildew (Erysiphe necator Sch.) In caucasian grapes (Vitis vinifera L.). BMC Plant Biol 21:528. https://doi.org/10.1186/s12870-021-03174-4
Ramsdell DC, Milholland RD (1988) Black rot. In: Pearson RC, Goheen AC (eds) Compendium of Grape Diseases. APS Press, pp 15–16
Rathnayake R, Sahibdeen S, Udawela K et al (2020) Application of Pedimap: a pedigree visualization tool to facilitate the decisioning of rice breeding in Sri Lanka. Sci Rep 2020 101 10:1–14. https://doi.org/10.1038/s41598-020-71260-y
Read P, Gamet S, Evaluating Diseases on Nebraska’s Major Cultivars. https://viticulture.unl.edu/viticulture/Disease-Identification-on-Nebraskas-Major-Cultivars.pdf. Accessed 26 Jan 2022
Rex F, Fechter I, Hausmann L, Töpfer R (2014) QTL mapping of black rot (Guignardia bidwellii) resistance in the grapevine rootstock ‘Börner’ (V. riparia Gm183 × V. cinerea Arnold). Theor Appl Genet 127:1667–1677. https://doi.org/10.1007/s00122-014-2329-4
Ries SM (1999) Reports on Plant Diseases No. 703 Black Rot of Grape. http://ipm.illinois.edu/diseases/series700/rpd703/. Accessed 31 Jan 2022
Rossman AY, Crous PW, Hyde KD et al (2015) Recommended names for pleomorphic genera in Dothideomycetes. IMA Fungus 6:507–523. https://doi.org/10.5598/imafungus.2015.06.02.14
Roznik D (2019) Szent István Egyetem A feketerothadás (Guignardia bidwellii (Ellis) Viala et ravaz) elleni rezisztencia források azonosítása és felhasználhatósága a szőlő rezisztencia nemesítésében Doktori (PhD) értekezés Roznik Dóra Budapest. Szent István University, UN
Roznik D, Hoffmann S, Kozma P (2017) Screening a large set of grape accessions for resistance against black rot (Guignardia bidwellii/(Ell.)). Mitteilungen Klosterneuburg, Rebe und Wein, Obs und Früchteverwertung 67, pp 149–157
Shaw PD, Graham M, Kennedy J et al (2014) Helium: visualization of large scale plant pedigrees. BMC Bioinform 15:1–15. https://doi.org/10.1186/1471-2105-15-259/FIGURES/8
Shorinola O, Simmonds J, Wingen LU, Uauy C (2022) Trend, population structure, and trait mapping from 15 years of national varietal trials of UK winter wheat. G3 Genes|Genomes|Genetics 12. https://doi.org/10.1093/G3JOURNAL/JKAB415
Soursac L (1908) Recherches sur le black-rot. Coulet et Fils, Montpellier, France
Toffolatti SL, De Lorenzis G, Costa A et al (2018) Unique resistance traits against downy mildew from the center of origin of grapevine (Vitis vinifera). Sci Rep 8:12523. https://doi.org/10.1038/s41598-018-30413-w
Töpfer R, Maul E (2017) Grapevine genetic resources: evaluation and pre-breeding at the European level grapevine Breeding. In: Private Public Partnerships Workshop of the European Cooperative Programme for Plant Genetic Resources (ECPGR). Bonn, Germany
Ullrich CI, Kleespies RG, Enders M, Koch E (2009) Biology of the black rot pathogen, Guignardia bidwellii, its development in susceptible leaves of grapevine Vitis vinifera. J Für Kult 61:82–90
Van Heerden CJ, Burger P, Prins R (2018) Microsatellite-based DNA fingerprinting of selected grapevine cultivars. South Afr J Enol Vitic 39:58–66. https://doi.org/10.21548/39-1-2053
Vezzulli S, Bianco L, Nicolini D et al (2022) FEMVitisDB: a FAIR data management system for data integration in grapevine. In: XIII. International Symposium on Grapevine Breeding and Genetics, Pioneering Wines (PIWIs) - Innovation and Tradition, Abstract Book: 10th – 15th July 2022, Institute for Grapevine Breeding Geilweilerhof I Siebeldingen/Germany. Quedlinburg, Germany: Julius Kühn-Institut. S. 104. (= Julius-Kühn-Archiv). Online unter: https://www.openagrar.de/receive/openagrar_mods_00080180
Voorrips RE, Bink MCAM, Van De Weg WE (2012) Pedimap: Software for the visualization of genetic and phenotypic data in pedigrees. J Hered 103:903–907. https://doi.org/10.1093/jhered/ess060
Ward NA, Kaiser CA (2012) Black Rot of Grape Plant Pathology Fact Sheet
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols. A guide to methods and applications. Academic Press, San Diego, CA, pp 315–322
White MR, Mikel MA, Leon N, Kaeppler SM (2020) Diversity and heterotic patterns in north american proprietary dent maize germplasm. Crop Sci 60:100–114. https://doi.org/10.1002/csc2.20050
Wicht B, Petrini O, Jermini M et al (2012) Molecular, proteomic and morphological characterization of the ascomycete Guignardia bidwellii, agent of grape black rot: a polyphasic approach to fungal identification. Mycologia 104:1036–1045. https://doi.org/10.3852/11-242
Wilkinson MD, Dumontier M, Aalbersberg IjJ et al (2016) The FAIR Guiding Principles for scientific data management and stewardship. Sci Data 3:160018. https://doi.org/10.1038/sdata.2016.18
Zini E, Dolzani C, Stefanini M et al (2019) R-Loci arrangement Versus Downy and Powdery Mildew Resistance Level: a Vitis Hybrid Survey. Int J Mol Sci 20:3526. https://doi.org/10.3390/IJMS20143526
Acknowledgements
The authors thank Dr. Daniele Prodorutti and Dr. Christian Cainelli (CTT-FEM) for the support in the characterization of the pathogen, and Dr. Pietro Franceschi for the assistance with the statistical analysis. They are also grateful to the personnel of the grapevine breeding unit (CRI-FEM) for greenhouse and field management.
Funding
Open access funding provided by Fondazione Edmund Mach - Istituto Agrario di San Michele all'Adige within the CRUI-CARE Agreement. This study was co-funded by the University of Trento - Center Agriculture, Food and Environment (Italy) and the Grapevine Genetics and Breeding Unit as part of the PhD programme of Paola Bettinelli.
Author information
Authors and Affiliations
Contributions
PB and SV contributed to the study conception and design. Material preparation, data collection and analysis were performed by PB, DN and OG. The draft of the manuscript was written by PB and improved by SV. LH and MS commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
10681_2023_3235_MOESM1_ESM.eps
Pedigree reconstruction of black rot resistance donors based on historical phenotypic data. Color code from very low resistance (1, violet) to very high (9, yellow). Graphical representation by Helium (Shaw et al. 2014).
Supplementary file1 (EPS 75423 kb)
10681_2023_3235_MOESM2_ESM.crdownload
Pedigree snapshot of V. labrusca subpopulation. Color code from very low resistance (1, violet) to very high (9, yellow). Graphical representation by Helium (Shaw et al. 2014).
Supplementary file2 (TIFF 23800 kb)
10681_2023_3235_MOESM3_ESM.png
Pedigree snapshot of the 24 genotypes evaluated in this work. Color code from very low resistance (1, violet) to very high (9, yellow). Graphical representation by Helium (Shaw et al. 2014).
Supplementary file3 (TIFF 1524 kb)
10681_2023_3235_MOESM4_ESM.tiff
Linear model prediction of conidia release from fresh leaf lesions. Based on the estimated standard error, conidia concentration changes significantly from 5 to ten minutes, reaching a plateau thereafter.
Supplementary file5 (TIFF 18511 kb)
10681_2023_3235_MOESM5_ESM.crdownload
Pedigree of black rot resistant French hybrids ‘Seyval blanc’ and ‘Villard blanc’. Color code from very low resistance (1, violette) to very high (9, yellow). Color code from very low resistance (1, violet) to very high (9, yellow). Graphical representation by Helium (Shaw et al. 2014).
Supplementary file4 (TIFF 11016 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bettinelli, P., Nicolini, D., Giovannini, O. et al. Breeding for black rot resistance in grapevine: advanced approaches for germplasm screening. Euphytica 219, 113 (2023). https://doi.org/10.1007/s10681-023-03235-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-023-03235-9