Abstract
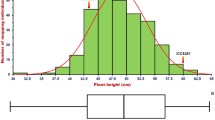
The tallness of the oil palm is a major hurdle for harvesting of ripened bunches after attaining the age of 25 years. Hence, the study conducted with an objective to identify and validate the major loci for dwarf trait using integrated genomic approaches viz., bulk segregant analysis (BSA), genome wide association mapping (GWAS), and bioinformatics. The average height increment of the germplasm was found to be 28.6 cm which ranges from 10.12 to 58.21 cm. The principal component analysis results showed that Zambian and Cameroon genotypes are having less height increment nature. The GWAS resulted in identification of six QTLs on chromosomes 4, 8, and 14. Out of the six QTLs, two SSRs (mEgCIR3350andmEgCIR0059) linked to two QTLs (qtlH1 and qtlH2 respectively) were significant. The BSA analysis also showed that the marker mEgCIR0059 is tightly linked to less height increment. Interestingly, the qtlH2 (mEgCIR0059) is found to be linked by both the BSA and GWAS analysis. The bioinformatics analysis of qtlH2 (mEgCIR0059) also confirmed that it is linked to the asparagine synthetase candidate gene. The marker was validated on large number of populations consisting of African germplasm, progeny derived from the cross between dwarf and tall germplasm, and dwarf palm which is available at our experimental farm. This marker can be effectively used in molecular breeding programmes to select dwarf germplasm at an early stage.







Similar content being viewed by others
Data availability
The genotypic data generated from the microsatellites have been submitted to the institute as per the mandate. It can’t be open to public as per the ICAR rules.
References
Alonso R, Oñate-Sánchez L, Weltmeier F, Ehlert A, Díaz I, Dietrich K, Vicente-Carbajosa J, Droge-Laser W (2009) A pivotal role of the basic leucine zipper transcription factor bZIP53 in the regulation of Arabidopsis seed maturation gene expression based on heterodimerization and protein complex formation. Plant Cell 21:1747–1761
Babu BK, Mathur RK, Kumar PN, Ramajayam D, Ravichandran G, Venu MVB (2017) Development, identification and validation of CAPS marker for SHELL trait which governs dura, pisifera and tenera fruit forms in oil palm (Elaeis guineensis Jacq). PLoS ONE 12(2):e0171933. https://doi.org/10.1371/journal.pone.0171933
Babu BK, Mathur RK, Ravichandran G et al (2019) Genome-wide association study (GWAS) for stem height increment in oil palm (Elaeis guineensis) germplasm using SNP markers. Tree GenetGenom. https://doi.org/10.1007/s11295-019-1349-2
Babu BK, Mathur RK, Ravichandran G, Anita P, Venu MVB (2020) Genome wide association study (GWAS) and identification of candidate genes for yield and oil yield related traits in oil palm (Eleaeis guineensis) using SNPs by genotyping-based sequencing. Genomics. https://doi.org/10.1016/j.ygeno.2019.06.018
Babu BK, Mathur RK, Venu MVB, Sandip S, Ravichandran G, Anitha P, Bhagya HP (2021) Genome-wide association study (GWAS) of major QTLs for bunch and oil yield related traits in Elaeis guineensis L. Plant Sci 305:110810
Bai B, Wang L, Lee M, Zhang Y, Rahmadsyah YA, Ye BQ, Zi YW, Lim CH, Suwanto A, Chua NH, Yue GH (2017) Genome-wide identification of markers for selecting higher oilcontent in oil palm. BMC Plant Biol 17:93
Bhagya HP, Kalyana Babu B, Gangadharappa PM, Mahantesha Naika BN, Satish D, Mathur RK (2020) Identification of QTLs in oil palm (Elaeis guineensis Jacq.) using SSR markers through association mapping. J. Genet. 99 (19): 1–10. https://doi.org/10.1007/s12041-020-1180-4
Billotte N, Risterucci AM et al (2005) Microsatellite based high density linkage map in oil palm (Elaeis guineensis Jacq). Theor Appl Genet 110:754–765
Billotte N, Jourjon MF, Marseillac N, Berger A, Flori A et al. (2010) QTL detection by multi-parent linkage mapping in oil palm (Elaeis guineensis Jacq). Theor Appl Genet 120:1673–1687
Bradbury P, Zhang Z, Kroon D, Casstevens T, Ramdoss Y et al (2007) TASSEL software for association mapping of complex traits in diverse samples. Bioinformatics 23(19):2633–2635
Corley RHV, Hardon JJ, Tan GY (1971) Analysis of growth of the oil palm (Elaeis guineensis Jacq) I. Estimation of growth parameters and application in breeding. Euphytica 20:307–315
Ithnin M, Xu Y, Marhalil M, Norhalida M, Amiruddin MD, Low L, Tan YC, Yap SJ, Li C, Rajanaidu N, Singh R, Xu S (2017) Multiple locus genome-wide association studies for important economic traits of oil palm. Tree Genet Genom 13:103
Kamal-Eldin A, Andersson R (1997) A multivariate study of the correlation between tocopherol content and fatty acid composition in vegetable oils. J Am Oil Chem Soc 74:375–380
Kumar B, Gomez SM, Boopathi NM, Kumar SS, Kumaresan D, Biji KR, Babu BK, Prasad NSR, Shanmugasundaram P, Babu RC (2005) Identification of microsatellite markers associated with drought tolerance in rice (Oryza sativa L.) using bulked line analysis. Trop Agric Res 17:39–47
Lee M, Xia JH, Zou Z, Ye J, Yuzer A et al (2015) A consensus linkage map of oil palm and a major QTL for stem height. Sci Rep 5:8232. https://doi.org/10.1038/srep08232
Morcillo F, Gagneur C, Adam H, Richaud F, Singh R, Cheah SC, Rival A, Duval Y, Tregear JW (2006) Somaclonal variation in micro propagated oil palm. Characterization of two novel genes with enhanced expression in epigenetically abnormal cell lines and in response to auxin. Tree Physiol 26:585–594
Mott R, Talbot CJ, Turri MG, Collins AC, Flint J (2000) A method for fine mapping quantitative trait loci in outbred animal stocks. Proc Natl Acad Sci USA 97:12649–12654
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4326
Ong PW, Maizura I, Marhalil M, Rajanaidu N, Abdullah NAP, Rafi MY, Ooi LCL, Low ETL, Singh R (2018) Association of SNP markers with height increment in MPOB-Angolan natural oil palm populations. J Oil Palm Res 30(1):61–70
Osorio-Guarín JA, Garzón-Martínez GA, Delgadillo-Duran P, Bastidas S, Moreno LP, Enciso-Rodríguez FE, Cornejo OE, Barrero LS (2019) Barrero Genome-wide association study (GWAS) for morphological and yield-related traits in an oil palm hybrid (Elaeis oleifera x Elaeis guineensis) population. BMC Plant Biol 19:533. https://doi.org/10.1186/s12870-019-2153-8
Pootakham W, Jomchai N, Areerate P, Shearman JR, Sonthirod C, Sangsrakru D, Tragoonrung S, Tangphatsornruang S (2015) Genome-wide SNP discovery and identification of QTL associated with agronomic traits in oil palm, using genotyping –bysequencing (GBS). Genomics 105:288–295
Pritchard JK, Wen W (2003) Documentation for the structure software, version 2 Department of Human Genetics, University of Chicago, Chicago. http://pritch.bsd.uchicago.edu/software.
Singh R, Tan SG, Panandam JM, Rahman RA, Ooi LC, Low ET et al (2009) Mapping quantitative trait loci (QTLs) for fatty acid composition in an inter specific cross of oil palm. BMC Plant Biol 9:114. https://doi.org/10.1186/1471-2229-9-114
Singh R, Ong-Abdullah M, Low ET, Manaf MA, Rosli R, Nookiah R et al (2013) Oil palm genome sequence reveals divergence of inter fertile species in old and new worlds. Nature 500:335–339. https://doi.org/10.1038/nature12309
Turner PD, Gillbanks RA (1974) Oil palm cultivation and management, 1st edn. The incorporated society of planter, Kuala Lampur
Acknowledgements
The authors thank Director, ICAR-IIOPR and DST-SERB/STAR (STR/2021/000037) for all the support.
Funding
Authors thank Indian Council of Agricultural Research for providing financial support.
Author information
Authors and Affiliations
Contributions
Conceptualization—BK, RKM. Experiments carried out—BK, RKM, MVB. Statistical analysis—BK. Writing—BK, RKM, GR, PA. Reviewing and editing—BK, RKM, HPB.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interests.
Human or animal rights
This article does not contain any studies with human or animal subjects.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Babu, B.K., Mathur, R.K., Ravichandran, G. et al. A novel QTL linked to asparagine synthetase gene for stem height increment in oil palm (E. guineensis Jacq.) identified and validated through integrated genomic approaches. Euphytica 219, 73 (2023). https://doi.org/10.1007/s10681-023-03201-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-023-03201-5