Abstract
East African highland bananas (Musa spp. AAA group) are an important staple in the Great Lakes region of East Africa. Their production has declined due to pests and diseases. Breeding for host plant resistance is a sustainable option for addressing this challenge. Understanding the relationships between growth parameters and bunch weight (i.e., yield) is crucial to guide breeding efforts for this crop. We investigated cause-effect relationships, through path analysis, in bunch weight of East African highland banana derived hybrids, their parents and grandparents. These family structures were planted in a 7 × 8 rectangular lattice design, replicated twice. Genetic gains for bunch weight (kg plant−1) and yield potential (t ha−1 year−1) were estimated. Significant increases of bunch weight and yield potential were noted from the landrace triploid germplasm, their derived primary tetraploid hybrids and secondary triploid bred-germplasm. Path analysis revealed that fruit length, circumference and number, number of hands and plant cycle number had a direct positive effect on the bunch weight. Days to fruit filling, days to maturity and index of non-spotted leaves had indirect effects on bunch weight. The average genetic gains for bunch weight and yield potential were 1.4% and 1.3% per year, respectively. This is the first report about genetic gains in banana breeding. Our findings may be useful for assessing progress and directing future breeding efforts in banana breeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
East African highland bananas (Musa spp., AAA group) are grown mainly by smallholder farmers in sub-Saharan Africa, who eat and sell their extra fruit to local rural and urban markets (Ochola et al. 2013). Because they are unpalatable when raw, plant fruits are usually boiled or steamed and pounded before consumption. Uganda is the largest producer and consumer of bananas in Africa, with an estimated per capita consumption between 400 to 600 kg per year (Tushemereirwe et al. 2000; Karamura et al. 2008), which is ranked as the highest in the world. Bananas cover about 38% of land under crops and they are grown by 75% of farmers in Uganda (Nowakunda and Tushemereirwe 2004). The East African highland bananas (EAHB) constitute at least 85% of these bananas. Despite their importance, banana production has been declining since the 1970s due mainly to insect (weevils) and nematode pests, diseases (bacteria, fungi and viruses), poor soil fertility, population pressure, shifted resource allocation in favor of crops like; sweet potato (Ipomoea batatas), cassava (Manihot esculenta) and beans (Phaseolus spp.) which are most suited to satisfying household food needs, reduced labor supply, increase in non-farm income in the central region which reduced farmers’ need for cash income generated from farm production, and other socioeconomic considerations (Bagamba et al. 2010; Gold et al. 1999). The use of resistant banana cultivars is regarded as the most feasible solution to the biotic production constraints (Nowakunda and Tushemereirwe 2004).
The improvement of EAHB through crossbreeding involves crossing of the landrace germplasm (triploids) with diploid males (crop wild relative, cultivars or breeding clones) having target trait(s). Due to the triploid nature of the landraces, their cross with diploids yields progeny with different ploidy levels. The tetraploids generated with good bunch characteristics are selected and used as females in a subsequent cross with improved diploid male. This cross results in secondary triploid hybrids that are the aim for further release to farmers after a series of evaluations for target traits. However, this process requires a clear understanding of morphological and phenological characteristics contributing to high yield potential (Batte et al. 2019). This knowledge is essential in the selection of potential parents for hybrid development, deciding the type of data to be collected in evaluation trials, all of which are crucial in guiding the breeding process. The success of many yield-improvement programs hinges on the accurate selection of parents from which to generate superior breeding populations, and the selection of high-yielding genotypes from early-segregating generations (Nyine et al. 2018). Over the years, plant breeders and physiologists have focused on the identification of simple morphological characters which influence physiological processes determining yield. This can be further aided by the use of DNA markers as selection tools (Sharma et al. 2013). It is important to first establish the cause-effect relationships between the morphological characters and the performance of the crop for for example yield. Path analysis is one way of ascertaining the cause-effect relationships (Akintunde 2012). It involves a series of ‘layered’ multiple regression analyses, which lead to the establishment of the direct and indirect effects of the independent variables to the dependent variable (Nathans et al. 2012; Akintunde 2012).
Genetic gain is an important concept applied in quantitative genetics and plant breeding. It is a metric used to assess if a breeding program makes progress. Moose and Mumm (2008) defined genetic gain as the predicted change in the mean value of a trait within a population that occurs with selection thus, it serves as a simple universal expression for expected genetic improvement. They also defined the genetic gain equation and elaborated its terms in relation to the fundamental parameters of quantitative genetics as shown below.
where ΔG is the genetic gain, h2 is the heritability, σp is the phenotypic variability in the population, i is selection intensity and L is the length of the selection cycle.
Heritability (h2) is the probability that a trait phenotype will be transmitted from parent to offspring. Phenotypic variability in the population (σp) is a measure of the degree of phenotypic variation present in the population. Selection Intensity (i) is the proportion of the population selected as parents for the next generation. Length of selection cycle (L) is the length of time necessary to complete a cycle of selection.
From the genetic gain equation, it is evident that genetic gain increases with an increase in heritability, phenotypic variability in the population, selection intensity and decrease in length of the selection cycle. This information can be based to make informed decisions in resource allocation to achieve breeding objectives.
There are different approaches to estimate genetic gain. Wu et al. (2016), while estimating the performance of seven tree breeding strategies under conditions of inbreeding depression, determined genetic gain accumulated for a given population at a particular generation (g) by calculating the increase of average population genotypic value at that generation (g) and compared it to the mean value of the founders. Ortiz et al. (2002), determined the annual genetic gains of Nordic spring barley breeding over a period of sixty years by performing a linear regression analysis, using the year of release as the independent variable while the dependent variable was the mean phenotypic variation for each specific characteristic within each set of barley cultivars. The absolute rates of genetic improvement were divided by the mean of each characteristic to calculate the annual genetic gains. The absolute rates of genetic gains were obtained from the slope of the regression. This agrees with the CGIAR’s Excellence in Breeding approach that the slope of the regression gives the realized genetic gain. This is a measure of increase in a trait’s genetic value per unit increase in generation time. In our study, we used an approach as per Ortiz et al. (2002), to estimate the genetic gain in Matooke (EAHB) breeding over a period of 20 years.
Cause-effect relationships between the morphological characters and the performance of the crops have been investigated in some crops such as Coffea canephora (Cilas et al. 2006), forest trees like bush mango (Irvingia gabonensis) (Leakey and Page 2006), barley (Hordeum vulgare) (Rasmusson 1987), mango (Mangifera indica) (Dickman et al. 1994) and plantains (Ortiz and Langie 1997), among others. However, no such study has been done on the East African highland bananas. Also, the concept of genetic gain has not been applied in banana breeding especially the East African highland bananas. The objective of this study was therefore to determine the most important growth characteristics contributing to yield in cooking East African highland bananas and derived hybrids and to estimate the genetic gain realized in the East African highland banana breeding in the first 20 years of breeding Matooke bananas in Uganda.
The aim was to identify agronomic traits which contribute significantly to yield with the intention to maximize these in the breeding process and to assess the progress of the East African highland banana breeding programme.
Materials and methods
Location and experimental design
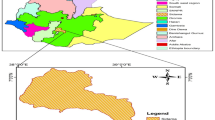
Fifty-six banana genotypes comprising hybrids (secondary triploids), their parents (primary tetraploids and improved diploid breeding clones) and grandparents (triploid matooke cultivars and diploid wild relative) were planted in the field at Namulonge-Sendusu in Uganda (00° 31′ 47″ N and 32o36′ 9″ E), following a 7 × 8 rectangular lattice design with two replications. The plant spacing for the field trials was 3 m between rows and 2 m among plants within a row, thereby having a plant density of 1667 plants ha−1.
Data collection
Data were recorded over three cycles on planting date, date of flowering, height of plant at flowering, number of standing leaves at flowering, youngest leaf with at least 10 necrotic spots at flowering (youngest leaf spotted or YLS), plant girth at 100 cm from the ground, harvest date, bunch weight, number of hands on a bunch, number of fruits on a bunch, fruit length and circumference of the middle fruit on the second hand from the top of the bunch. The plant stature was computed as the ratio of plant girth at 100 cm to plant height at flowering. The number of days to flowering was obtained by counting the number of days from planting or sucker emergence (for ratoons) to the appearance of inflorescence. Number of days to maturity or length of growth cycle were obtained by counting the number of days from planting or sucker emergence (for ratoons) to harvest of the bunch. The days to fruit filling were obtained by counting the number of days from flowering to harvest. The planting dates for the second and third cycles were obtained by recording the dates of sucker emergence from the soil.
The index of non-spotted leaves (INSL) –which indirectly measured host plant resistance to black leaf streak caused by the fungus Pseudocercospora fijiensis (Alakonya et al. 2018) was computed using the formula below:
where INSL is Index of non-spotted leaves, YLS is youngest leaf spotted, NSL is number of standing leaves at flowering.
When the YLS was 0, the above formula was modified in such a way that YLS = NSL + 1.
Data analysis
Path analysis was performed using IBM SPSS version 23 (IBM corporation 2015) on the agronomic and yield parameters to investigate the traits which contribute significantly to bunch weight and thus contributing significantly to yield potential (Ortiz and Langie 1997). Bunch weight was the dependent variable while, plant cycle number, number of hands on a bunch, number of fruits on a bunch, fruit length, fruit circumference, plant stature, number of days to flowering, number of days to maturity, number of days to fruit filling and index of non-spotted leaves were the independent variables. Path coefficients for direct effects and for indirect effects were calculated as follows: (a) the first step was to run a Pearson correlation of all variables together with bunch weight, to identify variables to include in path analysis. Only variables with significant phenotypic correlation (P ≤ 0.05) with bunch weight were selected for path analysis; (b) the second step was to test for multicollinearity—a phenomenon where two independent variables are highly correlated. Multicollinearity has an effect of making a significant variable insignificant by increasing its standard error. If the standard error goes up, the t-value goes down and the probability (P) value goes up, so that particular variable becomes insignificant but in reality, it is not. To achieve this, a Pearson correlation analysis was run using only the independent variables which had significant phenotypic correlation (P ≤ 0.05) with bunch weight. A correlation coefficient of 0.7 and above between two independent variables was regarded to indicate multicollinearity. In order to correct this effect, one of the two highly correlated variables had to be eliminated from the linear regression model. The selection of which variable to eliminate was based on the P-value after a linear regression model was run with all variables included. The variable with a higher P-value was eliminated. Then a linear regression of only the independent variables with no multicollinearity, on bunch weight was run.
Regression analysis on bunch weight was run with only variables that significantly (P ≤ 0.05) contributed to bunch weight, to get the direct effects (Table 1). The indirect effects on bunch weight were obtained by running regression models using variables that did not contribute to direct effects (number of days to maturity, number of days to fruit filling and Index of non-spotted leaves) on the variables with direct effects on bunch weight namely: number of fruits on bunch; plant cycle number; fruit length and number of hands on the bunch (Table 2). Error variance for the dependent variable was also calculated from the formula:
where R2 is the coefficient of determination, i.e. the square value of the correlation coefficient (r).
Genetic gain was estimated after a regression of selection cycle (C0 for landraces, C1 for their derived primary tetraploid hybrids and C2 for secondary triploid bred-germplasm) on bunch weight, number of days to maturity and yield potential was done, the slope of which was divided by the mean value of the trait considered and then divided by selection cycle period, all multiplied by 100 (Ortiz et al. 2002).
Yield potential (t ha−1 year−1) was defined as:
where YLD is yield potential (t ha−1 year−1) (Ortiz and Langie 1997).
Results and discussion
Agronomic traits with direct and indirect effects on yield
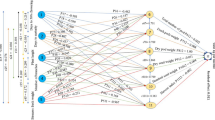
Plant stature was not significantly correlated to bunch weight (r = 0.003, P = 0.917) and therefore was removed from the variables for path analysis. Days to maturity and days to flowering had a high correlation coefficient of 0.86. Days to flowering was eliminated from the model on grounds that its tolerance value was below 0.1, which is indicative of multicollinearity. This implies that in designing agronomic traits in bananas and data to be collected, it is advisable not to include both days to maturity and days to flowering as characters to be measured since one will be redundant. Fruit length, fruit circumference, number of hands on a bunch, number of fruits on a bunch and plant cycle number were found to have a significant (P < 0.01) direct effect on bunch weight (Table 1). Fruit length had the highest direct effect (0.489) followed by number of fruits on a bunch (0.373), among main traits with significant direct effects on bunch weight (Fig. 1).
Path diagram showing traits with direct and indirect effects on bunch weight. FC, fruit circumference; NOHOB, number of hands on bunch; FL, fruit length; NOFB, number of fruits on bunch; Cycle, plant cycle number; DTM, days to maturity; DTFF, days to fruit filling; INSL, index of non-spotted leaves; BW, bunch weight
Ortiz and Langie (1997), investigated the cause-effect relationships between growth parameters and yield potential of six plantain landraces through path analysis. They found out that, characteristics with direct effects on yield potential were, number of fruits on a bunch, and number of hands per bunch in the French plantains ‘Obino l'Ewai’ and ‘Ntanga 2’. This result agrees with our findings. However, they also reported that days to harvest had a direct effect on yield potential in the French plantain ‘Bobby Tannap’, contrary to our findings. They also observed that bunch weight increased with ratooning for cultivars ‘Agbagba’, ‘Big Ebanga’, ‘Obino l'Ewai’ and ‘Ntanga 2’. This is in support of our findings that plant cycle has a direct and positive effect on bunch weight. Swennen and De Langhe (1985) observed that the rate of leaf production was fast in tall plants of ‘Agbagba’, which could be a contributing factor to their height and heavy bunches. Although in our study the plant stature had no effect on the bunch weight, it is desirable to have banana plants with short and robust stature to be able to support the heavy bunches and reduce the chances of falling especially when strong winds blow. They further pointed out that, consumer preference for taste, host plant resistance to pathogens and pests, and tolerance to abiotic stresses in the targeted agro-ecozone should be considered by breeders in developing new cultivars for further release to farmers. Stevens et al. (2020) used allometric studies to forecast yield in the Mchare cultivar ‘Huti-Green Bell’ and a Cavendish cultivar ‘Grande Naine’. They noted that the number of fruits on a bunch, number of hands on a bunch, fruit length and pseudostem volume were significantly correlated with fresh bunch weight and were good predictors of yield in the Mchare and Cavendish cultivar, thus supporting the results of our study.
Days to fruit filling, days to maturity and index of non-spotted leaves had significant indirect effects on bunch weight in our study. Days to fruit filling had an indirect effect on bunch weight (0.124) through the number of fruits on a bunch and through plant cycle number (−0.087) (Table 2). Days to maturity had an indirect effect on bunch weight (0.279) through the number of fruits on a bunch, plant cycle (0.339), fruit length (0.204) and number of hands on a bunch (0.121) (Table 2). Index of non-spotted leaves had an indirect effect on bunch weight (0.108) through the number of fruits on a bunch, plant cycle number (0.062), and fruit length (0.075) (Table 2). The coefficient of determination (R2) for the model defining the direct effects and the F-statistics were 0.639 and 315.956, P < 0.0001 respectively, thereby indicating that the model fits within both the sample and in the population.
The CGIAR Research Program on Roots, Tubers and Bananas (RTB) under the project “Improvement of Banana for Smallholder Farmers in the Great Lakes Region of Africa”, came up with a product profile for East African highland bananas (IITA 2019). Among the traits included is plant stature, which they recommend being at least 0.15, INSL of at least 70%, maturity period of between 300 and 390 days, and yield of 25% greater than that of cultivar ‘Mbwazirume’. The CGIAR’s Excellence in Breeding Platform (CGIAR 2019) also pointed out that a product profile describes a cultivar with the necessary characteristics to replace the less productive old cultivars still grown by farmers in the target population of environments. They, however, noted the low rates of adoption of breeding outputs in farmers’ fields thereby rendering the impact of breeding on poverty alleviation very limited especially in the developing world. They recommended an approach based on product profiles, whereby the new cultivars are focused on client needs rather than being a by-product of scientific research interests, and to incorporate cross-functional knowledge, such as breeding, market, crop management, socioeconomics, gender and farmer or consumer expertise.
It is necessary therefore to select as donor parents East African highland banana and their derived primary tetraploid parent without severe defects in agronomic fitness (like small bunches). Interrelationships among the traits and compensation among plant parts should be considered (Marshall 1991). Ying et al. (1998) reported that an increase in overall biomass production can break the negative relationship among traits and result in an improvement in yield potential. The development of a new cultivar may require concurrent modification of crop management such as fertilizer application, irrigation regime, and weed control in order to fully express its yield potential.
The mean number of hands (fruit clusters) for landraces used to produce primary tetraploids in our study was 8, while the mean number of hands of primary tetraploids that generated secondary triploid hybrids was 7, and the mean number of hands for the secondary triploid hybrids that were generated was 10. The mean number of fruits on bunch for landraces used to produce primary tetraploids was 106, while the mean number of fruits on bunch of primary tetraploids that generated secondary triploid hybrids was 126, and the mean number of fruits on bunch for the secondary triploid hybrids that were generated was 168. The mean fruit circumference for landraces used to produce primary tetraploids was 11 cm, while the mean fruit circumference of primary tetraploids that generated secondary triploid hybrids was 12 cm, and the mean fruit circumference for the secondary triploid hybrids that were generated was 12 cm. The mean fruit length for landraces used to produce primary tetraploids was 16 cm, while the mean fruit length of primary tetraploids that generated secondary triploid hybrids was 15 cm, and the mean fruit length for the secondary triploid hybrids that were generated was 20 cm. The mean bunch weight for landraces used to produce primary tetraploids was 11 kg, while the mean bunch weight of primary tetraploids that generated secondary triploid hybrids was 13 kg, and the mean bunch weight for the secondary triploid hybrids that were generated was 20 kg. The above shows that there was a progressive increase in yield as reflected in the main yield determining characteristics, from Matooke –landraces through their derived primary tetraploid hybrid parents to the secondary triploid hybrids– targeted for release to farmers.
The path coefficients for fruit circumference, number of hands on bunch, fruit length, number of fruits on bunch and cycle number were positive, thereby implying that bunch weight increases with increase in number of hands on bunch, number of fruits on bunch, fruit length and circumference, and cycle number of the banana plant. Large bunches are expected from a mat as cycles increase, therefore short cycling cultivars are preferred since they lead to more harvests in a short period. Tushemereirwe et al. (2015) noticed that bunch weights for NARITA cultivars (name used to refer to secondary triploid bred-germplasm, jointly bred by the National Agricultural Research Organization and the International Institute of Tropical Agriculture in Uganda) from cycle one (plant crop) were significantly lower than those of cycle two and three (first and second ratoons, respectively). They further recommended that selections of East African highland banana derived secondary triploid hybrids for bunch weight should be done in the first ratoon, which is in agreement with the results from this study.
Genetic gains
From the regression equation for cycle of selection on yield potential (Y = 13.28 + 5.95x) with an R2 of 0.199, a unit increase in selection cycle led to an increase in yield potential of 6 tons per hectare per year, while for bunch weight (Y = 10.14 + 4.88x, R2 of 0.187), it indicated that a unit increase in selection cycle resulted in an increase of 4.9 kg of a banana bunch. For both cases above, the P values were significantly different (P < 0.001). The regression of selection cycle on days to maturity (Y = 4.72E2 + 8.5x), had R2 = 0.003. However, the P-value (P = 0.104) was not significant at P ≤ 0.05, indicating that changes in selection cycles did not significantly influence the maturity period of the bananas (Fig. 2). The selection cycle time was 10 years, implying that from landrace germplasm (C0) to primary tetraploid parent (C1) it took 10 years and from the primary tetraploid parent (C1) to the secondary triploid bred-germplasm (C2) it also took 10 years. The mean bunch weight for landraces, primary tetraploid hybrids and secondary triploid bred-germplasm was 17.8 kg, whereas the mean yield for landraces, primary tetraploid hybrids and secondary triploid bred-germplasm was 22.6 tons per hectare per year. The average genetic gain (from landraces (C0) to secondary triploid bred-germplasm (C2)) for bunch weight was [1.4% per year, while the average genetic gain per year for yield potential was1.3%. This implies therefore that there were significant increases of bunch weight (kg plant−1) and yield potential (t ha−1 year−1) from landrace germplasm (C0) to secondary triploid bred-germplasm (C2). However, these values were computed based on data for three generations (C0, C1 and C2). In bananas, obtaining such information takes quite a long time and requires large fields, which makes it very costly. According to the results from this study, the landrace germplasm, the derived primary tetraploid hybrids and secondary triploid bred germplasm were not significantly different in terms of maturity period. Hence the breeding program should consider using in the crossing block parents having short maturity period among other traits, to further increase the genetic gains. Peltonen-Sainio & Karjalainen (1991) used long-term cultivar trial data to estimate the genetic gain in spring cereals in Finland during a period from 1920 to 1988. The genetic gains were evaluated by comparing widely grown cultivars with long-time check cultivars. They found out that the average annual genetic gain for yield of oats was 0.49–0.56% and for spring wheat 0.38–0.45%, while for barley, no clear yield increase was noticed. These results indicate that plant breeding successfully increased oat and spring wheat yields in Finland, but barley breeding was less productive. CGIAR’s Excellence in Breeding suggests that an annual genetic gain of 1% and above indicates that the breeding program is doing well. Here we demonstrate that fruit length, fruit circumference, number of fruits, number of hands and plant cycle number are the main traits contributing to bunch weight (yield) in bananas.
Conclusions
From our study, it was revealed that the main traits contributing to bunch weight (which is a proxy of yield) in East African highland bananas are fruit length, number of fruits on a bunch, fruit circumference and number of hands on a bunch. Bunch weight also increased with increase in plant cycle number. Hence these traits should be considered while selecting parents to be used in breeding for yield in East African highland bananas to ensure that they are passed on to the new hybrids targeted for release to farmers. However, it is important to study and elucidate the inheritance patterns of these traits in order to design appropriate crossing strategies to meet the breeding objectives. This has been the first time to study the cause-effect relationships between growth parameters and yield (bunch weight/ kg) through path analysis in East African highland bananas.
There was a significant increase in bunch weight (kg plant−1) and yield potential (t ha−1 year−1) from the cultigen matooke (C0), their derived primary tetraploid hybrids (C1) and secondary triploid bred-germplasm (C2) with genetic gains of 1.4% and 1.3%, respectively over 20 years. These genetic gains are reasonable but perhaps can still be improved. Targeting hybrids with a short maturity period by using early maturing parents in crosses is one approach to further increase the genetic gains since according to the results from the studies, the landrace germplasm, the derived primary tetraploid hybrids and secondary triploid bred-germplasm were not significantly different in terms of maturity period. Shortening the selection cycle by integrating marker-assisted breeding can also lead to increased genetic gain. This is the first report about genetic gain estimates in a banana breeding program.
Data availability
The datasets generated and analyzed during the current study are available from the corresponding author on reasonable request.
References
Akintunde AN (2012) Path analysis step by step using Excel. J Techni Scie Technol 1:9–15
Alakonya AE, Kimunye J, Mahuku G, Amah D, Uwimana B, Brown A, Swennen R (2018) Progress in understanding Pseudocercospora banana pathogens and the development of resistant Musa germplasm. Plant Pathol 67:759–770. https://doi.org/10.1111/ppa.12824
Bagamba F, Burger K, Tushemereirwe WK (2010) BANANA (MUSA SPP.) PRODUCTION CHARACTERISTICS AND PERFORMANCE IN UGANDA. Acta Hortic 879:187–198. https://doi.org/10.17660/ActaHortic.2010.879.17
Batte M, Swennen R, Uwimana B, Akech V, Brown A, Tumuhimbise R, Hovmalm HP, Mulatu Geleta M, Ortiz R (2019) Crossbreeding East African highland bananas: lessons learnt relevant to the botany of the crop after 21 years of genetic enhancement. Front Plant Sci 10:81. https://doi.org/10.3389/fpls.2019.00081
CGIAR (2019) Product profiles are a blueprint for breeding with impact. CGIAR Excellence in Breeding Platform, Texcoco, Mexico. http://www.excellenceinbreeding.org/blog/product-profiles-are-blueprint-breeding-impact (Accessed on 14th February 2019)
Cilas C, Bar-Hen A, Montagnon C, Godin C (2006) Definition of architectural ideotypes for good yield capacity in Coffea canephora. Ann Bot 97:405–411. https://doi.org/10.1093/aob/mcj053
Dickman DI, Gold MA, Flore JA (1994) The ideotype concept and the genetic improvement of tree crops. Plant Breed Rev 12:163–193
Gold CS, Karamura EB, Kiggundu A, Bagamba F, Abera AMK (1999) Geographic shifts in the highland cooking banana (Musa spp., group AAA-EA) production in Uganda. Int J Sustain Dev World Ecol 6:49–59. https://doi.org/10.1080/13504509.1999.9728471
IBM Corporation Released (2015) IBM SPSS Statistics for Windows Version 23.0. IBM Corp, Armonk, New York
IITA (2019) Breeding Better Bananas Work Package 1. International Institute of Tropical Agriculture, Arusha, Tanzania. http://breedingbetterbananas.org/index.php/2017/01/24/work-package-1-banana-breeding-pipeline/(Accessed on 14th February 2019)
Karamura EB, Turyagyenda FL, Tinzaara W, Blomme G, Ssekiwoko F, Eden-Green SJ, Molina A, Markham R (2008) Xanthomonas wilt of bananas in East and Central Africa: Diagnostic and Management Guide. Bioversity International, Kampala, Uganda
Leakey RRB, Page T (2006) The ‘ideotype concept’ and its application to the selection of cultivars of trees providing agroforestry tree products. For Trees Live 16:5–16. https://doi.org/10.1080/14728028.2006.9752542
Marshall DR (1991) Alternative approaches and perspectives in breeding for higher yields. Field Crops Res 26:171–190. https://doi.org/10.1016/0378-4290(91)90034-S
Moose SP, Mumm RH (2008) Molecular plant breeding as the foundation for 21st Century Crop Improvement. Plant Physiol 147:969–977. https://doi.org/10.1104/pp.108.118232
Nathans LL, Frederick OL, Kim N (2012) Interpreting multiple linear regression: A guidebook of variable importance. Pract Assess Resear Eval 17(9) http://pareonline.net/getvn.asp?v=17&n=9
Nowakunda K, Tushemereirwe W (2004) Farmer acceptance of introduced banana genotypes in Uganda. Afr Crop Sci J 12:1–6. https://doi.org/10.4314/acsj.v12i1.27656
Nyine M, Uwimana B, Blavet N, Hribova E, Vanrespaille H, Batte M, Akech V, Brown A, Lorenzen J, Swennen R, Dolezel J (2018) Genomic prediction in a multiploid crop: genotype by environment interaction and allele dosage effects on predictive ability in banana. Plant Genome. https://doi.org/10.3835/plantgenome2017.10.0090
Ochola D, Jogo W, Ocimati W, Rietveld A, Tinzaara W, Karamura D, Karamura E (2013) Farmers awareness and perceived benefits of agro-ecological intensification practices in banana systems in Uganda. Afric J Biotech 12:4603–4613. https://doi.org/10.5897/AJB12.2868
Ortiz R, Nurminiemi M, Madsen S, Odd Arne Rognli OA, Bjørnstad Å (2002) Genetic gains in Nordic spring barley breeding over sixty years. Euphytica 126:283–289. https://doi.org/10.1023/A:1016302626527
Ortiz R, Langie H (1997) Path analysis and ideotypes for plantain breeding. Agron J 89:988–994. https://doi.org/10.2134/agronj1997.00021962008900060023x
Peltonen-Sainio P, Karjalainen R (1991) Genetic Yield Improvement of Cereal Varieties in Northern Agriculture since 1920. Acta Agriculturae Scandinavica 41(3):267–273. https://doi.org/10.1080/00015129109439909
Rasmusson DC (1987) An evaluation of ideotype breeding. Crop Sci 27:1140–1146. https://doi.org/10.2135/cropsci1987.0011183X002700060011x
Sharma D, Sanghera GS, Sahu P, Sahu P, Parikh M, Sharma B, Bhandarkar S, Chaudhari PR, Jena BK (2013) Tailoring rice plants for sustainable yield through ideotype breeding and physiological interventions. Afr J Agric Res 8:5004–5019. https://doi.org/10.5897/AJAR2013.7499
Stevens B, Diels J, Brown A, Bayo S, Ndakidemi PA, Swennen R (2020) Banana Biomass Estimation and Yield Forecasting from Non-Destructive Measurements for Two Contrasting Cultivars and Water Regimes. Agronomy 10:1435. https://doi.org/10.3390/agronomy10091435
Swennen R and De Langhe E (1985) Growth parameters of yield of plantain (Musa cv. AAB). Ann Bot 56:197–204. https://www.jstor.org/stable/42764227. Accessed: 03–09–2019 08:50 UTC
Tushemereirwe WK, Holderness M, Gold CS, Deadman M (2000) Effects of the leaf spot complex and leaf pruning on growth and yield in highland bananas: Results of the first ratoon crop. Acta Hortic 450:335–341. https://doi.org/10.17660/ActaHortic.2000.540.39
Tushemereirwe W, Batte M, Nyine M, Tumuhimbise R, Barekye A, Ssali T, Talengera D, Kubiriba J, Lorenzen J, Swennen R, Uwimana B (2015) Performance of NARITA hybrids in the preliminary yield trial for three cycles in Uganda. NARO—IITA, Kampala, Uganda
Ying J, Peng S, He Q, Yang H, Yang C, Visperas RM, Cassman KG (1998) Comparison of high yield rice in a tropical and subtropical environment. I. Determinants of grain and dry matter yields. Field Crops Res 57:71–84. https://doi.org/10.1016/S0378-4290(98)00077-X
Wu HX, Hallingbäck HR, Sánchez L (2016) Performance of seven tree breeding strategies under conditions of inbreeding depression. G3 (genes Genome and Genetics) 6:529–540
Acknowledgements
This work was part of the PhD research of the first author (https://pub.epsilon.slu.se/16129/7/batte_m_190509.pdf), with funding from the International Institute of Tropical Agriculture coordinated Project “Improvement of Banana for Smallholder Farmers in the Great Lakes Region of Africa.” The authors would furthermore like to thank all donors who supported this work through their contributions to the CGIAR Fund (https://www.cgiar.org/funders/) and in particular to the CGIAR Research Program for Roots, Tubers and Bananas (CRP-RTB).
Funding
The funding of this work was by the International Institute of Tropical Agriculture and the GIAR Fund (https://www.cgiar.org/funders/).
Author information
Authors and Affiliations
Contributions
MB corresponding author, wrote the first manuscript draft, analyzed data; RS planned the study, reviewed the manuscript; BU planned the study, reviewed the manuscript; VA collected and cleaned data, reviewed the manuscript; AB planned the study, reviewed the manuscript; MG planned the study, reviewed the manuscript; RO planned the study, advised in data analysis, reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Consent to participate
All authors agreed to participate in this work willingly.
Consent for publication
All authors agreed to have this paper published in this journal by open access.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Batte, M., Swennen, R., Uwimana, B. et al. Traits that define yield and genetic gain in East African highland banana breeding. Euphytica 217, 193 (2021). https://doi.org/10.1007/s10681-021-02920-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-021-02920-x