Abstract
Allopolyploidization plays the special role in the evolution of many crops. Moreover, the evolution in early stage of some allopolyploidization events is predicted to be effected by nuclear-cytoplasmic interactions. Maize and teosintes are well model system for study of genetic recombination in allopolyploidization. In order to investigate the effects of genome organization and cytoplasm on genome evolution in newly synthesized allopolyploids (neoallopolyploids), a series of neoallopolyploids were produced by reciprocal crosses of maize and Zea perennis. By using dual-color genomic in situ hybridization, intra- and intergenomic meiosis pairing of these polyploids were quantified and compared with regard to its genome organization and cytoplasm background. In the four neoallopolyploids, the stability of maize genome is consistently lower than that of Z. perennis genome. Additional, the stability of maize genome is affected by genome ploidy. The cytoplasm, genome composition and their interaction do have the special role in chromosome paring and the meiosis behaviors in Zea allopolyploids vary significantly and showed non-diploidization. Z. perennis cytoplasm may give a relatively relaxed environment for maize genome.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Polyploid refers to the presence of three or more chromosome sets in one nucleus, is wide spread in plants and considered to be a major force in plant evolution (Comai 2000; Lavania 2013). Recent studies show that all of flowering plants undertake one or more ancient genome doubling events (Wood et al. 2009; Conant et al. 2014; Woodhousea et al. 2014; Moghe and Shiu 2014). Additional, there are also high frequency of polyploid events among most important crops such as maize, rape, coffee, cotton, wheat and so on (Hilu 1993; Chen 2007; Matsushita et al. 2012). Otherwise, as a result of ancient and recent polyploid events, it is interesting to study what happens during the process of diploidization following neoallopolyploids (Cheung et al. 2009).
In some neoallopolyploids, comparing with the parents, genome, which is not a simple accumulation, can change fast or in a long time (Mandakova et al. 2010; Schnable et al. 2012). The increased genes and genome dosages in neoallopolyploids can result in genome instability, such as chromosome imbalance, meiosis chaos, sequence loss and gene mutation (Wendel 2000; Levy and Feldman 2002; Liu and Wendel 2002; Chen 2007; Jackson and Chen 2010). Genome organization and cytoplasm background have a significant influence on genome instability in allopolyploids (Cheung et al. 2009, De Storme and Geelen 2013). During wheat polyploidization process, the genome tends to lose specific low copy sequences from D genome, which shows parental tendency and inherent difference of parent genome stability (Ozkan et al. 2001). Similar phenomenon is observed in synthetic brassica hybrids, comparing with A and C genome, B genome has more stable and less pairing variation (Cui et al. 2012). But for Brassica allotetraploids, which have highly differentiated cytoplasm, the genome coming from the male parent was significantly modified, compared to the genome coming from the female parent (Song et al. 1995), and other experimental results obtain the same conclusion (Cui et al. 2012). A possible explanation is that the maternal enzymatic system may treat the male chromosomes as exogenous intruder (Wang et al. 2010; Hu et al. 2013). The degree of cytoplasmic influence is decided by divergent relationship of parent genome in allopolyploid. B. juncea has higher genome change than that in B. napus with the explanation is that A and B genomes have higher degrees of divergence than that between A and C genome (Cui et al. 2013). In addition, some other reports find that genome instability of arabidopsis is controlled by both genetic and environmental factors (Armstrong et al. 2001). Poor homologous synapsis 1 (Phs1) and penetration and arbuscule morphogenesis 1 (Pam1) also play an important role in genome instability (Ronceret et al. 2009; Lukaszewski and Kopecky 2010; Feddermann et al. 2010). As a whole, the genome instability in the early stage of polyploidization is affected by multiple factors, in order to have a further understanding, more experimental models and more experimental methods should be explored.
Meiosis plays an important role in the life cycle of sexual species (Tiang et al. 2012). It insures the genetic stability of genome, increases the genetic diversification of gamete and enhances the adaptability of offspring (Santos et al. 1983; Bogdanov 2003; Bernstein and Bernstein 2010). Studies on meiosis not only address the question of genetic relationship of parental genome but also provide important clues on genome evolution in allopolyploids. Previous researchers have shown that crosses are possible between maize and Z. perennis (Tang et al. 2005). Therefore, allopolyploids which contain whole sets chromosomes of the two species may be created by artificial pollination (Zea allopolyploids). In past years, a series of studies about meiosis of Zea allopolyploids have been conducted, and lead to great advances in our understanding of the relationship between maize and Z. perennis genome, and a hypothesis has been offered that maize has a tetraploid origin and Z. perennis has an octaploid origin (Tito et al. 1991; Naranjo et al. 1994; Poggio et al. 1999; Molina et al. 2012). But, the genome evolution during the process of diploidization following Zea allopolyploid is still cryptic. As mentioned above, genome organization and cytoplasm background do influence the stability of parent genome in some allopolyploids. However, the effects on meiosis behavior of Zea allopolyploid are still uncertain, as well as, whether inherent difference of genome stability exists between male and female genome in Zea allopolyploids.
Here, different allopolyploids between maize and Z. perennis can be synthesized in the laboratory from unequivocal progenitors, which can provide a new efficient model for studying the early evolution of neoallopolyploids. In this study, genomic in situ hybridization (GISH) is used to authenticate the pattern of chromosome paring in a series of Zea allopolyploid hybrids which have maize and Z. perennis cytoplasm, respectively. Thus, this article will put forward a new insight into the influence of genome organization and cytoplasm background on chromosome paring in Zea allopolyploids and give a further understanding of the evolution in early stage of allopolyploidization events.
Materials and methods
Plant material and crosses
Diploid maize wf9 (2n = 20) and tetraploid maize Twf9 (2n = 40) derived from wf9 were provided by USDA, and Z. perennis (9475, 2n = 40) was provided by the International Maize and Wheat Improvement Center (CIMMYT). These materials were grown in the experimental field of Sichuan Agricultural University, Jinghong, China, in 2011. Two pairs of reciprocal crosses were performed by means of hand pollination. In 2012, pre-germination interspecific hybrid seeds were planted in the plastic pots (12 × 12 cm, inner diameter × height) filled with soil and placed in the experimental garden of Sichuan Agricultural University for early identification. Seedlings were transplanted into larger plastic pots (26.5 × 26.5 cm, inner diameter × height) after 5-leaf stage.
Chromosome and DNA preparation
Root tips collected from young seedlings of interspecific hybrids were pretreated in α-bromonaphthalene saturated solution for 2.5–3 h at room temperature, and then fixed in Carnoy’s solution I (3: 1, ethanol: glacial acetic acid, v/v) for at least 24 h, finally, transferred to 70 % ethanol solution and stored at 4 °C. Young anthers collected from tassels of interspecific hybrids were fixed directly in the Carnoy’s solution I for at least 12 h, and then transferred to 70 % ethanol solution and stored at 4 °C.
The treated root tips and anthers were washed in water to remove the ethanol solution, and subsequently transferred to an enzyme solution containing 6 % cellulase (R-10, Yakult, Japan) and 1 % pectinase (Y-23, Yakult, Japan) for 2.5–5.0 h at 37 °C. Root tips and anthers were washed in water again to clean the remaining enzyme solution, and finally squashed onto clean slides in a drop of Carnoy’s solution I and flame dried. Preparations showing well-spread metaphase cells and pollen mother cells (PMC) were selected by phase-contrast light microscopy (Olympus BX-41, Japan) and saved at −20 °C for in situ hybridization procedures. Total DNA was extracted from young leaves of wf9 and 9475 using the improved 2 × CTAB method (Fu et al. 2015).
Genomic in situ hybridization
Wf9 genomic DNA were labeled with DIG-Nick Translation Mix (Roche, Swiss) and 9475 genomic DNA were labeled with BIOTIN-Nick Translation Mix (Roche, Swiss) following the manufacturer’s protocols. The prepared slides were dried in air blowing thermostatic oven for 1 h at 60 °C and then pretreated with 50 ml RNase solution (0.1 μg/ml RNase in 2 × SSC, Solarbio) in thermostat water bath for 1 h at 37 °C, subsequently washed twice in 2 × SSC for 5 min each at room temperature. Chromosomal DNA was denatured for 2.5 min by immersing the slides in 70 % deionized formamide (FAD) in 2 × SSC at 70 °C, and then dehydrated quickly through an pre-cooling 70, 95, and 100 % ethanol series for 5 min each at −20 °C, respectively. The hybridization mixture contained 150 µl 50 % FAD, 60 µl 10 % dextran sulfate (DS), 30 µl 2 × SSC, 15 µl 0.5 % sodium dodecyl sulfate (SDS), 3 µl 10 μg/µl salmon sperm DNA (SSDNA) and 18 µl labeled probes for six slides. The hybridization mixture was denatured in thermostat for 5 min at 85 °C, and then chilled on ice for 10 min quickly. 45 µl denatured hybridization mixture was added on each air dried slide and hybridization was performed in incubator chamber for 18–24 h at 37 °C. After hybridization, slides were washed in 20 % FAD, 2 × SSC and 0.1 × SSC at 42 °C for 15 min each, and then washed in 0.1 % TritonX-100 for 5 min, subsequently washed in 1 × PBS three times for 5 min each at room temperature. The next steps were performed in the dark. After air drying the slides, 50 µl antibody diluent which contained anti-digoxigenin-fluorescein (0.6 µg/µl in 1 × PBS, Roche) and CY-3 fluorescein (0.6 % in 1 × PBS, sigma) were loaded onto slides, and hybridization was performed in incubator chamber for 1 h at 37 °C. Slides were washed with 1 × PBS three times at room temperature for 5 min each. After drying, chromosomes were counterstained with 4′,6-diamidino-2-phenylindole (DAPI), and then observed with fluorescence microscope (Olympus BX-61, Japan). Images were taken with Media Cybernetics CCD 700 (Charge Coupled Device) and Image Pro Plus 6.0 (Media Cybernetics, Inc.). Adobe Photoshop 5.1 was used for image processing.
Analysis of chromosomes composition
In order to analyze the chromosome paring with overall level, the chromosome calculation formula of autosyndesis and allosyndesis were carried out referring to the method of Cheng Cui (Cui et al. 2012) and modified slightly (Fig. 1). Additional, the relative chaotic coefficient (RCC) = (chromosomes number of bivalents)/(total chromosomes number—chromosomes number of bivalents).
The model diagram of chromosome calculation formula. Maize and Z. perennis chromosomes were labeled by green and pink, respectively. MAu represented autosyndesis chromosomes of maize and MAl represented allosyndesis chromosomes of maize, PAu represented autosyndesis chromosomes of Z. perennis and PAl represented allosyndesis chromosomes of Z. perennis
Results
Materials synthesis and cytology identification
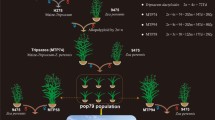
Two pairs of reciprocal crosses between maize and Z. perennis were carried out, and the following allopolyploid hybrids were produced (Fig. 2). Allotriploid MP30 and PM30 were obtained from the reciprocal crosses between wf9 and 9475, which contained the cytoplasm of wf9 and 9475, respectively. While allotetraploid MP40 and PM40 were obtained from the reciprocal crosses between Twf9 and 9475, which contained the cytoplasm of Twf9 and 9475, respectively.
Schematic illustration showing the production of allopolyploid hybrids from pair crosses of three parents (wf9, Twf9, 9475). The light green circle represents maize cytoplasm and the light pink circle represents Z. perennis cytoplasm. The maize and Z. perennis chromosomes are labeled by green strip and pink strip, respectively, and both of them have red centromere in the middle of chromosome
Generally, genome organization of MP30 and PM30 were stable. However, chromosome number instability occurred in MP40 and PM40, mainly due to aneuploidy individuals and abnormal meiosis behavior existed in tetraploid maize population (unpublished). In order to accurately obtain the allopolyploid hybrids with whole sets of chromosomes from parents, dual-color GISH was used to identify the chromosome number and chromosome constitution. Allotriploids (MP30, PM30) with one set of maize (10) chromosomes and one set of Z. perennis (20) chromosomes, and allotetraploids (MP40, PM40) with two sets of maize (20) chromosomes and one set of Z. perennis (20) chromosomes were selected for further meiotic research (Fig. 3), in addition, there were no chromosome recombination detected in the interspecific hybrids.
Chromosome pairing analysis in synthetic allotriploids
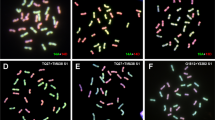
In order to study the chromosome origin of different configuration and analyze the meiotic behavior, especially the differential chromosome paring affected by genome composition and cytoplasm, dual-color GISH was carried out (Figs. 4, 5). Both allotriploids (MP30, PM30) with 30 chromosomes showed irregular meiosis (Fig. 4a, c), lagging chromosomes were observed frequently in meiosis anaphase I (Fig. 4b, d). The most frequent meiotic configuration of MP30 was 5III + 5II + 5I (16.9 %), with an average of 4.56I + 5.44II + 4.73III + 0.07IV, while the most frequent meiotic configuration of PM30 was 4I + 6II + 5III (8.47 %), with an average of 4.04I + 6.13II + 4.31III + 0.20IV (Table 1S). The results were not consistent to previous research (Molina et al. 2012). Most univalents in MP30 and PM30 belonged to the chromosomes from maize (IM), with the frequent number of five, and most bivalents were formed by autosyndetic pairings between homologous chromosomes from Z. perennis (IIPP), with the frequent number of five; most trivalents in MP30 and PM30 were formed by allosyndetic pairings consisted of one maize chromosome and two Z. perennis chromosomes (IIIMPP), with the frequent number of five (Fig. 4a, c; Table 1S). The results suggested that five chromosomes of maize in allotriploids were homologous to ten chromosomes of Z. perennis.
GISH analysis of PMCs in MP30 and PM30. a and b represent PMCs of MP30 at diakinesis and anaphase I, c and d represent PMCs of PM30 at diakinesis and anaphase I. Blue triangle represents maize univalent, white triangle represents autosyndetic bivalent of Z. perennis genome, red triangle represents allosyndetic trivalent which consisted of one maize chromosome and two Z. perennis chromosomes, white arrow represents lagging chromosome. Yellow signals are from maize genome and pink signals are from Z. perennis genome. All bars 10 μm
Cytology and GISH analysis of tetraploid hybrids. a and b represent PMCs of MP40 at diakinesis and anaphase I, c and d represent PMCs of PM40 at diakinesis and anaphase I. Blue triangle represents maize univalent, red triangle represents Z. perennis univalent, tangerine triangle represents autosyndetic bivalent of maize genome, white triangle represents autosyndetic bivalent of Z. perennis genome, purple triangle represents allosyndetic bivalent which consisted of one maize chromosome and one Z. perennis chromosome, yellow triangle represents allosyndetic quadrivalent which consisted of two maize chromosomes and two Z. perennis chromosomes, white arrow represents lagging chromosome. Yellow signals are from maize genome and pink signals are from Z. perennis genome. All bars 10 μm
In MP30 and PM30, the average number of IM (3.79, 2.73) was higher than that of IP (0.77, 1.31), while the average number of IIMM (0.25, 0.58) was lower than that of IIPP (4.34, 4.24) (Table 1). In order to further analyze the meiosis behavior, chromosome pairing within each genome (autosyndesis) and between different genomes (allosyndesis) were quantified and compared. The rates for autosyndesis within maize genome (abbreviated as maize autosyndesis) in MP30 and PM30 were 0.07 and 0.17 respectively, while the rates for autosyndesis within Z. perennis genome (abbreviated as Z. perennis autosyndesis) were 0.71 and 0.69, which were much higher than the former. On the other hand, the rate for allosyndesis between maize and Z. perennis genome (abbreviated as maize allosyndesis) in MP30 (0.56) was identical to that in PM30, while the rates for allosyndesis between maize and Z. perennis genome (abbreviated as Z. perennis allosyndesis) in MP30 and PM30 were 0.25 and 0.26, respectively, which were lower than the former (Table 2). The results suggested that maize genome in allotriploids had lower ability of chromosome paring than that of Z. perennis genome. The RCC of maize genome in MP30 and PM30 (9.00, 9.30) were much higher than that of Z. perennis chromosome (1.56, 1.86) (Table 1), probably due to the different genome ploidy level of two parents.
Quadrivalents were barely seen in MP30 and PM30,and the average number of IM and IIIMPP in MP30 was higher than that in PM30, while the average number of IP, IIMM, IIMP and IIIMMP in MP30 was lower than that in PM30, and the average number of other valent types had no difference between MP30 and PM30 (Table 1). As a whole, the rate for maize autosyndesis in MP30 was lower than that in PM30, but there were no significant difference between MP30 and PM30 in the rates for maize allosyndesis, Z. perennis autosyndesis and Z. perennis allosyndesis (Table 2). The RCC of maize chromosome in meiosis of MP30 was higher than that of PM30, however there was no significant difference between MP30 and PM30 in the RCC of Z. perennis chromosome (Table 1). The results suggested that cytoplasm probably had influence on chromosome paring in allotriploids, and Z. perennis cytoplasm might enhance chromosome paring and stability of maize genome.
Chromosome pairing analysis in synthetic allotetraploids
The allotetraploids (MP40 and PM40) had balanced chromosome numbers of two parent genomes (Fig. 3), and lagging chromosomes were also observed in MP40 (Fig. 5b). However, lagging chromosomes were rarely observed in PM40 (Fig. 5d). The most frequent meiotic configuration of MP40 were 12II + 4IV (15.94 %) and 8II + 6IV (15.94 %), with an average of 1.17I + 9.97II + 0.13III + 4.62IV, while in PM40, the most frequent meiotic configuration was 12II + 4IV (19.70 %), with an average of 0.56I + 11.74II + 0.29III + 3.78IV (Table 1S). The results were not consistent to previous research (Molina et al. 2012). Most bivalents in MP40 and PM40 were formed by autosyndetic pairings within homologous chromosomes from maize (IIMM) or Z. perennis (IIPP), both of them with the frequent number of five; although MP40 and PM40 had the balanced genome, allosyndesis and multivalent still formed. Quadrivalents were common in the two allotetraploids, and most quadrivalents formed by allosyndetic pairings and consisted of two maize chromosomes and two Z. perennis chromosomes (IVMMPP), with the frequent number of five (Fig. 5a, c; Table 1S). The meiosis chromosome configuration in both allotetraploids also suggested ten chromosomes of maize were homologous to ten chromosomes of Z. perennis.
In MP40 and PM40, the average number of IM (0.81, 0.39) was higher than that of IP (0.36, 0.17), while the average number of IIMM (4.30) in MP40 was lower than that of IIPP (4.71) and the average number of IIMM (4.45) in PM40 was a little higher than that of IIPP (4.41) (Table 1), probably due to the influence of cytoplasm. The rates for maize autosyndesis in MP40 and PM40 were 0.69 and 0.78 respectively, while the rates for Z. perennis autosyndesis were 0.71 and 0.79, and there were no significant difference in maize autosyndesis and Z. perennis autosyndesis between MP40 and PM40. The rates for maize allosyndesis in MP40 and PM40 were 0.27 and 0.20 respectively, and the rates for Z. perennis allosyndesis were also 0.27 and 0.20, additional, the rates for maize and Z. perennis autosyndesis were both higher than that of maize and Z. perennis allosyndesis (Table 2). The RCC of maize genome (1.60, 1.05) in PM40 and PM40 was a little higher than that of Z. perennis genome (1.47, 1.00) (Table 1), suggested the stability of both genomes were enhanced, especially for maize genome. However, Z. perennis genomes in both allotetraploids were also more stable than maize genome.
The average number of IM, IP and IVMMPP in PM40 was lower than that in MP40, while the average number of IIMM, IIPP and IVPPPP in PM40 was higher than that in MP40 and there were no significant difference on other valent types (Table 1). As a whole, the rates for maize autosyndesis and Z. perennis autosyndesis in PM40 were higher than that in MP40, but the rates for maize autosyndesis and Z. perennis autosyndesis in PM40 showed an opposite situation (Table 2). Additional, comparing with MP40, the stability of maize and Z. perennis genome were both enhanced in PM40, and the average of RCC in PM40 was lower than that in MP40 (Table 1). The results suggested Z. perennis cytoplasm in allotetraploids could enhance the stability of both parent genomes and the ability of homologous paring in both genomes.
Discussion
Maize, Z. perennis and their hybrids is a good model system for allopolyploid study
Allopolyploid, which plays a crucial role in plant speciation, probably undergoes genome change after formed (Mandakova et al. 2010). Interestingly, in the newly synthesized allopolyploid, corresponds to a nature one, genome will change rapidly, if there is no corresponding nature allopolyploid, whereas genome will change slowly (Comai 2000). The process of genome changes can be best researched in the newly synthetic allopolyploids or the nature one with unambiguous source. As one of the main crops in the word, maize and closely related species which called teosintes become a well genetic model in the world (Doebley et al. 2006; Weber et al. 2007; Vigouroux et al. 2008; Van Heerwaarden et al. 2010, 2011; Wills et al. 2013). In this study, reciprocal crosses were successfully made between maize (diploid or tetraploid) and Z. perennis which is the polyploid species in teosintes. These materials which contain different ploid and different cytoplasm will be a good model system to study the genome changes, chromosomal behaviors as well as genome affinities, and probably provide a further insight into the genome of maize and Z. perennis.
Different genome effects on meiotic pairing
In allotriploids, haploid chromosomes of maize could not pair themselves, maize univalents and maize allosyndesis were frequently observed in chromosome synapsis at meiosis I. While in allotetraploids, with diploid chromosomes of maize, there were more bivalents and the rate of maize autosyndesis was improved. Different ploidy levels did influence the meiosis behavior especially in chromosome pairing, and balanced parental chromosome number of allotetraploid may enhance the ability of proper chromosome pairing in meiosis.
Two or more genomes are sharing one nucleus in allopolyploids. For some allopolyploids, new genetic combinations are created which are beyond the addition of parent genomes and show non-Mendelian inheritance (Levy and Feldman 2002). In some allopolyploids, the different genomes from parents may occur divergent changes and different degrees of diplodization (Liu and Wendel 2002), probably due to the discrepant stability of parental genome. In this study, Z. perennis genome is more stable than that of maize in four allopolyploids, and possible explanations for our observations include: (a) the phylogenetic analysis shows that Z. perennis is more primitive than maize (Wang et al. 2011), therefore, the inherent stability probably exist in Z. perennis genome as a result from long-term evolution; (b) recent studies demonstrate maize genome has tetraploid origin (Gonzalez and Poggio 2011), “A” and “B” genome probably exist partial homology (Molina et al. 2012); (c) for the two allotriploids, the haploid genome of maize can hardly perform the homologous chromosome paring; (d) different genomes in one nucleus may lead to the disordered expression of Phs1 and Pam1, which play important roles on chromosome paring (Molina et al. 2012).
Although the genome stability is improved in allotetraploids, allosyndetic valents also exist and the most frequent allosyndetic valent is IVMMPP, with the frequent number of five (Table 1S). Additional, the most frequent allosyndetic valent in allotriploids is IIIMPP, with the frequent number of five (Table 1S). All of the results suggest that high level of homology exists between maize and Z. perennis and ten chromosomes from maize genome are homologous to twenty chromosomes from Z. perennis genome, additional, the homologous chromosomes belong to “A” genome (Gonzalez et al. 2006; Swanson-Wagner et al. 2010; Molina et al. 2012).
Nuclear-cytoplasmic interactions effects on meiotic pairing
Cytoplasm is predicted to play a vital role in meiosis behavior (Cifuentes et al. 2010). In this study, although a close relationship exists between maize and Z. perennis (Tang et al. 2005), chromosome paring is significantly different in two allotriploids or two allotetraploids, which reveals parental cytoplasm has distinct contribution for meiosis behavior. Comparing MP30 with PM30, Z. perennis cytoplasm in PM30 can reduce the number of maize univalents and improve the maize autosyndetic bivalents, thus, enhancing the stability of meiosis pairing in maize genome. However, Z. perennis cytoplasm has little influence on Z. perennis genome and only increases the number of univalents in PM30. Comparing MP40 with PM40, Z. perennis cytoplasm in PM40 can reduce the number of univalents and allosyndesis, improve the number of autosyndesis, enhance the stability of meiosis pairing in both maize and Z. perennis genome. The results suggest Z. perennis cytoplasm may give a relatively relaxed environment especially for maize genome. The possible reasons include: (a) the presence of foreign nuclear genome in cytoplasm from the male parent can result in nuclear-cytoplasm incompatibility (Wang et al. 2010; Cui et al. 2012) and maternal enzymatic system may treat the male chromosomes as exogenous intruders (Hu et al. 2013), thus, gene structure variation, aneuploidy and chromosomes structure variation as well as meiosis confusion can potentially occur, comparing to Z. perennis cytoplasm, maize cytoplasm probably has a stronger defense system; (b) Z. perennis is more primitive than maize (Wang et al. 2011), and the cytoplasm may have a greater containment than maize; (c) Z. perennis cytoplasm probably has a less negative influence on Phs1 and Pam1 than maize cytoplasm. Additional, interactions can occur between nuclear and cytoplasm followed by loss of genes from both nuclear genome and plasmon (Rand et al. 2004), which suggests genome stability also may be affected by interactions between different genome composition and cytoplasm (Cui et al. 2012), different results in allotriploids and allotetraploids conform to this speculation.
References
Armstrong SJ, Franklin FC, Jones GH (2001) Nucleolus-associated telomere clustering and pairing precede meiotic chromosome synapsis in Arabidopsis thaliana. J Cell Sci 114:4207–4217
Bernstein H, Bernstein C (2010) Evolutionary origin of recombination during meiosis. Bioscience 60:498–505
Bogdanov YF (2003) Variation and evolution of meiosis. Russ J Genet 39:453–473
Chen ZJ (2007) Genetic and epigenetic mechanisms for fene expression and phenotypic variation in plant polyploids. Annu Rev Plant Biol 58:377–406
Cheung F, Trick M, Drou N, Lim YP, Park JY, Kwon SJ, Kim JA, Scott R, Pires JC, Paterson AH, Town C, Bancroft L (2009) Comparative analysis between homeologous genome segments of brassica napus and its progenitor species reveals extensive sequence-level divergence. Plant Cell 21:1912–1928
Cifuentes M, Eber F, Lucas MO, Lode M, Chevre AM, Jenczewski E (2010) Repeated polyploidy drove different levels of crossover suppression between homoeologous chromosomes in brassica napus allohaploids. Plant Cell 22:2265–2276
Comai L (2000) Genetic and epigenetic interactions in allopolyploid plants. Plant Mol Biol 43:387–399
Conant GC, Birchler JA, Pires JC (2014) Dosage, duplication, and diploidization: clarifying the interplay of multiple models for duplicate gene evolution over time. Curr Opin Plant Biol 19:91–98
Cui C, Ge X, Gautam M, Kang L, Li Z (2012) Cytoplasmic and genomic effects on meiotic pairing in Brassica hybrids and allotetraploids from pair crosses of three cultivated diploids. Genetics 191:725–738
Cui C, Ge X, Zhou Y, Li M, Li Z (2013) Cytoplasmic and genomic effects on non-meiosis-driven genetic changes in Brassica hybrids and allotetraploids from pairwise crosses of three cultivated diploids. PLoS One 8:e65078
De Storme N, Geelen D (2013) Cytokinesis in plant male meiosis. Plant Signal Behav 8:e23394
Doebley JF, Gaut BS, Smith BD (2006) The molecular genetics of crop domestication. Cell 127:1309–1321
Feddermann N, Muni RR, Zeier T, Stuurman J, Ercolin F, Schorderet M, Reinhardt D (2010) The PAM1 gene of petunia, required for intracellular accommodation and morphogenesis of arbuscular mycorrhizal fungi, encodes a homologue of VAPYRIN. Plant J 64:470–481
Fu J, Yang XY, Cheng MJ, LÜ GH, Wang P, Wu YQ, Zheng MM, Zhou SF, Rong TZ, Tang QL (2015) Perennial aneuploidy as a potential material for gene introgression between maize and Zea perennis. J Integr Agric 14:839–846 (in Chinese)
Gonzalez GE, Poggio L (2011) Karyotype of Zea luxurians and Z. mays subsp. mays using FISH/DAPI, and analysis of meiotic behavior of hybrids. Genome 54:26–32
Gonzalez G, Comas C, Confalonieri V, Naranjo CA, Poggio L (2006) Genomic affinities between maize and Zea perennis using classical and molecular cytogenetic methods (GISH-FISH). Chromosome Res 14:629–635
Hilu K (1993) Polyploidy and the evolution of domesticated plants. Am J Bot 80:1494–1499
Hu LP, Huang XT, Mao JX, Wang CD, Bao ZM (2013) Genomic characterization of interspecific hybrids between the scallops Argopecten purpuratus and A. irradians irradians. PLoS One 8:e62432
Jackson S, Chen ZJ (2010) Genomic and expression plasticity of polyploidy. Curr Opin Plant Biol 13:153–159
Lavania UC (2013) Polyploidy, body size, and opportunities for genetic enhancement and fixation of heterozygosity in plants. Nucleus 56:1–6
Levy AA, Feldman M (2002) The impact of polyploidy on grass genome evolution. Plant Physiol 130:1587–1593
Liu B, Wendel JF (2002) Non-mendelian phenomena in allopolyploid genome evolution. Curr Genomics 3:1–17
Lukaszewski AJ, Kopecky D (2010) The Ph1 locus from wheat controls meiotic chromosome pairing in autotetraploid rye (Secale cereale L.). Cytogenet Genome Res 129:117–123
Mandakova T, Joly S, Krzywinski M, Mummenhoff K, Lysak MA (2010) Fast diploidization in close mesopolyploid relatives of Arabidopsis. Plant Cell 22:2277–2290
Matsushita SC, Tyagi AP, Thornton GM, Pires JC, Madlung A (2012) Allopolyploidization lays the foundation for evolution of distinct populations: evidence from analysis of synthetic arabidopsis allohexaploids. Genetics 191:535–547
Moghe GD, Shiu SH (2014) The causes and molecular consequences of polyploidy in flowering plants. Ann NY Acad Sci 1320:16–34
Molina MC, López CG, Staltari S, Chorzempa SE, Moreno FV (2012) Cryptic homoeology analysis in species and hybrids of genus Zea. Biol Plant 57:449–456
Naranjo CA, Poggio L, Molina MDC, Bernatene EA (1994) Increase in multivalent frequency in F1 hybrids of Zea diploperennis × Z. perennis by colchicine treatment. Hereditas 120:241–244
Ozkan H, Levy AA, Feldman M (2001) Allopolyploidy-induced rapid genome evolution in the wheat (Aegilops–Triticum) group. Plant Cell 13:1735–1747
Poggio L, Confalonieri V, Comas C, Gonzalez G, Naranjo CA (1999) Genomic affinities of Zea luxurians, Z. diploperennis, and Z. perennis: meiotic behavior of their F1 hybrids and genomic in situ hybridization (GISH). Genome 42:993–1000
Rand DM, Haney RA, Fry AJ (2004) Cytonuclear coevolution: the genomics of cooperation. Trends Ecol Evol 19:645–653
Ronceret A, Doutriaux MP, Golubovskaya IN, Pawlowski WP (2009) PHS1 regulates meiotic recombination and homologous chromosome pairing by controlling the transport of RAD50 to the nucleus. PNAS 106:20121–20126
Santos JL, Orellana J, Giraldez R (1983) Pairing competition between identical and homologous chromosomes in rye and grasshoppers. Genetics 104:677–684
Schnable JC, Freeling M, Lyons E (2012) Genome-wide analysis of syntenic gene deletion in the grasses. Genome Biol Evol 4:265–277
Song K, Lu P, Tang K, Osborn TC (1995) Rapid genome changes in synthetic polyploids of Brassica and its implications for polyploid evolution. Proc Natl Acad Sci USA 92:7719–7723
Swanson-Wagner RA, Eichten SR, Kumari S, Tiffin P, Stein JC, Ware D, Springer NM (2010) Pervasive gene content variation and copy number variation in maize and its undomesticated progenitor. Genome Res 20:1689–1699
Tang QL, Rong TZ, Song YC, Yang JP, Pan GT, Li WC, Huang YB, Cao MJ (2005) Introgression of perennial teosinte genome into maize and identification of genomic in situ hybridization and microsatellite markers. Crop Sci 45:717–721
Tiang CL, He Y, Pawlowski WP (2012) Chromosome organization and dynamics during interphase, mitosis, and meiosis in plants. Plant Physiol 158:26–34
Tito CM, Poggio L, Naranjo CA (1991) Cytogenetic studies in the genus zea : 3. DNA content and heterochromatin in species and hybrids. Theor Appl Genet 83:58–64
Van Heerwaarden J, Ross-Ibarra J, Doebley J, Glaubitz JC, Gonzalez JJ, Gaut BS, Eguiarte LE (2010) Fine scale genetic structure in the wild ancestor of maize (Zea mays ssp. parviglumis). Mol Ecol 19:1162–1173
Van Heerwaarden J, Doebley J, William HB, Glaubitzd JC, Goodmane MM, Jesus JD, Sanchez G, RossI-barraa J (2011) Genetic signals of origin, spread, and introgression in a large sample of maize landraces. PNAS 108:1088–1092
Vigouroux Y, Glaubitz JC, Matsuoka Y, Goodman MM, Sanchez GJ, Doebley J (2008) Population structure and genetic diversity of New World maize races assessed by DNA microsatellites. Am J Bot 95:1240–1253
Wang S, Zhang L, Hu J, Bao Z, Liu Z (2010) Molecular and cellular evidence for biased mitotic gene conversion in hybrid scallop. BMC Evol Biol 10:6
Wang P, Lu YL, Zheng MM, Rong TZ, Tang QL (2011) RAPD and internal transcribed spacer sequence analyses reveal Zea nicaraguensis as a section Luxuriantes species close to Zea luxurians. PLoS One 6:e16728
Weber A, Clark RM, Vaughn L, Sanchez-Gonzalez Jde J, Yu J, Yandell BS, Bradbury P, Doebley J (2007) Major regulatory genes in maize contribute to standing variation in teosinte (Zea mays ssp. parviglumis). Genetics 177:2349–2359
Wendel JF (2000) Genome evolution in polyploids. Plant Mol Biol 42:225–249
Wills DM, Whipple CJ, Takuno S, Kursel LE, Shannon LM, RossIbarra J, Doebley JF (2013) From many, one: genetic control of prolificacy during maize domestication. PLoS Genet 9:e1003604
Wood TE, Takebayashi N, Barker MS, Mayrose I, Greenspoon PB, Rieseberg LH (2009) The frequency of polyploid speciation in vascular plants. PNAS 106:13875–13879
Woodhousea MR, Cheng F, Pires JC, Lischa D, Freelinga M, Wang X (2014) Origin, inheritance, and gene regulatory consequences of genome dominance in polyploids. PNAS 111:5283–5288
Acknowledgments
This work was supported by the National Program on Key Basic Research Project of China (973 Program, 2014CB138705) and the National Natural Science Foundation of China (31371640).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Cheng, M., Zheng, M., Yang, S. et al. The effect of different genome and cytoplasm on meiotic pairing in maize newly synthetic polyploids. Euphytica 207, 593–603 (2016). https://doi.org/10.1007/s10681-015-1552-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-015-1552-7