Abstract
Omega gliadin proteins are one of the most allergenic components of wheat gluten. Proteins of the ω-5 subgroup are recognized as main allergens causing wheat dependent exercise induced anaphylaxis—the most dangerous, life-threatening IgE mediated food allergy. A set of wheat genotypes lacking all ω-gliadins has been developed by cumulating inactive gene variants in three gliadin coding loci (Gli A1, Gli B1 and Gli D1), using traditional plant breeding methods. Endosperm proteins of ω-gliadin-free genotypes were compared to a control genotype containing all ω-gliadins by A-PAGE, SDS-PAGE and RP-HPLC. A considerable decrease (about 30 %) of gliadin immunoreactivity as a consequence of ω-gliadin elimination was demonstrated by ELISA, using sera of ten patients allergic to gluten. Preliminary evaluation of the technological properties of the ω-gliadin-free genotype by the SDS sedimentation test suggests that elimination of all ω-gliadins may also significantly improve wheat bread making quality.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
It is well documented that wheat gliadins—one of the main gluten protein fractions—may also be important allergens involved in pathogenesis of skin, gastrointestinal and respiratory tract diseases. Allergies to wheat may cause, among others diseases, dermatitis herpetiformis, urticaria, food allergy, rhinitis, or asthma (Inomata 2009).
There are close relationships between allergenicity, chemical composition and structures of allergenic proteins. Gliadins are a highly complex and polymorphic group of proteins. Acid polyacrylamide gel electrophoresis (A-PAGE) separates gliadins into four groups (α, β, γ and ω) based on electrophoretic mobility (Bushuk 1993). Alpha and β gliadins have similar protein structures and are often classified as one group of α/β gliadins. On the other hand, slower-moving ω-gliadins are separated into ω-1.2 and ω-5 subgroups (Wieser 1991). Proteins of the α, β, γ and ω groups differ in composition of their polypeptide domains. All gluten proteins contain both repetitive and specific sequence regions. Gliadins of the α/β and γ classes contain about 100 amino acids in repetitive sequences at the N ends and about 200 amino acids in specific sequences at the C ends of the polypeptides (Tatham 1995). The N-terminal sequences of α/β and γ gliadins contain 5 and 12 amino acid residues, respectively, while C-terminal sequences are not clearly separated from the specific region. The ω-gliadins have long, repetitive central regions of more than 200 residues, flanked by short N- and C-terminal sequences of 13 and 10 amino acids, respectively.
Repetitive sequences of gliadin proteins are closely related to their allergenic properties (Sandiford et al. 1997). In IgE mediated allergies a specific IgE class antibodies (sIgE), present in elevated amounts in sera of allergic people, bind with epitopes—short amino acid sequences—of allergenic proteins. Resulting IgE-protein complexes coat surfaces of basofiles causing their degranulation. Basofiles are immunological system cells containing chemical mediators, mainly histamine. Histamine released to the bloodstream increases permeability of blood vessels which in turn causes inflammation and, later, allergic symptoms (Ebo and Stevens 2001). In gluten proteins, epitopes are composed of 4–8 amino acids. Gliadins’ repetitive sequences frequently overlay sequences of epitopes. For example, the most immunoreactive epitopes in ω-gliadins are the heptapeptides QQIPQQQ, QQFPQQQ, and QQSPQQQ, and the tetrapeptide QQQP. These partly overlap commonly observed repetitive sequences in ω-gliadins such as PQQPFPQQ (the motif typical for ω-1.2 gliadins), PQQF, FPQQQ or QQIPQQ (present in ω-5 gliadins) (Wieser 1991; Maruyama et al. 1998; Matsuo et al. 2004). Epitope-like sequences occurring several times in allergenic proteins may increase the efficiency of immunological reaction and, consequently, aggravate allergic symptoms.
Gliadins of the ω-class are the most allergenic among gluten proteins (Battais et al. 2005; Morita et al. 2003). Among them, proteins of the ω-5 subgroup are especially important, since they are the main allergens causing wheat dependent exercise induced anaphylaxis (WDEIA), the most dangerous, life-threatening wheat allergy. Palosuo et al. (2001a, b) found ω-5 gliadins to be important allergens in immediate allergic reactions to wheat while examining allergic children with symptoms of atopic dermatitis and gastrointestinal and respiratory symptoms. In the follow up study, these scientists showed that ω-5 gliadins and related proteins from rye and barley (γ-35/γ-70 secalins and γ-3 hordein) are the main allergens in WDEIA of adult patients (Palosuo et al. 2001a, b). People who suffer from WDEIA show, in most cases, elevated levels of IgE and IgA antibodies against both ω-5 gliadins and against the entire gliadin complex, and react by increasing the histamine release as compared to other allergies (Lehto et al. 2003; Daengsuwan et al. 2005). A more intense immunological reaction of ω-5 proteins, as compared to the α, β and γ groups, has been evidenced by the skin prick tests and dot-blot experiments (Morita et al. 2003). Studies of the ω-gliadin amino acid sequences showed the heptapeptide QQIPQQQ as the most immunoreactive epitope, with Q1, P4, Q5, Q6, Q7 playing key roles in binding the IgE antibodies (Matsuo et al. 2004). Other gliadin classes, however, especially γ-gliadins, may also be active in anaphylactic reactions (Palosuo et al. 1999; Morita et al. 2000) but to a lesser extent. Jacquenet et al. (2009) concluded that elimination of ω-5 gliadins may completely eliminate the risk associated with consumption of foodstuffs made from wheat flour.
Recognition of the biochemical basis of immunoreactive properties and identification of epitope sequences in gluten proteins have inspired scientists to model a hypoallergenic wheat genotype, postulating gene technology as the main tool in the wheat genome modification (Matsuo et al. 2004). It is possible, however, that natural sources of reduced immunoreactivity of gliadin proteins may also occur among some wheat populations. One such example is wheat in which some gluten protein genes are down-regulated or completely inhibited by natural processes. Natural inactivation of HMW glutenin genes was first reported by Bietz et al. (1975). Biotypes lacking different HMW glutenin subunits were identified in the variety Nap Hal from Nepal. Authors hypothesized that deletions of individual subunits would reduce bread making quality of mutants, which was later confirmed (Bustos et al. 2000; Lawrence et al. 1988; Rogers et al. 2001). Two major genetic mechanisms may lead to inactivation of gluten protein genes. In one, chromatin fragments may be lost generating null alleles. In another, spontaneous mutations may change individual nucleotides in DNA sequences (Anderson et al. 1997). As a result, codons of particular amino acids may turn into stop codons, down-regulating structural gene expression leading to pseudogenes. An example is glutamine codon—CAA or CAG—that may change to stop codons UAA or UAG by a C > T transition (Hsia and Anderson 2001). Such mutations are more probable for gluten protein genes, especially those coding ω-gliadins, because of their extremely high glutamine contents (average 40 %). Matsuo et al. (2005) identified two stop codons among ω-5 gliadin DNA sequences of the Japanese wheat cultivar Norin 61.
Genes for gliadin proteins are closely linked in clustered loci Gli A1, Gli B1, Gli D1, Gli A2, Gli B2 and Gli D2 on chromosomes 1A, 1B, 1D, 6A, 6B and 6D of hexaploid wheat genome (Metakovsky and Graybosch 2006). Genes within clusters are inherited together as single Mendelian units. The products of their expression—groups of polypeptides called gliadin blocks—are carried unchanged through generations, as evidenced by A-PAGE or RP-HPLC (Payne 1987; Bietz and Waga 1996). Sometimes, however, mutations may inhibit expression of individual, several, or all genes of a cluster (Metakovsky and Graybosch 2006). Thus, it seems likely that inhibition of protein blocks containing the most immunoreactive gliadin fractions may significantly decrease immunoreactivity of the entire gliadin complex.
In our research on the gliadin polymorphism, a set of wheat genotypes containing inactive ω-gliadin genes on the 1A, 1B or 1D chromosomes was developed. The aim of the present work was to:
-
(1)
accumulate three inactive ω-gliadin gene variants in one genotype and develop a wheat lacking all ω-gliadins;
-
(2)
characterize the gluten protein composition of ω-gliadin-free wheat; and
-
(3)
determine whether elimination of the most allergenic ω-gliadins significantly decreases wheat allergenic properties.
We have also undertaken to evaluate whether and how, elimination of this group of gluten proteins may affect wheat’s technological properties.
Materials and methods
Plant materials
Two closely related winter wheat genotypes, very different in ω-gliadin composition, were used in this study. The first (control) genotype, 3×C, carried active ω-gliadin loci on chromosomes 1A, 1B and 1D encoding nine ω-gliadin fractions, as evidenced by A-PAGE. The second genotype, 3×N, contained inactive genes in ω-gliadin loci that code no proteins. Each genotype was represented by ten lines, five with Glu D1-5+10 and five with Glu D1-2+12 HMW glutenin subunits. Analyzed materials were developed from crosses of two winter wheat lines, B1N and Nep1, which we had developed and analyzed in previous years. B1N originated by a spontaneous mutation among hybrids from a cross of a spelt cv. Oberkummler Rotkorn with a Polish breeding line LAD 480. It contains one inactive gene in the Gli B1 locus and lacks one ω-5 gliadin relative to the wild genotype. Nep1 is a gliadin biotype selected from heterogenic wheat from Nepal, and contained inactive genes at gliadin loci Gli A1 and Gli D1. Nep1 lacked several gliadins of the ω-1.2 group. B1N and Nep1 were crossed in 2009. In 2010, about 15,000 brush halves of the F2 generation kernels were analyzed by A-PAGE to identify gliadins, while germ ends were stored for selection of genotypes containing desirable ω-gliadin combinations. Thirty-three 3×C and twenty-one 3×N germ ends were identified in the analyzed hybrid population. The selected germ ends were vernalized 30 days at 4 °C under the 8 h day/16 h night regime and grown in a greenhouse. Kernels from F2 plants were analyzed by A-PAGE and SDS-PAGE to determine gliadin and HMW glutenin compositions, as described below. Based on the electrophoregrams, harvested plants were divided into four groups of five lines each. Based on the ω-gliadins/GluD1 composition the groups were: 3×C/5+10, 3×N/5+10, 3×C/2+12, 3×N/2+12. In 2011/12, kernels of each line were sown in two-row 1 m-long plots. Plants harvested in 2012 were analyzed by A-PAGE, SDS-PAGE, RP-HPLC, ELISA and the SDS-sedimentation test.
Electrophoresis: A-PAGE and SDS-PAGE
Gliadins were separated using the acidic polyacrylamide gel electrophoresis (A-PAGE) (Metakovsky and Novoselskaya 1991). Gliadins were extracted overnight from individual grains or grain brush halves in 70 % ethanol, and concentrated with saturated sucrose solution in the aluminum lactate buffer (pH 3.1). Total monomer concentration in gels was T = 8 % (w/v), and the crosslinker (methylene bisacrylamide) concentration was C = 0.29 % (w/v). Electrophoresis was at constant voltage (U = 500 V) in the aluminum lactate buffer for 3 h.
Glutenin subunits with along with other wheat kernel proteins (total proteins, TP) were separated by the sodium dodecylsulphate polyacrylamide gel electrophoresis (SDS-PAGE). Proteins were extracted from individual grains, from grain brush halves, or from wheat flour using 400 μl sample buffer (6 M urea, 2 % SDS and 1.5 % mercaptoethanol). The total monomer concentration (T) of the separating gel was 10 %, while crosslinker concentration (C) was 1.5 %. In the stacking gel, monomer concentration was 4 % and crosslinker concentration was 0.5 %. Electrophoresis was run in the Tris–HCl buffer at a constant current I = 90 mA for 4 h at 4 °C.
Both gliadins and glutenins were separated using a Desaphor apparatus (Desaga GmbH, Heidelberg, Germany). Gels were stained overnight in a Commasie Brillant Blue (R250 + G250) solution with methanol (17 %) and acetic acid (5 %), and de-stained in distilled water for ca. 24 h.
Wheat kernel proteins were also analyzed on the Mini Protean apparatus using “ready to use” 10 % TGX gels and 10× diluted Tris–HCl concentrate. The apparatus and reagents for mini gel SDS-PAGE were from BioRad, USA. Albumins and globulins (A + G) were extracted for 2 h with 0.15 M NaCl. Gliadins were then extracted from the pellet for 5 h with 70 % ethanol. Gliadin extracts were added to sample buffer containing mercaptoethanol (reduced gliadins), and also to the same buffer lacking mercaptoethanol (non-reduced gliadins). Pellets obtained after removing salt soluble and gliadin proteins were treated overnight with sample buffer containing mercaptoethanol, yielding mainly HMW and LMW glutenin subunits. Electrophoresis was run for 40 min at a constant voltage U = 200 V.
Chromatography RP-HPLC
RP-HPLC was performed as described by Bietz (2002) and modified in our laboratory. Gliadins were extracted using 70 % ethanol for 12 h and supernatants were condensed for 60 min in a Labconco CentriVap concentrator to remove ethanol. Acetonitrile (ACN) and trifluoroacetic acid (TFA) were added to extracts to final concentrations of 20 % ACN and 0.1 % TFA. Gliadins were separated using the Agilent 1100 apparatus on C-18 type Zorbax 300SB columns; 10 μl of extracts were injected from the autosampler to the column. Separations were done at 70 °C, at the eluent flow of 1 ml/min. A mixture of ACN, water and 0.1 % TFA was used as the mobile phase. Analyses were performed in a 25–50 % ACN gradient for 60 min. Proteins in the column effluent were detected by UV at 210, 254 and 280 nm. Resulting chromatograms were compared for the retention times (RT) and peak heights. Peak heights were expressed as absorbance units (mAU).
ELISA
Gliadin’s immunoreactive properties were evaluated by the “sandwich ELISA” using the modified method of Battais et al. (2003). Gliadins for ELISA were extracted from flour overnight using 70 % ethanol, after previously removing water and aqueous saline soluble proteins (A + G) by three consecutive extractions with 0.15 M NaCl for 2 h each. Gliadin concentration in the ethanol extracts was estimated by the Bradford method using the “ready to use” kit from BioRad, USA. Concentrations of extracts were adjusted to the value of the lowest gliadin extract concentration. Extracts diluted 1:1,000 in the phosphate buffered saline plus 0.05 % tween (PBS-T) were introduced to microtiter plate wells (100 μl per well). The microtiter plate coated with gliadins was incubated overnight at 4 °C, and blocked for 2 h with 1 % defatted milk in PBS-T. After removing the blocker, human sera of ten patients and three healthy controls diluted 1:5 were added to each well and incubated at 37 °C. Human sera numbers 1–4 (according to Table 1) were obtained from people with wheat allergy, treated at the Department of Clinical and Environmental Allergology, Collegium Medicum Jagiellonian University, Krakow, Poland. Sera 5-10 were kindly supplied by Dr. Yuri Lebedin (Xema Medical, Moscow, Russia). The level of sIgE antibodies to gluten in the obtained sera ranged from 0.06 to 23.00 U/ml. After removing the sera, the conjugate solution (IgE antihuman antibodies linked with horseradish peroxidase, Sigma Chemical Co., USA) diluted 1:1,000 was added to each well and incubated for 1.5 h at 37 °C. The “ready to use” ortophenylenediamine (OPD, Sigma Chemical, Co., USA) dissolved in water according to manufacturer’s instructions was used as substrate for HRP, detecting the activity of marking enzyme. After stopping the reaction with 0.5 M H2SO4, optical density (OD) of products was calculated using an ELISA test reader (Opsys MR, Thermolab System, USA) at 490 nm; obtained values were accepted as gliadin’s immunoreactivity indicators. Each sample was analyzed in triplicate.
Quality evaluation
Total protein content of wheat grains was determined by the near-infrared spectroscopy using the Infratec 1241 (Foss Tecator, Sweden) grain analyzer. Technological properties were estimated by the SDS-sedimentation test (Axford et al. 1979), with flour weight of analyzed samples reduced to 3.2 g and the volume of lactic acid containing 2 % SDS reduced to 20 ml.
Statistics
Mean values of the OD and the SDS-sedimentation were compared using the Student’s t test.
Results
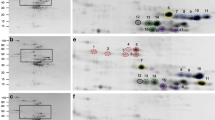
A-PAGE of gliadins extracted from flour of the 3×C and 3×N wheats showed nine protein bands in the ω-zone of 3×C control line; these ω-gliadin bands are almost completely absent in the 3×N genotype (Fig. 1). SDS-PAGE showed considerable differences between protein bands of the 3×C and 3×N wheats in the D-zone, where both ω-gliadins and the D-type LMW glutenin subunits are located (Fig. 2). Two intensive, adjacent bands were present in 3×C ω-5 gliadins. In 3×N, however, the slower moving band had a greatly reduced intensity, while two new minor bands appeared. The faster major band in 3×C, however, completely disappeared in 3×N. Three proteins bands were present among ω-1.2 gliadins of 3×C; in 3×N, the fastest of these three bands was absent.
SDS-PAGE on BioRad mini gel slabs (Fig. 3) compared total wheat kernel proteins (TP) of 3×C and 3×N with proteins obtained by sequential extraction [salt soluble proteins (mainly albumins and globulins), ethanol soluble proteins (mainly gliadins, analyzed in native and reduced forms), and residual proteins (mainly HMW and LMW glutenins)]. This permitted identification of proteins in the D-zone of the 3×N line.
SDS-PAGE of wheat kernel proteins sequentially extracted from control (3×C) and ω-gliadin-free (3×N) genotypes. a total proteins, b proteins remaining after extraction of salt- and ethanol-soluble proteins, c ethanol soluble proteins reduced by mercaptoethanol, d unreduced ethanol-soluble proteins; and e salt soluble proteins. MW molecular weight standards. Masses of ω-gliadin fractions are shown on the left
Three highest molecular weight ω–gliadin bands (59, 56 and 54 kDa) were present among TP and both native and reduced gliadins in 3×C. Two of these bands (59 and 56 kDa), however, were also present among the D-type LMW glutenins and A + G of 3×C. This indicates that 59 and 56 kDa proteins are composed of A + G, gliadins and LMW glutenins, while the third band (54 kDa) is a typical ω-5 gliadin. These three bands were completely absent in native and reduced 3×N gliadin. Two other protein bands of MW 51 and 50 kDa were present among TP and native and reduced gliadins of 3×C, but were completely absent among LMW glutenins and A + G of 3×C. They were also completely absent in all analyzed protein fractions of 3×N. This indicates that the 51 and 50 kDa bands are ω-gliadins specific for the 3×C wheat line. The last protein band among ω-gliadins, of MW 46 kDa, was present in TP, LMW glutenins, native and reduced gliadins of 3×C, but not in A + G of 3×C. In 3×N, however, this band was present in TP and LMW glutenins, but not in gliadins and A + G. This suggests that the 46 kDa protein band is a typical ω-gliadin of 3×C, which co-elutes with the LMW glutenin subunits of identical MW.
RP-HPLC separates gliadins into fractions of lowest (RT 20-27), medium (RT 27-40) and highest (RT 40-55) hydrophobicity (Fig. 4). These correspond to the ω, α/β, and γ fractions, respectively, as determined by RP-HPLC of purified gliadins (results not shown). This classification agrees with other authors (Becker et al. 2006). RP-HPLC of gliadins from 3×C and 3×N hybrids revealed that the heights of two peaks of the lowest hydrophobicities (RT 22.133 and 24.871 min), corresponding to ω-gliadins in 3×C, were strongly reduced (about 90 %) in 3×N (Table 1). The height of three other peaks in 3×N (RT 42.426, 44.141, and 53.889 min), corresponding to γ-gliadins of high hydrophobicity, were also reduced by about 90 % compared with 3×C. At the same time, α/β gliadin peaks (RT 32.119, 32.702, 34.204, 34.995, 36.546, 37.333, 37.895 min) and one γ-gliadin peak (RT = 49.028 min) increased in 3×N from about 40–240 %. Quantification of gliadins expressed as areas under the peaks suggests that all α-, β-, γ- and ω-peaks do not differ considerably in 3×C and 3×N (Table 2). However, in some gliadin groups major changes have taken place. Large reductions in total areas in 3×N were observed for ω- and γ-gliadins (86 and 13 %, respectively). However, for the α/β peaks, total areas increased in 3×N by almost 30 % as compared with 3×C.
Immunoreactive properties of gliadins from 3×C and 3×N were evaluated by the “sandwich ELISA” using individual sera of ten patients exhibiting symptoms of allergies after consuming foods containing wheat flour (Table 3). The gliadin immunoreactivity was expressed as the potential of gliadins to bind IgE antibodies, specific to gluten, present in patients’ sera. The results showed a considerable reduction of the OD values for 3×N as compared to 3×C, ranging from 26.5 to 35.7 % for individual patients. The level of the immunoreactivity reduction was independent of the sIgE concentration. The difference between average OD values calculated for the entire group of ten patients estimated for 3×C and 3×N separately was statistically significant (P < 0,001) using the Student’s t test.
In a preliminary experiment, technological properties of 3×C and 3×N were compared using the total protein content and the SDS-sedimentation test. Ten lines of each ω-gliadin genotypes were divided into groups based on the 5+10 and 2+12 HMW glutenin subunits they carry (coded by the Glu D1 locus, and correlated with higher and lower quality, respectively). Four groups (3×C-5+10, 3×N-5+10, 3×C-2+12, and 3×N-2+12) were compared pairwise to evaluate differences of total protein content and of the SDS sedimentation volumes between 3×C and 3×N of different Glu D1 alleles background, and, vice versa, between 5+10 and 2+12 subunits of different ω-gliadin backgrounds. Total protein contents of 3×C and 3×N hybrid lined ranged from 13.8 to 15.4 % but no significant differences between genotypes were observed (results not shown). However, average SDS sedimentation volumes for 3×N were significantly higher than those for 3×C, independently of the Glu D1 allele composition (Table 4). Lines with Glu D1-5+10 subunits had significantly higher SDS sedimentation volumes than lines with Glu D1-2+12 subunits (for both 3×C and 3×N ω-gliadin type backgrounds).
Discussion
The main goal of this work is to eliminate ω-gliadins—the most allergenic gluten proteins—from the wheat proteome by accumulation of inactive ω-gliadin gene variants in a single wheat genotype. This goal appears to have been achieved: we developed a set of wheat lines containing inactive ω-gliadin loci on each homoeologous chromosomes, 1A, 1B and 1D. These genotypes lack all ω-gliadin fractions. The combination of inhibited genes was stable in three consecutive generations. Elimination of ω-gliadins was demonstrated by A-PAGE, which showed no ω-gliadin protein bands. The SDS-PAGE of sequential protein extracts showed several protein bands in the D-zone (where ω-gliadins are usually present) to be albumins, globulins, and LMW glutenins, but not gliadins. Co-inhibition of γ and ω-gliadins was also shown by the RP-HPLC. Elimination of two peaks corresponding to the least hydrophobic ω-gliadins was associated with the elimination of three highly hydrophobic γ-gliadin peaks. This is due to common inheritance of closely-linked ω- and γ-gliadin genes grouped in clusters on chromosomes 1A, 1B and 1D (Metakovsky and Graybosch 2006). Co-inhibition of γ- and ω-gliadins revealed by the RP-HPLC showed that mutations of Gli A1, Gli 1, and Gli D1 loci inactivate entire gene clusters, rather than individual genes. The present work shows that elimination of ω- and some γ-gliadin peaks is accompanied by an increase in the α/β zone peaks and one γ-gliadin peak. This confirms that down-regulation of one group of plant protein genes is often accompanied by up-regulation of another (Becker et al. 2006). This has also been observed among gluten proteins. Wieser et al. (2006) showed that silencing α-gliadin genes by the interference RNA (RNAi) technique caused overexpression of albumin and globulin genes, and also those for γ- and ω-gliadins and HMW glutenins. Such a “defense” reaction of plants upon genome modification shows a tendency to compensate for missing proteins, and preserving total protein content at a constant level.
Medical research has shown that the frequency of allergies, including food allergies, increases with societies’ technological progress (Ebo and Stevens 2001). This problem affects not only highly developed industrialized countries; it has become global in character. Disease protection in allergies consists mainly of medical treatment and elimination of allergens from the diet. However, alternative strategies of allergy control, based on eliminating allergens from the human environment, have also been developed (Maruyama et al. 1998). Recently some toxic or allergenic proteins in cultivated plant species have been eliminated or their levels strongly reduced. Much of this research used antisense or, more recently, RNAi technologies. Tada et al. (1996) strongly reduced expression of 14–16 kDa allergenic proteins in transgenic rice plants. In soybean, the P34 pathogen resistance protein, a member of the papain superfamily of cysteine proteases, is the most allergenic seed protein. Herman et al. (2003) developed soybean transgenic plants in which protein P34 was eliminated.
Some gluten protein genes have also been the objects of the gene silencing research. Alvarez et al. (2000) transformed wheat cultivar Federal (HMW glutenin subunit composition 2, 7+9, 5+10) with genes coding for A×1 and D×5 subunits, and developed genetically modified genotypes either overexpressing D×5 or silencing most HMW glutenin subunits. Gil-Humanes et al. (2008) created an RNA construct composed of a D-hordein promoter and two γ-gliadin sequences (165 bp) in reverse position divided by an intron. Transformation of wheat with this construct effectively silenced γ-gliadin genes, as confirmed by A-PAGEced. More recently, Altenbach and Allen (2011) were able to down-regulate ω-5 gliadin genes and eliminate the ω-5 gliadin fractions using the RNAi technology.
Contrary to these examples, our ω-gliadin-free wheat genotypes were developed by classical plant breeding. Crossing and selection were aided by A-PAGE, SDS-PAGE and RP-HPLC—the main tools for identifying desirable genotypes. As a consequence, the produced lines are not transgenic, and are not GMO according to European Union law. To the best of our knowledge, our ω-gliadin-free hybrids are the first wheat genotypes where all ω-gliadins were eliminated by plant breeding. As expected, ELISA showed a statistically significant reduction of about 30 % in immunoreactivity, as compared to the ω-gliadin-carrying control lines. This decrease in immunoreactivity occurred in spite of overexpression of α/β and some γ-gliadin genes. α/β and γ-gliadins are less allergenic than ω-gliadins (Battais et al. 2003; Bouchez-Mahiout et al. 2010; Waga et al. 2011). However, higher amounts of low immunoreactivity α/β or γ-gliadins might, theoretically, have a greater effect than lower-amounts of more immunoreactive ω-gliadin proteins. ELISA results, however, did not confirm this hypothesis.
Elimination of an important gluten protein fraction may be expected to considerably alter wheat’s technological properties. The obtained amounts of the F4 seed was insufficient for comprehensive quality tests. However, a preliminary evaluation for the total grain protein content and using the SDS-sedimentation test suggested that wheats lacking of all ω-gliadins, may have considerably higher values of SDS sedimentation test than typical genotypes, despite similar total protein contents. This increased sedimentation values of the 3×N lines were observed in two sets of genotypes grouped by the HMW subunits 5+10 and 2+12, which affect wheat quality in different ways (Popineau et al. 1994). In other words, quality differences between 3×C and 3×N lines were independent of the glutenin background. Moreover, ω-gliadins more strongly affected the sedimentation values of hybrids than did the HMW glutenin subunits. The observed increase in quality of ω-gliadin-free hybrid lines agrees with results of other authors. Branlard and Metakovsky (2006) analyzed wheat hybrids with regard to their Gli D1 alleles, which code the ω-1.2 gliadin fractions. Progenies with inactive genes at the Gli D1 locus had better quality: significantly higher dough strength and reduced dough extensibility. The most probable reason for quality improvement with ω-gliadins absent is an increased ratio of the HMW polymeric proteins to monomeric gliadins. Polymeric proteins play a most important role in gluten’s rheological properties, mainly its elasticity and, consequently, the proportion of polymeric to monomeric proteins relates to wheat quality (Huebner et al. 1997).
Our results suggest that elimination of ω-gliadins from the wheat proteome may be beneficial in two ways—by reducing immunoreactivity, and by improving technological properties. Further studies of the 3×C and 3×N hybrid genotypes are in order, using more advanced immunological and technological methods, with larger quantities of plant materials, cultivated in different environmental conditions.
References
Altenbach SB, Allen PV (2011) Transformation of the US bread wheat “Butte 86” and silencing of omega-5 gliadin genes. GM Crops 2–1:66–73
Alvarez ML, Guelman S, Halford NG, Lustig S, Reggiardo MI, Ryabushkina N, Shewry P, Stein J, Vallejos RH (2000) Silencing of HMW glutenins in transgenic wheat expressing extra HMW subunits. Theor Appl Genet 100:319–327
Anderson OD, Litts JC, Green FC (1997) The α-gliadin gene family. Characterization of ten new wheat α-gliadin genomic clones, evidence for limited sequence conservation of flanking DNA, and southern analysis of the gene family. Theor Appl Genet 95:50–58
Axford DWE, McDermott EE, Redman DG (1979) Note on the sodium dodecyl sulfate test of breadmaking quality: comparison with Pelshenke and Zeleny tests. Cereal Chem 56(6):582–584
Battais F, Pineau F, Popineau Y, Aparicio C, Kanny G, Guerin L, Moneret-Vautrin DA, Denery-Papini S (2003) Food allergy to wheat: identification of immunoglobulin E and immunoglobulin G-binding proteins with sequential extracts and purified proteins from wheat flour. Clin Exp Allergy 33:962–970
Battais F, Mothes T, Moneret-Vautrin DA, Pineau F, Kanny G, Popineau Y, Bodinier M, Denery-Papini S (2005) Identification of IgE-binding epitopes on gliadins for patients with food allergy to wheat. Allergy 60:815–821
Becker D, Folck A, Wieser H (2006) Inhibierung der α-gliadingen expression in hexaploidem Brotweizen. Getreidetechnologie 60:153–156
Bietz JA (2002) HPLC of cereal endosperm storage proteins. In: Gooding KM, Regnier FE (eds) HPLC of biological macromolecules. 2nd ed, revised and expanded. Marcel Dekker Inc, New York, pp 547–587
Bietz JA, Waga J (1996) Gliadin block analysis by RP-HPLC and capillary electrophoresis for genotype identification and quality prediction. In: Wrigley CW (ed) Gluten’96—Proceedings 6th Int. Gluten Workshop. Am. Assoc. Cereal Chem. St. Paul, MN, pp 362
Bietz JA, Shepherd KW, Wall JS (1975) Single-kernel analysis of glutenin: use in wheat genetics and breeding. Cereal Chem 52:513–532
Bouchez-Mahiout I, Snegaroff J, Tylichova M, Pecquet C, Branlard G, Lauriere M (2010) Low molecular weight glutenins in wheat dependant, exercise-induced anaphylaxis: allegenicity and antigenic relationships with omega 5-gliadins. Int Arch Allergy Immunol 153:35–45
Branlard GP, Metakovsky EV (2006) Some Gli alleles related to common wheat dough quality. In: Wrigley C, Bekes F, Bushuk W (eds) Gliadin and glutenin: the unique balance of wheat quality. AACC International, St. Paul, pp 115–139
Bushuk W (1993) Wheat flour proteins: composition, structure and functionality in breadmaking. Pol J Food Nutr Sci 2(43):5–23
Bustos AD, Rubio P, Jouve N (2000) Molecular characterization of the inactive allele of the gene Glu A1 and the development of a set of AS-PCR markers for HMW glutenins of wheat. Theor Appl Genet 100:1085–1094
Daengsuwan T, Palosuo K, Phankingthongkum S, Visitsunthorn N, Jirapongsananuruk O, Alenius H, Vichyanond P, Reunala T (2005) IgE antibodies to ω-5 gliadin in children with wheat-induced anaphylaxis. Allergy 60:506–509
Ebo DG, Stevens WJ (2001) IgE-mediated food allergy—extensive review of the literature. Acta Clin Belg 56(4):234–247
Gil-Humanes J, Piston F, Hernando A, Alvarez JB, Shewry PR, Barro F (2008) Silencing of γ-gliadins by RNA interference (RNAi) in bread wheat. J Cereal Sci 48:565–568
Herman EM, Helm RM, Jung R, Kinney AJ (2003) Genetic modification removes an immunodominant allergen from soybean. Plant Physiol 132:36–43
Hsia CC, Anderson OD (2001) Isolation and characterization of wheat ω-gliadin genes. Theor Appl Genet 103:37–44
Huebner FR, Nelsen TC, Chung OK, Bietz JA (1997) Protein distributions among hard red winter wheat varieties as related to environment and baking quality. Cereal Chem 74:123–128
Inomata N (2009) Wheat allergy. Curr Opin Allergy Clin Immunol 9:238–243
Jacquenet S, Morissert M, Battais F, Denery-Papini S, Croizier A, Baudouin E, Bihain B, Moneret-Vautrin DA (2009) Interest of immunoCAP system to recombinant ω-5 gliadin for the diagnosis of exercise-induced wheat allergy. Int Arch Allergy Immunol 149:74–80
Lawrence GJ, MacRitchie F, Wrigley CW (1988) Dough and baking quality of wheat lines deficient in glutenin subunits controlled by the Glu-A1, Glu-B1 and Glu-D1 loci. J Cereal Sci 7:109–112
Lehto M, Palosuo K, Varjonen E, Majuri M-L, Andersson U, Reunala T, Alenius H (2003) Humoral and cellular responses to gliadin in wheat-dependent, exercise-induced anaphylaxis. Clin Exp Allergy 33:90–95
Maruyama N, Ichise K, Katsube T, Kishimoto T, Kawase S, Matsumura Y, Takeuchi Y, Sawada T, Utsumi S (1998) Identification of major wheat allergens by means of the Escherichia coli expression system. Eur J Biochem 255:739–745
Matsuo H, Morita E, Tatham AS, Morimoto K, Horikawa T, Osuna H, Ikezawa Z, Kaneko S, Kohno K, Dekio S (2004) Identification of the IgE-binding Epitope in ω-5 Gliadin, a major Allergen in wheat-dependent exercise-induced anaphylaxis. J Biol Chem 279(13):12135–12140
Matsuo H, Kohno K, Morita E (2005) Molecular cloning, recombinant expression and IgE-binding epitope of ω-5 gliadin, a major allergen in wheat dependent exercise-induced anaphylaxis. FEBS J 272:4431–4438
Metakovsky EV, Graybosch RA (2006) Gliadin alleles in wheat: identification and applications. In: Wrigley C, Bekes F, Bushuk W (eds) Gliadin and glutenin: the unique balance of wheat quality. AACC International, St. Paul, pp 85–114
Metakovsky EV, Novoselskaya AY (1991) Gliadin allele identification in common wheat. I. Methodological aspects of the analysis of gliadin pattern by one-dimensional polyacrylamide-gel electrophoresis. J Genet Breed 45:317–324
Morita E, Yamamura Y, Mihara S, Kameyoshi Y, Yamamoto S (2000) Food-dependent excecise-induced anaphylaxis: a report of two cases and determination of wheat γ-gliadin as the presumptive allergen
Morita E, Matsuo H, Mihara S, Morimoto K, Savage AWJ, Tatham AS (2003) Fast ω-gliadin is a major allergen in wheat-dependent exercise-induced anaphylaxis. J Dermatol Sci 33:99–104
Palosuo K, Alenius H, Varionen E, Koivuluhta M, Mikkola J, Keskinen H, Kalkkinen N, Reunala T (1999) A novel wheat gliadin as a cause of exercise-induced anaphylaxis. J Allergy Clin Immunol 103(5):912–917
Palosuo K, Alenius H, Varionen E, Kalkkinen N, Reunala T (2001a) Rye γ-70 and γ-35 secalins and barley γ-3 hordein cross-react with ω-5 gliadin, a major allergen in wheat dependent, exercise-induced anaphylaxis. Clin Exp Allergy 31:466–473
Palosuo K, Varionen E, Kekki OM, Klemola T, Kalkkinen N, Alenius H, Reunala T (2001b) Wheat ω-5 gliadin is a major allergen in children with immediate allergy to ingested wheat. J Allergy Clin Immunol 108(4):634–638
Payne PI (1987) Genetics of wheat storage proteins and the effect of allelic variation on bread-making quality. Annu Rev Plant Physiol 38:141–152
Popineau Y, Cornec M, Lefebre J, Marchylo B (1994) Influence of high Mr glutenin subunits on gluten polymers and rheological properties of glutens and gluten sub-fractions of near-isogenic lines of wheat Sicco. J Cereal Sci 19:231–241
Rogers WJ, Sayers EJ, Ru KL (2001) Deficiency of individual high molecular glutenin subunits affords flexibility in breeding strategies for bread-making quality in wheat Triticum aestivum L. Euphytica 117:99–109
Sandiford CP, Tatham AS, Fido R, Welch JA, Jones MG, Tee RD, Shewry PR, Newman Taylor AJ (1997) Identification of the major water/salt insoluble wheat proteins involved in cereal hypersensitivity. Clin Exp Allergy 27:1120–1129
Tada Y, Nakase M, Adachi T, Nakamura R, Shimada H, Takahashi M, Fujimura T, Matsuda T (1996) Reduction of 14–16 kDa allergenic proteins in transgenic rice plants by antisense gene. FEBS Lett 391:341–345
Tatham AS (1995) The structures of wheat proteins. In: The proceedings of a Conference wheat structure, biochemistry and functionality. Royal Society of Chemistry, Food Chemistry Group. 10–12 April 1995, Reading, pp 53–62
Waga J, Obtułowicz K, Zientarski J, Czarnobilska E, Skoczowski A (2011) Purified wheat gliadin proteins as immunoglobulin E bindings factors in wheat mediated allergies. Am J Plant Sci 2:476–483
Wieser H (1991) Chemistry of gliadins. Eur J Gastroenterol Hepatol 3(2):102–107
Wieser H, Kohler P, Folck A, Becker D (2006) Characterization of wheat with strongly reduced α-gliadin content. In: Lookhart GL, Ng PKW (eds) Gluten Proteins. AACC, St Paul, pp 13–16
Acknowledgments
This work was carried out with financial support from The Polish National Center for Research and Development by research Grant NR12-0001-10/2011.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Waga, J., Skoczowski, A. Development and characteristics of ω-gliadin-free wheat genotypes. Euphytica 195, 105–116 (2014). https://doi.org/10.1007/s10681-013-0984-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-013-0984-1