Abstract
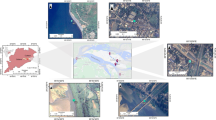
Faecal microorganisms represent a key threat to human health. Potential origins of faecal microbial contamination in a typical urban-representative micro-scale were evaluated. The quantitative polymerase chain reaction (qPCR) method was used in this study. The Bacteroidetes is selected as the indicative microorganism in runoff samples that are collected during four representative stormwater events in north China. The principal component analysis (PCA) method indicated the distribution feature of the environmental factors. The largest contributor is dog, followed by bird and human to the faecal pollution in stormwater runoff. The output of human and dog faecal pollutants in response to the first flush effect of nonpoint source pollution while the transmit time of bird faecal pollutant is relatively longer. In addition, the number of antecedent drying days represents the key factor for dog faecal pollution, while human faecal pollution is impacted by more factors. The results of this study will provide sound evidence for the tracking and management of nonpoint source faecal pollution in urban catchment areas.






Similar content being viewed by others
References
Ahmed, W., Payyappat, S., Cassidy, M., Besley, C., & Power, K. (2018). Novel crAssphage marker genes ascertain sewage pollution in a recreational lake receiving urban stormwater runoff. Water Research, 145, 769–778.
Boehm, A. B., Graham, K. E., & Jennings, W. C. (2018). Can we swim yet? Systematic review, meta-analysis, and risk assessment of aging sewage in surface waters. Environmental Science & Technology, 52, 9634–9645.
Brooks, Y. M., Spirito, C. M., Bae, J. S., Hong, A., Mosier, E. M., Sausele, D. J., et al. (2020). Fecal indicator bacteria, fecal source tracking markers, and pathogens detected in two Hudson River tributaries. Water Research, 171, 115342.
Bucci, J. P., Shattuck, M. D., Aytur, S. A., Carey, R., & Mcdowell, W. H. (2017). A case study characterizing animal fecal sources in surface water using a mitochondrial DNA marker. Environmental Monitoring and Assessment, 189, 406–416. https://doi.org/10.1007/s10661-017-6107-z.
Cho, K. H., Pachepsky, Y. A., Kim, M., Pyo, J., Park, M.-H., Kim, Y. M., et al. (2016). Modeling seasonal variability of fecal coliform in natural surface waters using the modified SWAT. Journal of Hydrology, 535, 377–385.
Cizek, A. R., Characklis, G. W., Krometis, L. A., Hayes, J. A., Simmons, O. D., Lonardo, D., et al. (2008). Comparing the partitioning behaviour of Giardia and Cryptosporidium with that of indicator organisms in stormwater runoff. Water Research, 42, 4421–4438.
Derx, J., Schijven, J., Sommer, R., Zoufal-Hruza, C. M., van Driezum, I. H., Reischer, G., et al. (2016). QMRAcatch: human-associated fecal pollution and infection risk modeling for a river/floodplain environment. Journal of Environmental Quality, 45, 1205.
Eregno, F. E., Tryland, I., Tjomsland, T., Myrmel, M., Robertson, L., & Heistad, A. (2016). Quantitative microbial risk assessment combined with hydrodynamic modelling to estimate the public health risk associated with bathing after rainfall events. Science of the Total Environment, 548-549, 270–279.
Fan, L., Zhang, X., Zeng, R., Wang, S., Jin, C., He, Y., & Shuai, J. (2019). Verification of Bacteroidales 16S rRNA markers as a complementary tool for detecting swine fecal pollution in the Yangtze Delta. Journal of Environmental Sciences. https://doi.org/10.1016/j.jes.2019.11.016.
Fauvel, B., Cauchie, H.-M., Gantzer, C., & Ogorzaly, L. (2016). Contribution of hydrological data to the understanding of the spatio-temporal dynamics of F-specific RNA bacteriophages in river water during rainfall-runoff events. Water Research, 94, 328–340.
Fleisher, J. M., Fleming, L. E., Solo-Gabriele, H. M., Kish, J. K., Sinigalliano, C. D., Plano, L., Elmir, S. M., Wang, J. D., Withum, K., Shibata, T., Gidley, M. L., Abdelzaher, A., He, G., Ortega, C., Zhu, X., Wright, M., Hollenbeck, J., & Backer, L. C. (2010). The BEACHES study: health effects and exposures from non-point source microbial contaminants in subtropical recreational marine waters. International Journal of Epidemiology, 39, 1291–1298.
Fock-Chow-Tho, D., Topp, E., Ibeagha-Awemu, E. A., & Bissonnette, N. (2017). Comparison of commercial DNA extraction kits and quantitative PCR systems for better sensitivity in detecting the causative agent of paratuberculosis in dairy cow fecal samples. Journal of Dairy Science, 100, 572–581.
Harwood, V. J., Staley, C., Badgley, B. D., Borges, K., & Korajkic, A. (2013). Microbial source tracking markers for detection of fecal contamination in environmental waters: relationships between pathogens and human health outcomes. FEMS Microbiology Reviews, 38, 1–40.
Haugland, R. A., Siefring, S., Lavender, J., & Varma, M. (2012). Influences of sample interference and interference controls on quantification of enterococci fecal indicator bacteria in surface water samples by the qPCR method. Water Research, 46, 5989–6001.
Kebede, G., Mushi, D., Linke, R. B., Dereje, O., Lakew, A., Hayes, D. S., et al. (2020). Macroinvertebrate indices versus microbial fecal pollution characteristics for water quality monitoring reveals contrasting results for an Ethiopian river. Ecological Indicators, 108, 105733.
Kildare, B. J., Leutenegger, C. M., McSwain, B. S., Bambic, D. G., Rajal, V. B., & Wuertz, S. (2007). 16S rRNA-based assays for quantitative detection of universal, human-, cow-, and dog-specific fecal Bacteroidales: a Bayesian approach. Water Research, 41, 3701–3715.
Kirschner, A. K. T., Reischer, G. H., Jakwerth, S., Savio, D., Ixenmaier, S., Toth, E., Sommer, R., Mach, R. L., Linke, R., Eiler, A., Kolarevic, S., & Farnleitner, A. H. (2017). Multiparametric monitoring of microbial faecal pollution reveals the dominance of human contamination along the whole Danube River. Water Research, 124, 543–555.
Lee, D.-Y., Lee, H., Trevors, J. T., Weir, S. C., Thomas, J. L., & Habash, M. (2014). Characterization of sources and loadings of fecal pollutants using microbial source tracking assays in urban and rural areas of the Grand River Watershed, Southwestern Ontario. Water Research, 53, 123–131.
Lu, J., Domingo, J. W. S., Lamendella, R., Edge, T., & Hill, S. (2008). Phylogenetic diversity and molecular detection of bacteria in gull feces. Applied and Environmental Microbiology, 74, 3969–3976.
Malla, B., Ghaju Shrestha, R., Tandukar, S., Bhandari, D., Inoue, D., Sei, K., Tanaka, Y., Sherchand, J. B., & Haramoto, E. (2018). Validation of host-specific Bacteroidales quantitative PCR assays and their application to microbial source tracking of drinking water sources in the Kathmandu Valley, Nepal. Journal of Applied Microbiology, 125, 609–619.
McCarthy, D. T., Deletic, A., Mitchell, V. G., & Diaper, C. (2011). Development and testing of a model for micro-organism prediction in urban stormwater (MOPUS). Journal of Hydrology, 409, 236–247.
McIntyre, J. K., Davis, J. W., Hinman, C., Macneale, K. H., Anulacion, B. F., Scholz, N. L., et al. (2015). Soil bioretention protects juvenile salmon and their prey from the toxic impacts of urban stormwater runoff. Chemosphere, 132, 213–219.
McQuillan, J. S., & Robidart, J. C. (2017). Molecular-biological sensing in aquatic environments: recent developments and emerging capabilities. Current Opinion in Biotechnology, 45, 43–50.
Memon, S., Paule, M. C., Park, S.-J., Lee, B.-Y., Kang, S., Umer, R., et al. (2013). Monitoring of land use change impact on stormwater runoff and pollutant loading estimation in Yongin watershed Korea. Desalination and Water Treatment, 51, 4088–4096.
Odagiri, M., Schriewer, A., Hanley, K., Wuertz, S., Misra, P. R., Panigrahi, P., & Jenkins, M. W. (2015). Validation of Bacteroidales quantitative PCR assays targeting human and animal fecal contamination in the public and domestic domains in India. Science of the Total Environment, 502, 462–470.
Paule-Mercado, M. A., Ventura, J. S., Memon, S. A., Jahng, D., Kang, J. H., & Lee, C. H. (2016). Monitoring and predicting the fecal indicator bacteria concentrations from agricultural, mixed land use and urban stormwater runoff. Science of the Total Environment, 550, 1171–1181.
Petrucci, G., Gromaire, M.-C., Shorshani, M. F., & Chebbo, G. (2014). Nonpoint source pollution of urban stormwater runoff: a methodology for source analysis. Environmental Science & Pollution Research, 21, 10225–10242.
Rajal, V. B., McSwain, B. S., Thompson, D. E., Leutenegger, C. M., & Wuertz, S. (2007). Molecular quantitative analysis of human viruses in California stormwater. Water Research, 41, 4287–4298.
Ryu, H., Elk, M., Khan, I. U. H., Harwood, V. J., Molina, M., Edge, T. A., et al. (2014). Comparison of two poultry litter qPCR assays targeting the 16S rRNA gene of Brevibacterium sp. Water Research, 48, 613–621.
Sauer, E. P., VandeWalle, J. L., Bootsma, M. J., & McLellan, S. L. (2011). Detection of the human specific Bacteroides genetic marker provides evidence of widespread sewage contamination of stormwater in the urban environment. Water Research, 45, 4081–4091.
Schoen, M. E., & Ashbolt, N. J. (2010). Assessing pathogen risk to swimmers at non-sewage impacted recreational beaches. Environmental Science & Technology, 44, 2286–2291.
Sidhu, J. P. S., Hodgers, L., Ahmed, W., Chong, M. N., & Toze, S. (2012). Prevalence of human pathogens and indicators in stormwater runoff in Brisbane, Australia. Water Research, 46, 6652–6660.
Sinha, S. N., & Paul, D. (2015). Density of pollution indicator bacteria in relation to physicochemical factors during diel cycle of river ganga at Ichapore, West Bengal, India. Frontiers in Environment Microbiology, 1, 9–13.
Sivaganesan, M., Haugland, R. A., Chern, E. C., & Shanks, O. C. (2010). Improved strategies and optimization of calibration models for real-time PCR absolute quantification. Water Research, 44, 4726–4735.
Somnark, P., Chyerochana, N., Mongkolsuk, S., & Sirikanchana, K. (2018). Performance evaluation of Bacteroidales genetic markers for human and animal microbial source tracking in tropical agricultural watersheds. Environmental Pollution, 236, 100–110.
Sterk, A., Schijven, J., de Roda Husman, A. M., & de Nijs, T. (2016). Effect of climate change on runoff of Campylobacter and Cryptosporidium from land to surface water. Water Research, 95, 90–102.
Tran, N. H., Gin, K. Y.-H., & Ngo, H. H. (2015). Fecal pollution source tracking toolbox for identification, evaluation and characterization of fecal contamination in receiving urban surface waters and groundwater. Science of the Total Environment, 538, 38–57.
Walker, D. I., McQuillan, J., Taiwo, M., Parks, R., Stenton, C. A., Morgan, H., Mowlem, M. C., & Lees, D. N. (2017). A highly specific Escherichia coli qPCR and its comparison with existing methods for environmental waters. Water Research, 126, 101–110.
Wu, B., Chunwei, W., Zhang, C., Sadowsky, M. J., Dzakpasu, M., & Wang, X. C. (2019). Source-associated gastroenteritis risk from swimming exposure to aging fecal pathogens. Environmental Science & Technology. https://doi.org/10.1021/acs.est.9b01188.
Zhi, X., Chen, L., & Shen, Z. (2018). Impacts of urbanization on regional nonpoint source pollution: case study for Beijing, China. Environmental Science and Pollution Research, 25, 9840–9860.
Acknowledgements
The authors want to thank the logistics department of Beijing Normal University for their support during monitoring and other basic data collection.
Funding
This research was funded by the Fund for Innovative Research Group of the National Natural Science Foundation of China (No. 51721093), the State Key Program of National Natural Science of China (No. 41530635), and the Interdisciplinary Research Funds of Beijing Normal University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
A.Electronic supplementary material
ESM 1
(DOCX 1816 kb)
Rights and permissions
About this article
Cite this article
Chen, L., Zhang, X., Zhi, X. et al. Tracking faecal microorganisms using the qPCR method in a typical urban catchment in China. Environ Monit Assess 192, 158 (2020). https://doi.org/10.1007/s10661-020-8130-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10661-020-8130-8