Abstract
Water quality impairment by fecal waste in coastal watersheds is a public health issue. The present study provided evidence for the use of a mitochondrial (mtDNA) marker to detect animal fecal sources in surface water. The accurate identification of fecal pollution is based on the notion that fecal microorganisms preferentially inhabit a host animal’s gut environment. In contrast, mtDNA host-specific markers are inherent to eukaryotic host cells, which offers the advantage by detecting DNA from the host rather than its fecal bacteria. The present study focused on sampling water presumably from non-point sources (NPS), which can increase bacterial and nitrogen concentrations to receiving water bodies. Stream sampling sites located within the Piscataqua River Watershed (PRW), New Hampshire, USA, were sampled from a range of sites that experienced nitrogen inputs such as sewer and septic systems and suburban runoff. Three mitochondrial (mtDNA) gene marker assays (human, bovine, and canine) were tested from surface water. Nineteen sites were sampled during an 18-month period. Analyses of the combined single and multiplex assay results showed that the proportion of occurrence was highest for bovine (15.6%; n = 77) compared to canine (5.6%; n = 70) and human (5.7%; n = 107) mtDNA gene markers. For the human mtDNA marker, there was a statistically significant relationship between presence vs. absence and land use (Fisher’s test p = 0.0031). This result was evident particularly for rural suburban septic, which showed the highest proportion of presence (19.2%) compared to the urban sewered (3.3%), suburban sewered (0%), and agricultural (0%) as well as forested septic (0%) sites. Although further testing across varied land use is needed, our study provides evidence for using the mtDNA marker in large watersheds.



Similar content being viewed by others
References
Andreasson, H., Gyllensten, U., & Allen, M. (2002). Real-time DNA quantification of nuclear and mitochondrial DNA in forensic analysis. BioTechniques, 33, 407–411.
AWWA, APHA, and WEF. 2012. Standard Methods for the Examination of Water and Wastewater, 22th ed.
Baker-Austin, C., Rangdale, R., Lowther, J., & Lees, D. (2010). Application of mitochondrial DNA analysis for microbial source tracking purposes in shellfish harvesting waters. Water Science and Technology, 61, 1–7.
Bernhard, A. E., & Field, K. G. (2000). Identification of nonpoint sources of fecal pollution in coastal waters by using host-specific 16S ribosomal DNA genetic markers from fecal anaerobes. Applied and Environmental Microbiology, 66, 1587–1594.
Bowen, J. L., & Valiela, I. (2001). The ecological effects of urbanization of coastal watersheds: historical increases in nitrogen loads and eutrophication of Waquoit bay estuaries. Canadian Journal of Fisheries and Aquatic Sciences, 58(8), 1489–1500.
Bustin, S. A., Benes, V., Garson, J. A., Hellemans, J., Huggett, J., Kubista, M., Mueller, R., Nolan, T., Pfaffl, M. W., Shipley, G. L., Vandesompele, J., & Wittwer, C. T. (2009). The MIQE guidelines: Minimum Information for Publication of Quantitative real-time PCR Experiments. Clinical Chemistry, 55, 611–622.
Cabelli, V. J., Dufour, A. P., McCabe, L. I., & Levin, M. A. (1982). Swimming-associated gastroenteritis and water quality. American Journal of Epidemiology, 115, 606–616.
Caldwell, J. M., & Levine, J. F. (2009). Domestic wastewater influent profiling using mitochondrial real-time PCR for source tracking animal contamination. Journal of Microbiological Methods, 77, 17–22.
Caldwell, J. M., Raley, M. E., & Levine, J. F. (2007). Mitochondrial multiplex real-time PCR as a source tracking method in fecal-contaminated effluents. Environmental Science & Technology, 41, 3277–3283.
Caldwell, J. M., Patment, P., & Villemur, R. (2011). Mitochondrial DNA as source tracking markers of fecal contamination. Microbial source tracking: methods, applications, and case studies. In C. Hagedorn, A. R. Blanch, & V. J. Harwood (Eds.), Microbial source tracking: methods, applications, and case studies (pp. 229–250). New York: Springer.
Carey RO, et al. (2013). Evaluating nutrient impacts in urban watersheds: challenges and research opportunities.
Carey, R. O., Wollheim, W. M., Mulukutla, G. K., & Mineau, M. M. (2014). Characterizing storm-event nitrate fluxes in a fifth order suburbanizing watershed using in situ sensors. Environmental Science & Technology, 48(14), 7756–7765.
Daley, M.L. (2002). Export of dissolved organic carbon, dissolved organic nitrogen and nitrate from the Lamprey River Watershed, New Hampshire: examining relationships with watershed characteristics. Thesis, University of New Hampshire, Durham, NH.
Dufour, A.P., Schaub, S. (2007). The evolution of water quality criteria in the United States, 1922-2003. In: L. J. Wymer (Ed.) Statistical framework for recreational water quality monitoring. John Wiley & Sons.
Fremaux, B., Gritzfeld, J., Boa, T., & Yost, C. K. (2009). Evaluation of host-specific Bacteroidales 16S rRNA gene markers as a complementary tool for detecting fecal pollution in a prairie watershed. Water Research, 43(19), 4838–4849.
Gaffield, S. J., Goo, R. L., Richards, L. A., & Jackson, R. J. (2003). Public health effects of inadequately managed stormwater runoff. American Journal of Public Health, 9, 1527–1533.
Gameroff, M. (2002). Using the proportional odds model for health-related outcomes. SAS SUGI paper 205–30. Retrieved from http://www2.sas.com/proceedings/sugi30/205-30.pdf.
Garcia, M., Darzacq, X., Delaveau, T., Jourdren, L., Singer, R. H., & Jac, C. (2007). Mitochondria-associated yeast mRNAs and the biogenesis of molecular complexes. Molecular Biology of the Cell, 18, 362–336.
Gerber, A. S., Loggins, R., Kumar, S., & Dowling, T. E. (2001). Does non-neutral evolutions shape observed patterns of DNA variation in animal mitochondrial genomes? Annual Review of Genetics, 35, 539–566.
Griffith, J., Wisberg, S., & McGee, C. D. (2003). Evaluation of microbial source tracking methods using mixed fecal sources in aqueous test samples. Journal of Water and Health, 1, 4.
Harwood, V. J., Brownell, M., Wang, S., et al. (2009). Validation and field testing of library-independent microbial source tracking methods in the Gulf of Mexico. Water Research, 43, 4812–4819.
Harwood, V. J., Staley, C., Badgley, B. D., Borges, K., & Korajkic, A. (2014). Microbial source tracking markers for detection of fecal contamination in environmental waters: relationships between pathogens and human health outcomes. FEMS Microbiology Reviews, 38, 1–40.
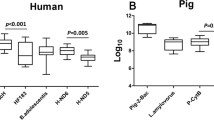
He, Xiwei, Liu, Peng, Zheng, Guolu, Chen, Huimei, Shi, Wei, Cui, Yibin, Ren, Hongqiang, Zhang, Xu-Xiang. (2016). Evaluation of five microbial and four mitochondrial DNA markers for tracking human and pig fecal pollution in freshwater. 6:35311.
Jent, J. R., Ryu, H., Toledo-Hernández, C., Santo Domingo, J. W., & Yeghiazarian, L. (2013). Determining hot spots of fecal contamination in a tropical watershed by combining land-use information and meteorological data with source-specific assays. Environmental Science & Technology, 47, 5794–5802.
Kaushal, S. S., Groffman, P. M., Band, L. E., Elliott, E. M., Shields, C. A., & Kendall, C. (2011). Tracking nonpoint source nitrogen pollution in human impacted watersheds. Environmental Science & Technology, 19, 8225–8232.
Kortbaoui, R., Locas, A., Imbeau, M., Payment, P., & Villemur, R. (2009). Universal mitochondrial PCR combined with species-specific dot-blot assay as a source-tracking method of human, bovine, chicken, ovine, and porcine in fecal-contaminated surface water. Water Research, 43, 2002–2010.
Layton, A., McKay, L., Williams, D., Garrett, V., Gentry, R., & Sayler, G. (2006). Development of Bacteroides 16S rRNA gene TaqMan-based real-time PCR assays for estimation of total, human, and bovine fecal pollution in water. Applied and Environmental Microbiology, 72, 4214–4224.
Lee, D. Y., Lee, H., Trevors, J. T., Weir, S. C., Thomas, J. L., & Habash, M. (2014). Characterization of sources and loadings of fecal pollutants using microbial source tracking assays in urban and rural areas of the Grand River watershed, Southwestern Ontario. Water Research., 53, 123–131.
Marsalek, J., & Rochfort, Q. (2004). Urban wet-weather flows: sources of fecal contamination impacting on recreational waters and threatening drinking-water sources. Journal of Toxicology and Environmental Health, 67, 1765–1777.
Martellini, A., Payment, P., & Villemur, R. (2005). Use of eukaryotic mitochondrial DNA to differentiate human, bovine, porcine and ovine sources in fecally contaminated surface water. Water Research, 39, 541–548.
McDowell, W. H., Bucci, J. B., Hobbie, E., French, C., Daley, M., Potter, J., & Miller, S. (2014). Nitrogen sources and transport pathways: science and management collaboration to reduce nitrogen loads in the Great Bay Reserve ecosystem. NOAA/NERRS Science Collaborative Research Project, 2010–2014.
Meays, C. L., Broersma, K., Nordin, R., & Mazumder, A. (2004). Source tracking fecal bacteria in water: a critical review of current methods. Journal of Environmental Management, 73, 71–79.
Neave, M., Luter, H., Padovan, A., Townsend, S., Schobben, X., & Gibb, K. (2014). Multiple approaches to microbial source tracking in tropical northern Australia. Microbial Open, 3, 860–874.
New Hampshire Department of Environmental Services (NHDES). (2013). Great Bay nitrogen non-point source study. NH: Concord.
Nguyet-Minh, V., Villemur, R., Payment, P., Topp, E., & Masson, L. (2012). Fecal source tracking in water using a mitochondrial DNA microarray. Water Research, 47, 16–30.
Olson, S.A. (2009). Estimation of flood discharges at selected recurrence intervals for streams in New Hampshire: U.S. Geological Survey Scientific Investigations Report 2008–5206, 57 p.
Pandey, P. K., Kass, P. H., Soupir, M. L., Biswas, S., & Singh, V. P. (2014). Contamination of water resources by pathogenic bacteria. AMB Express, 4, 51.
Piscataqua Region Estuaries Partnership (PREP). (2013). State of our estuaries. Durham: University of New Hampshire http://prep.unh.edu/resources/pdf/2013%20SOOE/SOOE_2013_FA2.pdf.
Purdue University (2010). Fisher procedure demonstrated with an example. Retrieved from http://www.stat.purdue.edu/tqin/system101/method/method_fisher_sas.htm.
Schill, W. B., & Mathes, M. V. (2008). Real-time PCR detection and quantification of nine potential sources of fecal contamination by analysis of mitochondrial cytochrome b targets. Environmental Science & Technology, 42, 5229–5234.
Scott, T. M., Rose, J. B., Jenkins, T. M., Farrah, S. R., & Lukasik, J. (2002). Microbial source tracking: current methodology and future directions. Applied and Environmental Microbiology, 68, 5796–5803.
Semenza, J. C., Herbst, S., Rechenburg, A., Suk, J. E., Hoser, C., & Schreiber, C. (2012). Climate change impact assessment of food and waterborne diseases. Critical Reviews in Environmental Science and Technology, 42, 857–890.
Shanks, O. C., Kelty, C. A., Sivaganesan, M., Varma, M., & Haugland, R. A. (2009). Quantitative PCR for genetic markers of human fecal pollution. Applied and Environmental Microbiology, 75, 5507–5513.
Simpson, J. M., Santo Domingo, J. W., & Reasoner, D. J. (2002). Microbial source tracking: state of the science. Environmental Science & Technology, 36, 5279–5288.
Stea, E. C., Purdue, L. M., Jamieson, R. C., Truelstrup, H.,. L., & Yost, C. K. (2015). Fecal contamination in the surface waters of a rural and an urban-source watershed. Journal of Environmental Quality, 44(5), 1556–1567.
Stoeckel, D. M., & Harwood, V. J. (2007). Performance, design, and analysis in microbial source tracking studies. Applied and Environmental Microbiology, 73, 2405–2415.
Toledo-Hernandez, C., Ryu, H., Gonzalez-Nieves, J., Huertas, E., Toranzos, G. A., & Santo Domingo, J. W. (2013). Tracking the primary sources of fecal pollution in a tropical watershed in a one-year study. Applied and Environmental Microbiology, 79(5), 1689–1696.
Trowbridge, P. R., & Jones, S. H. (2009). Detecting water quality patterns in New Hampshire’s estuaries using National Coastal Assessment probability-based survey data. Journal of Environmental Monitoring and Assessment, 150, 129–142. doi:10.1007/s10661-008-0683-x.
Trowbridge, P., Wood, M., & Underhill, J. (2014). NH DES Great Bay non-point source study report. New Hampshire: Concord.
University of California Los Angeles (UCLA) (2017). Statistical Analysis Using SAS. Retrieved from https://stats.idre.ucla.edu/sas/whatstat/what-statistical-analysis-should-i-usestatistical-analyses-using-sas/.
United States Department of Agriculture, (USDA) Natural Resources Conservation Service. (2010). Field indicators of hydric soils in the United States, version 7.0. In: L.M. Vasilas, G.W. Hurt, and C.V. Noble (eds.). USDA, NRCS, in cooperation with the National Technical Committee for Hydric Soils.
United States Environmental Protection Agency. US EPA. (2001). Nutrient criteria technical guidance manual: estuarine and coastal marine waters. Washington, DC: U.S. Environmental Protection Agency.
US EPA (2009). Review of published studies to characterize relative risks from different sources of fecal contamination in recreational water. U. S. Envirnmental Protection Agency, Office of Science and Technology. EPA 822-R-09-001. Washington, D.C.: United States Environmental Protection Agency.
Verhougstraete, M. P., Martin, S. L., Kendall, A. D., Hyndman, D. W., & Rose, J. B. (2015). Linking fecal bacteria in rivers to landscape, geochemical, and hydrologic factors and sources at the basin scale. PNAS, 112(33), 10419–10424.
Wade, T. J., Pai, N., Eisenberg, N. S., & Colford, J. M. (2003). Do U.S. Environmental Protection Agency water quality guidelines for recreational waters prevent gastrointestinal illness? A systematic review and meta-analysis. Environmental Health Perspectives, 111, 1102–1109.
Wade, T., Calderon, R., Sams, E., Beach, M., Brenner, K., Williams, A., & Dufour, A. (2006). Rapidly measured indicators of recreational water quality are predictive of swimming-associated gastrointestinal illness. Environmental Health Perspectives, 114(1), 24–28.
Waller, J. (2012). How to perform and interpret chi-square and t-tests. SAS Institute Global Forum Report, Paper 155-2012. http://support.sas.com/resources/papers/proceedings12/155-2012.pdf.
Wang, Y., Liu, V. W., Xue, S., Tsang, W. C., Cheung, P. K., & Ngan, H. S. (2005). The increase of mitochondrial DNA content in endometrial adenocarcinoma cells: a quantitative study using laser-captured micro-dissected tissues. Gynecologic Oncology, 98, 104–110.
Wood, M. A. & Trowbridge, P. (2014). "Nitrogen, phosphorus, and suspended solids concentrations in tributaries to the Great Bay Estuary Watershed in 2013". PREP Publications. Paper 252.
Acknowledgments
The authors sincerely thank Jody Potter for his assistance in sample collection, processing, and laboratory analysis and Inga Sidor for her expertise with qPCR assay development. Special thanks are due to Jay Levine at the School of Veterinary Medicine, North Carolina State University and Jane Caldwell at TransAgra International for their input.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding information
Funds supporting this study were provided in part by a NOAA National Estuarine Research Reserve System Science Collaborative grant.
Appendices
Appendix 1
Appendix 2
Rights and permissions
About this article
Cite this article
Bucci, J.P., Shattuck, M.D., Aytur, S.A. et al. A case study characterizing animal fecal sources in surface water using a mitochondrial DNA marker. Environ Monit Assess 189, 406 (2017). https://doi.org/10.1007/s10661-017-6107-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10661-017-6107-z