Abstract
Pythium isolates from diseased and dead bait plants of maize and cress grown in compost or various soils (maize fields, parkland under deciduous trees, grassland) were characterised and tested for pathogenicity to maize (Zea mays L.). In pot tests performed under controlled conditions, pathogenicity of the isolates to maize was apparent by reduction of root and shoot growth, whereas damping-off of maize seedlings was less frequent. Contrarily, pea seedlings were killed by pathogenic Pythium isolates. Pythium isolates from diseased maize seedlings and pathogenic strains from other gramineous plants (P. phragmitis, P. aff. phragmitis, P. catenulatum) were not necessarily more virulent to maize compared to isolates originating from dicotyledonous plants (cress). The most virulent isolates originated from compost and caused a reduction of maize shoot growth of up to 60%. Phylogenetic analysis revealed that they were very closely related to P. ultimum var. ultimum and P. arrhenomanes, respectively. Isolates originating from maize fields, grassland and parkland under deciduous trees, a reference culture of P. arrhenomanes and strains of P. phragmitis, P. aff. phragmitis and P. catenulatum with known pathogenicity on reed were non-pathogenic on maize. Isolates from compost, and from maize fields generally had a higher temperature optimum for mycelial growth (30 °C) and a faster growth rate (1.5–2.0 mm h−1) compared to the isolates from parkland under deciduous trees and grassland soil (20–25 °C, ~1.0 mm h−1), respectively. This study indicates a potential impact of pathogenic Pythium on maize plants even in the absence of visible symptoms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oomycetes of the genus Pythium are important soil-borne pathogens that cause seedling losses worldwide due to pre- and post-emergence damping-off and root rot. Pythium species are necrotrophs that are usually unspecialised in their host range. However, the bulk of studies on the role of Pythium as plant pathogen emphasises disease in dicotyledonous plants, where pre-emergence damping-off occurs more often, and ensuing economic losses tend to be more severe. Fewer studies have focussed on the role of Pythium species in monocotyledonous crops, including maize, where they have been identified as a cause of pre-emergence damping-off (Matthiesen et al. 2016) and root rot (Deep and Lipps 1996; Chamswarng and Cook 1985, Ingram and Cook 1990). The reports cited above originate from the U.S., and the role of Pythium as a maize pathogen in the U.S. has been further summarised by Munkvold and White (2016). Pythium has also been reported as pathogen of gramineous crops in Japan (Toda et al. 2015) and as the cause of stalk rot on maize in China (Song et al. 2015). Reports of Pythium infecting maize in Europe are scarce, but anti-oomycete seed treatments for maize are available. The extent of damage due to Pythium is unknown.
Morphological identification of Pythium species is difficult and often ambiguous due to intraspecific variation and overlap of morphological traits between species (McLeod et al. 2009). Identification based on morphology becomes increasingly difficult if the species do not regularly form sexual structures which exhibit important taxonomic traits (Van der Plaats-Niterink, 1981). Molecular characterisation has emerged as a powerful tool to complement morphology-based taxonomic approaches. Lévesque and de Cock (2004) provided the first comprehensive molecular phylogeny of the genus Pythium based on sequences of the ribosomal RNA operon genes and used an analysis of the internal transcribed spacer (ITS) region to separate the taxonomic genus into 11 major well-defined clades (termed A through K). Nuclear ITS sequences and mitochondrial cytochrome c oxidase subunit I (COI) have been evaluated as complementary DNA barcoding markers for oomycete species delineation with particular emphasis on the genera Pythium and Phytophthora (Robideau et al. 2011). One of the main outcomes of this and further studies using the ITS marker has been the reorganisation of Pythium clade K into the independent new genus Phytopythium (Schroeder et al. 2013, and references therein).
The aim of the present study was to determine the potential of Pythium species to cause disease on maize, one of the most important crops in Central Europe. Apart from agricultural soils, natural habitats are potential sources of Pythium species with unknown properties; some of these species may be aggressive pathogens (Nechwatal and Mendgen 2009). The campus area of the JKI Institute for Biological Control, Darmstadt contained a variety of promising edge habitats close to the experimental areas (compost, grasslands and parklands) that were sampled alongside agricultural soils from maize fields. We expected soil samples from such different habitats to yield a relatively comprehensive representation of Pythium isolates for characterisation and pathogenicity screening. Additionally, European isolates of the reed pathogens P. phragmitis and related hybrids that had already been shown to be pathogenic on maize leaves in in-vitro tests (Nechwatal et al. 2005) were screened for their potential to cause disease in maize. Stock cultures from species known to be pathogenic to maize were also included for comparison.
Material and methods
Maintenance of isolates
Once obtained, all Pythium isolates were maintained on carrot juice agar (100 ml carrot juice in 1 L water agar; EDEKA Bio-Wertkost, EDEKA AG & Co. KG, Hamburg). Pythium isolates were stored as soil or water cultures.
Source of isolates
Pythium isolates (Table 1) were obtained from diseased maize (Zea mays L.) and cress (Lepidium sativum L.) seedlings used as bait plants in soils from various sources. Cress was chosen as an additional bait plant due to its fast growth and early onset of damping-off symptoms, allowing to capture a potentially greater number of potentially maize pathogenic Pythium isolates with a broad host range within a short time. Two soils were sampled from maize fields in a nearby village (Roßdorf, Germany). Further soils were obtained from the experimental campus area of the Julius Kühn-Institut, Institute for Biological Control in Darmstadt, Germany (termed “JKI” in Table 1; location 49°52′3.864”N, 8°40′58.835″E) which included parkland (under oak trees), grassland under an apple orchard and compost as habitats close to experimental areas. “Compost JKI “was approximately 2-years old, unheated and prepared from greenhouse waste (mainly peat-rich horticultural potting substrates and plant residues).
Trays (40 × 60 × 6 cm) were filled with the respective source soils and sown with maize seeds of cultivars “Ricardinio”, “P8000”, “Pr38A79”, “KWS Diplomat”, “Expert”, “Loreto” and “Marcello”; one set of trays was incubated at 10 °C, another set of trays was incubated at room temperature (15–20 °C) to capture isolates with diverse optimal growth temperatures. JKI compost was also sown with cress. Bait plants were grown until disease symptoms (damping-off in cress and retarded growth or dying of shoots in maize) appeared. Diseased seedlings were washed 3 × with sterile tap water, the roots, seeds and shoots dissected and placed on water agar (WA), synthetic low nutrient agar (SNA; Nirenberg and O’Donnell 1998) or potato dextrose agar (PDA). Outgrowing mycelia were transferred to carrot extract agar. Pythium isolations were identified by mycelial morphology; in most isolates, identity could also be confirmed by the presence of oogonia and oospores.
Pythium catenulatum 1-1bM Type D, P. phragmitis P13 Mü1s and P. aff. phragmitis P11 MT4s (putative hybrid of P. phragmitis and species related to P arrhenomanes (Nechwatal and Mendgen 2009) were obtained from the research group of Prof. Mendgen (University of Constance). Pythium catenulatum 1-1bM Type D was isolated by Wielgoss et al. (2009). P11 MT4s and P13 Mü1s originate from strains P11, P13, described by Nechwatal et al. (2005, 2008). Pythium oligandrum was isolated from a commercial preparation Polyversum ® (BIOPREPARÁTY, spol. s r. o., Únětice, Czech Republic).
Effect of temperature on growth of isolates
Pythium isolates were grown on carrot juice agar (10 ml medium per 9-cm Petri plate) to determine the optimal temperature for growth (5 °C, 10 °C, 15 °C, 20 °C, 25 °C, 30 °C, 40 °C). Colony diameters were measured daily until colonies covered the plate and growth rates were determined at their maximum during this time course. There were three replicate plates at each temperature. For a subset of isolates (CC-1, CC-3, CM-3, CM-8, PM-1, PM-5), the experiment was repeated.
Amplification, determination and analysis of internal transcribed spacer (ITS) sequences of Pythium isolates
DNA was isolated from aerial mycelium using the DNeasy Plant Mini Kit (QIAGEN GmbH, Hilden, Germany). PCR primers ITS1–0 (5’-CGGAAGGATCATTACCAC-3′; Bachofer 2004) and ITS4-H (5’-TCCTCCGCTTATTAATATGC-3′; Göker et al. 2004) were used to amplify a partial sequence of the ribosomal RNA operon comprising the complete 5.8S rRNA gene together with the flanking internal transcribed spacer regions ITS1 and ITS2. Red Hot Taq polymerase (ABgene, ThermoScientific) was employed for amplification in reaction mixtures containing 0.5 μM of each PCR primer, 0.2 mg ml−1 bovine serum albumin (New England Biolabs), and 0.25 mM of each dATP, dCTP, dGTP, and dTTP. The PCR programme consisted of an initial denaturation step of 2 min at 94 °C, followed by 30 cycles of denaturation for 30 s at 94 °C, annealing for 30 s at 55 °C, and elongation for 1 min at 72 °C, and a final elongation step of 5 min at 72 °C. Alternatively, ITS1f and ITS4r primers (White et al. 1990) and Phusion High-Fidelity DNA Polymerase with GC buffer (New England Biolabs) were used under a modified PCR programme (3 min initial denaturation at 98 °C, 35 cycles of denaturation for 10 s at 98 °C, annealing for 30 s at 64 °C, elongation for 25 s at 72 °C and a final elongation step of 10 min at 72 °C). PCR products were separated and visualised by conventional agarose gel electrophoresis using ethidium-bromide or Midori Green Direct (Nippon Genetics Europe) staining, purified using the QIAquick PCR purification kit (QIAGEN GmbH) and sequenced using the reverse PCR primer ITS-4-H by Seqlab GmbH (Göttingen, Germany).
For each isolate, raw sequence data were combined into a consensus sequence using the BioEdit software tool (Hall 1999) and the sequence compared against the Barcode of Life ITS sequence database (http://www.boldsystems.org). Assembled ITS sequences have been deposited in the Genbank database under Accession Nos. KY829003 - KY829015 and MK253663 (Table 1).
Preliminary screening for pathogenicity on seedlings
Preliminary screening of Pythium isolates for pathogenicity was done on maize seedlings in Petri dishes. As maize mostly did not develop very pronounced symptoms in this in-vitro test, pea (Pisum sativum L.) was additionally selected as a plant for preliminary screening as it displayed pronounced damping-off symptoms, making it suitable to screen for potentially maize pathogenic isolates with a broad host range. Seeds were surface sterilised (1% NaOCl 3 min, 3 washes with sterile water) and germinated between sterile moistened filter papers within Petri dishes; three maize seeds (cultivars “Ricardinio”, “P8000”, “Pr39A79”) and two pea seeds (cultivars “Lancet” and “Kleine Rheinländerin”) were placed in each Petri dish. One mycelial plug (1.2 cm diameter) was placed on the emerging root with symptoms assessed after 9–13 days. There were at least three replicate Petri dishes per isolate.
Pathogenicity testing in pot experiments with maize
The first series of pot experiments with maize (cultivars “Ricardinio” and “P8000”) was performed in 11 × 11 × 11 cm pots. Horticultural substrate (Fruhstorfer Erde Type T; HAWITA Gruppe GmbH, Vechta, Germany) was incubated at 60 °C for 3 days before use to eliminate potentially suppressive microflora or soil-borne pathogens (Koch 1999). This substrate was sieved (12 mm), mixed with sand (substrate:sand = 80:20 w/w) and adjusted to a gravimetric water content (GWC) of 0.73 (60% water holding capacity) before inoculation. Preparation of Pythium inoculum on pea mash and inoculation of the re-wetted substrate were performed as previously described (Schmidt et al. 2004). Substrate for controls was inoculated with 2.5% (w/w related to substrate fresh weight) sterilised pea mash only. Each pot was filled with 650 g of the substrate/inoculum mixture. Sixteen maize kernels per pot were sown at a depth of 2 cm. To minimise evaporation from the substrate it was covered with a 1 cm layer of vermiculite (7 g dry weight), which was moistened with a sprayer. Seedlings were grown for 14 days in a growth room at 15 °C under metal halide lamps (OSRAM Power Star HQI-T 400 W/N; Osram GmbH, München; 16 h / 8 h day/night). The distance between pot surface and lamps was 80 cm. At the end of the experiment, emergence, fresh weight (FW) and dry weight (DW) of roots and shoots, as well as shoot length were determined. For the determination of root FW and DW, all loosely adhering soil was removed. The large number of Pythium isolates could not be tested in one experimental run and therefore, sub-sets of isolates were tested in three consecutive runs. Isolates CM-3, MS-1, P. aff. phragmitis P11 MT4s, P. phragmitis P13 Mü1s, P. catenulatum 1-1bM2 Type D were tested in the first experimental run, isolates P. ultimum JKI12/03, CM-1, MM-1, PM-5 and P. arrhenomanes CBS 430.86 were tested in the second run and isolates CC-1, CM-8, GM-1, NM-1, PM-1, PM-2, PM-3, PM-4 were tested in the third experimental run. Controls without Pythium and the pathogenic isolate CC-3 as a positive control were included in every experimental run. The pathogenicity tests were performed with the two maize cultivars “P8000” and “Ricardinio” with three replicate pots for each isolate × cultivar combination.
Selected isolates (CC-1, CC-3, CM-3, CM-8, PM-1, PM-5) were re-tested in a repeat pot experiment following a modified procedure. Pythium isolates were grown in Erlenmeyer flasks on autoclaved buckwheat (Fagopyron tataricum): 50 g seeds with 50 ml demineralised H2O were autoclaved at 121 °C for 21 min and inoculated with five mycelial plugs; flasks were incubated at 20 °C for 72 h in darkness. It proved not to be necessary to autoclave the buckwheat repeatedly to eliminate potential contamination. Horticultural substrate (Fruhstorfer Erde Type P; HAWITA Gruppe GmbH, Vechta, Germany) prepared as described above was mixed with sand (60: 40 w/w) and adjusted to a GWC of 0.45. Pythium inoculum grown on buckwheat was mixed into this substrate (1.5% of soil fresh weight); controls received no inoculum. Pots (8 × 8 × 8.5 cm, 0.544 L) were filled with 150 g of this substrate, sown with five maize seeds (cultivar Movanna) each, and covered with vermiculite. Five replicate pots were set up per treatment. The pots were then placed for 13 days in a growth room at 23 °C, ~45% rH, 7300 lx, 16 h/8 h day/night. In all pot experiments, pots were re-watered gravimetrically every 2–3 days. Treatments were randomised.
Data analysis
The data from the in planta screening were analysed using SAS® version 9.4 (SAS Institute Inc., Cary NC, US). The data from the first series of pot experiments consisting of three experimental runs were pooled into one dataset. To analyse data originating from several experimental runs, the experimental runs were considered a series with repeated measures to take into account the covariance structure and were analysed using a mixed model two-way analysis of variance (ANOVA) with Pythium isolate as factor A and maize cultivar as factor B. The repeat pot experiment was analysed as a separate data set. The differences between the Pythium isolates were calculated using LSMEANS Isolate / adjust = simulate according to Schumacher and Weimer (2006). Dunnett’s test was employed to assess whether there was a significant difference between the control treatment and single Pythium isolates, i.e. whether the reduction in growth due to a specific Pythium isolate compared to the control was significant. Differences between the single treatments were also analysed using adjust = simulate.
Results
Pre-screening and physiological characterisation of Pythium isolates
Among the source soils, compost from the JKI campus yielded a far higher number of Pythium isolates compared to the other substrates (Table 1). Cress sown in compost showed nearly 100% mortality due to damping-off and yielded 24 Pythium isolates while maize plants sown in compost yielded a further 16 isolates. Due to the high number of isolates from this one substrate, with a potential high risk of multiple, clonal isolations originating from the same mycelium, isolates from compost were pre-screened for plant pathogenicity in vitro on maize and pea seedlings. Pea was included as a susceptible indicator plant for screening of potentially maize-pathogenic isolates with a broad host range, as it developed damping off symptoms very rapidly within 9 days. Nearly 80% of isolates from cress (20 out of 24) and half of the isolates from maize (nine out of 16) killed pea seedlings whereas only four isolates produced pronounced brown root discolouration or growth retardation in maize, but only rarely killed maize seedlings. These were isolates CC-1 and CC-3 from cress, and the isolates CM-3, and CM-8 from maize (Table 1). Among the isolates from compost, only those four maize-pathogenic isolates and a non-pathogenic isolate (CM-1) for comparison were included in further studies. As there were only a few isolates from the two maize fields and from the other soils (Table 1) they were all included in the pot experiments. This strategy ensured that isolates from a wide variety of substrates were included in subsequent investigations.
Effect of temperature on mycelial growth of isolates
With the exception of isolate CM-1, the isolates from maize fields and isolates from compost (MM-1, NM-1, MS-1, CC-1, CM-3) had higher temperature optima around 30 °C compared to those from parkland soil under deciduous trees (PM-1, PM-2, PM-3, PM-4, PM-5). Isolates with low temperature optima also had lower maximum temperatures around 30 °C (Table 1, Supplementary Fig. S1). Results of the first experiment were confirmed in a second run for the isolates CC-1, CM-3, CM-8, PM-1 and PM-5, whereas isolate CC-3 had a higher temperature optimum (30 °C) and faster growth rate (1.5 mm h−1) in the second run compared to the first run (1.0 mm−1) at a temperature range of 15–30 °C). Isolates with a high temperature optimum (CC-1, CM-3, CM-8 from compost, MS-1 from maize field soil) had an up to two-fold higher growth rate (1.5–2.0 mm h−1) compared to isolates with temperature optima around 20–25 °C (~1.0 mm h−1; Table 1). The temperature optimum of most isolates was rather broad and isolates with high temperature optima showed considerable growth also at lower temperatures (Supplementary Fig. 1).
Molecular-taxonomic characterisation of Pythium isolates
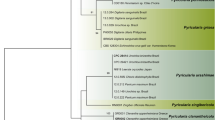
ITS1–5.8S rRNA gene and ITS2 consensus sequences from the different Pythium isolates ranged in length from 633 to 898 bp. When compared to Barcode of Life database entries, consensus sequences from all isolates had the greatest similarity (mostly >99%) with ITS barcodes representing one or several Pythium species (Table 2). For phylogenetic analysis, ITS sequences were aligned with a total of 115 reference sequences resulting in alignment of 1238 nucleotide positions, of which 931 were variable and 856 parsimony-informative. The reconstructed ML and NJ phylogenies (Supplementary Figs. S2A and B) had high bootstrap support and showed close agreement. Isolates in our study clustered within five of the main Pythium Clades A-K as defined by Lévesque and de Cock (2004) with bootstrap support approaching 100% for all clades, except for clade B. Systematic identification inferred from phylogenetics was highly consistent with the DNA barcoding results (Table 2). Both barcoding and phylogenetic analysis strongly support identification of isolates CM-8, CC-1, CC-3, and MS-1 as P. ultimum var. ultimum (Clade I), isolate PM-5 as P. perplexum (Clade J) and isolates PM-1, PM-3 and GM-1 as P. intermedium (Clade F) with no ambiguity (99–100% sequence homology, 89–100% bootstrap support; Table 2, Supplementary Fig. S2). Isolates PM-2 and PM-4 were isolated from the same habitat as PM-1 and PM-3, but were excluded from phylogenetic analysis due to poor sequence quality. However, Blast searches of PM-2 and PM-4 against the NCBI Genbank database yielded P. intermedium as the closest relative. The remaining isolates were related to groups and species where barcoding and phylogeny based on 16 s-rRNA do not allow a clear assignment to a single species (Robideau et al. 2011). NM-1 associated with Clade F (100% bootstrap support, 100% sequence homology to P. sylvaticum and P. terrestris; supplementary Fig. S2, Table 2). Isolates CM-3 and MM-1 had 99.8–100% sequence homology with P. arrhenomanes and P. aristosporum, and clustered within clade B (73% bootstrap support). Isolates CM-1 and 1-1bM Type D were associated with Clade B together with P. catenulatum and P. rhizo-oryzae whereas the isolate from Polyversum®, belonged to clade D comprising P. amasculinum, P. hydnosporum, P. lycopersicum, P. oligandrum, and P. ornamentatum (Table 2, Supplementary Fig. S2).
Pathogenicity to maize
In preliminary in vitro tests on maize and pea seedlings, most Pythium isolates killed pea seedlings, whereas pathogenicity to maize was only visible as a brown discolouration of roots. This was similar in the first series of pot experiments where Pythium never reduced the emergence of maize seedlings but selected isolates (CC-3, CM-8) caused 100% pre-emergence damping of pea (data not shown). Pathogenicity of Pythium isolates to maize was apparent in the reduction in shoot weight and shoot height. When data from the two maize cultivars “Ricardinio” and “P8000” from all three experimental runs were analysed together as a pooled data set (Table 3) the effect of isolate on shoot fresh weight (P = 0.023; Table 3) and shoot height (P = 0.049; Table 3) was significant. The effect on root fresh weight was close to significance (P = 0.052; Table 3), whereas there were no significant effects on dry weight (Table 3). The choice of maize cultivar affected shoot fresh weights (P = 0.009; Table 3). There was no significant interaction of the factors “Pythium isolate” and “maize cultivar” indicating that the effects of the Pythium isolates on maize growth were not cultivar specific, although differences existed (Supplementary Figs. S3 and S4). Thus, we averaged the effect of “Pythium isolate” over “maize cultivar” in Fig. 1, yielding six replicates for each isolate (18 replicates for the controls and isolate CC-3, which were included in every experimental run).
Effect of Pythium isolate on Ashoot fresh weight, B shoot height, and C root fresh weight of maize (first series of experiments). Growth time was 14 days at 15 °C in inoculated soil; pea mash overgrown with Pythium was used for inoculation of soil. Means and standard deviation indicated are from pooled results of three independent experimental runs with six replicate pots per isolate, each containing 16 maize seedlings (cvs. “Ricardinio and “P8000”). Isolate CC-3 and non-inoculated controls were included in all three experimental runs. Asterisks indicate significant differences to the non-treated control (P value of the post hoc Dunnett’s test in parentheses). Treatments with the same letter are not significantly different
Pythium isolate CM-8 caused the greatest reduction in shoot weight compared to the control (P = 0.015; Dunnett’s Test; Fig. 1a), followed by isolate CC-1 (P = 0.042; Dunnett’s Test; Fig. 1a); both isolates belonged to the species P. ultimum var. ultimum according sequence data. CM-8 also lowered shoot fresh weight compared to plants inoculated with any of the other isolates (multiple comparison at P < 0.05, Fig. 1a), and reduced shoot height (P = 0.028; Dunnett’s test Fig. 1b). While growth reduction by isolates CC-3 (P. ultimum var. ultimum) and CM-3 (P. arrhenomanes) was not significant when data for the two cultivars were pooled (Fig. 1), it was highly significant in cultivar “Ricardinio” (Fig. S3). Influence of Pythium isolates on root growth was less obvious, but a trend towards reduced root fresh weight was observed in maize inoculated with isolates CM-8 and CC-1 (Fig. 1c, Supplementary Figs. S3c and S3d, Fig. S4c and S4d). Unexpectedly, none of the isolates from the two maize fields had any significant effect on maize growth (Fig. 1, Supplementary Fig. S3). Isolates MS-1 and P. ultimum JKI12/03, and type strain P. arrhenomanes CBS 430.86, although closely related to the virulent isolates CC-1, CM-8 and CM-3, respectively, had no detectable effect on growth of maize seedlings. P. aff. phragmitis P11 MT4s, P. phragmitis P13 Mü1s also were non-pathogenic to maize, as were all isolates from parkland under deciduous trees and grassland soil (Fig. 1a and b, Supplementary Figs. S3, Fig. S4). The observed increase in shoot weight of maize cultivars Ricardinio by P. catenulatum 1-1bM Type D (Figs. S3a, S3b) could not be reproduced in repeated experiments (data not shown).
To confirm virulence of isolates CC-1, CC-3, CM-3 and CM-8 to maize, they were re-tested in a repeat pot experiment together with two isolates that had shown no effect (PM-1, PM-5) using a modified procedure (inoculum grown on buckwheat). Under these conditions, virulence of Pythium isolates was higher; contrary to the first series of experiments, significant pre-emergence damping-off was observed (Table 4). All four most virulent isolates (CC-1, CC-3, CM-3, CM-8) caused highly significant growth reduction of maize (P < 0.001; Table 4). General observations of the first series of experiments were reproduced: CM-8 was most virulent, followed by CC-1, CC-3 and CM-3, whereas PM-5 and PM-1 showed no or moderate virulence, respectively (Table 4).
Discussion
In this study, pathogenic Pythium species generally caused more severe symptoms on seedlings of dicotyledonous plants compared to maize. Broders et al. (2007) observed a similar trend with Pythium in soybean and maize, but a greater number of Pythium isolates were able to kill maize seedlings in their study. Pre-emergence damping-off in maize caused by Pythium was reported by other authors in the U.S. (Matthiesen et al. 2016) and in the Philippines (van Buyten et al. 2013). Although we screened a large number (54) of Pythium isolates originating from diverse soils and habitats, even the most virulent isolates did not cause significant pre-emergence damping-off in maize during the first screening, but only in the repeat pot experiment. One reason for these differing results might be the different inoculum type used (pea mash vs. buckwheat). On the other hand, inoculation of soil with overgrown pea mash produced 100% mortality in peas with selected isolates (CM-8, CC-3), and in sugar beet (Schmidt et al. 2004), indicating a higher virulence in dicotyledonous plants. Disease development caused by Pythium is generally promoted by high soil moisture content (Munkvold and White 2016; Schmidt et al. 2004), and consequently we adjusted conditions in our in planta-screening accordingly. Lower temperatures prolong germination times and thus extend the timespan where seedlings are susceptible to these soil-borne pathogens (Munkvold and White 2016). Thus, we chose moderate temperatures (15–23 °C) to mimic conditions in Central Europe more closely.
On the other hand, maize-pathogenic Pythium isolates found in this study (P. ultimum var. ultimum CM-8, CC-1 and CC-3, P. arrhenomanes CM-3) had high temperature optima for mycelial growth and the slightly elevated temperature in the second experiment (23 °C vs. 15 °C) may partially explain their higher virulence observed there. For P. arrhenomanes, the observed temperature optimum is consistent with prior observations reporting temperature optima of 25–30 °C (Lévesque and de Cock 2004) with a maximum temperature of 40 °C for both P. arrhenomanes and P. phragmitis (Nechwatal et al. 2005). Contrarily, maize-pathogenic Pythium species isolated in the U.S. had contrasting optimal temperatures with P. sylvaticum causing more severe rot at 18–23 °C and P. torulosum being more aggressive at 13 °C; this maximum virulence coincided with their temperature for optimal growth (Matthiesen et al. 2016). In our study, pathogenic isolates were able to reduce growth of maize seedlings already significantly at temperatures that were suboptimal for their mycelial growth (15 °C), possibly owing to their considerable growth rate already under suboptimal temperatures. This indicates a potential threat to the maize crop under a wider range of temperature conditions.
Although we isolated Pythium from a wide range of habitats, we were not able to detect novel species pathogenic to maize. The isolations we made were either closely related to P. ultimum var. ultimum or to P. arrhenomanes/aristosporum. While P. ultimum is more widely recognised as a pathogen of dicotyledonous plants, there is increasing evidence of its role as a pathogen of monocots. In the U.S., P. ultimum has been identified as the causal agent of pre-emergence damping-off in maize (Broders et al. 2007) and of root rot in maize and wheat (Munkvold and White 2016; Higginbotham et al. 2004; Ingram and Cook 1990; Chamswarng and Cook 1985). P. arrhenomanes has been reported as a causal agent of root rot in maize in the U.S. (Munkvold and White 2016; Deep and Lipps 1996), but also as a pathogen of other gramineous hosts (Toda et al. 2015; Van der Plaats-Ninterink 1981). It has been described as the most virulent species of Pythium causing disease in rice (Van Buyten et al. 2013). While we isolated a maize-pathogenic strain of this species in this study, a type strain of P. arrhenomanes (CBS 430.86) proved to be non-pathogenic to maize. The same was the case for the strains of P. phragmitis and P. aff. phragmitis, although the latter two are confirmed pathogens on reed (Nechwatal et al. 2005, 2008) and caused large leaf lesions on detached maize leaves, similar to P. arrhenomanes (Nechwatal et al. 2005). There is evidence that strains such as P. aff. phragmitis P11 MT4s are the result of hybridisation between P. phragmitis and an ancestor closely related to P. arrhenomanes, that possibly originated from nearby maize fields (Nechwatal and Mendgen 2009). Ability to cause lesions on detached leaves correlated with the ability to kill reed seedlings in infested substrate (Nechwatal et al. 2005) but the same may not necessarily be true for the isolates infecting maize. The observation that P. phragmitis could be recovered exclusively from reed leaves used as bait, but not from leaves of other grass species (Nechwatal et al. 2005) points to a greater specificity towards reed. Another possible aspect that might have contributed to the failure of P. arrhenomanes CBS 430.86, P. phragmitis and P. aff. phragmitis, and of some P. ultimum isolates (JKI12/03; two other isolates, unpublished data) to reduce the growth of maize seedlings may be loss of pathogenicity during long-term preservation in stock cultures. Temperature might be another relevant factor affecting pathogenicity, as the two isolates related to P. phragmitis had high temperature optima for mycelial growth (Nechwatal at al. 2005, 2008) and P. arrhenomanes CBS 430.86 was isolated from tropical rice fields (Van Buyten et al. unpublished; Genbank sequence HQ877870). The growth temperature of 15 °C in our in planta-screening may have been sub-optimal for the infection of maize. On the other hand, other Pythium isolates were pathogenic at this temperature, although it was substantially below their optimum for mycelial growth.
We could not assign the isolates related to P. arrhenomanes to this species with absolute certainty as ITS analysis does not separate P. arrhenomanes from P. aristosporum (Robideau et al. 2011), a Pythium species that has previously been isolated in the U.S. (Robideau et al. 2011) and in South Africa (McLeod et al. 2009). Cox II sequences also do not allow a differentiation between the two species (Nechwatal et al. 2005) which only differ in the number of antheridia per oospore (Van der Plaats-Ninterink 1981), and thus it has been put forward that there is little justification for P. aristosporum as a separate species (Nechwatal et al. 2005). We observed oogonia with antheridia attached only in one of the two isolates related to P. arrhenomanes (MM-1). This is consistent, however, with earlier observations of some isolates of P. arrhenomanes that lacked the ability to produce oospores (Deep and Lipps 1996).
Nechwatal et al. (2008) noted that bait plant species directly affected the species of Pythium that were isolated, as grass-associated Pythium species were isolated from reed, but not from oak tree leaves. Contrarily, in this study, isolates from maize were not necessarily more virulent on maize than those baited with cress; P. ultimum-related isolates originating from cress (CC-1, CC-3) were not only pathogenic to dicots (peas) but were also among the most virulent maize-pathogens. A similar broad host range was reported for isolates of P. sylvaticum, and P. torulosum in the U.S., which were pathogenic to both maize and soybean (Matthiesen et al. 2016). These two species, as well as P. irregulare and P. lutarium were also listed as important maize pathogens in the Midwestern U.S. by Munkvold and White (2016). We did not isolate these species.
A large number of Pythium isolates originating from apparently diseased or dead maize seedlings proved to be non-pathogenic on maize. One possible explanation is that other fungi and not Pythium were the primary cause of disease, with Pythium acting as a secondary coloniser. We could isolate Fusarium from maize seeds and also from diseased bait plants, and demonstrated the pathogenicity of some of these isolates on wheat (cultivar “Apogee”) and on maize (cultivar “P8000”) using the soil inoculation method of Bevivino et al. (1998) (unpublished results). In addition, the possible role of Pythium as secondary coloniser after infection by another pathogen is indicated by the co-isolation of non-pathogenic Pythium isolates alongside pathogenic Pythium from cress plants with severe damping-off symptoms in this study.
In conclusion, we demonstrated the presence of maize-pathogenic Pythium closely related or identical to P. ultimum var. ultimum and P. arrhenomanes in Central Europe. Although the symptoms were generally less pronounced than in dicotyledonous crops under identical test conditions, we could clearly show their potential to negatively impact the germination and early growth of maize seedlings. Broad temperature optima of maize-pathogenic Pythium strains indicate a potential to infect maize under a wide range of weather conditions. As the effects of Pythium spp. on seedling growth are non-specific and could be attributable to other causes, impact of pathogenic Pythium on early maize plant establishment might be underestimated and thus warrants further investigation.
References
Bachofer, M., (2004). Molekularbiologische Populationsstudien an Plasmopara halstedii, dem Falschen Mehltau der Sonnenblume. PhD thesis, University of Hohenheim, Stuttgart.
Bevivino, A., Sabrina Sarrocco, S., Claudia Dalmastri, C., Tabacchioni, S., Cantale, C., & Chiarini, L. (1998). Characterization of a free-living maize-rhizosphere population of Burkholderia cepacia: effect of seed treatment on disease suppression and growth promotion of maize. FEMS Microbiology Ecology, 27(3), 225–237.
Broders, K. D., Lipps, P. E., Paul, P. A., & Dorrance, A. E. (2007). Characterization of Pythium associated with corn and soybean seed and seedling disease in Ohio. Plant Disease, 91(6), 727–735.
Chamswarng, C., & Cook, R. J. (1985). Identification and comparative pathogenicity of Pythium species from wheat roots and wheat-field soils in the Pacific Northwest. Phytopathology, 75, 821–827.
Deep, I. W., & Lipps, P. E. (1996). Recovery of Pythium arrhenomanes and its virulence to corn. Crop Protection, 15(1), 85–90.
Göker, M., Riethmüller, A., Voglmayr, H., Weiß, M., & Oberwinkler, F. (2004). Phylogeny of Hyaloperonospora based on nuclear ribosomal internal transcribed spacer sequences. Mycological Progress, 3(2), 83–94.
Hall, T. A. (1999). BioEdit: A user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acids Symposium Series, 41, 95–98.
Higginbotham, R. W., Paulitz, T. C., & Kidwell, K. K. (2004). Virulence of Pythium species isolated from wheat fields in Eastern Washington. Plant Disease, 88(9), 1021–1026.
Ingram, D. M., & Cook, R. J. (1990). Pathogenicity of four Pythium species to wheat, barley, peas and lentils. Plant Pathology, 39, 110–117.
Koch, E. (1999). Evaluation of commercial products for microbial control of soil-borne diseases. Crop Protection, 18, 119–125.
Lévesque, C. A., & de Cock, A. W. A. M. (2004). Molecular phylogeny and taxonomy of the genus Pythium. Mycological Research, 108(12), 1363–1383.
Matthiesen, R. L., Ahmad, A. A., & Robertson, A. E. (2016). Temperature affects aggressiveness and fungicide sensitivity of four Pythium spp. that cause soybean and corn damping off in Iowa. Plant Disease, 100(3), 583–591.
McLeod, A., Botha, W. J., Meitz, J. C., Spies, C. F., Tewoldemedhin, Y. T., & Mostert, L. (2009). Morphological and phylogenetic analyses of Pythium species in South Africa. Mycolocial Research, 113(9), 933–951.
Munkvold, G. P., & White, D. G. (2016). Compendium of corn diseases. APS Press, St. Paul, Minnesota, U.S.A.: The American Phytopathological Society.
Nechwatal, J., & Mendgen, K. (2009). Evidence for natural hybridisation in reed associated Pythium species. Plant Pathology, 58(2), 261–270.
Nechwatal, J., Wielgoss, A., & Mendgen, K. (2005). Pythium phragmitis sp. nov., a new species close to P. arrhenomanes as a pathogen of common reed (Phragmites australis). Mycological Research, 109(12), 1337–1346.
Nechwatal, J., Wielgoss, A., & Mendgen, K. (2008). Diversity, host, and habitat specificity of oomycete communities in declining reed stands (Phragmites australis) of a large freshwater lake. Mycological Research, 112(6), 689–696.
Nirenberg, H. I., & O’Donnell, K. (1998). New Fusarium species and combinations within the Gibberella fujikuroi species complex. Mycologia, 90(3), 434–458.
Robideau, G. P., De Cock, A. W., Coffey, M. D., Voglmayr, H., Brouwer, H., Bala, K., Chitty, D. W., Desaulniers, N., Eggertson, Q. A., Gachon, C. M., Hu, C. H., Kupper, F. C., Rintoul, T. L., Sarhan, E., Verstappen, E. C., Zhang, Y., Bonants, P. J., Ristaino, J. B., & Lévesque, C. A. (2011). DNA barcoding of oomycetes with cytochrome c oxidase subunit I and internal transcribed spacer. Molecular Ecology Resources, 11(6), 1002–1011.
Schmidt, C. S., Agostini, F., Leifert, C., Killham, K., & Mullins, C. (2004). Influence of soil matric potential and soil temperature on sugarbeet seedling colonization and suppression of Pythium damping-off by antagonistic bacteria Pseudomonas fluorescens and Bacillus subtilis. Phytopathology, 94(4), 351–363.
Schroeder, K. L., Martin, F. N., De Cock, A. W. A. M., Lévesque, C. A., Spiess, C. F. J., Okubara, P. A., & Paulitz, T. C. (2013). Molecular detection and quantification of Pythium species: Evolving taxonomy, new tools, and challenges. Plant Disease, 97(1), 4–20.
Schumacher, E., & Weimer, M. (2006). Multiple Vergleiche mit der SAS-Prozedur MIXED. Proceedings of the 10. KSFE (Konferenz der SAS Anwender in Forschung und Entwicklung), Hamburg. http://de.saswiki.org/images/0/04/10.KSFE-2006-Schumacher-Multiple-Vergleiche-mit-der-SAS-Prozedur-MIXED.pdf. Accessed 24 November 2017.
Song, F.-J., Xiao, M. G., Duan, C. X., Li, H. J., Zhu, Z. D., Liu, B. T., Sun, S. L., Wu, X. F., & Wang, X.-M. (2015). Two genes conferring resistance to Pythium stalk rot in maize inbred line Qi319. Molecular Genetics and Genomics, 290(4), 1543–1549.
Toda, T., Iwasa, A., Fuji, S., & Furuya, H. (2015). Widespread occurrence of Pythium arrhenomanes pathogenic to rice seedlings around Japanese rice fields. Plant Disease, 99(12), 1823–1831.
Van Buyten, E., Banaay, C. G. B., Vera Cruz, C., & Höfte, M. (2013). Identity and variability of Pythium species associated with yield decline in aerobic rice cultivation in the Philippines. Plant Pathology, 62(1), 139–153.
Van der Plaats-Niterink, J. (1981). Monograph of the genus Pythium. Studies in Mycology, 21, 1–242.
White, T. J., Bruns, T. D., Lee, S., & Taylor, J. (1990). Analysis of phylogenetic relationship by amplification and direct sequencing of ribosomal RNA genes. In M. A. Innis, D. H. Gelfand, J. J. Sninsky, & T. J. White (Eds.), PCR protocols: A guide to methods and applications (pp. 315–322). New York: Academic Press.
Wielgoss, A., Nechwatal, A., Bogs, C., & Mendgen, K. (2009). Host plant development, water level and water parameters shape Phragmites australis-associated oomycete communities and determine reed pathogen dynamics in large lake. FEMS Microbiology Ecology, 69(2), 255–265.
Acknowledgements
We wish to thank Caroline Bogs (University of Constance, Germany) for the provision of the Pythium strains P. catenulatum 1-1bM Type D, P. aff. phragmitis P11 MT4s and P. phragmitis P13 Mü1s and Petra Zink (JKI Darmstadt, Germany) for skilful technical assistance.
Funding
Open Access funding provided by Projekt DEAL. This research was partially supported through employment of Christoph Stephan Schmidt in the long term project RVO 67985939.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Research involving human participants and/or animals
No animals or data from human participants were involved in the study.
Electronic supplementary material
ESM 1
(PDF 1747 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schmidt, C.S., Leclerque, A., Pfeiffer, T. et al. Pathogenicity of Pythium species to maize. Eur J Plant Pathol 158, 335–347 (2020). https://doi.org/10.1007/s10658-020-02076-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-020-02076-9