Abstract
The fungus Puccinia coronata Corda. is the causal agent of crown rust on oats (Avena sativa) and grasses and the disease is a major problem in oat production causing devastating yield losses. The population biology of P. coronata in oat fields and on the aecial host in central Sweden was studied to get a deeper understanding of the role of the aecial hosts in the epidemiology of the disease. Samples were collected from the aecial hosts common buckthorn (Rhamnus cathartica) and alder buckthorn (Frangula alnus), and three adjacent spring oat (Avena sativa) fields. Microsatellite markers were used to evaluate the relationships between populations sampled from the different hosts. According to our results F. alnus can be excluded as a part of the oat crown rust disease cycle. The results further show that samples collected from the aecial host were genetically separate from the population sampled in adjacent oat fields. Concurrently, the genotypic variation of P. coronata observed within oat fields was high. No population differentiation was observed within or between samples collected from different fields within the region, suggesting that airborne spores from other than the sampled specimens of the aecial hosts were contributing to the genetic diversity of P. coronata f. sp. avenae in the selected oat fields.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Oat crown rust, caused by the fungus Puccinia coronata f. sp. avenae has long been considered as one of the most devastating diseases on oats worldwide (Nazareno et al. 2018; Simons 1985). Like other cereal rusts, the life cycle of P. coronata includes two hosts and five spore stages (Simons 1970; Leonard and Szabo 2005; Jin et al. 2010), and an aecial host is a prerequisite for the fungi to complete the sexual part of their life cycle. The disease is named after the crown-like structures on the dark teliospores of P. coronate; however, it is the clonal, uredinial stage causing orange pustules on the oat leaves that causes the major yield losses.
The aecial hosts of P. coronata are Rhamnus spp. and Frangula spp. (Anikster and Wahl 1979; Hemmami et al. 2006; Dietz 1926; Gäumann 1959; Nazareno et al. 2018). In Sweden Rhamnus cathartica and Frangula alnus are prevalent in the agricultural landscape (Mossberg and Stenberg 2003) and both species harbor aecia annually. In spring, the teliospores on the remaining grass leaves germinate into basidiospores that can infect young leaves of the aecial hosts, where haploid pycnia are formed and fertilized by pycniospores from another pycnium. Fertilized pycnia form aecia that produce dikaryotic aeciospores. The aeciospores are dispersed to oats and other grass hosts in early summer (June in Sweden) where uredinia containing urediniospores are formed. The first disease symptoms of crown rust in oats are usually observed three to 4 weeks later.
Early studies by Eriksson and Henning (1896) showed that urediniospores from one grass host were unable to infect other grass hosts, and consequently divided the pathogen into different formae speciales. The type infecting oats was named P. coronata f. sp. avenae. Gäumann (1959) reported that the formae speciales infecting oats and some wild grasses (f. sp. avenae, f. sp. agropyri, f. sp. arrhenatheri, f. sp. bromi, f. sp. festucae, f. sp. lolii) infected the alternate host R. cathartica while other formae speciales (f. sp. agrostis, f. sp. calamagrostis, f. sp. holci and f. sp. phalaridis) infected the alternate host Frangula alnus. Phylogenetic studies of other cereal rusts have shown clear genetic differentiation between formae speciales (Abbasi et al. 2005; Liu and Hambleton 2010; Liu and Hambleton 2013). Leonard (2007) reported high phenotypic diversity in terms of numbers of virulence phenotypes in both aecial and uredinial populations of P. coronata in North America. Furthermore, the diversity in pathogenicity of a populations of P. coronata f.sp. avenae sampled in Minnesota, in proximity to the alternate host R. cathartica, was found higher than that of a population sampled in Texas, far away from the alternate host (Simons et al. 1979). In addition, studies of the genetic diversity of the rust fungi on their aecial host showed high genetic diversity within each aecial cluster, with each aecium (aecial cup) potentially producing spores of distinct genotypes (Anikster et al. 1999; Berlin et al. 2017; Rodriguez-Algaba et al. 2017), illustrating the diversity within cereal rusts in the presence of their alternate hosts.
The objective of this study was to investigate the role of the aecial hosts of P. coronata in the epidemiology of oat crown rust with the aim to improve the understanding of this pathosystem. We hypothesized that genotypes of P. coronata belonging to the same population can be found on both R. cathartica and oats, and that the inoculum initiating crown rust epidemics is dispersed from aecial hosts in close proximity to infested oat fields.
Material and method
Samples from the aecial hosts were collected at two locations (59.2011 N, 15.5988 E and 59.2922 N, 15.0651 E) in July 2015. To infer the population structure and diversity of P. coronata infecting oats, samples from the uredinal host were collected in three oat fields in close proximity to the aecial host locations, Field L (59.2009 N,15.5983 E), Field F (59.2837 N, 15.0642 E) and Field N (59.3017 N, 15.0566 E). At location L, R. cathartica was present 350 m from the sampled section of the field. Field F was located 950 m south of the F. alnus sampling site, and Field N 1.2 km North West of the F. alnus site. The distance between the two sampling locations (field F and field N) was 28 km. 30 oat leaves clearly showing uredinial infections were collected in each field in a regular grid design. Sampling was done along three rows, 10 m apart, and 10 samples were taken in each row at a distance of 10 m. All samples were kept in separate paper bags and were air-dried before further processing.
For DNA extractions, large aecial clusters were cut into three sections, while small aecial cluster were kept intact. This was done to reduce the risk of having more than one genotype within a sample (Berlin et al. 2017). The samples were put into 2-ml plastic tubes together with 30 glass beads and diatomaceous earth and extracted using the OmniPrep kit (GenoTech. St. Louis) according to manufacturer’s instructions for fungal tissues with minor modifications as described by Berlin et al. (2013). For samples collected from oats, single uredinia were carefully picked from the infected leaves. The DNA extractions were performed according to the same procedure as for the aecial samples. All samples were analyzed using 12 microsatellite markers specifically developed for Puccinia coronata f. sp. avenae (Dambroski and Carson 2008) (Table 1). Amplification of the microsatellites were performed in a PCR reaction including each of the following to the final concentration of 2 ng μl−1 DNA template, 0.02 mM dNTP, 0.2 m μM of each primer, 2.75 mM MgCl2, 0.05 U μl−1 DreamTaq DNA polymerase and DreamTaq buffer according to the manufacturer’s recommendation. The PCR program included an initial denaturation at 94 °C for 5 min, followed by 35 cycles of denaturation at 94 °C, annealing at 57 °C, and extension at 72 °C, each for 30 s, and a final extension step for 7 min at 72 °C. The PCR products were analysed by capillary electrophoresis (ABI 3730 XL DNA Analyzer). The length of the fragments was determined using GeneMarker (Softgenetics) and compiled to identify the multi locus genotype (MLG) of each sample.
The markers’ suitability to infer the population structure in this particular study was evaluated by calculating the expected heterozygosity (Hexp), evenness of the allelic distribution and possible deviation from Hardy-Weinberg equilibrium (Kamvar et al. 2014). Genotypic accumulation curves were calculated to confirm that adequate numbers of loci were included in this study (Kamvar et al. 2014). For the population biology analysis, samples were grouped based on host and location of the sampled oat fields. The genetic and genotypic diversity for each strata and strata combination were calculated in the R package Poppr (Kamvar et al. 2014), and the genotypic diversity is reported as the number of unique MLGs and expected heterozygosity (Hexp). The linkage disequilibrium was evaluated by calculating the \( \overline{r} \)D values (Agapow and Burt 2001) on clone corrected data to accommodate for unequal sample sizes (Lott et al. 2010). To determine the population differentiation between the different hosts, an analysis of molecular variance (AMOVA) between the different strata was performed using Poppr (Kamvar et al. 2014). The calculations of pairwise FST was performed using GenAlEx 6.51 (Peakall and Smouse 2012) and based on the heterozygosity between individuals within each population. The significance of each values was calculated by 9999 permutations. In addition, the Bayesian clustering software Structure version 2.3.4 (Pritchard et al. 2000) was used to infer the common ancestry among the different samples. The assumptions for this analysis were set to mixed populations with independent alleles. The range of presumed number of genotypic clusters (K) was 1–12 and the program was set to a burn-in period of 300,000 iterations followed by 300,000 iterations. The software calculates the Log likelihood for each K, and the highest likelihood was chosen as the most likely number of genotypic clusters. Five independent simulations were performed to test the consistency of the results. The most likely number of genetic groups was determined based on the Evanno method (Evanno et al. 2005) using STRUCTURE HARVESTER (Earl and von Holdt 2012).
Results
All 12 microsatellite loci used in this study were informative (Table 1) and found to be at Hardy-Weinberg equilibrium. The genotypic accumulation curve for the full data set was saturated at nine loci (data not shown). A total of 188 alleles were detected, ranging from 8 to 26 alleles by locus (Table 1).
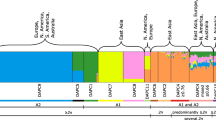
In total, 143 P. coronata samples were successfully genotyped and 142 unique MLGs detected (Table 2). Samples were omitted if there was a suspicion of mixture of genotypes or if the amplifications were unsuccessful. The samples collected from F. alnus clearly differed from the samples collected from R. cathartica and A. sativa. Two loci (PcaSSRA04 and PcaSSRA66) completely failed to amplify samples collected from F. alnus. For the loci PcaSSRB25, PcaSSRC26, PcaSSRC76 and PcaSSRC52, samples from one of the F. alnus sites failed to amplify the loci using these primers (Table 1). Result from the Bayesian cluster analysis showed that number of inferred clusters, K = 3 had the strongest support according to the Evanno method (Evanno et al. 2005), clearly dividing the samples into different populations based on the host from where it was collected. In Fig. 1, K 2 to 4 is presented, highlighting the robust population differentiation. The variation within and between the different hosts was examined by AMOVA. The largest variation was found within samples (71%), followed by the variation between hosts (25%) (data not shown). The pairwise FST between populations based on hots ranged from 0.296 to 0.401 and were all significant (Table 4). The large values reflects the differentiation between the populations visualized in the Bayesian cluster analysis (Fig. 1).
The population genetic data for the oat fields clearly shows a large genotypic diversity, as all MLGs were unique except for one MLG detected twice (Table 3). In addition, no linkage disequilibrium was observed in any of the three oat field populations, as the \( \overline{r} \)D values were not significant (Table 3). The variation within and between the three oat fields was examined by AMOVA. The largest variation was found within each MLG (99%), and no differentiation was found between fields (data not shown). The Bayesian clustering analysis confirms that all samples collected from the three different oat fields belonged to the same population when compared to samples from the other hosts (Fig. 1).
Discussion
In this study, P. coronata sampled from oat fields and from the two alternate hosts R. cathartica and F. alnus was shown to differentiate into genetically discrete groups (Tables 1 and 4, Fig. 1). A clear effect of host origin was observed as only the samples from F. alnus were split into two genetic clusters (Fig. 1), possibly caused by a large number of missing values due to the markers’ inability to amplify the targeted regions within that particular group of samples. Several formae speciales may simultaneously infect the same alternate hosts (Dracatos et al. 2010). The 12 microsatellite markers selected in our study were developed for P. coronata f. sp. avenae (Dambroski and Carson 2008), and all markers amplified samples collected on oats and most samples collected from R. cathartica, but performed less well on samples from F. alnus. The majority of alleles were shared between P. coronata infecting R. cathartica and oats (Table 1). Since the allele sizes detected in the samples from A. sativa and R. cathartica did not overlap with the ones from F. alnus, it is evident that these genetic groups are genetically different (Tables 1 and 4 ; Fig. 1). A reason for this could be that formae speciales infecting F. alnus are genetically more distant to P. coronata f. sp. avenae than the formae speciales infecting the same alternate host, R. cathartica. The established concept that P. coronata f. sp. avenae does not complete its sexual cycle on F. alnus (Gäumann 1959), thus excluding F. alnus as an alternate host in the oat crown rust epidemiology, is hereby supported. Concurrently, the absence of linkage disequilibrium suggests that P. coronata f. sp. avenae undergoes sexual reproduction in Sweden (Table 3).
Earlier studies show that common buckthorn, R. cathartica is the aecial host of P. coronata f. sp. avenae (Gäumann 1959; Anikster and Wahl 1979), and it was hypothesized that at least a proportion of the genotypes collected from R. cathartica and oats respectively would belong to the same population. This genetic link was not detected in this study, indicating that the importance of R. cathartica in the epidemiology of oat crown rust is unclear. The absence of a genetic link between the samples from oats and R. cathartica underlines that R. cathartica is host also for other formae speciales of P. coronata (Dracatos et al. 2010; Gäumann 1959). Both R. cathartica and F. alnus are prevalent within the studied agricultural area (Mossberg and Stenberg 2003), contributing to the possibility for the fungus to perform its sexual cycle between seasons. Increased prevalence of oat crown rust in recent years is likely linked to changes in agricultural practices such as increased practice of no tillage and an extensive use of oats as cover crop when establishing grass and legumes for forage ley, leaving infected crop residues in the field. In spring, the teliospores germinate into short-lived basidiospores that are released under wet conditions such as dew or rain showers, and infect the alternate hosts in the close surroundings (Zhao et al. 2016). Possibly, the sampled bushes of aecial hosts were not surrounded by oats the year prior to sampling, limiting the infections from P. coronata f. sp. avenae basidiospores on the alternate hosts. The missing genetic link between samples collected on R. cathartica and P. coronata found in the field in close proximity to the alternate host suggests that the inoculum originated from a more distant source. Thus, only a fraction of the large diversity within P. coronata aeciospores released from alternate hosts will constitute the initial source of inoculum causing oat crown rust. In a previous study of the stem rust pathogen P. graminis, Berlin et al. (2012) reported that samples of P. graminis collected from the aecial host barberry, harboured a range of different populations, and only few samples grouped in the same populations as samples collected from oats and rye (Secale cerealis). To determine the role of the aecial host for P. coronata f. sp. avenae, an extensive number of samples from potential aecial hosts need to be genotyped, in order to detect genotypes shared between populations collected from aecial hosts on one side, and populations of P. coronata f. sp. avenae collected from its uredinial host on the other side.
Most formae speciales of P. coronata have R. cathartica as aecial host (Gäumann 1959), and the aecial genotypes found in this study on R. cathartica and on F. alnus probably belonged to formae speciales other than P. coronata f. sp. avenae. From our results, it is also evident that the aecial hosts adjacent to an infested oat field are not the only source of inoculum for a particular field. Other aecial hosts within the area, or even within a region, will contribute to the pathogen diversity within a field. Basidiospores cannot disperse far because they are colorless, sensitive to UV light, and to dry conditions (Zhao et al. 2016). Aeciospores have a limited capacity of dispersal and primarily infect grass hosts in close proximity to the aecial host (Roelfs 1985). The urediniospores may spread for hundreds of kilometers (Nagarajan and Singh 1990), as they are colored and adapted to long distance dispersal (Simons 1970), hence, at the uredinial stage, genotypes may further spread between distant oat fields. Since the samples collected from different oat fields in this study belonged to the same P. coronata f. sp. avenae population, we suggest that the dispersal of inoculum initiating crown rust epidemics is not limited to fields in close proximity to the aecial hosts.
References
Abbasi, M., Goodwin, S. B., & Scholler, M. (2005). Taxonomy, phylogeny, and distribution of Puccinia graminis, the black stem rust: New insights based on rDNA sequence data. Mycoscience, 46, 241–247.
Agapow, P.-M., & Burt, A. (2001). Indices of multilocus linkage disequilibrium. Molecular Ecology Notes, 1, 101–102.
Anikster, Y., & Wahl, I. (1979). Coevolution of the rust fungi on gramineae and liliaceae and their hosts. Annual Review of Phytopathology, 17, 367–403.
Anikster, Y., Eilam, T., Mittelman, T., Szabo, L. J., & Bushnell, W. R. (1999). Pycnial nectar of rust fungi induces cap formation on pycniospores of opposite mating type. Mycologia, 91(5), 858–870.
Berlin, A., Djurle, A., Samils, B., & Yuen, J. (2012). Genetic variation in Puccinia graminis collected from oat, rye and barberry. Phytopathology, 102, 1006–1012.
Berlin, A., Samils, B., Djurle, A., Wirsén, H., Szabo, L., & Yuen, J. (2013). Disease development and genotypic diversity of Puccinia graminis f. sp. avenae in Swedish oat fields. Plant Pathology, 62(1), 32–40.
Berlin, A., Samils, B., & Andersson, B. (2017). Multiple genotypes within aecial clusters in Puccinia graminis and Puccinia coronata: Improved understanding of the biology of cereal rust fungi. Fungal Biology and Biotechnology, 4(1), 3.
Dambroski, H. R., & Carson, M. L. (2008). Development and characterization of novel, polymorphic microsatellite markers for oat crown rust, Puccinia coronata. Molecular Ecology Resources, 8, 1395–1398.
Dietz, S. M. (1926). The alternate hosts of crown rust, Puccinia coronata Corda. Journal of Agricultural Research, 33, 953–970.
Dracatos, P. M., Cogan, N. O. I., Keane, P. J., Smith, K. F., & Forster, J. W. (2010). Biology and genetics of crown rust disease in ryegrasses. Crop Science, 50(5), 1605–1624.
Earl, D. A., & von Holdt, B. M. (2012). Structure Harvester: a website and program for visualizing structure output and implementing the evanno method. Conservation Genetics Resources, 4(2), 359–361.
Eriksson, J., & Henning, E. (1896). Die Getrideroste ihre Geschichte und Natur sowie Massregeln gegen dieselben. Stockholm: P. A. Nordstedt & Söner.
Evanno, G., Regnaut, S., & Goudet, J. (2005). Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Molecular Ecology, 14(8), 2611–2620.
Gäumann, E. (1959). Die Rostpilze Mitteleuropas. Bern: Buchdruckeri Büchler & Co.
Hemmami, I., Allagui, M. B., Chakroun, M., & Gazzah, M. E. (2006). Rhamnus lycioides in Tunisia is a new aecial host of oat crown rust. European Journal of Plant Pathology, 115(3), 357–361.
Jin, Y., Szabo, L. J., & Carson, M. (2010). Century-old mystery of Puccinia striiformis life history solved with the identification of Berberis as an alternate host. Phytopathology, 100, 432–435.
Kamvar, Z. N., Tabima, J. F., & Grünwald, N. J. (2014). Poppr: An R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ, 2, e281.
Leonard, K. J. (2007). Persistent virulence associations in sexual populations of Puccinia coronata. Plant Pathology, 56, 35–45.
Leonard, K. J., & Szabo, L. J. (2005). Stem rust of small grains and grasses caused by Puccinia graminis. Molecular Plant Pathology, 6(2), 99–111.
Liu, M., & Hambleton, S. (2010). Taxonomic study of stripe rust, Puccinia striiformis sensu lato, based on molecular and morphological evidence. Fungal Biology, 114(10), 881–899.
Liu, M., & Hambleton, S. (2013). Laying the foundation for a taxonomic review of Puccinia coronata s.l. in a phylogenetic context. Mycological Progress, 12(1), 63–89.
Lott, T. J., Frade, J. P., Lockhart, S. R., & R, S. (2010). Multilocus sequence type analysis reveals both clonality and recombination in populations of Candida glabrata bloodstream isolates from U.S. surveillance studies. Eukaryotic Cell, 9(4), 619–625.
Mossberg, B., and Stenberg, L. (2003). Den nya nordiska floran, Wahlström och Widstrand.
Nagarajan, S., & Singh, D. V. (1990). Long-distance dispersion of rust pathogens. Annual Review of Phytopathology, 28, 139–153.
Nazareno, E. S., Li, F., Smith, M., Park, R. F., Kianian, S. F., & Figueroa, M. (2018). Puccinia coronata f. sp. avenae: A threat to global oat production. Molecular Plant Pathology, 19, 1047–1060.
Peakall, R., & Smouse, P. E. (2012). GenAlEx 6.5: Genetic analysis in excel. Population genetic software for teaching and research—An update. Bioinformatic, 28(19), 2537–2539.
Pritchard, J., Stephens, M., & Donnelly, P. (2000). Inference of population structure using multilocus genotype data. Genetics, 15, 945–959.
Rodriguez-Algaba, J., Sørensen, C. K., Labouriau, R., Justesen, A. F., & Hovmøller, M. S. (2017). Genetic diversity within and among aecia of the wheat rust fungus Puccinia striiformis on the alternate host Berberis vulgaris. Fungal Biology, 121(6), 541–549.
Roelfs, A. P. (1985). Wheat and rye stem rust. Orlando, Florida: Academic Press.
Simons, M. D. (1970). Crown rust of oats and grasses. Monographs. American Phytopathology Society, 5, 47.
Simons, M. D. (1985). Crown rust. In A. P. Roelfs & W. R. Bushnell (Eds.), The cereal rusts (Vol. II, pp. 131–172). St.Paul: Academic Press Inc.
Simons, M. D., Rothman, P. G., & Michel, L. J. (1979). Pathogenicity of Puccinia coronata from buckthorn and from oats adjacent to and distant from buckthorn. Phytopathology, 69, 156–158.
Zhao, J., Wang, M., Chen, X., & Kang, Z. (2016). Role of alternate hosts in epidemiology and pathogen variation of cereal rusts. Annual Review of Phytopathology, 54(1), 207–228.
Acknowledgements
We would like to thank Maria Jonsson at the Dept. Forest Mycology and Plant Pathology, SLU for technical assistance. This research was supported by Carl Tryggers Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The manuscript complies with the ethical standards of the journal.
Conflict of interest
The authors all agreed on this manuscript and declare no conflict of interest.
Rights and permissions
OpenAccess This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Berlin, A., Wallenhammar, AC. & Andersson, B. Population differentiation of Puccinia coronata between hosts –implications for the epidemiology of oat crown rust. Eur J Plant Pathol 152, 901–907 (2018). https://doi.org/10.1007/s10658-018-01605-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-018-01605-x