Summary
Melanoma has a high degree of malignancy and mortality. While there are some hopeful clinical trials for melanoma treatment in progress, they have not yet to yield significant long-term cure rates. Cancer vaccines including mRNA are currently one of the most promising strategy for tumor immunotherapy. The aim of this study was to analyze the potential tumor antigens in melanoma that could be used to develop mRNA vaccines and identify suitable vaccine populations. The gene expression data and complete clinical information of 471 melanoma samples and 1 normal tissue were retrieved from TCGA. Then, 812 samples of normal skin and their corresponding gene expression data were obtained from GTEx. Overexpressed genes, mutated genes and IRDEGs are used to identify potential tumor antigens. The relationship between the expression level of potential antigen and prognosis was analyzed in GEPIA, and then the immune cell infiltration was estimated based on TIMER algorithm. The expression profiles of IRDEGs were used to identify consensus clusters and immune subtypes of melanoma. Finally, mutational status and immune microenvironment characterization in immune subtypes were analyzed. Five tumor antigens (PTPRC, SIGLEC10, CARD11, LILRB1, ADAMDEC1) were identified as potential tumor antigens according to overexpressed genes, mutated genes and immune-related genes. They were all associated with OS, DFS and APCs. We identified two immune subtypes of melanoma, named IS1 and IS2, which exhibit different clinical features and immune landscapes. Based on the different immune landscape, we may conclude that IS1 is immunophenotypically “cold”, while IS2 is "hot". The present research implicates that PTPRC, SIGLEC10, CARD11, LILRB1 and ADAMDEC1 may be the antigenic targets for melanoma mRNA vaccines and IS2 patients may be more effective to these vaccines.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Melanoma has a high degree of malignancy and mortality, it is produced by malignant transformation of melanocytes. The incidence of melanoma has steadily increased in the past few years. Although melanoma is not as common in tumors, it accounts for the majority of skin cancer deaths. Ultraviolet exposure [1] is the most important risk factor for cutaneous melanoma, while other risk factors often include a personal history [2] or family history of cutaneous melanoma [3], multiple benign naevi or atypical naevi [4], phenotypic characteristics including freckling [5], blue eyes [6] and so on. So far, the primary treatment for early primary melanoma is surgery. Other adjunctive therapies include immune checkpoint blockade and RAF and MEK kinase inhibitors. As for metastatic melanoma, effective systemic therapies are still lacking. Whatsmore, due to the little progress made in medical treatment for metastatic melanoma, its prognosis and survival rate are poor. Therefore, identifying novel strategies to improve the prognosis of melanoma patients are of great clinical significance.
As an ideal immunotherapy target, tumor antigen has received more and more attention in recent years. Tumor antigen vaccine has many advantages, including multi-target, safety and broad spectrum. In addition, the tumor immune response it triggers in the body was dynamic and continuous. The sources of tumor neoantigens include single nucleotide variation, insertion and deletion, gene fusion, frameshift mutation, structural variation and so on [7]. Other tumor antigens include tumor-associated antigens, cancer-germline antigens [8]. Among the above tumor antigen vaccine, mRNA vaccines are a more novel class of vaccines and may be a major breakthrough in the future. In fact, the concept of genetic (DNA and RNA) vaccines has been around for decades [9, 10]. But in the last few years, mRNA vaccines have regained widespread interest due to the major technological innovation and research investment. It has several beneficial characteristics compared to other types of vaccines, such as safety, efficacy and high efficiency [11,12,13]. In addition, it has the potential for fast, cheap and scalable manufacturing.
Currently, some clinical trials have confirmed the positive effect of novel antigen vaccine in the treatment of melanoma. Ugurel et al. [14] found that part of melanoma patients who were treated with the neoantigen vaccine experienced significant tumor reduction or even complete remission. While there are some hopeful clinical trials for melanoma treatment in progress, they have not yet to yield significant long-term cure rates. So, the development of novel treatments still remains both essential and a major challenge. In this article, we aimed to identify potential cancer vaccine against melanoma and determined immune subtypes to identify candidate populations for mRNA cancer vaccination. We hope our findings might lead to new ideas for tumor immunotherapy and provide some new views for tumor vaccine research and development.
Methods and materials
Dataset and processing
The gene expression and full clinical information of 471 melanoma samples and 1 normal tissue were retrieved from The Cancer Genome Atlas(TCGA). Then, 812 samples of normal skin and their corresponding gene expression data were downloaded from Genotype-Tissue Expression (GTEx). And R package "limma" was used to merge the mRNA data in the two databases. To make the data more accurate, the merged data was further normalized. The data of somatic mutation in the VarScan2 platform was downloaded from TCGA.
We defined the genes in the merged cohort which were log2 fold change > 2 and adjustable p-value < 0.001 were upregulated. The mutated genes in melanoma and their chromosomal localization were performed with the R package “maftools”. Whatsmore, the ESTIMATE algorithm estimates the immune infiltration level (including Immunescore, Stromalscore and ESTIMATEScore) of each melanoma patient. All patients were divided into two groups(high score group and low score group) based on the median cutoff of Immunescore and Stromalscore, respectively. The differentially expressed genes of the two groups were defined as immune-related differentially expressed genes (IRDEGs). All these were caculated by R package. We defined the intersection of upregulated genes, IRDEGs, mutant genes as the possible mRNA tumor antigens in melanoma.
GEPIA
Gene Expression Profiling Interactive Analysis (GEPIA) contains a large amount of RNA sequencing data from the TCGA and the GTEx datasets. We use this tool to calculate each selected antigen's gene expression and patients’ survival information. The OS (overall survival) and disease-free survival (DFS) were performed by Kaplan–Meier method.
TIMER
Tumor Immune Estimation Resource (TIMER) is a resource to analyze gene expression level and its relationship with tumor-infiltrating immune cells (TIICs). Here, we focused on analyzing the association between the antigen-presenting cell(APCs) and the potential mRNA antigens.
Identification of immune subtypes
Based on the intersection of stromal score and immune score related DEGs, 910 genes were obtained, which defined as IRDEGs. Based on these IRDEGs’ expression profiles, the melanoma patients were clustered into different groups by the optimal k-means clustering. Cluster sets varied from 2 to 9. ConsensusClusterPlus R package was performed to identify a robust cluster, the number of cycle computation times was set to 1,000 to ensure stability and reliability.
Immune landscape and differential analysis of ICPs and ICD
ESTIMATE was performed to caculated the infiltration of 22 immune cells between different immune subtypes.
Then, considering the important role of immune checkpoint (ICP) and immunogenic cell death modulators (ICD) related genes in modulating the host anti-tumor immunity, we further analyzed the expression of ICPs and ICDs related genes in the two immune subtypes. The results were all filtered with p value < 0.05.
WGCNA
The coexpression modules of the IRDEGs was identified by “WGCNA” package of R software. The bottom-up algorithm and dynamic hybrid cut method were performed to examined coexpression gene modules. Then, the association between genes from different module and immune subtypes was recognize. In the same way, the results were also filtered with p value < 0.05.
Results
Screen five genes as possible antigens in melanoma
Firstly, we screened out 736 upregulated genes in melanoma compared to normal skin samples with log2 fold change > 2 and p-value < 0.001. Figure 1C showed the distribution of over-expressed genes of melanoma on chromosomes. Subsequently, 941 mutated genes were screened (the number of mutations in melanoma patients was greater than 50). The chromosomal locations of the mutated gene (the number of mutations is greater than or equal to 100) were shown in Fig. 1A and waterfall plot (Fig. 1B) showed the top 20 genes with the highest mutation frequency. Then, according to the intersecting of stromal related DEGs(1398 genes) and immune related DEGs(1142 genes), we screened out 910 genes as IRDEGs. 10 genes were obtained based on the intersection of IRDEGs, mutated genes, upregulated genes. To further optimize the results, the 10 genes were incorporated into the univariate cox regression analysis. Finally, five tumor antigens (PTPRC, SIGLEC10, CARD11, LILRB1, ADAMDEC1) were defined as possible cancer antigens for the development of mRNA vaccines.
Identification of potential tumor antigens in melanoma. A. The corresponding chromosome positions of the mutated genes(the number of mutations is greater than or equal to 100). B. Waterfall plot of the distribution of the top 20 mutant genes in melanoma. C. The distribution of over-expressed genes of melanoma on chromosomes according to GEPIA dataset
Identification of the five tumor antigens associated with melanoma prognosis and APCs
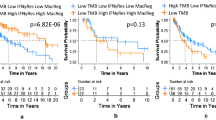
In order to identify the relationship between the five possible tumor antigens and prognosis, we analyzed their association with OS and DFS in melanoma patients. As shown in Fig. 2A-J, higher expressions of PTPRC, SIGLEC10, CARD11, LILRB1 and ADAMDEC1 were all associated with higher OS and DFS. These resuts showed that in melanoma, the higher the level of expression of these genes, the better the patient's clinical prognosis.
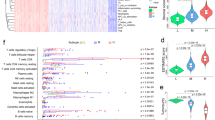
Then, immune cell infiltration estimation analysis was performed according to the TIMER algorithm, the results were shown in Fig. 3A-E. Dendritic cells, macrophages, and B cells are the main APCs. The expression levels of PTPRC, SIGLEC10, CARD11, LILRB1 and ADAMDEC1 were positively correlated with infiltration of macrophages and dendritic cells. As for B cells, the higher expressions of these five hub genes were also positively correlated with the levels of it, although the tendency were more variable. Taken together, these results demonstrate that the five tumor antigens can be presented to T cells by antigen-presenting cells(APCs) and initiate further immune responses.
Immune subtypes of melanoma
The expression profiles of 910 IRDEGs in 471 melanoma patients from TCGA database were used to identify consensus clusters. Based on their accumulative distribution functions as well as function delta area of K value, we selected k = 2. Two immune subtypes were abtained, which defined as immune subtype 1 (IS1) and immune subtype 2 (IS2) respectively (Fig. 4A-C). As shown in Fig. 4d, the patients in IS1 had a lower probability of survival than patients in the IS2. The heatmap in Fig. 4E shown the clinicopathological features and the expression level of the five tumor antigens among the two subtypes. Subsequently, we analyzed the specific proportions of different clinicopathological characteristics in the two subtypes. The expression levels of PTPRC, SIGLEC10, CARD11, LILRB1 and ADAMDEC1 were higher in samples of IS2 than in IS1 (Fig. 4E). The proportion of dead patients and male patients in IS1 were higher than in IS2 (Fig. 4F, H), while the differences were not significant in the proportion of patients over 65 years old and stage I-II patients (Fig. 4G, I).
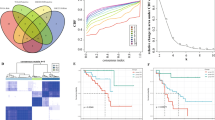
Identification of potential immune subtypes of melanoma. A-B. Consensus clustering CDF and Relative change in area under CDF curve when k = 2 to k = 9. C. Consensus matrix k = 2. D. Kaplan–Meier curves survival based on the two immune subtypes. E. The heatmap of expression levels of the five genes. F. Distribution ratio of fustat among IS1 and IS2. G. Distribution ratio of age among IS1 and IS2. H. Distribution ratio of gender among IS1 and IS2. I. Distribution ratio of stage among IS1 and IS2
Mutational status and immune microenvironment characterization in immune subtypes
We analyzed mutation for every patient. Among the two subtypes, IS1 had the higher mutation rate (91.75%), while the IS2 was 87.88%. In addition, the TTN mutation proportion was the highest in both subtypes, which is 72%(IS1) and 67%(IS2), respectively (Fig. 5A-B). Besides, TTN, MUC16 and BRAF accounted for the top three mutations in the two immune types. Therefor, we speculate that they might be the potential targets for mRNA vaccine.
Mutational status and immune microenvironment characterization in immune subtypes. A. Waterfall diagram of the top 20 mutated genes in IS1. B. Waterfall diagram of the top 20 mutated genes in IS2. C-E. Different expression of immune score, stromal score and ESTIMATE score in the two subtypes. F. Distribution of ICP genes among the two immune subtypes. G. Distribution of ICD genes among the two immune subtypes. H. Violin plot visualizing the differentially infiltrated immune cells among the two immune subtypes. ***P < 0.001; **P < 0.01; *P < 0.05
Next, we investigated whether there were differences in immune landscape between the two subtypes. The results revealed that compared with IS1, IS2 had higher immune score, stromal score and ESTIMATE score(Fig. 5C-E). Then, 22 different immune cell types among the two immune subtypes were performed by CIBERSORT algorithm. The proportion of 22 kinds of immune infiltrating cells were generally low. As shown in Fig. 5H, comparing with IS1, T cells CD4 memory resting, Macrophages M0, Macrophages M2 and mast cells resting were down-regulated in the IS2, while B cells memory, T cells CD8, NK cells resting, T cells CD4 memory activated, T cells follicular helper, Macrophages M1 were significantly up-regulated (p < 0.05). Therefor, we may conclude that IS1 may be an immune “cold”phenotype, while IS2 may be an immune “hot”phenotype.
ICPs and ICD play an important role in tumor immunization. Thus, we investigated whether there were differences in the expression of ICP and ICD related genes between the two immune subtypes. Among 47 ICP-related genes, 29 genes were distinctly expressed in the two subtypes. The specific results were that IS2 had higher expression of CD40LG, CD244, IDO1, TNFRSF18, TNFSF14, LAIR1, PDCD1, HAVCR2, CD86, CD274, LGALS9, TNFRSF25, ICOS, BTLA, CD27, TNFRSF8, LAG3, PDCD1LG2, CD80, TNFRSF4, CD28, TNFRSF9, CD48, CD40, TNFSF15, TNFSF18, CD200R1, TMIGD2, TIGIT(Fig. 5F). As for ICD genes, fourteen genes were differentially expressed in the two subtypes, IFNE, IFNAR2, EIF2AK2, IFNB1, CXCL10, CALR, ANXA1, TLR3, IFNAR1, EIF2AK3 and TLR4 in the IS2 group were higher than those in the IS1 group, while MET, EIF2AK1, IFNW1 were higher in the IS1 group(Fig. 5G).
Detection of immune gene coexpression modules of melanoma
910 IRDEGs coexpression modules were identified by the WGCNA. As shown in Fig. 6A, the soft threshold was set at seven based on the scale-free fit index. Different gene clusters were shown in different colors in the hierarchy clustering dendrogram, the modules determined by dynamic tree cuting and merged modules were shown on the bottom of the tree diagram. Similar modules were merged into a new one according to the following criteria: minimum module size = 30 and deep split = 4. Eventually, all IRDEGs were divided into five modules(brown, yellow, blue, green, grey) and the number of genes in each module varies from 49 in the green module to 496 genes in the blue module. We further conducted correlation analysis for modules and immune subtypes. The results were shown in Fig. 6C, MEbrown, MEyellow, MEblue and MEgreen were all negatively correlated with IS1 and the correlation coefficients were -0.64, -0.68, -0.62, -0.33, respectively(P values were all less than 0.05). While the results for IS2 were the opposite, which means the five module were positively correlated with IS2 and the correlation coefficients were the same as above.
Discussion
Melanoma has a high degree of malignancy and mortality. Malignant melanoma is derived from transformed melanocytes and the skin melanocytes are mainly located in the basal layer of the epidermis and in the hair follicles. At present, there is no long-term effective treatment for malignant melanoma. Immunotherapy has revolutionized oncology in recent years. mRNA vaccines, in particular, have been shown in many studies to be a promising immunotherapy. And therapeutic cancer vaccines are mainly aimed to stimulate cellular immunity. As for melanoma, they are particularly immunogenic due to their high mutation load. These characteristics have led to some advances in tumor immunotherapy for melanoma. For instance, Sahin U [15] introduced the concept of individualized mutanome vaccines and pioneered its use in melanoma. They designed a complete set of procedures for the development of personalized mRNA vaccines, including mutanome identification, novel epitope prediction and selection and the result was also encouraging. Although some clinical trials of melanoma mRNA cancer vaccines are ongoing, the clinical benefits remain limited. Therefore, the potential role of mRNA cancer vaccine in melanoma patients still needs to be explored.
In this study, five hopeful melanoma mRNA vaccine antigens were identified from overexpressed genes, mutated genes and immune-related genes, which were PTPRC, SIGLEC10, CARD11, LILRB1 and ADAMDEC1, respectively. The function of these several mRNA vaccine antigens(PTPRC, SIGLEC10, CARD11, LILRB1 and ADAMDEC1) needs to be further verified, but the role of some genes in immune response has been reported in previous studies. For example, Marc Rosenbaum [16] reported that CARD11-BCL10-MALT1 signaling mediates T cell receptor-induced NF-κB activation in Tregs and controls the conversion of resting Tregs to effector Tregs under homeostatic conditions. Major Histocompatibility Complex (MHC) class I plays a central role in the control of phagocytosis of macrophages by mediating inhibitory receptor LILRB1, which may be one of the targets of cancer immunotherapy [17]. Upregulation of these five genes’ expression were all associated not only with higher OS, but also with DFS, which means that these genes are significantly associated with the prognosis of melanoma. Subsequently, we evaluated the effectiveness and feasibility of these mRNA tumor vaccine antigens by exploring the relationship between antigens and APCs. APCs play an important role in the initiation of antigen-specific immune response. During this process, naive T cells must physically interact with APCs in order to eventually lead to T cell activation, mother cell formation and proliferation [18]. And our results shown that higher expression levels of the five antigens were significantly associated with increased tumor infiltration of macrophages, dendritic cells and B cells, which means the five antigens could be presented to T cells by APCs and initiate further immune responses.
To analyzed the most suitable population for mRNA vaccination, the melanoma patients were divided into two immune subtypes based on immune-related gene expression profiles. The clinical characteristics, mutation, survival prognosis and immune landscape of the two subtypes were further analyzed. Patients with IS2 have longer survival than those with IS1, suggesting that immunotype may be predictive of prognosis in patients with melanoma. We further analyzed the immune landscape in the two subtypes. The results of immune microenvironment characterization indicated that IS1 is an immune “cold” phenotype, while IS2 is an immune “hot” phenotype. Subsequently, the expression level of ICPs and ICD in the two subtypes were further analyzed. Immune checkpoint inhibitors are a new and effective strategy in recent years, PD-1/PD-L1 and CTLA-4 inhibitors have showed promising therapeutic effects and some have been approved for clinical use [19]. Our results showed that PDCD1 (PD-1) and CD274 (PD-L1) were highly expressed in IS2 compared with IS1. These immune characteristics suggest that different immune subtypes may respond differently to mRNA vaccines. IS1 was associated with low expression of B cells memory, T cells CD8, NK cells resting, T cells CD4 memory activated and was characterized by low infiltration of immune cells, therefore represents a tumor microenvironment with low inflammation. In contrast, IS2 is characterized by increased infiltration of immune cells and thus represents an extremely inflammatory microenvironment. Therefore, we speculate that these two subtypes may represent different immune mechanisms and corresponding therapeutic strategies should be different. As for IS1, to ameliorate the hypoimmunogenicity of this subtype, mRNA vaccines that stimulate the immune system by triggering immune cell infiltrating may be effective for these types of patients. The tumor microenvironment of the immune "hot" phenotype(IS2) is more complex. Previous studies have shown a close relationship between inflammation and melanoma. Melanoma cells express a variety of cytokines, such as IL-1, IL-6, IL-8, TNF-α, TGF-β, and they are essential for tumor progression [20]. Our study showed that the prognosis of IS2 is superior to that of IS1, which further suggests that triggering immune cell infiltration to stimulate the immune system may be a better therapeutic strategy for tumor vaccines.
Finally, WGCNA reveals the modules closely related to each immune subtype, which is important for exploring the underlying biological mechanisms of these subtypes.
Conclusion
The present research implicates that PTPRC, SIGLEC10, CARD11, LILRB1 and ADAMDEC1 may be the antigenic targets for melanoma mRNA vaccines and patients in IS2 may be more effective to these vaccines. And this research provides a theoretical basis for mRNA vaccine development of melanoma.
Data availability statement
This article are available from TCGA (https://portal.gdc.cancer.gov/); GEPIA (http://gepia.cancer-pku.cn/); GTEx (https://www.gtexportal.org/home); TIMER (https://cistrome.shinyapps.io/timer).
Change history
27 August 2022
A Correction to this paper has been published: https://doi.org/10.1007/s10637-022-01296-6
Abbreviations
- SNVs:
-
Single nucleotide variation
- INDEL:
-
Insertion and deletion
- TAAs:
-
Tumor-associated antigens
- CGAs:
-
Ancer-germline antigens
- GTEx:
-
Genotype-Tissue Expression GTEx
- IRDEGs:
-
Immune-related differentially expressed genes
- GEPIA:
-
Gene Expression Profiling Interactive Analysis
- OS:
-
Overall survival; DFS: disease-free survival
- TIMER:
-
Tumor Immune Estimation Resource
- TIICs:
-
Tumor-infiltrating immune cells
- APCs:
-
Antigen-presenting cell
- ssGSEA:
-
Single-sample gene set enrichment analysis
- ICP:
-
Immune checkpoint
- ICD:
-
Immunogenic cell death modulators
- WGCNA:
-
Weighted Gene Coexpression Network Analysis
- IS1:
-
Immune subtype 1
- IS2:
-
Immune subtype 2
- Tregs:
-
Regulatory T cells
- MHC:
-
Major Histocompatibility Complex
References
Gilchrest BA, Eller MS, Geller AC et al (1999) The pathogenesis of melanoma induced by ultraviolet radiation. N Engl J Med 340(17):1341–1348. https://doi.org/10.1056/NEJM199904293401707
van der Leest RJ, Flohil SC, Arends LR et al (2015) Risk of subsequent cutaneous malignancy in patients with prior melanoma: a systematic review and meta-analysis. J Eur Acad Dermatol Venereol 29(6):1053–1062. https://doi.org/10.1111/jdv.12887
Berwick M, Erdei E, Hay J (2009) Melanoma epidemiology and public health. Dermatol Clin 27(2):205–14, viii. https://doi.org/10.1016/j.det.2008.12.002
Tucker MA, Halpern A, Holly EA et al (1997) Clinically recognized dysplastic nevi. A central risk factor for cutaneous melanoma. JAMA 277(18):1439–44
Bliss JM, Ford D, Swerdlow AJ et al (1995) Risk of cutaneous melanoma associated with pigmentation characteristics and freckling: systematic overview of 10 case-control studies. The International Melanoma Analysis Group (IMAGE). Int J Cancer 62(4):367–76. https://doi.org/10.1002/ijc.2910620402
Marrett LD, King WD, Walter SD et al (1992) Use of host factors to identify people at high risk for cutaneous malignant melanoma. CMAJ 147(4):445–453
Schumacher TN, Schreiber RD (2015) Neoantigens in cancer immunotherapy. Science 348(6230):69–74. https://doi.org/10.1126/science.aaa4971
Simpson AJ, Caballero OL, Jungbluth A et al (2005) Cancer/testis antigens, gametogenesis and cancer. Nat Rev Cancer 5(8):615–625. https://doi.org/10.1038/nrc1669
Wolff JA, Malone RW, Williams P et al (1990) Direct gene transfer into mouse muscle in vivo. Science 247(4949 Pt 1):1465–1468. https://doi.org/10.1126/science.1690918
Jirikowski GF, Sanna PP, Maciejewski-Lenoir D et al (1992) Reversal of diabetes insipidus in Brattleboro rats: intrahypothalamic injection of vasopressin mRNA. Science 255(5047):996–998. https://doi.org/10.1126/science.1546298
Karikó K, Muramatsu H, Welsh FA et al (2008) Incorporation of pseudouridine into mRNA yields superior nonimmunogenic vector with increased translational capacity and biological stability. Mol Ther 16(11):1833–1840. https://doi.org/10.1038/mt.2008.200
Thess A, Grund S, Mui BL et al (2015) Sequence-engineered mRNA Without Chemical Nucleoside Modifications Enables an Effective Protein Therapy in Large Animals. Mol Ther 23(9):1456–1464. https://doi.org/10.1038/mt.2015.103
Karikó K, Muramatsu H, Ludwig J et al (2011) Generating the optimal mRNA for therapy: HPLC purification eliminates immune activation and improves translation of nucleoside-modified, protein-encoding mRNA. Nucleic Acids Res 39(21):e142. https://doi.org/10.1093/nar/gkr695
Ugurel S, Uhlig D, Pföhler C et al (2004) Down-regulation of HLA class II and costimulatory CD86/B7-2 on circulating monocytes from melanoma patients. Cancer Immunol Immunother 53(6):551–559. https://doi.org/10.1007/s00262-003-0489-1
Sahin U, Derhovanessian E, Miller M et al (2017) Personalized RNA mutanome vaccines mobilize poly-specific therapeutic immunity against cancer. Nature 547(7662):222–226. https://doi.org/10.1038/nature23003
Rosenbaum M, Gewies A, Pechloff K et al (2019) Bcl10-controlled Malt1 paracaspase activity is key for the immune suppressive function of regulatory T cells. Nat Commun 10(1):2352. https://doi.org/10.1038/s41467-019-10203-2
Barkal AA, Weiskopf K, Kao KS et al (2018) Engagement of MHC class I by the inhibitory receptor LILRB1 suppresses macrophages and is a target of cancer immunotherapy. Nat Immunol 19(1):76–84. https://doi.org/10.1038/s41590-017-0004-z
Friedl P, Gunzer M (2001) Interaction of T cells with APCs: the serial encounter model. Trends Immunol 22(4):187–191. https://doi.org/10.1016/s1471-4906(01)01869-5
Darvin P, Toor SM, Sasidharan Nair V et al (2018) Immune checkpoint inhibitors: recent progress and potential biomarkers. Exp Mol Med 50(12):1–11. https://doi.org/10.1038/s12276-018-0191-1
Moretti S, Pinzi C, Spallanzani A et al (1999) Immunohistochemical evidence of cytokine networks during progression of human melanocytic lesions. Int J Cancer 84(2):160–168. https://doi.org/10.1002/(sici)1097-0215(19990420)84:2%3c160::aid-ijc12%3e3.0.co;2-r
Funding
This work were supported by the Natural Science Foundation of China(31760263) and the Non-profit Central Research Institute Fund of the Chinese Academy of Medical Sciences(2020-PT320-004).
Author information
Authors and Affiliations
Contributions
Haiqin Ping and Wenjun Yu designed and collected the data. Haiqin Ping, Wenjun Yu, Xiaoming Gong, Xin Tong, Zhaojun Chen, Caiyun Cai, Kai Guo and Cheyu Lin designed the research. Haiqin Ping drafted the manuscript, Hengning Ke revised the manuscript. All authors collected patient data and read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Conflict of inetrests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: Author information was missing and will be added.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ping, H., Yu, W., Gong, X. et al. Analysis of melanoma tumor antigens and immune subtypes for the development of mRNA vaccine. Invest New Drugs 40, 1173–1184 (2022). https://doi.org/10.1007/s10637-022-01290-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10637-022-01290-y