Abstract
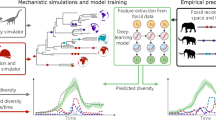
Fossils are the remains organisms from earlier geological periods preserved in sedimentary rock. The global fossil record documents and characterizes the evidence about organisms that existed at different times and places during the Earth’s history. One of the major directions in computational analysis of such data is to reconstruct environmental conditions and track climate changes over millions of years. Distribution of fossil animals in space and time make informative features for such modeling, yet concept drift presents one of the main computational challenges. As species continuously go extinct and new species originate, animal communities today are different from the communities of the past, and the communities at different times in the past are different from each other. The fossil record is continuously increasing as new fossils and localities are being discovered, but it is not possible to observe or measure their environmental contexts directly, because the time is gone. Labeled data linking organisms to climate is available only for the present day, where climatic conditions can be measured. The approach is to train models on the present day and use them to predict climatic conditions over the past. But since species representation is continuously changing, transfer learning approaches are needed to make models applicable and climate estimates to be comparable across geological times. Here we discuss predictive modeling settings for such paleoclimate reconstruction from the fossil record. We compare and experimentally analyze three baseline approaches for predictive paleoclimate reconstruction: (1) averaging over habitats of species, (2) using presence-absence of species as features, and (3) using functional characteristics of species communities as features. Our experiments on the present day African data and a case study on the fossil data from the Turkana Basin over the last 7 million of years suggest that presence-absence approaches are the most accurate over short time horizons, while species community approaches, also known as ecometrics, are the most informative over longer time horizons when, due to ongoing evolution, taxonomic relations between the present day and fossil species become more and more uncertain.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Fossils are remains of organisms from earlier geological periods preserved in sedimentary rock. The global fossil record broadly refers to all the geological evidence documenting the history of life on Earth. Analyzing this evidence to understand evolutionary change and its causal mechanisms is one of the fundamental questions in science, reaching back to Charles Darwin (1859) and beyond. The fossil record is the only knowledge we have about the world when it was in other circumstances than now. Analyzing how species have been distributed in space and how they have changed over time not only provides a basis for developing and testing advanced evolutionary theories, but also presents solid material for reconstructing how the environment has been changing over millions of years.
Understanding environmental change processes is attracting massive public attention, particularly in the context of rapidly changing world today (Barnosky et al. 2012). The broad interest has dramatically increased in the last few years to an extent that even creating a new geological epoch—Anthropocene—has been proposed, which conceptually begins when human activities started to have a significant global impact on Earth’s geology and ecosystems.
Similarly to software suites and benchmark data collections in other domains, a lot of domain knowledge and evidence have been put into rapidly growing fossil databases over the last few decades (Uhen et al. 2013). Such databases accumulate evidence and knowledge that geoscientists acquired over many years, and form the “big” data of paleontology. Datasets are compiled from multiple sources, they are to an extent subjective, include specific biases and uncertainties, data sparseness and quality varies over time and space. These datasets are complemented by satellite observations of the present day world, which present the ground truth for training predictive models. While not excessively large in storage volume, these datasets are covering a large part of all that is globally known of life in the past.
In this paper we discuss computational methodologies for building predictive models over fossil data for analyzing biospheric and environmental change. Such models are becoming increasingly popular for large scale reconstruction of the paleoclimate (Eronen et al. 2010b; Fortelius et al. 2016; Polly and Head 2015; Meloro and Kovarovic 2013), understanding evolution of faunal communities (Eronen et al. 2009; Sukselainen et al. 2015), analyzing contexts of human evolution (Fortelius et al. 2016), and providing quantitative reasoning and future insights for the ongoing efforts to mitigate climate change (Barnosky et al. 2017). We do not present conceptually new methods, but for the first time discuss from the machine learning, transfer learning and concept drift perspectives computational methods that have been engineered in paleobiology. In addition, for the first time we directly experimentally compare the performance of three main alternative methodological approaches. The particular instantiations of the approaches (a combination of dental traits for ecometric modeling, as well as predicting productivity based on the fossil assemblages of the Turkana Basin) reported in this paper are new. In addition to methodological contributions, the predictions themselves are of great interest for the paleoanthropology research community due to significance of the Turkana Basin for understanding early hominin evolution (Fleagle and Leakey 2011).
2 Fossils, fossil data and computational tasks
Life on the Earth leaves a track record of fossils. Fossilization of an organism is a very unlikely event, but over many millions of years even very small chances cumulatively produce a lot of fossils. Most typically fossils look like obscurely shaped pieces of rock, since the original parts of an organism have been replaced by minerals. Fossilization is a complex process, there is a whole discipline that studies the process itself, it is called taphonomy (Behrensmeyer et al. 2000). For fossilization to start, the body of a dead organism needs to be buried quickly and cut of interaction with oxygen, otherwise the body would disintegrate before fossilization process could start.
Not all the types of depositional environments are equally likely to yield fossils. Perhaps the most likely circumstances for fossilization are for an animal to fall into a body of water or a swamp and be quickly covered by sediments at the bottom (Mirsky 1998). Another quite likely situation is for an animal to fall into a cave and be buried under deposits there. A mature dense forest, on the other hand, is very unlikely to yield fossils, since the soil is rich in microorganisms which immediately recycle organic materials.
Millions of years later fossils can be found when the sediment, in which organisms have been buried and fossilized, gets exposed to the surface of the Earth by geological processes. Typically such exposures occurs at geological faults, where rocks get displaced due to tectonic forces affecting on the plate boundaries. The age of fossils is typically inferred from the age of the geological formation in which they were found. Thus, the locality from which a fossil comes from determines spatial and temporal information for that fossil.
It is rather unusual to find a complete organism as a fossil, typically only fragments remain. Mineralized body structures, such as teeth, bones or shells, are the most likely to be preserved, because in a way these parts are already somewhat like fossils. Even if an organism has initially fossilized as a whole, its parts can be lost later when tectonic processes, also water, wind and gravity, move sediments. Each fossil remain is assigned a taxonomic identification, aiming to identify it to the species level. Identification is based on expert opinions, and often is uncertain. The smaller the fragment, the more challenging it is for experts to identify the species. The difficulty of identification also depends on which body parts are preserved in the fragment. In case of mammals, teeth are generally easier to attribute to species than bones.
Fossil assemblage data have a large degree of uncertainty not only because of uncertainties in species identification, but also because fossilization is a very rare and unlikely event (Kidwell and Behrensmeyer 1988). Out of millions of individuals that lived at a particular time and place only very few or none actually turn into fossils. Therefore, datasets from fossil localities almost certainly provide incomplete species lists, lacking species that might have lived there, but no representatives have been found as fossils there (yet). A range of statistical methods has been developed to estimate and partially correct for such incompleteness (Foote and Sepkoski 1999).
Over long history of palaeontological research, the focus has shifted from primarily describing individual fossils, towards analyzing fossil assemblages and their ecological, climatic and evolutionary circumstances. The mainstream palaeontology research today is very computationally oriented. Aggregated data on fossil occurrences, including their location of origin, age, taxonomic identification, characteristics of fossil finds, fossil images and more are increasingly commonly stored in curated databases. There is no single database recording all the global fossil finds, the same way as there is no database recording all the available algorithms in data mining or machine learning, but there is a number of databases specializing in different aspects of the life history.Footnote 1 Such databases (Uhen et al. 2013) record data from many sources, including scientific articles, textbooks, expedition reports and museum collections around the world. There are many angles in which such data can be analyzed, and methodologies very often extend beyond classical statistical analyses or straightforward application of comparative methods. Many advanced computational methods have been developed over the last decades, as a outlined in a recent survey (Zliobaite et al. 2017). While more data sources and information types are aggregated, data are becoming more and more heterogeneous, more noisy and less structured, there is more and more room for tailor-made computational methods integrating formation of computational tasks, defining computational proxies (in a narrow sense this can be thought of as feature extraction), and designing algorithmic solutions.
From the data mining and machine learning perspective computational tasks in fossil data analysis fall under the following major categories:
-
Discovering macroevolutionary patterns and processes (originations extinctions), statistically correcting for potential biases and missing observations.
-
Constructing ancestry trees, identifying relationships between groups of organisms based on their physical characteristics and genetic information.
-
Reconstructing diets and ways of life from physical characteristics of fossil remains, analyzing how various parts of organisms and their ways of life scale with body size.
-
Identifying raise and fall of (faunal, floral or microbial) communities and tracking them over time and space, tracking human ancestors.
-
Inferring past climate (paleoclimate) and environmental change from fossil communities, reconstructing change processes.
Our study focuses on the last category—paleoclimate reconstruction. Approaches for reconstructing paleoclimate in general can be divided into two main groups: chemical and observational analyses. Chemical approaches, such as stable isotope tests (Cerling et al. 2015, 2011; Blumenthal et al. 2017), require taking destructive samples from fossil specimens. As a result, application of such approaches is limited by access to fossil collections and permits to sample materials from fossil specimens. Our account is focused on observational data analyses, that is, predictive modeling over data of fossil finds, without the need to access or sample the actual fossil specimens. Such analyses typically rely on large scale compilations of fossil data, such as data stored in fossil databases.
Predictive modeling for paleoclimate reconstruction has methodological similarities to ecological niche modeling. The goal there is to predicting or characterize environmental conditions at which a given species can occur (Myers et al. 2015; Warton et al. 2015). In paleoclimate reconstruction, which is the subject of our study, the task is to predict climate given species occurrences with or without complementary data, describing characteristics of those species. Conceptually, the task is similar to user modeling (Rich 1979): given, for instance, movies watched, books read or items bought the goal is to reconstruct user profile. In our setting users are pieces of land in space and time, and items are species that have occurred there.
3 Predictive modeling task
From the computational perspective the main objective is to infer past climatic and environmental conditions from data about fossil finds. Here we focus on the mammalian fossil record, particularly on that of plant eating mammals, but this modeling principle broadly applies to any animals or plants. Living mammals well understood from the physiological and ecological perspectives, therefore, this group provides a good reference for inferring the past conditions. Predictive models are trained on the present day for which climatic characteristics are known. Those predictive models are then applied to the fossil records to estimate and analyze the climate of the past.
The mammalian fossil record is composed of a set of localities (places) all over the world. A locality can be thought of as a pit in the ground from which fossils have been excavated, or an area of up to several square kilometers, where fossils have been collected from the surface, or a mixture of both. Each locality has a geological age or an age range assigned, typically determined by geologists from the sediment. From the data mining perspective these are spatiotemporal data with time stamps and geographic coordinates.
The modeling setting is as follows, schematically illustrated in Fig. 1. Environmental conditions of the past are generally unknown, there is typically no direct way to obtain labels at all. Therefore, models are built and calibrated on present day data of living species, where climate conditions can be measured, for instance, by weather stations. Predictive models built on present day data can then be applied to the fossil record, assuming that the fundamental relationships how animal communities interact with their environments stay the same over time.
Predictive modeling for paleoclimate reconstruction, a schematic illustration. In the present day world animal occurrences, their features and environmental conditions are known, the goal is to build predictive models relating distribution and/or features of animals to environmental conditions. In the fossil data animal occurrences and their features are known, environmental conditions are unknown—the goal is to predict them using models trained on the present day data, and use the predictions for analyzing environmental change over geological times and across space
The unit of analysis and an observation for predictive modeling is a site, which refers to a physical space. Sites can be existing units, such as a national parks, fossil excavation localities, or they can be arbitrary units obtained by placing a grid over the world map. Each site can be described by occurrence of animals or plants, as a vector of binary variables \(X = (x_1,\ldots ,x_m)^T\), where \(x_i=1\) denotes that species i is present at the site, and \(x_i=0\) means that it is absent. The setting resembles text processing in machine learning, where a site can be thought as a document, and the vector can be thought of as presence or absence of words. The fossil setting in principle can be extended to term frequency-inverse document frequency (TFIDF) setting, taking the frequency of species occurrences into account, but for clarity of exposition of the concept drift and transfer learning challenge within the focus of this study, we resort to presence-absence.
Each species x can be described by a vector of features \(H = (h_1,\ldots ,h_k)^T\). These features can be numerical, categorical, or a mixture of both. All m species considered in the presence-absence vector X are described by these features, for example, average body mass of the species, number of legs, type of teeth, presence of horns and such. In the document domain these features could be, for example, the length of the word, whether it is a verb, language of origin of the word, and such.
Each site can be described by physical climate or environmental characteristics, such as mean annual rainfall, mean temperature, whether it is a forest, a grassland, and alike, which is the prediction or classification target. For simplicity in this study we assume that the target is one dimensional, denote y. In the document classification example outlined earlier this can be, for example, the topic of the text.
The predictive modeling task is given occurrences of species and their characteristics to predict climate, that is to build a model \(\hat{y} = f(X,H),\) where \(\hat{y}\) denotes prediction, such that the expected deviation from the true value y is minimized. Deviation from the true target can be measured by least squares or any other optimization criteria of convenience.
One of the main challenges for such modeling is that present species are not the same as species in the past. The further to the past, the less overlap between the species lists at present and in the past is expected due to continuously ongoing evolution, origination of new species and extinction of past species, as explained by Darwin (1859) and immense body of evolutionary theory thereafter. Thus, from the data perspective there is a continuously ongoing concept drift (Gama et al. 2014), as illustrated in Fig. 2. The closer to the present day (left side of the timeline), the larger is the overlap with the genera that are alive today. No genera are the same as genera observed eight million years ago. If we were to use presence or absence of species as features for predictive model, there would be a gradual, but catastrophic drift in the feature space (Zliobaite and Gabrys 2014). Feature drift is the main instantiation of concept drift in this setting.
Summary of concept drift in the Turkana Basin fossil data (used in our experimental analysis later). Fraction of genera alive indicate which share of genera observed at that time are the same as genera alive at the present day. The horizontal axis indicates age of fossil sites in millions of years (1 is close to the present day, and 8 is far from the present day)
In the document example evolution of language presents an analogy. Suppose some documents today are describing the weather, and in the past they did as well. The challenge is that the words used to describe the weather phenomena now are not the same as they were in the past. If the modeling goal is to infer from the documents in the past how cold were the winters, we cannot directly rely on word occurrence as features. An analogy can also be made to different languages today. Suppose a corpus of documents describing winter activities is available in English, and independent temperature measurements corresponding to each document are available, and suppose there is another corpus of documents in German describing winter activities, and the goal is to predict temperature measurements from those documents. Some words describing winters in English and German will overlap, but many will not. A computational challenge will be to build a mapping from English to German for the purpose of such predictions.
In a similar spirit, in the fossil record setting the challenge is to construct feature mapping from present day species to the past species. The link might be explicit, where each species from the target domain is mapped to one species from the source domain \(g: \mathbf {X}^\star \longmapsto \mathbf {X}\), or implicit via distribution of descriptive features, to be detailed lated under term ecometrics. Explicit mapping may come from external sources, for example domain knowledge-based ancestry relationships, or such a mapping may be inferred from descriptive features in H, such that \(x_i^\star \longmapsto x_j: j = \arg \min _{j=1,\ldots ,m} dist (H_i,H_j)\).
From the machine learning perspective this predictive modeling setting falls under transfer learning (Pan and Yang 2010). Yet, in conventional transfer learning labels in the target domain are generally possible to obtain, but are costly. In the fossil analysis no labels for the target domain are available, they are impossible to obtain. If there was a way to obtain reliable labels, the predictive modeling approaches would not be needed. Analysis can and do cross-validate models on the present day, of course, expecting them to work for the past. But in such an extreme absence of feedback qualitative domain-based validation plays a critical role. We will first discuss three alternative computational approaches from the computational perspective, and at the end we will showcase a qualitative validation. We hope to open a study domain for the data mining and machine learning community, especially in the context of ongoing major efforts (Barnosky et al. 2017) calling for understanding the past in order to better navigate the changing world.
4 Transfer learning for paleoenvironment reconstruction
4.1 Modeling principles
Existing approaches for quantitative reconstruction of paleoclimate based on observational data fall into three main categories: co-occurrence, taxon assemblage and ecometric approaches.
Reasoning about paleoclimate based on fossil occurrences has been around for nearly as long as paleontology itself. Early qualitative approaches relied on parallels with living species. For example, if a fossil of an elephant has been found, one could, to an approximation, argue that the environmental conditions should have been similar to those where elephants live today. Quantitative approaches to climate estimation based on fossils go back to the nineteenth century when Oswald Heer estimated the yearly temperatures from the climatic requirements of the living relatives of the pre-Quaternary Cenozoic trees.Footnote 2 Recent approaches (Mosbrugger and Utescher 1997; Utescher et al. 2014) derive estimates from assemblages of species using similar principles. For each fossil species its climatic range is inferred by considering habitats of its living relatives. The climates of a fossil locality is then estimated as an intersection of the climatic ranges of all fossil species encountered at that locality. Since inference is based on intersection of ranges, it is not required that all living relatives share a locality in the present day. These approaches fall under lazy machine learning (Aha et al. 1991) since they do not produce a model in a functional form, instead generalization is delayed until a query about a particular locality is made.
And alternative is to consider as units of analysis real localities in the present day, where occurrence of taxa and climate variables are known, and fit a model in a functional form relating the two. Such approaches, known as taxon assemblage models, rely on species or higher taxa indicating particular environmental conditions. The relationships between occurrence of taxa and climate are modeled computationally by fitting, for example, a regression model (Fernandez and Vrba 2006). The main challenge is to find a mapping between the past species (features) and the present species, such mapping is most commonly done manually from domain knowledge. To the best of our knowledge computational approaches for discovering such mapping have not yet been reported, but there is a potential for discovering such mappings from data, for example, using information-theory approaches (Tatti and Vreeken 2012).
The third type of approaches does not explicitly link past and present species, but instead transform the feature space into a representation that is common for past and present species. In paleobiology such approaches are known as ecometrics,Footnote 3 and primarily refer to capturing functional relationships between faunal communities and their environments (Fortelius et al. 2002; Liu et al. 2012; Eronen et al. 2010a; Polly et al. 2011; Vermillion et al. 2017), as opposed to relying on taxonomic links in earlier two types of approaches. This computational methodology can be used for analyzing evolutionary contexts (Eronen et al. 2009; Schnitzler et al. 2017; Sukselainen et al. 2015), global scale relationships between animals, their environments (Liu et al. 2012; Zliobaite et al. 2016; Eronen et al. 2010a; Polly and Head 2015; Barr 2017; Lawing et al. 2012), reconstructing past climates and environmental change (Fortelius et al. 2016; Saarinen 2015; Eronen et al. 2010b; Meloro and Kovarovic 2013). Different traits have been explored for ecometric analysis. For plants, leaf shapes have been considered (Wolfe 1995; Traiser et al. 2005). For animals, considered traits include teeth (Eronen et al. 2010a; Fortelius et al. 2016; Liu et al. 2012; Zliobaite et al. 2016; Meloro and Kovarovic 2013; Polly and Head 2015), limbs and locomotion (Barr 2017; Polly and Head 2015), skeletal traits (Lawing et al. 2012), as well as body mass (Meloro and Kovarovic 2013). Traditionally, ecometrics refers to the analysis of animals. Conceptually similar computational approaches for the analysis of plant traits are referred to as transfer functions (Wolfe 1995; Traiser et al. 2005) or as species distribution models in ecology (Elith and Leathwick 2009; Ovaskainen et al. 2017). Generic statistical approaches, such as ordinary least squares regression, have served as core computational techniques in ecometrics until quite recently (Zliobaite et al. 2016).
Ecometric approaches link the present and the past via functional traits of species disregarding their shared origin. Even though most of present day species are likely to be different from those in fossil record, the functional traits of taxa, governed by the laws of physics, chemistry, and physiology, are likely to be similar in the present and in the past. For example, animals that run tend to leave the same pattern of skeletal architecture, e.g. long limbs (Reed 2013). Similar patterns of adaptation within communities would indicate similar habitats, therefore, reconstructing them can be approached as a predictive modeling task.
4.2 Relation to transfer learning and concept drift
In machine learning a situation where labeled training data is lacking is quite common, particularly in text or image processing domains. The branch of reusing models built on one problem for another but related problem is known as transfer learning (Pan and Yang 2010). There are three major categories of transfer learning: inductive transfer, where the source and target domains are the same, but the prediction tasks are different; transductive transfer, where the source and target domains are different, but prediction task is the same; and unsupervised transfer, where both are different. Environment reconstruction from the fossil record falls under transductive transfer. The prediction task is the same in the present and the past, but there is very little direct overlap in the feature space between the present, as the source domain, and the past, as the target domain. Within transductive learning our task falls under feature representation transfer approaches, where the goal is to find a feature representation that reduces difference between the source and the target domains and the error of classification or regression model (Pan and Yang 2010).
Feature representation transfer approaches have became increasingly popular about a decade ago in connection to natural language processing. There are two general types of solutions that can roughly be divided into supervised and unsupervised. The first type of solutions transforming (Raina et al. 2007; Argyriou et al. 2007; Davis et al. 2007), augmenting (Daume III 2007; Li et al. 2014) feature space, or inferring correspondence between features (Blitzer et al. 2006) in a supervised way. The second type of solutions assume some overlap between feature spaces and try to find a joint representation or links between the two domains in an unsupervised way, for instance, by means of co-clustering (Dai et al. 2007; Gopalan et al. 2014; Hoffman et al. 2012). The main difference from the fossil setting is that here at least some labeled data is assumed to be available for the target domain, while in the fossil setting there is none. Available instead is some auxiliary data that describes features, but not the instances in the source and target domain. This auxiliary data allows establishing links between the source and the target domain even at complete absence of labels in the target domain.
Concept drift is another closely connected research area in machine learning. Concept drift refers to changing data distribution over time, and as a result, predictive models having to detect changes and update themselves automatically to adapt to ever ongoing changes as new data keeps arriving (Gama et al. 2014). From the concept drift perspective transfer learning only deals with abrupt changes, changes are known in advance. That is, transfer learning does not need to detect change, the fact that the source and the target domains are different is known and the main algorithmic challenge is how to do adaptation when the target labels are scarce. Fossil data has a time dimension, like in the concept drift settings, but fossil data does not arrive in a stream over time. Therefore, there is no need for real time processing and no particular pressure for incremental algorithms. Unless a very coarse spatial resolution is necessary or climate models (Stute et al. 2001) need to be run in parallel to provide additional feedback for the analysis, it is reasonable and practical to have algorithms access and operate on all the historical data. In fact, data of the past is rather scarce, and the most recent data, for which labels are available, is more abundant.
Concept drift in fossil data can be of varying intensity, depending on how severe climate change has taken place, but most of the time an incremental and continuous drift is taking place, as can be seen from Fig. 2. The plot shows an aggregated statistics of the fossil record in the Turkana Basin area in Africa over the last 8 million years, from which a decline in feature overlap between the present and the past times can be seen. Standard adaptive preprocessing methods that have been developed to deal with drifts in feature space (Zliobaite and Gabrys 2014) do not directly apply here, since labeled data is only available at one point in time, at the end point of the time series, while those feature space adaptation methods require continuous feedback. This setting, thus, is a novel combination of transfer learning and concept drift settings, and therefore presents interesting algorithmic challenges that have not been considered before in either of the areas. We do not claim that the approaches formulated here solve the problem from the algorithmic point of view, but rather present simple baselines and a conceptual task setting for further research. These approaches can be relevant not only for geological data, but any historical data where we cannot go back to the past and obtain labels, for example, analysis of old texts, old demographic or social data.
4.3 Algorithmic approaches
Based on the existing work in palaeontology we formulate three baseline approaches for paleoclimate reconstruction. We analyze the performance experimentally with the focus on the transfer learning aspects to address the challenge that data is subject to a persistent concept drift over time. By concept drift here we primarily refer to drift in the feature space, as illustrated in Fig. 2. The traditional incremental concept drift methods (see e.g. Gama et al. 2014) do not apply to this setting, because there are no possibilities for online model updates. Standard concept drift handling methods require continuous feedback via arriving true labels, and in the fossil setting there are no labels for the past, but there is auxiliary data which can be used to link present and the past, and that is where transfer learning perspective comes into the solution. Transfer learning here refers to mechanisms for training predictive models that can be applied and are expected to perform well on data with different characteristics or different distribution from that of the training data. There are no model updates, as would be in the concept drift setting. But there are internal model training mechanisms that produce models tailored for the target data right away.
The algorithmic approaches presented here conceptually follow the three types of methods discussed in the related work section, but they are not direct replicas of those methods. We have simplified algorithmic approaches to make their transfer learning mechanisms comparable to each other, and therefore we refer to them as baselines. Our goal is to introduce the principle to the data mining and machine learning community, and leave computational choices open for future improvement. We therefore make the datasets and the code for our experimentally analysis publicly available.Footnote 4
The mean habitat approach works as follows. For each reference species from the present day the mean of the target variable representing environmental conditions is computed. Then for each fossil species the nearest living relative is found (based on domain expertise), this is the transfer learning aspect. The mean over environmental conditions of the nearest living relative is computed and assigned to the fossil species. A prediction for a given locality is a simple average over environments of all occurring species.
For example, if a fossil site has a giraffe and an elephant, we look where these animals or their nearest living relatives occur in the present day. Suppose we find three sites where elephants occur in the present day: with the mean annual temperatures correspondingly 20, 19 and 21 \({^{\circ }}\)C. The average environmental condition for occurrence of an elephant is thus 20. Similarly, suppose an average condition for occurrence of a giraffe is 24. Since the fossil locality has an elephant and a giraffe, we average over the mean environmental conditions of each animal occurrence and get a temperature estimate for the fossil locality equal to 22. The catch is that we find species in the fossil record that do not exist today, for example, a mammoth. Then we have to link mammoth with an elephant of the present day. This can be done computationally, but for now we use existing taxonomic trees to map an approximate relation. The nearest relative mapping has been assembled specifically for this paper and is given in the “Appendix” section.
The mean habitat approach is a simplified version of co-occurrence approach (Mosbrugger and Utescher 1997). The main difference is that instead of picking a range of overlapping habitats we take a simple average over all the habitats. The mean habitat approach does not explicitly build a predictive model, but works as an instance based learning approach. The approach is summarized in Algorithm 1.

The taxon assemblage approach works in a regular machine learning task setting, where the goal is to learn a predictive model over a set of binary features, where each feature describes presence or absence of a particular species. The learning instances are geographic areas (known as localities in the fossil record). The transfer learning element is that each species in the fossil record needs to be mapped to the species at the present day in order to match the input feature space of the training data (the present day) with the application data (fossils). The mapping is based on the identification of the nearest living relative, which requires domain expertise. We use the same mapping as for the mean habitat approach, the mapping is given in the “Appendix” section.
The taxon assemblage approach tested here is a simplified version of Fernandez and Vrba (2006). The main difference is that our simplified approach uses presence and absence data, while taxon-based approaches applied in paleobiology typically work on relative abundances of taxa (the proportion of each taxa in each locality). Here we use simple occurrence data in order for this approach to be experimentally comparable to the other two approaches, using the same occurrence information. The approach is summarized in Algorithm 2.

The functional approach (ecometrics) (Fortelius et al. 2002) works as follows. For each locality average traits can be computed over occurring species. This way the input space of the present day data becomes comparable to the input space of the fossil data. Any traditional machine learning model can be fit on the new input space. The approach is summarized in Algorithm 3.

This computational task setting closely relates to multiple instance learning (Zhou et al. 2012), where a bag of instances is a unit of analysis and predictive modeling. In our setting units of analysis are geographic areas, in the fossil record they are called localities. Each locality contains a different number of animal remains with their species identifications, and each species can be described by a set of quantitative features, as illustrated by a toy example in Fig. 3. The left panel illustrates a situation of extreme concept drift, where there is no overlap between the present day species and the fossil species. In such a case, instead of using species occurrence as input features directly, one can use traits as proxies, which can be measured for any species. A trait can be, for instance, body mass, number of legs, or height of teeth, as specified in Algorithm 3.
For example, if at one locality we have a giraffe and a zebra at a locality, we can compute the average length of a neck over these two occurring animals. If at another locality we have a hippo, and an elephant, we can compute average length of a neck over those two as well. Then instead of using animal occurrence as inputs to the models we can use average traits. This makes localities computationally comparable even if there are no overlap in species.
The mean habitat and taxon assemblage approaches use the nearest living relative as the transfer link between the present and the past. The transfer function is not learned computationally, but inferred from the domain knowledge. The latter, ecometric approach, learns the transfer function computationally using extra features that can describe species in the fossil record and at present day in the common feature space. The features for the taxon assemblage are binary, indicating presence or absence of a particular species at a particular site. The features for the eccometric approach are numeric, indicating average functional characteristics of species found at each site, for example, their body mass.
In this study we use eight features of mammalian teeth, described by Zliobaite et al. (2016): relative height of molar teeth, relative length of molar teeth, the number of longitudinal cutting edges, presence of sharp edges, presence of blunt edges, flatness, presence of rounded structural elements and presence of cement. Dental characteristics of plant eating mammals are reflective of their environments, because teeth act as an interface for obtaining energy from the environment. Different types of plant food require different properties of teeth, and different types of plants grow in different climatic conditions. Therefore, plants provide a functional link between plant eating mammals and climatic conditions. Plant types do not need to be known, they work as hidden variables in this predictive modeling. In palaeontology this approach is called dental ecometrics (Vermillion et al. 2017).
5 Computational experiments: predicting productivity in present day Africa
The goal of our first experiment is to test the effectiveness of the tree transfer learning strategies (mean habitat, taxon assemblage and ecometric approach) at the present day before proceeding to experiments with the fossil record. The task is to predict Net Primary Production (NPP) of environment, which is a rate at which plants produce biomass that is available for other participants of the ecosystem as food (energy) (see. e.g. Hairston and Hairston 1993). NPP, along with temperature or precipitation, is one of the key characteristics for describing terrestrial ecosystems. In present day ecosystems it is relatively straightforward to estimate NPP via remote sensing and/or numerical models of biosphere (Cramer et al. 1999), but remote sensing data is, of course, not available for the geological times. Instead, climate parameters of the past can be inferred from animal occurrences, documented in the fossil record. Therefore, the main intention behind building such models on the present day data is to be able to deploy them for analysis of the past.
5.1 Data and study region
We experimentally analyze the performance of three baseline approaches, described earlier. The task is to predict productivity using features derived from occurrence information and functional traits of large plant eating mammals (orders: Artiodactyla, Perissodactyla, Proboscidea and Primates). We work with animal occurrences at the Genus level, which is the next higher level in the taxonomic hierarchy after species. One of the reasons for this choice is that in the fossil record many specimens are not identified to the species level, leading to a lot of missing data otherwise.
The occurrence information for the present day is based on International Union for Conservation of Nature (IUCN) animal occurrence ranges, as compiled by Lawing et al. (2015). The world is divided into a grid of squares of approximately 50 \(\times \) 50 km, each grid cell makes one learning instance, for which animal occurrences and climate parameters are known. The target variable—observed NPP is computed from remote sensed and interpolated temperature and precipitation following Lieth (1975) as cited by Liu et al. (2012), as follows:
where \({\textit{MAT}}\) is the annual mean temperature in Celsius, \({\textit{MAP}}\) is annual precipitation in millimeters, and \({\textit{NPP}}\) is the net primary production in grams carbon per square meter per year of dry matter. MAT and MAP data comes from WorldClimFootnote 5 (version 1). All the datasets used for this analysis as well as our code will be made publicly available on Github upon publication of this study.
Our area of interest in the fossil record is the Turkana Basin in Africa, containing a rich fossil record of the last 7 million years, including key fossil evidence for current understanding of human evolution (Fleagle and Leakey 2011). Reconstructing climatic context of that area is, thus, of primary interest for understanding context of early hominin environments.
To roughly match expected range of environmental conditions of the fossil localities that we will analyze, for training the predictive models we use data from the present day tropical Africa, which is between − 25 and 25\(^{\circ }\) of latitude, that covers the majority of Africa below the Sahara desert. We exclude sites with extremely high precipitation (over 2000 mm per year) and sites more than half covered in agricultural land or water. The selected geographic region, along with the target values, is shown in Fig. 4.
5.2 Experimental protocol
We further partition the selected region of present day data into sub-regions, as illustrated in Fig. 5 and iteratively use them as training and testing datasets. Version (D) includes all three areas and is meant for building models to be applied on the fossil data later. Potential for overfitting is a serious concern, as in any predictive modeling study, and here in particular spacial autocorrelation is a concern, which means that sites nearby are likely to have similar animal occurrences and therefore cannot be treated as fully independent observations. To overcome potential spatial data leakage we introduce a safety margin between the testing and training data in this experiment. We leave out about 200 km of data from in between of training and testing areas, selecting areas for training and testing as depicted in the figure. In addition, we leave out the North East part of Africa from the experiment, since Sub-Sahel area is under strong pressure of agricultural and other human activities (Haberl et al. 2007). For this reason wild plant eating animal occurrences in this area are likely to be fewer than would be expected by the climate parameters of that area under regular relationships between animal occurrence and productivity of the ecosystems (Zliobaite et al. 2018).
Training and testing localities from the present day Tropical Africa: a east as training data, b south as training data, c diagonal training data, d all training data. Blue color indicates training data, orange color indicates testing data regions. Data dimensionality is given in Table 1 (Color figure online)
First we analyze how the prediction accuracy would be affected due to the nearest living relative transfer. That is, in the regular machine learning setting one would expect the input features (species) to be the same for training and testing data. In the fossil record analysis the same species are rarely available, instead surrogate features are used for training (the nearest living relatives of the fossil species). Here we identify those surrogate features following publicly available phylogenetic information. Table 3 in the “Appendix” section presents the identified relative species. As a toy example, if a mammoth was present in the fossil record but not available in the present day, instead of using a binary feature “is mammoth present at this locality”, in the training data one would replace the feature with “is elephant present at this locality”. Since in this section we are experimenting on the present day data (to be able to measure prediction accuracies), we assume that the testing dataset is fossil data and exactly the same
Nearest living relative based transfer is only applicable to mean habitat and taxon assemblage approaches, the ecometrics approach use a different transfer learning mechanism. The taxon assemblage and the ecometric approaches use partial least squares regression (Wold 1966) (linear model) with four components. Our first experiment in this section compares testing accuracies with the nearest living relative transfer to baseline accuracies that could be achieved if the same features were available in the training and in the testing data. We report prediction accuracies as the coefficient of determination (R2), which is the prediction accuracy normalized by the variance of the target variable. The best performance is when R2 = 1, and R2 = 0 means that the predictor is doing no better than a constant baseline.
5.3 Results
Table 1 summarizes the performance of the models. The lazy learning training accuracies are typically a bit higher than the transfer learning accuracies when comparing for the same modeling approach, since transfer learning may generate duplicate features (if two or more species in the fossil record are mapped into one species in the present day). This way the training data of the transfer would have a lower effective dimensionality. In most of the cases we can see substantial drops in testing accuracies due to having to use surrogate features, yet in all the cases the accuracies are positive and reasonably high for potential application of these models to the fossil record. Note that the baseline approaches are for illustration purposes only, they are not assumed to be applicable to the fossil record. On an absolute scale, the taxon assemblage transfer learning approach substantially outperforms the mean habitat approach, as it can be expected due to higher complexity of the model in the latter.
Table 2 compares all three transfer learning approaches, adding Ecometric approach. Ecometric approach is a transfer learning approach inherently and does not have a non-transfer learning counterpart. The transfer is via projection of the occurrence information to distribution of functional traits. As an example, a locality can be then described by the proportion of animals on that site possessing a give trait. For example, a site can be described by a share of animals that have high crowned teeth. In this analysis we describe each genera by eight dental traits, following dental trait characteristics such as presence of cutting edges, structural elements, topography of teeth and such, described in more detail in Zliobaite et al. (2016) and Fortelius et al. (2016). We build the ecometric model (later referred to as model I) using five manually selected dental variables to minimize potential overlap between variables capturing similar information. The variables we use for model I are: height of molar teeth (hypsodonty), the number of longitudinal cutting edges (lophs), structural fortification of cusps, flat occlusal topography and presence of cement.
We can see from the table that in two out of three testing cases the ecometric approach gives the best testing accuracy. When training on the South (B), taxon assemblage performs the best. The ecometric approach performs worse in this situation, because of known anomalies in the crown height distribution at the footsteps of the Sahara desert, where heigh crowned animals that could be expected to be there for a given climate are missing presumably due to high levels of agricultural activities. The ecometric transfer has been preferred from the evolutionary perspective (Fortelius et al. 2014), because it takes into account functional relationships between animals and their environment, instead of relying on the persistence of relations between animals and environment based on their common origin.
To the best of our knowledge this is the first computational comparisons of taxon-based and ecometric approaches to transfer learning. The difference is in how concept drift in the feature space (changing genera over time) is handled. The taxon-based approach relies on a direct matching of each feature from the source with a feature from the target. The ecometric approach relies on feature extraction, it projects source and target features into a common representation. Our computational analysis confirms expectations arising from paleobiology, that is, the ecometric approach performs well when the source and target features differ, but the underlying functional relationships stay fixed.
6 Case study: predicting net primary productivity from fossil data
The goal of our next set of experiments is to analyze how the these approaches perform on the fossil data. We will never know the ground true productivity corresponding to this fossil data. As in any predictive modeling, there is always a potential for overfitting, especially if we use very powerful models with a lot of degrees of freedom on small datasets. Our training datasets on the present day are not particularly small (in the order of thousands of observations) and the feature space is not particularly large (in the order of tens of observations). Our models are linear. Therefore, the risk of overfitting is not particularly high. A higher risk is that of concept drift, meaning that our models trained on the present day would fail to reflect patterns in the past accurately, and we would not know that, because there is no quantitative ground truth for the past. Therefore, this section emphasizes qualitative evaluation of performance and largely relies on visual inspection. Our goal with this section is not only compare the performance of the approaches, but also illustrate what qualitative aspects can be used for judging model performance.
The main criteria at hand is to inspect trends of predictions over time. Good predictions should not vary much up and down over short time periods time periods, but rather capture steady long term trends. In addition, the directionality of trends may be known from external sources. In the case of the Turkana Basin, a lot of research argues that the aridity in the area has been increasing towards present time (Cerling 1992; deMenocal 2004; Bobe 2006, 2011; Fernandez and Vrba 2006; Leakey et al. 2011; Cerling et al. 2015; Fortelius et al. 2016; Blumenthal et al. 2017), implying lower NPP closer to the present day, and we expect to see such a trend in our predictions.
Figure 7 shows predictions of the net primary productivity by the three models trained on the present day data (D) in the previous section. The last panel in the figure gives predictions of the ecometric model reported in Fortelius et al. (2016), which was made for precipitation and fit at the species level. We apply this model to the Genus level data here expecting that it would work to a reasonable approximation despite different data resolution, on which that model has been trained. We convert precipitation estimates produced by the model to net primary productivity estimates using equations from Lieth (1975) as reported by Liu et al. (2012).
Predictions of net primary productivity for fossil localities: comparison of trends obtained by three transfer learning models with linear predictors and a non-linear ecometric model developed by Fortelius et al. (2016) (Color figure online)
The two lines in each panel of the figure represent the eastern (blue) and the western (orange) regions of the Turkana Basin, which are expected to have somewhat different local environmental conditions (Fortelius et al. 2016), and therefore are better analyzed separately. Each dot represents a geological time unit (the name next to the dot is the reference to that time unit). Each time unit may have one or more localities, typically in the order of ten localities. The bars show the standard deviations of the estimates in cases where there are more than one locality per time unit.
Based on our qualitative performance criteria, the predictions of the last ecometric model look the most plausible. The model gives a visible drying trend, especially in the western part (orange). This model uses the same transfer learning principle as our baseline model, it transforms the initial feature spaces of the present and the past into a common representation using dental traits of species. The functional form used to fit this best performing model is quadratic (Fortelius et al. 2016), which might be better capturing underlying relationships that our linear baselines. Perhaps linearity of the baseline models is one of the reasons why we do not quite capture long term drying trends, as expected.
Interestingly, the average level of productivity is different in the three linear predictions: taxon assemblage has the lowest absolute values of NPP, the ecometric approach has the highest, while mean habitat is in between. We do not know the true values, but judging from the non-linear ecometric model, which we consider being the most plausible in this figure, taxon assemblage has more accurate values towards the present, while ecometric model I has more accurate values further to the geological past. From the domain perspective this is a reasonable and believable finding, since the closer to the present day, the more similar taxa is expected to be to the present day taxa on which the model is trained. The further to the past, the more concept drift, the more powerful the functional transfer learning (ecometrics) becomes, and we see evidence of that in this case study (Fig. 6).
The mean habitat approach lacks sensitivity, as could be seen also by the poor quantitative performance in the earlier section on the present day data. Its predictions are nearly flat in Fig. 7, suggesting a robust baseline, but not very informative one. The predictions of taxon assemblage and the ecometric approaches are similar in trends, except that the ecometric approach picks up an increasing trend at the very recent times (from 2 to 1 million years). This is primarily due to structural fortification and hypsodonty traits (for reference the raw dental trait features are plotted in the “Appendix” section, Fig. 8), which in that context increase productivity estimates. It may be that the model is too sensitive to those features, which hints overfitting the present day data.
7 Bottom-up feature selection
This last experiment illustrates a simple model building and validation process, using a combination of computational validation criteria and domain knowledge. Using the same fossil data from the Turkana Basin as in the previous section, we build alternative ecometric models, hoping to increase robustness of predictions, reduce the chance of overfitting by trend-based feature selection. The features we are inspecting are in the transformed feature space that is common for the present and the past data. The idea is to select features from the fossil data that show steady trends and low variability over time, go back to the present day data and train models using these features.
From the feature options given in Fig. 8 we select dental traits that conceptually capture similar information as hypsodonty and lophedness, which have been traditionally used for predicting productivity (Liu et al. 2012; Fortelius et al. 2016). We pick the traits that show steady long term trends and have relatively little short term oscillations. Two traits appear the most roust: flat occlusal topography and presence of acute lophs, which both are binary and have trends, as expected (as can be seen in Fig. 8).
Using the selected features we fit a model on present day data. Interestingly, the testing accuracies of the fit are below zero, indicating that this model would not have been selected if an automated feature selection have been used. For the interest of exposition, we plot the predictions of this model in Fig. 7. When comparing to the predictions in, this model looks the plausible. Yet validation on the present day data would not have identified this model as worth even considering. This case demonstrated the challenge of transfer learning with extreme lack of overlap between features and not available ground truth for validating the predictions. Yet, this analysis suggests that a combination of cross-validation on the present day data and inspection of trends can produce informative models, even if not to be trusted blindly. Similar challenges can arise and similar methodologies could potentially be applied to analysis of other long-term collections of information, such as historical documents, old pictures or statistical indicators spanning many decades.
8 Conclusion
Fossil data presents interesting challenges for data science and, particularly, for concept drift in machine learning research. Due to evolution and natural selection species (or higher taxa), which are the main units of analysis, originate and go extinct over geological times. The ground truth for paleoenvironment reconstruction model building is only available at the present day, but the further to the past predictions are needed, the less overlap is with the present species. There are at least two main transfer learning principles to address this predictive modeling challenge: replace extinct species with nearest living relatives in the model training phase, or redescribe characteristics of the past and present species in a common space—via functional traits.
We have analyzed both approaches computationally in the same setting and on the same training and fossil datasets to directly compare their performance, which to the best of our knowledge has not been attempted before. We have also analyzed the patterns of predictions via a case study of the fossil record of the Turkana Basin, the area which is of interest to a broad community of paleoanthropology due to its major significance for human evolution. Based on predictive accuracy and visual analysis ecometric methods appear most promising for paleoenvironment reconstruction, particularly in more distant times from the present, when less and less species overlap with the species today. We have demonstrated that a combination of numerically-driven model selection and domain expertise can produce robust and plausible models. Our investigation opens a new avenue for methodological research in combining transfer learning and concept drift. The benchmark datasets used are available on GitHub.Footnote 6
Even though there is no particular pressure on computational resources to make the algorithms incremental, it would be interesting from the research perspective particularly to extend transformation of features to an incremental setting. In would be even more interesting from the algorithmic perspective to generalize over feature transformation such that instead of taking an average, any arbitrary aggregation function could be used, including non-linear transformations, minimums, maximums and more. In principle, a kernel function could be there, and ideally the shape of the feature transfer function could be learned from data. This, to the best of my knowledge, has not been tried and it could have potentially interesting applications beyond fossil data analysis, for instance, in user modeling, personalization, or recommender systems.
Notes
NOW Database of fossil mammals http://www.helsinki.fi/science/now/ Paleobiology Database https://paleobiodb.org/ Fossil Calibration Database http://fossilcalibrations.org/ Paleobotany project http://www.paleobotanyproject.org/ EDNA Insect Database https://fossilinsectdatabase.co.uk/ Miocene Mammal Mapping Project http://www.ucmp.berkeley.edu/miomap/ The New Zealand Fossil Record https://fred.org.nz/ Morphobank https://morphobank.org/ Repository of phylogenetic information https://www.treebase.org/.
Not to be mixed with econometrics, which is a branch of financial mathematics.
References
Aha DW, Kibler D, Albert MK (1991) Instance-based learning algorithms. Mach Learn 6(1):37–66
Argyriou A, Evgeniou T, Pontil M (2007) Multi-task feature learning. In: Proceedings of the 19th annual conference on neural information processing systems, pp 41–48
Barnosky A, Hadly E, Bascompte J, Berlow E, Brown J, Fortelius M, Getz W, Harte J (2012) Approaching a state shift in Earth’s biosphere. Nature 486:52–58
Barnosky A, Hadly E, Gonzalez P, Head J, Polly D, Lawing M et al (2017) Merging paleobiology with conservation biology to guide the future of terrestrial ecosystems. Science 355:594–595
Barr AW (2017) Bovid locomotor functional trait distributions reflect land cover and annual precipitation in sub-Saharan Africa. Evol Ecol Res 18:253–269
Behrensmeyer AK, Kidwell SM, Gastaldo RA (2000) Taphonomy and paleobiology. Paleobiology 26(4):103–147
Blitzer J, McDonald R, Pereira F (2006) Domain adaptation with structural correspondence learning. In: Proceedings of the conference on empirical methods in natural language, pp 120–128
Blumenthal SA, Levin NE, Brown FH, Brugal J-P, Chritz KL, Harris JM, Jehle GE, Cerling TE (2017) Aridity and hominin environments. PNAS 114(28):7331–7336
Bobe R (2006) The evolution of arid ecosystems in eastern Africa. J Arid Environ 66:564–584
Bobe R (2011) Fossil mammals and paleoenvironments in the Omo-Turkana Basin. Evol Anthropol 20:254–263
Cerling TE (1992) Development of grasslands and savannas in East Africa during the Neogene. Palaeogeogr Palaeoclimatol Palaeoecol 97:241–247
Cerling TE, Wynn JG, Andanje SA, Bird MI, Korir DK, Levin NE, Mace W, Macharia AN, Quade J, Remien CH (2011) Woody cover and hominin environments in the past 6 million years. Nature 476(7358):51–6
Cerling TE, Andanjec SA, Blumenthal SA, Brown FH, Chritz KL, Harris JM, Hart JA, Kirera FM, Kaleme P, Leakey LN, Leakey MG, Levin NE, Manthi FK, Passey BH, Uno KT (2015) Dietary changes of large herbivores in the Turkana basin. Kenya from 4 to 1 ma. PNAS 112(37):11467–11472
Cramer W, Kicklighter DW, Bondeau A, Moore B, Churkina G, Nemry B, Ruimy A, Schloss AL (1999) Comparing global models of terrestrial net primary productivity (NPP): overview and key results. Glob Change Biol 5(S1):1–15
Dai W, Xue G, Yang Q, Yu Y (2007) Co-clustering based classification for out-of-domain documents. In: Proceedings of the 13th ACM SIGKDD international conference on knowledge discovery and data mining, pp 210–219
Darwin C (1859) On the origin of species by means of natural selection, or the preservation of favoured races in the struggle for life. John Murray, London
Daume III, H (2007) Frustratingly easy domain adaptation. In: Proceedings of the 45th annual meeting of the association of computational linguistics, pp 256–263
Davis J, Kulis B, Sra S, Dhillon I (2007) Information-theoretic metric learning. In: Proceedings of the 24th international conference on machine learning, ICML, pp 209–216
deMenocal PB (2004) African climate change and faunal evolution during the Pliocene–Pleistocene. Earth Planet Sci Lett 220:3–24
Elith J, Leathwick JR (2009) Species distribution models: ecological explanation and prediction across space and time. Annu Rev Ecol Evol Syst 40:677–697
Eronen JT, Mirzaie Ataabadi M, Micheels A, Karme A, Bernor RL, Fortelius M (2009) Distribution history and climatic controls of the Late Miocene Pikermian chronofauna. Proc Natl Acad Sci 106:11867–11871
Eronen JT, Puolamäki K, Liu L, Lintulaakso K, Damuth J, Janis C, Fortelius M (2010a) Precipitation and large herbivorous mammals, part I. Evol Ecol Res 12:217–233
Eronen JT, Puolamäki K, Liu L, Lintulaakso K, Damuth J, Janis C, Fortelius M (2010b) Precipitation and large herbivorous mammals, part II. Evol Ecol Res 12:235–248
Fernandez MH, Vrba ES (2006) Plio-pleistocene climatic change in the Turkana basin (East Africa): evidence from large mammal faunas. J Hum Evol 50:595–626
Fleagle JG, Leakey M (2011) The Turkana basin. Evol Anthropol 20:201
Foote M, Sepkoski JJ (1999) Absolute measures of the completeness of the fossil record. Nature 398:415–417
Fortelius M, Eronen J, Jernvall J, Liu L, Pushkina D, Rinne J et al (2002) Fossil mammals resolve regional patterns of eurasian climate change over 20 million years. Evol Ecol Res 4:1005–1016
Fortelius M, Eronen JT, Kaya F, Tang H, Raia P, Puolamaki K (2014) Evolution of neogene mammals in Eurasia: environmental forcing and biotic interactions. Annu Rev Earth Planet Sci 42:579–604
Fortelius M, Zliobaite I, Kaya F, Bibi F, Bobe R, Leakey L et al (2016) An ecometric analysis of the fossil mammal record of the Turkana basin. Philos Trans R Soc B 371(1698):1–13
Gama J, Zliobaite I, Bifet A, Pechenizkiy M, Bouchachia A (2014) A survey on concept drift adaptation. ACM Comput Surv 46(4):44
Gopalan R, Li R, Chellappa R (2014) Unsupervised adaptation across domain shifts by generating intermediate data representations. IEEE Trans Pattern Anal Mach Intell 36(11):2117
Haberl H, Heinz Erb K, Krausmann F, Gaube V, Bondeau A, Plutzar C, Gingrich S, Lucht W, Fischer-Kowalski M (2007) Quantifying and mapping the human appropriation of net primary production in earth’s terrestrial ecosystems. PNAS 104(31):12942–12947
Hairston NG Jr, Hairston NG Sr (1993) Cause–effect relationships in energy flow, trophic structure, and interspecific interactions. Am Nat 142(3):379–411
Hoffman J, Kulis B, Darrell T, Saenko K (2012) Discovering latent domains for multisource domain adaptation. In: Proceedings of the 12th European conference on computer vision, ECCV, pp 702–715
Kidwell SM, Behrensmeyer AK (1988) Overview: ecological and evolutionary implications of taphonomic processes. Palaeogeogr Palaeoclimatol Palaeoecol 63(1–3):1–13
Lawing A, Head J, Polly D (2012) The ecology of morphology: the ecometrics of locomotion and macroenvironment in North American snakes. In: Louys J (ed) Paleontology in ecology and conservation. Springer, Berlin, pp 117–146
Lawing AM, Eronen JT, Blois JL, Graham CH, Polly PD (2015) Community functional trait composition at the continental scale: the effects of nonecological processes. Ecography 40(5):651–663
Leakey MG, Grossman A, Gutierrez M, Fleagle JG (2011) Faunal change in the Turkana basin during the late oligocene and miocene. Evol Anthropol 20:238–253
Li W, Duan L, Xu D, Tsang IW (2014) Learning with augmented features for supervised and semi-supervised heterogeneous domain adaptation. IEEE Trans Pattern Anal Mach Intell 36(6):1134
Lieth H (1975) Modelling the primary productivity of the world. In: Lieth H, Whittaker RH (eds) Primary productivity of the biosphere. Springer, New York, pp 237–263
Liu L, Puolamäki K, Eronen JT, Mirzaie Ataabadi M, Hernesniemi E, Fortelius M (2012) Dental functional traits of mammals resolve productivity in terrestrial ecosystems past and present. Proc R Soc [Biol] 279:2793–2799
Meloro C, Kovarovic K (2013) Spatial and ecometric analyses of the plio-pleistocene large mammal communities of the Italian peninsula. J Biogeogr 40:1451–1462
Mirsky S (1998) I shall return. Earth 278:48–53
Mosbrugger V, Utescher T (1997) The coexistence approach—a method for quantitative reconstructions of tertiary terrestrial palaeoclimate data using plant fossils. Palaeogeogr Palaeoclimatol Palaeoecol 134:61–86
Myers CE, Stigall AL, Lieberman BS (2015) PaleoENM: applying ecological niche modeling to the fossil record. Paleobiology 41(2):226–244
Ovaskainen O, Tikhonov G, Norberg A, Blanchet FG, Duan L, Dunson D et al (2017) How to make more out of community data? a conceptual framework and its implementation as models and software. Ecol Lett 20:561–576
Pan SJ, Yang Q (2010) A survey on transfer learning. IEEE Trans Knowl Data Eng 22(10):1345–1359
Polly D, Eronen J, Fred M, Dietl G, Mosbrugger V, Scheidegger C et al (2011) History matters: ecometrics and integrative climate change biology. Proc R Soc [Biol] 278(1709):1131–1140
Polly D, Head J (2015) Measuring earth-life transitions: ecometric analysis of functional traits from late cenozoic vertebrates. In: Earth-life transitions. The Paleontological Society, Baltimore, pp 21–46
Raina R, Battle A, Lee H, Packer B, Ng AY (2007) Self-taught learning: transfer learning from unlabeled data. In: Proceedings of the 24th international conference on machine learning, ICML, pp 759–766
Reed K (2013) Multiproxy paleoecology: reconstructing evolutionary context in paleoanthropology. In: A companion to paleoanthropology. Wiley-Blackwell, Oxford, pp 204–225
Rich E (1979) User modeling via stereotypes. Cogn Sci 3:329–354
Saarinen J (2015) Ecometrics of large herbivorous land mammals in relation to climatic and environmental changes during the Pleistocene. Ph.D. thesis, University of Helsinki
Schnitzler J, Theis C, Polly P, Eronen J (2017) Fossils matter understanding modes and rates of trait evolution in musteloidea (Carnivora). Evol Ecol Res 18:187–200
Stute M, Clement A, Lohmann G (2001) Global climate models: past, present, and future. PNAS 98(19):10529–10530
Sukselainen L, Fortelius M, Harrison T (2015) Co-occurrence of pliopithecoid and hominoid primates in the fossil record: an ecometric analysis. J Hum Evol 84:25–41
Tatti N, Vreeken J (2012) Comparing apples and oranges measuring differences between data mining results. Data Min Knowl Discov 25(2):173–207
Traiser C, Klotz S, Uhl D, Mosbrugger V (2005) Environmental signals from leaves—a physiognomic analysis of European vegetation. New Phytol 166(2):465–484
Uhen M, Barnosky A, Bills B, Blois J, Carrano M, Carrasco M, Erickson G et al (2013) From card catalogs to computers: databases in vertebrate paleontology. J Vert Paleontol 33(1):13–28
Utescher T, Bruch AA, Erdei B, Francois L, Ivanov D, Jacques FMB, Kern AK, Liu YSC, Mosbrugger V, Spicer RA (2014) The coexistence approach—theoretical background and practical considerations of using plant fossils for climate quantification. Palaeogeogr Palaeoclimatol Palaeoecol 410(15):58–73
Vermillion W, Head J, Polly D, Eronen J, Lawing M (2017) Ecometrics: a trait-based approach to paleoclimate and paleoenvironmental reconstruction. In: Methods in paleoecology. Springer, Berlin
Warton DI, Shipley B, Hastie T (2015) Cats regression a model-based approach to studying trait-based community assembly. Methods Ecol Evol 6(4):389–398
Wold H (1966) Estimation of principal components and related models by iterative least squares. In: Krishnaiaah PR (ed) Multivariate analysis. Academic Press, New York, pp 391–420
Wolfe JA (1995) Paleoclimatic estimates from tertiary leaf assemblages. Annu Rev Earth Planet Sci 23:119–142
Zhou ZH, Zhang ML, Huang SJ, Li YF (2012) Multi-instance multi-label learning. Artif Intell 176(1):2291–2320
Zliobaite I, Gabrys B (2014) Adaptive preprocessing for streaming data. IEEE Trans Knowl Data Eng 26(2):309–321
Zliobaite I, Rinne J, Toth A, Mechenich M, Liu LP, Behrensmeyer AK, Fortelius M (2016) Herbivore teeth predict climatic limits in Kenyan ecosystems. PNAS 113(45):12751–12756
Zliobaite I, Puolamaki K, Eronen J, Fortelius M (2017) A survey of computational methods for fossil data analysis. Evol Ecol Res 18:477–502
Zliobaite I, Tang H, Saarinen J, Fortelius M, Rinne J, Rannikko J (2018) Dental ecometrics of tropical Africa: linking vegetation types and communities of large plant-eating mammals. Evol Ecol Res 19:127–147
Acknowledgements
Open access funding provided by University of Helsinki including Helsinki University Central Hospital. The author thanks Mikael Fortelius for help in scoring dental traits of the Turkana fossil species. Research leading to these results was partially funded the Academy of Finland (ECHOES Grant 274779). This is a contribution from the Valio Armas Korvenkontio Unit of Dental Anatomy in Relation to Evolutionary Theory.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Alípio Jorge, Rui L. Lopes, German Larrazabal.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Žliobaitė, I. Concept drift over geological times: predictive modeling baselines for analyzing the mammalian fossil record. Data Min Knowl Disc 33, 773–803 (2019). https://doi.org/10.1007/s10618-018-0606-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10618-018-0606-6