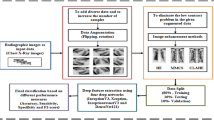
We consider the automatic hardness determination of a chest X-ray image and the effect of pre-filtering of the training and validation samples on the performance of the classification algorithm of tuberculosis diagnosis from chest X-rays. Convolutional neural networks are used in automatic hardness determination and tuberculosis diagnosis. The results of the present study are compared with those from different datasets, including datasets pruned by image hardness criteria.
Article PDF
Similar content being viewed by others
References
S. G. Finlayson et al., “Adversarial attacks on medical machine learning,” Science, 363, No. 6433, 1287–1289 (2019).
G. A. Chuiko and V. M. Tsvetkov, “Effects of X-ray hardness on fluorogram informativeness,” Biomedical Engineering, 16, No. 4, 117–119 (1982).
L. A. Timofeeva, T. N. Aleshina, and A. V. Bykova, Main X-ray Syndromes of Lung-Tissue Pathology: a Textbook, Izd. Chuvash. Univ., Cheboksary (2013).
A. U. Sidorov, A. A. Shcherbatykh, and L. N. Pokrovskaya, Methodology of Radiograph Analysis: a Textbook, IGMU, Irkutsk (2012).
K. Nousiainen et al., “Automating chest radiograph imaging quality control,” Physica Medica, 83, 138–145 (2021).
J. von Berg et al., “Robust chest x-ray quality assessment using convolutional neural networks and atlas regularization,” Medical Imaging 2020: Image Processing, SPIE, 11313, 391–398 (2020).
J. I. A. Xiao-Qian et al., “Application value of convolutional neural network in quality control of direct digital chest X-ray images,” Xi’an Jiao Tong da Xue Xue Bao. Yi Xue Ban, No. 5, 784 (2019).
R. Sadre et al., “Validating deep learning inference during chest X-ray classification for COVID-19 screening,” Scientific Reports, 11, No. 1, 1–10 (2021).
A. A. Dovganich, A. V. Khvostikov, Y. A. Pchelintsev et al., “Automatic out-of-distribution detection methods for improving the deep learning classification of pulmonary X-ray images,” Journal of Image and Graphics, 10, No. 2, (2022).
M. Oloko-Oba and S. Viriri, “A systematic review of deep learning techniques for tuberculosis detection from chest radiograph,” Frontiers in Medicine, 9 (2022).
S. Jaeger et al., “Automatic tuberculosis screening using chest radiographs,” IEEE Transactions on Medical Imaging, 33, No. 2, 233–245 (2013).
S. Candemir et al., “Lung segmentation in chest radiographs using anatomical atlases with nonrigid registration,” IEEE Transactions on Medical Imaging, 33, No. 2, 577–590 (2013).
Y. Liu et al., “Rethinking computer-aided tuberculosis diagnosis,” Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2646–2655 (2020).
K. He et al., “Deep residual learning for image recognition,” Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 770–778 (2016).
G. Huang et al., “Densely connected convolutional networks,” Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (2017), pp. 4700–4708.
M. Tan and Q. Le, “EfficientNetV2: Smaller models and faster training,” International Conference on Machine Learning, PMLR (2021), pp. 10096–10106.
O. Russakovsky et al., “ImageNet large scale visual recognition challenge,” International Journal of Computer Vision, 115, No. 3, 211–252 (2015).
J. D. M. Rennie and N. Srebro, “Loss functions for preference levels: Regression with discrete ordered labels,” Proceedings of the IJCAI Multidisciplinary Workshop on Advances in Preference Handling, vol. 1, AAAI Press, Menlo Park, CA (2005).
S. M. Pizer et al., “Adaptive histogram equalization and its variations,” Computer Vision, Graphics, and Image Processing, 39, No. 3, 355–368 (1987).
S. Baccianella, A. Esuli, and F. Sebastiani, “Evaluation measures for ordinal regression,” 2009 Ninth International Conference on Intelligent Systems Design and Applications, IEEE (2009), pp. 283–287.
K. H. Brodersen et al., “The balanced accuracy and its posterior distribution,” 2010 20th International Conference on Pattern Recognition, IEEE (2010), pp. 3121–3124.
I. Loshchilov and F. Hutter, “Decoupled weight decay regularization,” arXiv preprint arXiv:1711.05101 (2017).
D. Zwillinger and S. Kokoska, CRC Standard Probability and Statistics Tables and Formulae, Chapman & Hall, New York (2000).
V. Thambawita et al., “An extensive study on cross-dataset bias and evaluation metrics interpretation for machine learning applied to gastrointestinal tract abnormality classification,” ACM Transactions on Computing for Healthcare, 1, No. 3, 1–29 (2020).
Author information
Authors and Affiliations
Corresponding author
Additional information
Translated from Prikladnaya Matematika i Informatika, No. 70, 2022, pp. 52–70.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pchelintsev, Y.A., Khvostikov, A.V., Krylov, A.S. et al. Hardness Analysis of X-Ray Images for Neural-Network Tuberculosis Diagnosis. Comput Math Model 33, 230–243 (2022). https://doi.org/10.1007/s10598-023-09568-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10598-023-09568-3