Abstract
Adaptive genetic and neutral variations are essential for maintaining population viability in changing environmental conditions. Habitat loss and fragmentation can be reflected in the patterns of genetic variation in the populations. White-lipped peccaries (WLPs, Tayassu pecari) are wide-ranging Neotropical ungulates with important ecological roles in the ecosystem suffering local extinctions worldwide. Here, we used a RAD-seq protocol to genotype 192 individuals. After filtering, we identified sets of SNP markers (ranging from 147 to 151,792 SNPs) to assess the genetic diversity and population structure of WLPs from Pantanal, Cerrado, and Atlantic Forest in Brazil. We found signals of loss (θw < θπ) and lower genetic diversity (allelic richness, nucleotide diversity, and observed and expected heterozygosities) in the Central Cerrado and Atlantic Forest populations. Principal Component Analysis (PCA) and admixture analyses (NGSAdmix) using genome-wide and neutral SNP data sets showed three major genetic clusters according to the biomes. Multiple matrix regression with randomization (MMRR) analysis found an isolation-by-distance pattern explaining the neutral genetic differentiation. We used Latent Factor Mixed Models (LFMM) and Redundancy Analysis (RDA) to identify candidate SNPs involved in different biological processes, such as metabolism and immune and neuronal responses, mainly associated with temperature and precipitation variables. We found an adaptive population genetic structure, suggesting three adaptive units with significant patterns of isolation-by-distance and isolation-by-environment. Our results highlighted the importance of conservation strategies for maintaining the genetic diversity of WLP populations. Furthermore, conservation plans and translocation programs should preserve and consider the adaptive variation.

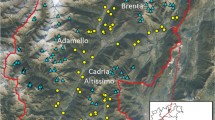
(Source: http://mapbiomas.org, Mapbiomas collection 4.1, 2016). Non-forest natural formations: wetland, grassland, and other non-forest natural formations



Similar content being viewed by others
Data availability
Raw sequence data are available at NCBI (SRA accession: Bioproject PRJNA1062824). Other data, such as genotypes and environmental data, are available at Figshare (https://doi.org/10.6084/m9.figshare.25438330).
References
Adamack AT, Gruber B (2014) PopGenReport: simplifying basic population genetic analyses in R. Methods Ecol Evol 5:384–387. https://doi.org/10.1111/2041-210X.12158
Ali OA, O’Rourke SM, Amish SJ, Meel MH, Luikart G, Jeffre C, Miller MR (2016) RAD capture (rapture): flexible and efficient sequence-based genotyping. Genetics 202:389–400. https://doi.org/10.1534/genetics.115.183665
Altrichter M, Carrillo E, Saenz J, Fuller T (2001) White-lipped peccary (Tayassu pecari) diet and fruit availability in a Costa Rican rain forest. Biol Trop 49:1105–1114
Altrichter M, Drews C, Sáens JC, Carrilo E (2002) Presupuesto De tiempo del chancho cariblanco (Tayassu pecari) em un bosque humedo de Costa Rica. Biotropica 34:136–143. https://doi.org/10.1111/j.1744-7429.2002.tb00249.x
Altrichter M, Taber A, Beck H, Reyna-Hurtado R, Lizarraga L, Keuroghlian A, Sanderson EW (2012) Range-wide declines of a key neotropical ecosystem architect, the Near threatened white-lipped peccary Tayassu pecari. Oryx 46:87–98. https://doi.org/10.1017/S0030605311000421
Baptista MSP, Keuroghlian A, Tambosi LR, Côrtes MC, Maciel FG, Cirino DW, Schmaedecke G, Biondo C (2024) Agriculturally developed areas reduce genetic connectivity for a keystone neotropical ungulate. https://doi.org/10.1101/2024.01.22.576460. bioRxiv
Beck H, Thebpanya P, Filiaggi M (2010) Do neotropical peccary species (Tayassuidae) function as ecosystem engineers for anurans? J Trop Ecol 26:407–414. https://doi.org/10.1017/S0266467410000106
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. Source J Royal Stat Soc Ser B (Methodological) 57:289–300. https://doi.org/10.2307/2346101
Biondo C, Keuroghlian A, Gongora J, Miyaki CY (2011) Population genetic structure and dispersal in white-lipped peccaries (Tayassu pecari) from the Brazilian pantanal. J Mammal 92:267–274. https://doi.org/10.1644/10-MAMM-A-174.1
Bodmer RE (1989) Frugivory in amazonian Artiodactyla: evidence for the evolution of the ruminant stomach. J Zool Lond 219:457–467. https://doi.org/10.1111/j1469-7998.1989.tb02593.x
Caillet-Boudin ML, Buée L, Sergeant N, Lefebvre B (2015) Regulation of human MAPT gene expression. Mol Neurodegeneration 10:1–14. https://doi.org/10.1186/s13024-015-0025-8
Carrillo E, Saenz JC, Fuller TK (2002) Movements and activities of white-lipped peccaries in Corcovado National Park, Costa Rica. Biol Conserv 108:317–324. https://doi.org/10.1016/S0006-3207(02)00118-0
Catchen J, Hohenlohe PA, Bassham S, Amores A, Cresko WA (2013) STACKS: an analysis tool set for population genomics. Mol Ecol 22:3124–3140. https://doi.org/10.1111/mec.12354
Clozato CL, Mazzoni CJ, Moraes-Barros N, Morgante JS, Sommer S (2015) Spatial pattern of adaptive and neutral genetic diversity across different biomes in the lesser anteater (Tamandua tetradactyla). Ecol Evol 5:4932–4948. https://doi.org/10.1002/ece3.1656
Cushman SA, Shirk A, Landguth EL (2012) Separating the effects of habitat area, fragmentation and matrix resistance on genetic differentiation in complex landscapes. Landsc Ecol 27:369–380. https://doi.org/10.1007/s10980-011-9693-0
Delputte PL, Van Gorp H, Favoreel HW, Hoebeke I, Delrue I, Dewerchin H, Verdonck F, Verhasselt B, Cox E, Nauwynck HJ (2011) Porcine Sialoadhesin (CD169/Siglec-1) is an endocytic receptor that allows targeted delivery of toxins and antigens to macrophages. PLoS ONE 6:e16827. https://doi.org/10.1371/journal.pone.0016827
Dib I, Khalil A, Chouaib R, El-Makhour Y, Noureddine H (2021) Apolipoprotein C‐III and cardiovascular diseases: when genetics meet molecular pathologies. Mol Biol Rep 48:875–886. https://doi.org/10.1007/s11033-020-06071-5
Dotsev AV, Deniskova TE, Okhlopkov IM, Mészáros G, Sölkner J, Reyer H, Wimmers K, Brem G, Zinovieva NA (2018) Genome-wide SNP analysis unveils genetic structure and phylogeographic history of snow sheep (Ovis nivicola) populations inhabiting the Verkhoyansk Mountains and Momsky Ridge (northeastern Siberia). Ecol Evol 8:1–11. https://doi.org/10.1002/ece3.4350
Dotsev A, Koshkina O, Kharzinova V, Deniskova T, Reyer H, Kunz E, Mészáros G, Shakhin A, Petrov S, Medvedev D et al (2023) Genome-wide insights into Intraspecific Taxonomy and Genetic Diversity of Argali (Ovis ammon). Diversity 15:627. https://doi.org/10.3390/d15050627
Durigan G, Franco GADC (2006) Vegetação. In: Faria HH, Pires AS (eds.) Parque Estadual do Morro do Diabo: plano de manejo. Editora Viena, pp. 111–118
Eaton DP, Keuroghlian A, Maria do Carmo AS, Desbiez AL, Sada DW (2017) Citizen scientists help unravel the nature of cattle impacts on native mammals and birds visiting fruiting trees in Brazil’s southern Pantanal. Biol Conserv 208:29–39. https://doi.org/10.1016/j.biocon.2016.09.010
Fick SE, Hijmans RJ (2017) WorldClim 2: new 1km spatial resolution climate surfaces for global land areas. Int J Climatol 37:4302–4315. https://doi.org/10.1002/joc.508
Forester BR, Lasky JR, Wagner HH, Urban DL (2018) Comparing methods for detecting multilocus adaptation with multivariate genotype-environment associations. Mol Ecol 27:2215–2233. https://doi.org/10.1111/mec.14584
Francis RM (2016) POPHELPER: an R package and web app to analyse and visualize population structure. Mol Ecol Resour 17:27–32. https://doi.org/10.1111/1755-0998.12509
François O, Martins H, Caye K, Schoville SD (2016) Controlling false discoveries in genome scans for selection. Mol Ecol 25:454–469. https://doi.org/10.1111/mec.13513
Frankham R (1995) Conservation genetics. Annu Rev Genet 29:305–327. https://doi.org/10.1146/annurev.ge.29.120195.001513
Frichot E, Francois O (2015) LEA: an R package for Landscape and Ecological Association studies. Methods Ecol Evol 6:925–929. https://doi.org/10.1111/2041-210X.12382
Frichot E, Schoville SD, Bouchard G, François O (2013) Testing for associations between loci and environmental gradients using latent factor mixed models. Mol Biol Evol 30:1687–1699. https://doi.org/10.1093/molbev/mst063
Fumagalli M, Vieira FG, Korneliussen TS, Linderoth T, Huerta-Sánchez E, Albrechtsen A, Nielsen R (2013) Quantifying population genetic differentiation from next-generation sequencing data. Genetics 195:979–992. https://doi.org/10.1534/genetics.113.154740
Funk WC, McKay JK, Hohenlohe PA, Allendorf FW (2012) Harnessing genomics for delineating conservation units. Trends Ecol Evol 9:489–496. https://doi.org/10.1016/j.tree.2012.05.012
Funk WC, Forester BR, Converse SJ, Darst C, Morey S (2019) Improving conservation policy with genomics: a guide to integrating adaptive potential into U.S. Endangered species Act decisions for conservation practitioners and geneticists. Conserv Genet 20:115–134. https://doi.org/10.1007/s10592-018-1096-1
Goudet J (2005) Hierfstat, a package for R to compute and test hierarchical F-statistics. Mol Ecol Notes 5:184–186. https://doi.org/10.1111/j.1471-8286.2004.00828.x
Hagiwara N (2011) Sox6, Jack of all trades: a versatile regulatory protein in vertebrate development. Dev Dyn 240:1311–1321. https://doi.org/10.1002/dvdy.22639
Harrison KA, Pavlova A, Telonis-Scott M, Snucks P (2014) Using genomics to characterize evolutionary potential for conservation of wild populations. Evol Appl 7:1008–1025. https://doi.org/10.1111/eva.12149
Hoekstra HE (2006) Genetics, development and evolution of adaptive pigmentation in vertebrates. Heredity 97:222–234. https://doi.org/10.1038/sj.hdy.6800861
Jácomo ATA, Furtado MM, Kashivakura CK, Marinho-Filho J, Sollmann R, Tôrres NM, Silveira L (2013) White-lipped peccary home-range size in a protected area and farmland in the central Brazilian grasslands. J Mammal 94:137–145. https://doi.org/10.1644/11-MAMM-A-411.1
Joly CA, Aidar MPM, Klink CA, McGrath DG, Moreira AC, Moutinho P, Nepstad DC, Oliveira AA, Pott A, Rodal MJN, Sampaio EVSB (1999) Evolution of the Brazilian phytogeography classification systems: implications for biodiversity conservation. Ciência E Cultura 51:331–348
Jorge MLSP, Galetti M, Ribeiro MC, Ferraz KMPMB (2013) Mammal defaunation as surrogate of trophic cascades in a biodiversity hotspot. Biol Conserv 163:49–57. https://doi.org/10.1016/j.biocon.2013.04.018
Jorge MLSP, Keuroghlian A, Bradham J, Oshima JEF, Ribeiro MC (2019) White-lipped peccary movement and range in agricultural lands of Central Brazil. In: Reyna-Hurtado R, Chapman CA (eds) Movement Ecology of Neotropical Forest Mammals, 1st edn. Springer, pp 39–55
Jorge MLSP, Bradham JL, Keuroghlian A, Oshima JEF, Ribeiro MC (2021) Permeability of neotropical agricultural lands to a key native ungulate – are well-connected forests important? Biotropica 53:201–212. https://doi.org/10.1111/btp.12861
Junk WJ, Cunha CN, Wantzen MK, Petermann P, Strüssmann C, Marques MI, Adis J (2006) Biodiversity and its conservation in the Pantanal of Mato Grosso, Brazil. Aquat Sci 68:278–309. https://doi.org/10.1007/s00027-006-0851-4
Kamvar ZN, Tabima JF, Grünwald NJ (2014) Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2:e281. https://doi.org/10.7717/peerj.281
Kanai TH, Tanioka Y, Tanigawa M, Matsumoto Y, Ueda S, Onodera T, Matsumoto Y (1999) Allelic diversity at class II DRB1 and DQB loci of the pig MHC (SLA). Immunogenetics 50:295–300. https://doi.org/10.1007/s002510050605
Keuroghlian A, Eaton DP (2008a) Fruit availability and Peccary Frugivory in an isolated Atlantic Forest Fragment: effects on Peccary ranging Behavior and Habitat Use. Biotropica 40:62–70. https://doi.org/10.1111/j.1744-7429.2007.00351.x
Keuroghlian A, Eaton DP (2008b) Importance of rare habitats and riparian zones in a tropical forest fragment: preferential use by Tayassu pecari, a wide-ranging frugivore. J Zool Lond 275:283–293. https://doi.org/10.1111/j.1469-7998.2008.00440.x
Keuroghlian A, Eaton DP (2009) Removal of palm fruits and ecosystem engineering in palm stands by white-lipped peccaries (Tayassu pecari) and other frugivores in an isolated Atlantic Forest fragment. Biodivers Conserv 18:1733–1750. https://doi.org/10.1007/s10531-008-9554-6
Keuroghlian A, Eaton DP, Longland WS (2004) Area use by white-lipped and collared peccaries (Tayassu pecari and Tayassu tajacu) in a tropical forest fragment. Biol Conserv 120:411–425. https://doi.org/10.1016/j.biocon.2004.03.016
Keuroghlian A, Eaton DP, Desbiez ALJ (2009) The response of a landscape species, white-lipped peccaries, to seasonal resource fluctuations in a tropical wetland, the Brazilian pantanal. Int J Biodivers Conserv 1:87–97
Keuroghlian A, Desbiez ALJ, Beisiegel BM, Medici EP, Gatti A, Mendes Pontes AR, Campos CB, Tófoli CF, Moraes EA Jr, Azevedo FC, Pinho GM, Cordeiro LP, Santos TS Jr, Morais AA, Mangini PR, Flesher K, Rodrigues LF, Almeida LB (2012) Avaliação do Risco de Extinção do queixada, Tayassu pecari (Link, 1795), no Brasil. Biodiversidade Brasileira 384 – 102
Keuroghlian A, Desbiez ALJ, Reyna-Hurtado R, Altrichter M, Beck H, Taber A, Fragoso JMV (2013) Tayassu pecari. The IUCN Red List of Threatened Species 2013. https://doi.org/10.2305/IUCN.UK.2013-1.RLTS.T41778A44051115.en
Keuroghlian A, Santos MCA, Eaton DP (2014) The effects of deforestation on white-lipped peccary (Tayassu pecari) home range in the southern Pantanal. Mammalia 79:491–497. https://doi.org/10.1515/mammalia-2014-0094
Kiltie RA (1981) Stomach contents of rain forest peccaries (Tayassu tajacu and T. pecari). Biotropica 13:234–236. https://doi.org/10.2307/2388133
Kiltie RA, Terborgh J (1983) Observations on the behavior of rain forest peccaries in Peru: why do white-lipped peccaries form herds? Z Tierpsychol 62:241–255. https://doi.org/10.1111/j.1439-0310.1983.tb02154.x
Kim SY, Lohmueller KE, Albrechtsen A, Li Y, Korneliussen T, Tian G, Grarup N, Jiang T, Andersen G, Witte D, Jorgensen T, Hansen T, Pedersen O, Wang J, Nielsen R (2011) Estimation of allele frequency and association mapping using next-generation sequencing data. BMC Bioinformatics 12:1–16. https://doi.org/10.1186/1471-2105-12-231
Korneliussen TS, Moltke I, Albrechtsen A, Nielsen R (2013) Calculation of Tajima’s D and other neutrality test statistics from low depth next-generation sequencing data. BMC Bioinformatics 14:289. https://doi.org/10.1186/1471-2105-14-289
Korneliussen TS, Albrechtsen A, Nielsen R (2014) ANGSD: analysis of next generation sequencing data. BMC Bioinformatics 15:356. https://doi.org/10.1186/s12859-014-0356-4
Lacy RC (1997) Importance of genetic variation to the viability of mammalian populations. J Mammal 78:320–335. https://doi.org/10.2307/1382885
Lahiri D, Nurnberger J (1991) A rapid non-enzymatic method for the preparation of HMW DNA from blood for RFLP studies. Nucleic Acids Res 19:5444. https://doi.org/10.1093/nar/19.19.5444
Lee JB, Yoo CK, Jung EJ, Hwang JH, Seo BY, Kim BW, Lim YT, Lee JG, Cho IC, Park HB (2012) A missense mutation (c.1963A < G) of the complementary component 2 (C2) gene is associated with serum ca + + concentrations in pigs. Mol Biol Rep 39:9291–9297. https://doi.org/10.1007/s11033-012-1679-8
Leite DA, Keuroghlian A, Rufo DA, Miyaki CY, Biondo C (2018) Genetic evidence of promiscuity in a mammal without apparent sexual dimorphism, the white-lipped peccary (Tayassu pecari). Mammal Biol 92:111–114. https://doi.org/10.1016/j.mambio.2018.05.005
Li H (2011) A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27:2987–2993. https://doi.org/10.1093/bioinformatics/btr509
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Li H, Handsaker B, Wysoker A et al (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25:2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Maciel FG, Rufo DA, Keuroghlian A, Russo AC, Brandt NM, Vieira NF, Nóbrega BM, Nava A, Nardi MS, Jácomo ATA, Silveira L, Furtado MM, Torres NM, Miyaki CY, Tambosi LR, Biondo C (2019) Genetic diversity and population structure of white-lipped peccaries (Tayassu pecari) in the Pantanal, Cerrado and Atlantic Forest from Brazil. Mamm Biol 95:85–92. https://doi.org/10.1016/j.mambio.2019.03.001
Martin ACRC (2018) Diversidade genética e estrutura populacional de queixadas (Tayassu pecari) da Floresta Atlântica da região do Pontal do Paranapanema, SP. Dissertation, Universidade Federal do ABC
Mayer J, Wetzel R (1987) Tayassu pecari. Mammalian Species 293:1–7
Mi H, Muruganujan A, Huang JX, Ebert D, Mills C, Guo X, Thomas PD (2019) Protocol update for large-scale genome and gene function analysis with PANTHER classification system (v.14.0). Nat Protoc 14:703–721. https://doi.org/10.1038/s41596-019-0128-8
Mi H, Ebert D, Muruganujan A, Mills C, Albou LP, Mushayamaha T, Thomas PD (2021) PANTHER version 16: a revised family classification, tree-based classification tool, enhancer regions and extensive API. Nucl Acids Res 49:D394–D403. https://doi.org/10.1093/nar/gkaa1106
Nava AFD (2008) Espécies sentinelas para a Mata Atlântica: as consequências epidemiológicas da fragmentação florestal no Pontal do Paranapanema. PhD Thesis, Universidade de São Paulo
Nielsen R, Korneliussen T, Albrechtsen A, Li Y, Wang J (2012) SNP calling, genotype calling, and sample allele frequency estimation from new-generation sequencing data. PLoS ONE 7:e37558. https://doi.org/10.1371/journal.pone.0037558
Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D, Minchin PR, O’Hara RB, Simpson GL, Solymos P, Stevens MHH, Szoecs E, Wagner H (2019) vegan: Community Ecology Package
Oshima JEF, Jorge MLSP, Sobral-Souza T, Börger L, Keuroghlian A, Peres CA, Vanscine MH, Collen B, Ribeiro MC (2021) Setting Priority Conservation Management regions to reverse Rapid Range decline of a key Neotropical Forest Ungulate. Global Ecol Conserv 31:1–12. https://doi.org/10.1016/j.gecco.2021.e01796
Peek RA, O’Rourke SM, Miller MR (2021) Flow modification associated with reduced genetic health of a river-breeding frog, Rana boylii. Ecosphere 12(5):e03496. https://doi.org/10.1002/ecs2.3496
Peterson M, Jorge MLSP, Jain A, Keuroghlian A, Oshima JEF, Richard-Hansen C, Berzins R, Ribeiro MC, Eaton E (2021) Temperature induces activity reduction in a neotropical ungulate. J Mammal 102:1524–1524. https://doi.org/10.1093/jmammal/gyab09
Prince DJ, O’Rourke SM, Thompson TQ, Ali OA, Lyman HS, Saglam IK, Hotaling TJ, Spidle AP, Miller MR (2017) The evolutionary basis of premature migration in Pacific salmon highlights the utility of genomics for informing conservation. Sci Adv 3:e1603198
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, Maller J, Sklar P, de Bakker PIW, Daly MJ, Sham PC (2007) PLINK: a toolset for whole-genome association and population-based linkage analysis. Am J Hum Genetic 81:559–575. https://doi.org/10.1086/519795
Rasmussen AH, Rasmussen HB, Silahtaroglu A (2017) The DLGAP family: neuronal expression, function and role in brain disorders. Mol Brain 10:43. https://doi.org/10.1186/s13041-017-0324-9
Reyna-Hurtado R (2009) Conservation status of the white-lipped peccary (Tayassu pecari) outside the Calakmul Biosphere Reserve in Campeche, Mexico: a synthesis. Trop Conserv Sci 2:159–172. https://doi.org/10.1177/194008290900200204
Reyna-Hurtado R, Beck H, Altrichter M, Chapman CA, Bonneli TR, Keuroghlian A, Desbiez AL, Moreira-Ramirez JF, O’Farrils G, Fragoso J, Naranjo EJ (2016) What ecological and anthropogenic factors affect Group size in White-lipped peccaries (Tayassu pecari)? Biotropica 8:246–254. https://doi.org/10.1111/btp.12269
Ribeiro MC, Metzger JP, Martensen AC, Ponzoni FJ, Hirota MM (2009) The Brazilian Atlantic Forest: how much is left, and how is the remaining forest distributed? Implications for conservation. Biol Conserv 142:1141–1153. https://doi.org/10.1016/j.biocon.2009.02.021
Ringler M, Hödl W, Ringler E (2015) Populations, pools, and peccaries: simulating the impact of ecosystem engineers on rainforest frogs. Behav Ecol 26:340–349. https://doi.org/10.1093/beheco/aru243
Roffler GH, Amish SJ, Smith S, Cosart T, Kardos M, Schwartz MK, Luikart G (2016) SNP discovery in candidate adaptive genes using exon capture in a free-ranging alpine ungulate. Mol Ecol Resour 16:1147–1164. https://doi.org/10.1111/1755-0998.12560
Rufo DA (2012) Parentesco e diferenciação genética em queixadas (Tayassu pecari) do Pantanal Matogrossense (MS). Dissertation, Universidade de São Paulo
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular Cloning: a Laboratory Manual. Cold Spring Harbor Laboratory Press, New York
Scarano FR, Ceotto P (2015) Brazilian Atlantic Forest: impact, vulnerability, and adaptation to climate change. Biodivers Conserv 24:2319–2331. https://doi.org/10.1007/s10531-015-0972-y
Sica GL, Choi IH, Zhu G, Tamada K, Wang SD, Tamura H, Chapoval AI, Flies DB, Bajorath J, Chen L (2003) B7-H4, a molecule of the B7 family, negatively regulates T cell immunity. Immunity 18:849–861. https://doi.org/10.1016/S1074-7613(03)00152-3
Skotte L, Korneliussen TS, Albrechtsen A (2013) Estimating individual admixture proportions from next generation sequencing data. Genetics 195:693–702. https://doi.org/10.1534/genetics.113.154138
Slatkin M (1987) Gene Flow and the Geographic structure of natural populations. Science 236:787–792. https://doi.org/10.1126/science.3576198
Smaldone S, Ramirez F (2016) Fibrillin microfibrils in bone physiology. Matrix Biol 52–54:191–197. https://doi.org/10.1016/j.matbio.2015.09.004
Tajima F (1983) Evolutionary relationship of DNA sequences in finite populations. Genetics 105:437–460. https://doi.org/10.1093/genetics/105.2.437
Taranda J, Maison SF, Ballestero JA, Katz E, Savino J, Vetter DE, Boulter J, Liberman MC, Fuchs PA, Elgoyhen AB (2009) A point mutation in the hair cell nicotinic cholinergic receptor prolongs cochlear inhibition and enhances noise Protection. PLoS Biol 7:e1000018. https://doi.org/10.1371/journal.pbio.1000018
Wang IJ (2013) Examining the full effects of landscape heterogeneity on spatial genetic variation: a multiple matrix regression approach for quantifying geographic and ecological isolation. Evol 67 – 12 3403–3411. https://doi.org/10.1111/evo.12134
Watterson GA (1975) On the number of segregating sites in genetical models without recombination. Theor Popul Biol 7:256–276. https://doi.org/10.1016/0040-5809(75)90020-9
Wickham H (2016) ggplot2: elegant graphics for data analysis. Springer, New York
Acknowledgements
We want to thank Tatiana P. Freitas, Ezidio Arruda, Celso Vicente, Maria do Carmo A. Santos, Paulino A. O. Santos, Renata R. Rocha, Cícero Sebastião, Cyntia K. Kashivakura, Maria Luisa S. P. Jorge, Donald P. Eaton, and Julia Oshima for their assistance capturing, collaring, collecting biological samples and monitoring white-lipped peccaries. We thank the support from Earthwatch Institute and Global Ecotours volunteers, Biofaces, WCS Brazil, and the landowners from the Upper Paraguay Basin in the southern Pantanal that allowed us to study on their property. We want to thank Anna Carolina R. C. Martin, Nataly F. Vieira, and Danilo A. Rufo for their help with the DNA extraction from the samples and members of Miller’s Lab – UC Davis for their assistance with the bioinformatic analyses. We thank Instituto Florestal de São Paulo (IF) and Instituto de Pesquisas Ecológicas (IPE) for the logistical support. We are also grateful to two anonymous reviewers for their comments on earlier versions of this manuscript.
Funding
This work was funded by FAPESP (Grant number 2015/20133-0), CNPq (Grant number 479760/2012-8), Fundação Manoel Barros, Pantanal and Cerrado ranchers, and Earthwatch Institute volunteers. Author F.G.M. has received support from CAPES (Grant numbers PhD – Funding code 001 and PDSE – 88881.134372/2016-01).
Author information
Authors and Affiliations
Contributions
F.G.M. and C.B. designed the study. M.R.M. contributed to study design, advised, and assisted with bioinformatics. A.K., A.F.D.N., M.S.N., A.T.A.J., L.S., M.M.F., and N.M.T. performed the sample collection. F.G.M. and S.O’R. performed the laboratory work. M.J. and W.H. assisted with bioinformatic analyses. G.S., L.R.T., and M.S.P.B. helped to obtain the environmental data. F.G.M. performed the data analyses and wrote the manuscript. C.B. helped write the manuscript and advised and assisted with the laboratory work and genetic analyses. M.J., W.H., G.S., L.R.T., M.S.P.B., A.K., A.F.D.N., M.M.F., and C.B. revised and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
de Góes Maciel, F., O’Rourke, S., Jones, M. et al. Loss of genetic diversity and isolation by distance and by environment in populations of a keystone ungulate species. Conserv Genet (2024). https://doi.org/10.1007/s10592-024-01614-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10592-024-01614-w