Abstract
The Arizona Toad (Anaxyrus microscaphus) is restricted to riverine corridors and adjacent uplands in the arid southwestern United States. As with numerous amphibians worldwide, populations are declining and face various known or suspected threats, from disease to habitat modification resulting from climate change. The Arizona Toad has been petitioned to be listed under the U.S. Endangered Species Act and was considered “warranted but precluded” citing the need for additional information – particularly regarding natural history (e.g., connectivity and dispersal ability). The objectives of this study were to characterize population structure and genetic diversity across the species’ range. We used reduced-representation genomic sequencing to genotype 3,601 single nucleotide polymorphisms in 99 Arizona Toads from ten drainages across its range. Multiple analytical methods revealed two distinct genetic groups bisected by the Colorado River; one in the northwestern portion of the range in southwestern Utah and eastern Nevada and the other in the southeastern portion of the range in central and eastern Arizona and New Mexico. We also found subtle substructure within both groups, particularly in central Arizona where toads at lower elevations were less connected than those at higher elevations. The northern and southern parts of the Arizona Toad range are not well connected genetically and could be managed as separate units. Further, these data could be used to identify source populations for assisted migration or translocations to support small or potentially declining populations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Arizona Toad (Anaxyrus microscaphus) is one of several bufonids of the arid southwestern United States but is the only toad species restricted to riverine corridors and adjacent uplands. Despite these restrictions, the Arizona Toad’s range encompasses a variety of biotic communities and elevations, extending diagonally from southeastern Nevada and southwestern Utah across central Arizona to southwestern New Mexico (Schwaner and Sullivan 2005). As with numerous amphibians worldwide (Scheele et al. 2019), and many in the southwestern United States (e.g., Hossack et al. 2017, 2022; Howell et al. 2020; Savage et al. 2018), populations are facing multiple known or suspected threats.
Ongoing threats to the Arizona Toad include effects of expected changes in hydrology and increased drought associated with climate change, as well as wildfire, disease, and invasive species (Ault et al. 2016; Bradford et al. 2020; Ryan et al. 2014; U.S. Fish and Wildlife Service 2015), challenges that are common to virtually all riparian species in the southwestern United States. In a broad meta-analysis of potential threats to 35 southwestern amphibians, Mims et al. (2020) identified invasive species, changing community composition, land-use, and altered hydrology as potential threats to native amphibians, but provided no information specific to the Arizona Toad. Changes in hydrology in particular could affect movement and subsequent gene flow in Arizona Toads as dispersal is thought to be limited by water and the presence of mesic habitat and sufficient rainfall (Schwaner and Sullivan 2005). The pathogenic chytrid fungus (Batrachochytrium dendrobatidis, Bd), that negatively affects many amphibians, is present throughout the region (Beirne 2015; Christman and Jennings 2018; Lannoo et al. 2011; Schlaepfer et al. 2007; Sigafus et al. 2014) and has been identified on Arizona Toads in New Mexico. Despite Bd continuing to contribute to declines in other western bufonids and ranids (e.g., Russell et al. 2019), we are unaware of evidence that Arizona Toad populations are affected by disease from this fungal pathogen (Ryan et al. 2014).
The construction of water impoundments over the past 100 years has facilitated hybridization between Arizona Toads and native Woodhouse’s Toads (A. woodhousii), which may have contributed to the local decline of Arizona Toads (Schwaner and Sullivan 2005; Sullivan 1993). Along the Bill Williams and Agua Fria rivers of Arizona, putative declines from this threat appear restricted to areas adjacent to newly established impoundments where the altered aquatic habitat is unfavorable to the Arizona Toad and allows contact between it and Woodhouse’s Toads (Schwaner and Sullivan 2009; Sullivan 1986; Sullivan et al. 2015). The Arizona Toad depends on shallow, perennial river and stream reaches with sandy, open floodplain habitats. These requirements suggest a greater sensitivity to flow reductions or drying, due to drought or to anthropogenic modifications. This sensitivity is unlike most other toads in the region that use a variety of natural and artificial water bodies (e.g., Woodhouse’s Toads, Schwaner and Sullivan 2005; Sullivan 1993).
The Arizona Toad was petitioned for federal protection under the U.S. Endangered Species Act in 2015 (U.S. Fish and Wildlife Service 2015), and was considered to be “warranted but precluded” with a request for additional information – specifically about the range, population trends and biology of the species (Federal Register: Endangered and Threatened Wildlife and Plants; 90-Day Findings on 31 Petitions). The species is slated for assessment by the U.S. Fish and Wildlife Service (USFWS) in fiscal year 2026 (https://www.fws.gov/media/national-listing-workplan-fiscal-years-2022-2027) and although it is currently listed as a species of greatest conservation concern in the four states where it occurs (Arizona, Nevada, New Mexico, and Utah) (Arizona Game and Fish Department 2022; Bradford et al. 2005; Nevada Department of Wildlife 2015; New Mexico Department of Game and Fish 2016; Utah Division of Wildlife Resources 2015), it currently has no federal protection.
The Arizona Toad occurs throughout much of its historical range in the southwestern United States (e.g., Blair 1955; Price and Sullivan 1988; Schwaner and Sullivan 2005; Sullivan 1993, 1995; Fig. 1). Although there have been no formal, range-wide assessments of population status or threats, reductions in distribution or number of populations are suspected in New Mexico (Forzley et al. 2021; Ryan et al. 2015), Arizona (Schwaner and Sullivan 2005), and Nevada (Bradford et al. 2005). Recent studies have documented idiosyncratic breeding activity and breeding success. For example, between 2013 and 2018, consecutive years of unseasonal flooding or drought in New Mexico were associated with no breeding activity or loss of entire cohorts (Ryan et al. 2015, but see Forzley et al. 2021). Recently, between 2020 and 2022, similar patterns were observed in Arizona (M. Ryan pers. obs.).
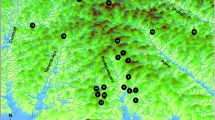
Location of Arizona Toad (Anaxyrus microscaphus) samples collected across the species’ range. Symbols represent samples belonging to the indicated drainages: orange diamond—Meadow Valley Wash, orange square—Beaver Dam Wash, light orange circle—Virgin River, light blue plus sign—Bill Williams River, light blue diamond—Verde River, light blue circle—Agua Fria River, blue X—Salt River, blue triangle—Gila River, blue square—Mimbres River, blue circle—Rio Grande (see Appendix 1 for sample details). The bold dotted line that coincides with the Grand Canyon separates the two genetically distinct groups. Within the southern group, more subtle genetic structure is present with separation (smaller dashed line) between eastern and western groups. AZ Arizona; NV Nevada, NM New Mexico, UT Utah
A central question underpinning all putative threats to the Arizona Toad is the degree of connectivity among populations across the species’ range. Animal dispersal has important implications for population demography, metapopulation dynamics, and species persistence as well as shaping genetic structure and patterns of diversity (Cosgrove et al. 2018; Fedy et al. 2017; Garant et al. 2005; Row et al. 2010, 2016; Tucker et al. 2018). Connectivity among populations of Arizona Toads is perhaps uniquely tied to components of biogeography (e.g., elevation and aridity) and may be negatively affected by any of the aforementioned threats. Information addressing connectivity among populations of interest for conservation planning is especially relevant to the USFWS’ species status assessment framework of resiliency, redundancy and representation (Smith et al. 2018). For instance, well-connected populations are typically considered more redundant than isolated populations, as high connectivity leads to genetic similarity among populations (i.e., populations are more genetically similar and redundant of one another). The ability to move easily among populations also buffers against a stochastic, catastrophic event as individuals are better able to recolonize after such an event. On the other hand, low connectivity can lead to genetically divergent populations which may harbor unique genetic variation associated with that specific location (e.g., local adaptation to temperature, habitat, or diet). This is relevant to USFWS’ representation concept as it involves ensuring distinct, genetic groups are considered for protection because of the potential for those populations to be locally adapted and the idea that protecting all unique genetic variation may improve the species’ ability to adapt to changing environments. The degree of connectivity among Arizona Toad populations is unknown despite its importance for management and conservation. Molecular methods are particularly useful for estimating gene flow, and improvements in technology and laboratory methods over the last decade have facilitated such analyses (Beja-Pereira et al. 2009; Haig et al. 2011; Oyler-McCance et al. 2021).
The objectives of this study were to characterize population structure and genetic diversity across this species’ range. Given the variable topography and hydrological patterns across the range of the Arizona Toad, we predicted that some parts of the range would be disconnected from others. Further, we predicted that some of these disconnects would be related to water availability and the aridity of the environment, which are associated with elevation (Schwaner and Sullivan 2005). Specifically, we predicted that low elevation sites in southern and central Arizona, that are much drier, would have less gene flow than sites at higher elevations in central and eastern Arizona and New Mexico.
Methods
Sampling methods
Tissues from Arizona Toads were collected opportunistically and during targeted surveys to assess the species’ range in Nevada (2020), Arizona (2018–2021), and New Mexico (2011–2019) (no targeted surveys were implemented in Utah). In Arizona, Nevada and New Mexico, samples were collected during surveys of historical localities and new, undocumented sites in appropriate riparian habitats within the Arizona Toad’s range (Forzley et al. 2021). Tissue samples (i.e., liver, toe clips, tadpoles) were obtained from either live animals that were subsequently released (some toe clips) or from freshly euthanized animals prior to preservation. Toe clips from live, adult toads were sampled by removing a portion of toe IV (i.e., longest toe) to just above the proximal tubercle to the toe pad with sterile scissors, placing each toe clip in 95% ethanol prior to releasing the animal at its point of capture. All adult voucher specimens were euthanized with either Benzocaine (administered as Oragel™ ) (Underwood and Anthony 2020, ASIH_HACC_FinalDraft.PDF (squarespace.com) or MS222 (ASIH_HACC_FinalDraft.PDF (squarespace.com) and then preserved (McDiarmid 1994). Livers were immediately placed in 95% ethanol (McDiarmid 1994; Beaupre et al. 2004). Tadpoles were euthanized in 95% ethanol (Simmons 2015). All tissues were stored frozen (minimum − 20 oC) until analysis. All specimens were catalogued at the Museum of Southwestern Biology (MSB) University of New Mexico.
Laboratory and analysis methods
Genomic DNA was extracted from all tissue samples using an ammonium acetate protocol (modified from the PUREGENE kit (Qiagen) to reduce reaction volume). Some samples could not be identified to species in the field, so we used the restriction-digest methods described in Lamb et al. (2000) to assign final taxonomic identities. We also sequenced the uncut polymerase chain reaction (PCR) products for portions of both cytochrome b (using primers from Lamb et al. 2000) and the 16 S rRNA regions (using primers from Bickham et al. 1996) following Oyler-McCance et al. (2013) and compared those sequences to the sequences available in GenBank. Only those samples identified as Arizona Toad were retained. We performed double digest restriction associated DNA (ddRAD) sequencing for 123 samples following the protocol outlined in Langin et al. (2018). Briefly, we used 1 µg of starting DNA (13 µL of 77ng/µL genomic DNA), used the enzymes Spe1 and Sau3A, and size selected for 300–500 bp. Library quality was checked on a BioAnalyzer High Sensitivity DNA Assay (Agilent Technologies) and quantified with a Qubit HS DNA Assay (Life Technologies). Libraries were sent to Genewiz LLC for sequencing on a NovaSeq (Illumina).
Locus clustering
Only the first read of each pair was used in this analysis because single nucleotide polymorphism (SNP) discovery on the paired read would be redundant for the current purposes, since tightly linked, unphased SNPs add little additional information to genetic-structure analysis while increasing bioinformatic complexity and run times (Falush et al. 2003). Raw reads were evaluated with FastQC (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/) and bbduk (https://jgi.doe.gov/data-and-tools/software-tools/bbtools/) to evaluate base call quality, the frequency of the expected Sau3A1 restriction site, and to check for exogenous sequence. Reads were cleaned with bbduk (Bushnell B. BBMap: sourceforge.net/projects/bbmap/) by first trimming to a length modulus of 5 to remove artifactual extra bases of poor quality (Bushnell B. BBMap: sourceforge.net/projects/bbmap/ no date), then trimming low-quality bases below a phred-scaled quality threshold of 10, and finally trimming exogenous sequences identified using kmer thresholds of 15 (read interior) and 11 (read edge). Reads less than 140 bases in length after trimming were discarded.
For locus discovery only, we down-sampled to at most five million reads per sample to mitigate bias that could arise from variation in read number among samples. Reads were then clustered within each sample at 97% with vsearch (Rognes et al. 2016) with iddef set to 1, representing a reasonable maximum within-individual variation per tag for downstream SNP discovery. Clusters were then denoised with the unoise algorithm implemented in vsearch specifying a minimum size of 2. Clusters smaller than five were discarded at this stage for computational efficiency and retained clusters from each sample were then pooled and reclustered at 95%, representing a reasonable maximum of inter-individual variation for downstream SNP discovery. A final round of clustering was performed to identify and remove very similar sets of tags (i.e., keeping only singleton clusters at 85% identity).
Evaluation of coverage distribution
Cleaned reads from each sample were mapped to the retained loci with bowtie2, specifying the “very-sensitive” and “end-to-end” parameter switches. Counts of reads mapping at a quality threshold of 20 were then obtained with SAMtools (Li et al. 2009), from which we plotted two empirical distributions of the loci: the average per-sample coverage and the coefficient of variation (CV) in coverage across samples. By inspection, we selected two sets of empirical coverage bounds. The first was between a minimum coverage of six (below which heterozygous sites would be frequently missed) and a maximum coverage of 20 (above which the distribution inflected away from the mean, suggesting repetitive sequences). The second was a CV between 0.7 and 1.3, given that the expected value is approximately 1 for Poisson-distributed counts (Anders and Huber 2010), and strong divergence from this expectation suggests copy-number variation among samples. Approximately 102,000 tags were retained after coverage filtering.
Variant calling
Variant calling was performed with the biallelic model of BCFtools (Li et al. 2009), following standard quality-control measures such as avoiding sites near indels, at edges of tags, and sites exhibiting base-call bias. Variants were required to have a reported quality score of 999 (the programmatic maximum), exhibit both heterozygous and homozygous genotypes in the total sample, and have a minor allele frequency of at least 0.05. Sites with a base-quality bias P-value less than 0.1, and sites within 5 bases of either end of a tag were excluded. We then removed samples with missing data at more than 10% of the variant sites.
Genetic diversity and population structure
To characterize genetic diversity, we calculated expected and observed heterozygosity in GenAlEx ver 6.5 (Peakall and Smouse 2012). We used several methods to investigate population differentiation and structure. First, we conducted a principal coordinates analysis by individual in GenAlEx ver. 6.5 (Peakall and Smouse 2012). We also assessed population structure using the Bayesian clustering algorithm implemented in STRUCTURE (Pritchard et al. 2000). We used a hierarchical approach, that first included all individuals in the analyses and later included subsets of samples that clustered together in prior analyses to identify finer-level clusters. All analyses used a model with no location prior, correlated allele frequencies and admixture. We ran STRUCTURE 10 times for each value of K from 1 to 5 using 300,000 Markov chain Monte-Carlo (MCMC) repetitions and a burn in of 150,000. We examined plots showing trends in estimates to ensure that they were stable following the burn in period. The best-supported value of the number of unique genetic groups (K) was determined using two methods implemented in STRUCTURE HARVESTER (Earl and vonHoldt 2012). First, we plotted the mean and standard deviation of ln Pr(X|K) at each value of K and evaluated the point at which those values plateaued. Second, we calculated and plotted ΔK following Evanno et al. (2005). STRUCTURE runs were summarized and visualized using CLUMPAK (Kopelman et al. 2015). Finally, we quantified the genetic differentiation among drainages by calculating pairwise differentiation metrics among all pairs of drainages. We estimated both Jost’s D (Jost 2008) and Nei’s GST (Nei 1973, 1978) in package mmod in R (Winter 2012).
We investigated the spatial distribution of population structure using the Estimated Effective Migration Surfaces (EEMS) program (Petkova et al. 2016). The EEMS program fits a model that estimates the spatial distribution of genetic similarity and then uses the rate of decay in genetic similarity to identify areas that deviate from the null expectation of isolation by distance (IBD) to identify gene flow corridors (less decay than expected) and barriers (more decay than expected). We fit the model assuming 300 demes in 4 independent chains for 5,000,000 MCMC iterations, discarding the first 2,000,000 iterations as a burn-in period, then sampling every 3,000 iterations, and retaining 1,000 for inference. We checked convergence through inspection of the log posterior trace plots and calculation of the Gelman-Rubin diagnostic (Gelman and Rubin 1992).
We then formally tested the prediction that higher elevation populations in the eastern watersheds in the southern genetic group (Salt River, Gila, Mimbres, Rio Grande) had relatively higher effective migration than the lower elevation western watersheds (Bill Williams, Verde, Aqua Fria). We used sampling coordinates to extract elevation from the WorldClim data (Fick and Hijmans 2017) and effective migration rates from the rasterized EEMS output generated with the eems.plots R script. We tested for a difference of means between the elevation and effective migration of the high and low elevation sites with a two-sample t-test in R.
Data generated during this study are available as a U.S. Geological Survey data release (Cornman et al. 2023).
Results
We analyzed 123 samples from ten watersheds and 37 waterbodies across the range of the Arizona Toad. After filtering and removing samples with high levels of missing data, our data set consisted of 99 individuals genotyped at 3,601 SNPs grouped by drainages (Table 1; Fig. 1). Analyzed samples were from New Mexico (MSB, n = 30); Arizona (Arizona Game and Fish, n = 13; BKS private collection, n = 12); Nevada (Nevada Department of Natural Resources, n = 4; MSB, n = 20); and Utah (Utah Division of Wildlife Resources, n = 20) (Table 1, Appendix 1). Samples consisted of 43 liver portions, 24 toe clips, and 32 tadpole tail clips. Liver samples and eight toe clips were acquired during specimen voucher preparation; 16 toe clips and 32 tadpole tail clips were collected in the field.
Heterozygosity values were generally similar among most drainages, yet samples from the Virgin River had the lowest observed heterozygosity (0.211) and the highest inbreeding coefficient (0.130) (Table 2). Principal coordinates analysis revealed three groups (Fig. 2), with the first axis explaining 23.35% of the variation and the second explaining an additional 5.32%. The first axis separated the Meadow Valley, Beaver Dam, and Virgin River drainages from the remaining drainages, while the second axis split the Agua Fria, Bill Williams, and Verde River drainages from the remaining drainages in the south (Salt, Mimbres, Rio Grande, and Gila rivers).
In our initial STRUCTURE analysis that included the full data set (n = 99), the most likely number of unique genetic groups (K) was 2 or 3 (Fig. 3). The two most common approaches for selecting the most appropriate K yielded different results. The point at which the mean ln Pr(X|K) began to plateau was at K = 3 (Fig. S1), whereas the ΔK indicated K = 2 (Fig. S2). Because there was a clear east/west break among the samples (particularly at K = 2), we separated our full data set into two groups (Group 1, north: Virgin River, Beaver Dam, and Meadow Valley washes, and Group 2, south: Rio Grande, Mimbres, Gila, Salt, Verde, Agua Fria, and Bill Williams rivers) and reran STRUCTURE for each of those two groups. Within Group 2, the point at which the mean ln Pr(X|K) began to plateau was K = 3 and ΔK suggested that the most supported K was 2 (Figs S3 and S4). The Rio Grande, Mimbres, and Gila river drainages formed one sub-group, the Verde, Agua Fria, and Bill Williams rivers formed another sub-group with the Salt River being an intermediate between them (Fig. 4). Within Group 1, the point at which the mean ln Pr(X|K) began to plateau was K = 3 (Fig. S5), whereas the ΔK indicated K = 2 (Fig. S6). At K = 2, samples within the Virgin River drainage were split into two groups, while the Beaver Dam and Meadow Valley samples aligned with the western Virgin River samples (Fig. 5). At K = 3, the three groups are comprised of Meadow Valley, Beaver Dam/western Virgin River, and eastern Virgin River (Fig. 5).
Estimated population genetic structure based on genetic variation at 3,601 single nucleotide polymorphisms as calculated in STRUCTURE at K = 2 (top) and K = 3 (bottom). Samples are ordered from northwest (far left) to southeast (far right). Each genetic group is represented by a unique color and each bar represents an individual Arizona Toad. The colors on the bars represent each individual’s estimated membership in each of the unique genetic groups
Estimated population genetic structure of the southern group of Arizona Toads based on genetic variation at 3,601 single nucleotide polymorphisms as calculated in STRUCTURE at K = 2 (top) and K = 3 (bottom). Samples are ordered from west (far left) to east (far right). Each genetic group is represented by a unique color and each bar represents an individual Arizona Toad. The colors on the bars represent each individual’s estimated membership in each of the unique genetic groups
Estimated population genetic structure of the northern group of Arizona Toads based on genetic variation at 3,601 single nucleotide polymorphisms as calculated in STRUCTURE at K = 2 (top) and K = 3 (bottom). Samples are ordered from west (far left) to east (far right). Each genetic group is represented by a unique color and each bar represents an individual Arizona Toad. The colors on the bars represent each individual’s estimated membership in each of the unique genetic groups
The pairwise genetic distances were consistent with the patterns found in our other analyses of population structure (Table 3). The highest values of differentiation using both Jost’s D and Nei’s GST occurred in comparisons that involved populations from both the northern and southern groups. These values were > 0.2 in both metrics, suggesting substantial differentiation. Differentiation was lower in comparisons between drainages within the northern and southern groups.
Our estimates of effective migration generally agreed with patterns of population structure (Fig. 6a). The sampling locations are divided into three main groups by a combination of IBD (white) and barriers to gene flow (lower than expected under IBD; red): group 1 = Meadow Valley, Beaver Dam, and Virgin River; group 2 = Bill Williams, Verde, and Agua Fria rivers; group 3 = Salt River, Gila, Mimbres, and Rio Grande rivers. The mean elevation was significantly higher (t18.91 = −5.72, p = 1.65e-05; Fig. 6b) for the eastern watersheds (mean = 1,873.08, se = 44.99; Salt, Gila, Mimbres, Rio Grande rivers) in the southern genetic group, than for the western watersheds (mean = 972.35, se = 150.82; Bill Williams, Verde, Aqua Fria rivers). As predicted, the higher elevation sites had significantly higher mean effective migration than the lower elevation sites, as predicted (t25.39 = −2.76, p = 0.01; Fig. 6c).
Estimated Effective Migration Surface (EEMS) for Arizona Toad. a The EEMS displays relative rates of effective migration: high = blue, low = red, white = consistent with isolation by distance. b Within the southern genetic cluster, the western watersheds (Bill Williams, Verde, Aqua Fria rivers) were at lower mean elevation than the eastern watersheds (Salt River, Gila, Mimbres, Rio Grande rivers). c The western watersheds also had lower mean effective migration than the eastern watersheds
Discussion
The most parsimonious interpretation of population structure across the range of the Arizona Toad indicates two genetically distinct groups. The first group includes populations in drainages in the northwest of the toad’s range in Utah and Nevada (i.e., Virgin River, Beaver Dam and Meadow Valley washes). The second group includes populations in drainages in the rest of the toad’s range in central and southeastern Arizona (i.e., Bill Williams, Agua Fria, Verde, and Salt rivers) and all populations in New Mexico. Within the second group we also found more subtle genetic differentiation between central Arizona (Bill Williams, Verde, and Agua Fria rivers) and those drainages to the east (Salt, Gila, Mimbres, and Rio Grande rivers). In general, this finding suggests that across the range of the Arizona Toad, the distribution of genetic variation is not uniform, and the species may benefit from region-specific management. There are a number of explanations for the differences that we found.
Presently, the distribution of the Arizona Toad spans a variety of topographic features, including putative barriers to dispersal such as large rivers, mountains, and basins, with two major geographic breaks across the range. The first and largest barrier is between the watersheds of the Virgin River, and the Bill Williams River on opposite sides of the Grand Canyon. We found differentiation between Group 1 (i.e., populations in the western drainages of the lower Virgin River and its tributaries in southern Utah and Nevada), and Group 2 (i.e., populations in central and southeastern Arizona and populations in New Mexico) which aligns with the first geographic break in the Arizona Toad’s range (Figs. 1, 2 and 3). Group 1 is separated from Group 2 by the mainstem of the Colorado River and the absence of relatively mesic uplands (e.g., forest and woodland with adjacent riparian corridors) that likely limit movements between watersheds. Based on our STRUCTURE analysis for Group 1 alone, there was subtle differentiation among the three drainages, with the Beaver Dam and western Virgin River samples being intermediate between the samples collected in the eastern Virgin River tributaries near Mt Carmel Junction and the Meadow Valley drainage (Fig. 5). In Group 2, farther to the south, the western drainages (Bill Williams, Agua Fria, and Verde rivers) were more subtly differentiated from the drainages to the east including the Salt River drainage that is not too distant geographically (Figs. 2, 4 and 6). The Colorado River separates the mouths of the Gila and the Bill Williams rivers, but the headwaters of the Bill Williams River are closer to other riparian systems in central Arizona (e.g., Hassayampa and Agua Fria rivers) suggesting that under some conditions (e.g., summer rains) gene exchange may occur between Arizona Toads from the Bill Williams populations and those in central Arizona. The second geographic break in the Arizona Toad range occurs at the continental divide between the Mimbres and Rio Grande watersheds (Fig. 1, within the New Mexico portion of Group 2), but our results do not indicate genetic differentiation among populations in these watersheds (Figs. 4 and 6).
There are two gaps within the distribution of the Arizona Toad for which we lack samples, possibly affecting our results. Samples were unavailable from these areas because Arizona Toads were not found (i.e., inhospitable habitat) or we were unable to survey the habitat. The first sampling gap is on either side of the Colorado River east of Lake Mead, including the Grand Canyon. While Arizona Toads have been observed southeast of Lake Mead in the Music Mountains (B. Sullivan, pers. obs.), much of this area is part of the Hualapai Reservation. Surveys off the reservation, however, have not found Arizona Toads. This gap corresponds to the first geographic break, and we suspect that rugged topography and limited streams and springs in this area likely preclude widespread occurrence in this area. The second sampling gap is in east-central Arizona along the eastern Mogollon Rim and foothills. Much of this area belongs to the White Mountain Apache, San Apache, and Fort Apache Tribes and to our knowledge has not been surveyed recently for the Arizona Toad. Although we lacked genetic samples from this area there are some historic museum records (prior to 1991, but tissues were not available) suggesting that Arizona Toads occur in this area, but the extent of occurrence and current status is unknown. The Salt River samples (west of this region) largely align with the Gila River samples (east of this region), suggesting gene flow through this area. Further, our effective migration surface (Fig. 6) suggests that this alignment is likely the result of isolation by distance (compared to the more stringent barriers to gene flow from Salt River west toward Agua Fria). Collaboration with Tribal Nations on future surveys will help to determine if Arizona Toads are present in these two regions, connecting populations to the east and west as predicted by our genetic data.
Data on genetic population structure in other bufonids of the southwestern United States are relatively scarce, with a few exceptions. The Great Plains Toad (Anaxyrus cognatus) breeds primarily in rain-formed pools during the summer. Jungels et al. (2010) and Chan and Zamudio (2009) documented a low level of genetic differentiation among breeding populations of this toad at distances of 1-100 km. These authors argued that high levels of connectivity (i.e., gene flow) occur because of homogeneity in the desert flats where these toads breed, uncertainty in breeding site locations across years, and the species’ adaptations to desert existence that allow them to move larger distances than one might expect for an anuran inhabiting arid habitats. A range-wide phylogeographic analysis of the Arroyo Toad (A. californicus), a sister taxon of the Arizona Toad, found in California and Baja California, Mexico, found two well-supported clades for northern and southern populations, and a strong genetic connectivity within watersheds in both areas (Lovich 2009). Based on previous work on movements and habitat use in Arroyo Toads, Lovich (2009) suggested that gene flow may be higher because toads are able to move between drainages in the more mesic landscapes of the southern populations. This idea was corroborated by studies on movement patterns of Arroyo Toads in northern populations inhabiting more xeric environments (Atkinson et al. 2003; Barto 1999; Beaman et al. 1995; Brown and Fisher 2002; Campbell et al. 1996; Griffin and Case 2001; Holland and Sisk 2000; Ramirez 2000; Sweet and Sullivan 2005; U. S. Fish and Wildlife Service 1999).
Information from A. cognatus and A. californicus suggest that movement among populations can occur, but is likely limited to the availability of relatively mesic habitat corridors or seasonal rainfall events that facilitate connectivity and allow dispersal. We found gene flow opportunities in the Arizona Toad to be lower (i.e., less connectivity) in the low elevation populations in west and central Arizona (Bill Williams, Agua Fria, and Verde rivers, Fig. 6b,c). Breeding sites in this area are less connected due to the harsh, dry conditions between ephemeral or intermittent streams in the Sonoran Desert Scrub habitat. Conversely, we showed that gene flow opportunities are higher (i.e., more connectivity) among high elevation populations in central and eastern Arizona and New Mexico (i.e., Salt, Gila, Mimbres, and Rio Grande headwaters) where populations occur in more mesic forest biomes and experience increased summer rainfall during their activity periods (Schwaner and Sullivan 2005). Our finding, that connectivity among populations of the Arizona Toad is facilitated by mesic conditions during summer rains at high elevations, is the reverse of the “valley-mountain” model proposed by Funk et al. (2005) to describe the structure of population genetics for Columbia Spotted Frog (Rana luteiventris) in the Pacific Northwest. The model was based on their analysis of genetic differentiation among Columbia Spotted Frog populations at high and low elevations, and a literature review of other ranid frogs in the region. Funk et al. (2005) argued that high elevation ridges serve as barriers to gene flow for these riparian dwelling anurans between drainages, but at lower elevations, valley bottoms allow higher gene flow rendering populations genetically less divergent.
Differences in the abilities of non-desert dwelling frogs and desert-dwelling toads to tolerate low water environments may account for this hypothesis reversal. The difference in connectivity that we found between populations of the Arizona Toad in western Arizona (i.e., Verde, Agua Fria and Bill Williams rivers) and those to the east (i.e., Salt, Gila, Mimbres rivers) supports the view that populations inhabiting watersheds at high elevations with headwater streams in mesic forest biomes experience higher connectivity that may be lacking among drainages in more arid environments. However, additional investigation into dispersal in the Arizona Toad, and especially limitations to dispersal, would be useful.
Our results can be used to prioritize surveys and habitat assessments in under-studied areas, such as upper headwater stream reaches, with the aim of determining the functionality (e.g., availability of water) of these corridors in maintaining connectivity among populations and determining if toads are present in these habitats. If higher connectedness among populations at higher elevations is a driving force in the population structure of the Arizona Toad, the loss of headwater springs from a combination of regional drought, reduced snowpack, and severe fire may lead to population isolation. Loss of breeding habitats around headwaters may push the Arizona Toad further downslope toward more xeric habitats, thereby increasing dispersal distance, reducing the availability of mesic corridors, and decreasing connectivity. The genetic data presented here can inform conservation decisions and strategies (e.g., whether management is needed and whether different strategies are required for different groups of the Arizona Toad). These data may also aid in identifying areas for habitat restoration and in identifying source populations for assisted migration or translocation to support small or potentially declining populations of this species. Considering our evidence of genetic differentiation between the north and south and the existing geographic barriers to dispersal, other threats are likely to have an increased impact on this species, especially climate change-associated threats of regional drought - including reduced snowpack, anthropogenic alterations to hydrologic flow, and increased wildfire intensity.
Data availability
The genotype data and sequencing vcf files are available in a U.S. Geological Survey data release (Corman et al. 2023) and genomic sequencing data have been deposited in GenBank (biosample accession number: PRJNA995169).
References
Anders S, Huber W (2010) Differential expression analysis for sequence count data. Nat Prec. https://doi.org/10.1038/npre.2010.4282.1
Arizona Game and Fish Department (2022) Arizona Wildlife Conservation Strategy: 2022–2032. Arizona Game and Fish Department, Phoenix, Arizona. https://azgfd-wdw.s3.amazonaws.com/awcs-2022/documents/AWCS_Final_Approved_11-22.pdf
Atkinson AJ, Yang BS, Fisher RN, Ervin E, Case TJ, Scott N, Shaffer HB (2003) MCB Camp Pendleton Arroyo Toad Monitoring Protocol. U. S. Geological Survey Technical Report. p 42
Ault TR, Mankin JS, Cook BI, Smerdon JE (2016) Relative impacts of mitigation, temperature, and precipitation on 21st-century megadrought risk in the American Southwest. Sci Adv 2:e1600873. https://doi.org/10.1126/sciadv.1600873
Barto WS (1999) Predicting potential habitat for the arroyo toad (Bufo microscaphus californicus) in San Diego County using a habitat suitability model and digital terrain data. Master’s Thesis. San Diego State University, San Diego, California
Beaman K, Myers SJ, McGaugh C (1995) San Bernardino National Forest, Arrowhead and Big Bear Ranger districts: Report on surveys for Arroyo toads on Deep Creek. Tierra Madre Consultants, Inc. Report no. 95–023. p 16
Beaupre SJ, Jacobson ER, Lillywhite HB, Zamudio K (2004) Guidelines for the use of live amphibians and reptiles in field and laboratory research. Herpetological Animal Care and Use Committee (HACC) of the American Society of Ichthyologists and Herpetologists, 2nd edn.
Beirne SA (2015) An Overview of Batrachochytrium dendrobatidis in Utah, With a Focus on Boreal Toads and Their Changing Conservation Status. Utah State University, Undergraduate honors program. https://digitalcommons.usu.edu/honors/579/
Beja-Pereira A, Oliveira R, Alves PC, Schwartz MK, Luikart G (2009) Advancing ecological understandings through technological transformations in noninvasive genetics. Mol Ecol Res 9:1279–1301. https://doi.org/10.1111/j.1755-0998.2009.02699.x
Bickham JW, Lamb T, Minx P, Patton JC (1996) Molecular systematics of the genus Clemmys and the intergeneric relationships of emydid turtles. Herpetologica 52:89–97. https://www.jstor.org/stable/3892960
Blair AP (1955) Distribution, variation and hybridization in a relict toad (Bufo microscaphus) in southwestern Utah. Amer Mus Novit 1722:1–37
Bradford DF, Jaeger JR, Shanahan SA (2005) Distributional changes and population status of amphibians in the eastern Mojave Desert. West N Am Nat 65:462–472. https://www.jstor.org/stable/41717482
Bradford JB, Schlaepfer DR, Lauenroth WK, Palmquist KA (2020) Robust ecological drought projections for drylands in the 21st century. Glob Change Biol 26:3906–3919. https://doi.org/10.1111/gcb.15075
Brown C, Fisher RN (2002) Survey results for the arroyo toad (Bufo californicus) in the San Bernardino National Forest, 2001. Final report, U.S. Geological Survey, Western Ecological Research Center. p 53
Campbell LA, Graham TB, Thibault LP, Stine PA (1996) The arroyo toad (Bufo microscaphus californicus), ecology, threats, recovery actions, and research needs. U.S. Dept. Interior, NBS, California Science Center, p 46. Tech. Report (NBS/CSC-96-01)
Chan LM, Zamudio KR (2009) Population differentiation of temperate amphibians in unpredictable environments. Mol Ecol 18:3185–3200. https://doi.org/10.1111/j.1365-294X.2009.04273.x
Christman BL, Jennings RD (2018) Distribution of the Amphibian Chytrid Fungus Batrachochytrium dendrobatidis (bd) in New Mexico. New Mexico Department of Game and Fish, Santa Fe, New Mexico, USA
Corman RS, Fike JA, Oyler-McCance SJ, Muths EL (2023) Reduced representation genome sequencing of Arizona Toad (Anaxyrus microscaphus) for population genetics. U.S. Geological Survey data release. https://doi.org/10.5066/P9621XL1
Cosgrove AJ, McWhorter TJ, Maron M (2018) Consequences of impediments to animal movements at different scales: a conceptual framework and review. Divers Distrib 24:448–459. https://doi.org/10.1111/ddi.12699
Cornman RS, Fike J, Oyler-McCance SJ, Muths EL (2023) Reduced representation sequencing and genotyping of Arizona Toads (Anaxyrus microscaphus) from the southwestern United States: U.S. Geological Survey data release, https://doi.org/10.5066/P9621XL1.
Earl DA, VonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–336. https://doi.org/10.1007/s12686-011-9548-7
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genet 164:1567–1587. https://doi.org/10.1093/genetics/164.4.1567
Fedy BC, Row JR, Oyler-McCance SJ (2017) Integration of genetic and demographic data to assess population risk in a continuously distributed species. Conserv Genet 18:89–104. https://doi.org/10.1007/s10592-016-0885-7
Fick SE, Hijmans RJ (2017) Worldclim 2: new 1-km spatial resolution climate surfaces for global land areas. Intern J Climatology 37:4302–4315. https://doi.org/10.1002/joc.5086
Forzley MJ, Ryan MJ, Latella IM, Giermakowski JT, Muths E, Sigafus BH, Hossack BR (2021) Staggered-Entry Analysis of Breeding Phenology and Occupancy Dynamics of Arizona Toads from historically occupied habitats of New Mexico, USA. Ichthyol Herpetol 109:851–859. https://doi.org/10.1643/h2020133
Funk WC, Blouin MS, Corn PS, Maxell BA, Pilliod DS, Amish S, Allendorf FW (2005) Population structure of Columbia spotted frogs (Rana luteiventris) is strongly affected by the landscape. Mol Ecol 14:483–496. https://doi.org/10.1111/j.1365-294X.2005.02426.x
Garant D, Kruuk LE, Wilkin TA, McCleery RH, Sheldon BC (2005) Evolution driven by differential dispersal within a wild bird population. Nature 433:60–65. https://doi.org/10.1038/nature03051
Gelman A, Rubin DB (1992) Inference from iterative simulation using multiple sequences. Stat Sci 7:457–511. https://doi.org/10.1214/ss/1177011136
Griffin PC, Case TJ (2001) Terrestrial habitat preferences of adult arroyo southwestern toads. J Wildl Manag 65:633–644. https://www.jstor.org/stable/3803014
Haig SM, Bronaugh WM, Crowhurst RS, D’Elia J, Eagles-Smith CA, Epps CW et al (2011) Genetic applications in Avian Conservation. Auk 128:205–229. https://doi.org/10.1525/auk.2011.128.2.205
Holland DC, Sisk NR (2000) Habitat use and population demographics of the arroyo toad (Bufo californicus) on MCB Camp Pendleton, San Diego County, California: Final report for 1998–1999. AC/S Environmental Security, United States Marine Corps, Camp Pendleton, California
Hossack BR, Honeycutt RK, Sigafus BH, Muths E, Crawford CL, Jones TR, Sorensen JA, Rorabaugh JC, Chambert TA (2017) Informing recovery in a human-transformed landscape: drought-mediated coexistence alters population trends of an imperiled salamander and invasive predators. Biol Conserv 209:377–394. https://doi.org/10.1016/j.biocon.2017.03.004
Hossack BR, Howell PE, Owens AK, Cobos C, Goldberg CS, Hall D, Hedwall S, MacVean SK, MacCaffery M, McCall AH, Mosley CD (2022) Identifying factors linked with persistence of reintroduced populations: lessons learned from 25 years of amphibian translocations. Glob Ecol Conserv 35:e02078. https://doi.org/10.1016/j.gecco.2022.e02078
Howell PE, Hossack BR, Muths E, Sigafus BH, Chenevert-Steffler A, Chandler RB (2020) A statistical forecasting approach to metapopulation viability analysis. Ecol Apps 30:e02038. https://doi.org/10.1002/eap.2038
Jost L (2008) GST and its relatives do not measure differentiation. Mol Ecol 17:4015–4026. https://doi.org/10.1111/j.1365-294X.2008.03887.x
Jungels JM, Griffis-Kyle KL, Boeing WJ (2010) Low genetic differentiation among populations of the Great Plains Toad (Bufo cognatus) in southern New Mexico. Copeia 2010(3):388–396. https://doi.org/10.1643/CH-09-152
Kopelman NM, Mayzel J, Jakobsson M, Rosenberg NA, Mayrose I (2015) CLUMPAK: a program for identifying clustering modes and packaging population structure inferences across K. Mol Ecol Res 15:1179–1119. https://doi.org/10.1111/1755-0998.12387
Lamb T, Sullivan BK, Malmos K (2000) Mitochondrial gene markers for the hybridizing toads Bufo microscaphus and Bufo woodhousii in Arizona. Copeia 20001:234–237. https://doi.org/10.1643/0045-8511(2000)2000[0234:MGMFTH]2.0.CO;2
Langin KM, Aldridge CL, Fike JA, Cornman RS et al (2018) Characterizing range-wide divergence in an alpine-endemic bird: a comparison of genetic and genomic approaches. Conserv Genet 19:1471–1485. https://doi.org/10.1007/s10592-018-1115-2
Lannoo MJ, Petersen C, Lovich RL, Nanjappa P, Phillips C, Mitchell JC, Macallister I (2011) Do frogs still get their kicks on Route 66? Continental U.S. Transect Reveals Spatial and temporal patterns of Batrachochytrium dendrobatidis infection. PLoS ONE 6:e22211. https://doi.org/10.1371/journal.pone.0022211
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25:2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Lovich RE (2009) Phylogeography and Conservation of the Arroyo Toad (Bufo californicus). PhD. Dissertation. Loma Linda University. Loma Linda, California
McDiarmid RW (1994) Preparing amphibians as scientific specimens. In: Heyer WR, Donnelly MA, McDiarmid RW, Hayek LC, Foster MS (eds) Measuring and monitoring Biological Diversity, Standard methods for amphibians. Smithsonian Institution Press, Washington, DC, pp 289–297
Mims MC, Moore CE, Shadle EJ (2020) Threats to aquatic taxa in an arid landscape: knowledge gaps and areas of understanding for amphibians of the American Southwest. WIREs Water 7:e1449. https://doi.org/10.1002/wat2.1449
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Nat Acad Sci 70:3321–3323. https://doi.org/10.1073/pnas.70.12.3321
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genet 89:583–590. https://doi.org/10.1093/genetics/89.3.583
Nevada Department of Wildlife (2015) Nevada. State Wildlife Action Plan. Nevada Department of Wildlife, Reno, Nevada. http://www.ndow.org/uploadedFiles/ndoworg/Content/Nevada_Wildlife/Conservation/2013-NV-WAP-ID-SOCP-Wildlife-Landscape.pdf
New Mexico Department of Game and Fish (2016) State Wildlife Action Plan for New Mexico. New Mexico Department of Game and Fish, Santa Fe, New Mexico, USA. https://www.wildlife.state.nm.us/download/conservation/swap/New-Mexico-State-Wildlife-Action-Plan-SWAP-Final-2019.pdf
Oyler-McCance SJ, Valdez EW, O’Shea TJ, Fike JA (2013) Genetic characterization of the Pacific sheath-tailed bat (Emballonura semicaudata rotensis) using mitochondrial DNA sequence data. J Mammal 94:1030–1036. https://doi.org/10.1644/13-MAMM-A-006.1
Oyler-McCance SJ, Oh KP, Zimmerman SJ, Aldridge CL (2021) The transformative impact of genomics on sage-grouse conservation and management. Wildlife, Population Genomics, pp 523–546
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28:2537–2539. https://doi.org/10.1111/j.1471-8286.2005.01155.x
Petkova D, Novembre J, Stephens M (2016) Visualizing spatial population structure with estimated effective migration surfaces. Nat Genet 48:94–100. https://doi.org/10.1038/ng.3464
Price AH, Sullivan BK (1988) Bufo microscaphus. Cat Am Amphib Rept 415:1–3
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1093/genetics/155.2.945
R Core Team (2022) R: A language and environment for statistical computing Vienna, Austria. R Foundation for Statistical Computing. https://www.R-project.org/
Ramirez RS Jr (2000) Arroyo toad (Bufo californicus) radio telemetry study, Little Rock Creek, Los Angeles County, California. U.S.D.A. Forest Service, Arcadia, California
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584. https://doi.org/10.7717/peerj.2584
Row JR, Blouin-Demers G, Lougheed SC (2010) Habitat distribution influences dispersal and fine-scale genetic population structure of eastern foxsnakes (Mintonius Gloydi) across a fragmented landscape. Mol Ecol 9:5157–5171. https://doi.org/10.1111/j.1365-294X.2010.04872.x
Row JR, Oyler-McCance SJ, Fedy BC (2016) Differential influences of local subpopulations on regional diversity and differentiation for greater sage-grouse (Centrocercus urophasianus). Mol Ecol 25:4424–4437. https://doi.org/10.1111/mec.13776
Russell RE, Halstead BJ, Mosher BA, Muths E, Adams MJ, Grant EH, Fisher RN, Kleeman PM, Backlin AR, Pearl CA, Honeycutt RK, Hossack BR (2019) Effect of amphibian chytrid fungus (batrachochytrium dendrobatidis) on apparent survival of frogs and toads in the western USA. Biol Conserv 236:296–304. https://doi.org/10.1016/j.biocon.2019.05.017
Ryan MJ, Latella IM, Painter CW, Giermakowski JT, Christman B, Jennings RD, Voyles JL (2014) First record of Batrachochytrium dendrobatidis in the toad Anaxyrus microscaphus. Herpetol Rev 45:616–618
Ryan MJ, Latella IM, Giermakowski JT, Snell HL (2015) Final report: Status of the Arizona Toad (Anaxyrus microscaphus) in New Mexico. New Mexico Department of Game and Fish. Available at the University of New Mexico's Digital Repository. https://digitalrepository.unm.edu/
Ryan MJ, Giermakowski JT, Latella IM, Snell HL (2017) Final report: Status of the Arizona Toad (Anaxyrus microscaphus) in New Mexico. Submitted to New Mexico Department of Game and Fish. Available at the University of New Mexico’s Digital Repository. https://digitalrepository.unm.edu/
Savage AE, Mulder KP, Torres T, Wells S (2018) Lost but not forgotten: MHC genotypes predict overwinter survival despite depauperate MHC diversity in a declining frog. Conserv Genet 19:309–322. https://doi.org/10.1007/s10592-017-1001-3
Scheele BC, Pasmans F, Skerratt LF, Berger L, Martel A, Beukema W et al (2019) Amphibian fungal panzootic causes catastrophic and ongoing loss of biodiversity. Science 3636434:1459–1463. https://doi.org/10.1126/science.aav0379
Schlaepfer MA, Sredl MJ, Rosen PC, Ryan MJ (2007) High prevalence of Batrachochytrium dendrobatidis in wild populations of lowland leopard frogs Rana yavapaiensis in Arizona. EcoHealth 4:421–427. https://doi.org/10.1007/s10393-007-0136-y
Schwaner TD, Sullivan BK (2005) Bufo microscaphus Cope, 1867 ‘‘1866’’: Arizona toad. In: Lannoo MJ (ed) Amphibian declines: the Conservation Status of United States species. University of California, Berkeley, CA, pp 422–424
Schwaner TD, Sullivan BK (2009) Fifty years of hybridization: introgression between the Arizona Toad (Bufo microscaphus) and Woodhouse’s Toad (B. Woodhousii) along Beaver Dam Wash in Utah. Herpetol Conserv Biol 4:198–206
Sigafus BH, Schwalbe CR, Hossack BR, Muths E (2014) Prevalence of the amphibian chytrid fungus (batrachochytrium dendrobatidis) at Buenos Aires National Wildlife Refuge, Arizona, USA. Herpetol Rev 45:41–42
Simmons JE (2015) Herpetological Collecting and Collections Management, Third edition, Society for the Study of Amphibians and Reptiles Herpetological Circular No. 42, p 191
Smith DR, Allan NL, McGowan CP, Szymanski JA, Oetker SR, Bell HM (2018) Development of a species status assessment process for decisions under the US Endangered species Act. J Fish Wildl Manag 9:302–320. https://doi.org/10.3996/052017-JFWM-041
Sullivan BK (1986) Hybridization between the toads Bufo microscaphus and Bufo Woodhousei in Arizona: morphological variation. J Herpetol 20:11–21. https://www.jstor.org/stable/1564120
Sullivan BK (1993) Distribution of the southwestern toad (Bufo microscaphus) in Arizona. Great Basin Nat 53:402–406. https://www.jstor.org/stable/41712805
Sullivan BK (1995) Temporal stability in hybridization between Bufo microscaphus and Bufo woodhousii (Anura: Bufonidae): behavior and morphology. J Evol Bio 8:233–247. https://doi.org/10.1046/j.1420-9101.1995.8020233.x
Sullivan BK, Wooten J, Schwaner TD, Sullivan KO, Takahashi M (2015) Thirty years of hybridization between toads along the Agua Fria River in Arizona: part I. evidence from morphology and mtDNA. J Herpetol 49:150–156. https://doi.org/10.1670/14-011
Sweet SS, Sullivan BK (2005) Bufo californicus Camp, 1915 Arroyo Toad. In: Lannoo M (ed) Amphibian declines: the conservation status of United States species. University of California Press, Berkeley, California, pp 396–400
Tucker MA, Böhning-Gaese K, Fagan WF, Fryxell JM, Van Moorter B, Alberts SC et al (2018) Moving in the Anthropocene: global reductions in terrestrial mammalian movements. Science 359:466–469. https://doi.org/10.1126/science.aam9712
U.S. Fish and Wildlife Service (1999) Arroyo southwestern toad (Bufo microscaphus californicus) recovery plan. U. S. Fish and Wildlife Service, Portland. Oregon. vi + p 119
U.S. Fish and Wildlife Service (2015) 90-day finding on a petition to list the Arizona Toad under the endangered species Act. Fed Reg 80(126):37568–37579
Underwood W, Anthony R (2020) AVMA guidelines for the euthanasia of animals: 2020 ed. American Veterinary Medical Association. https://www.spandidos-publications.com/var/AVMA_euthanasia_guidelines_2020.pdf
Utah Division of Wildlife Resources (2015) Utah State Wildlife Action Plan. Utah Division of Wildlife Resources, Salt Lake City, Utah. https://wildlife.utah.gov/pdf/WAP/Utah_WAP.pdf
Winter DJ (2012) MMOD: an R library for the calculation of population differentiation statistics. Mol Ecol Resour 12:1158–1160. https://doi.org/10.1111/j.1755-0998.2012.03174.x
Acknowledgements
Any use of trade, firm, or product names is for descriptive purposes only and does not imply endorsement by the U.S. Government. This is contribution number 892 of the U.S. Geological Survey Amphibian Research and Monitoring Initiative (ARMI). Taylor Cotten, Sidney Riddle, Sarah Baker, and Keith and Elizabeth Sullivan assisted in the collection of tissue samples. Kevin Wheeler, Utah Department of Natural Resources, Division of Wildlife Resources, assisted with sample collection. This work was conducted under New Mexico Department of Game and Fish Scientific Collection Authorization #3329 and University of New Mexico Institutional Animal Care and Use Committee Protocol 13-100983-MC.
Funding
This research was funded by the New Mexico Department of Game and Fish State Wildlife Program Grants T-32-4 and T-32-P-3, and the U.S. Fish and Wildlife Service State Wildlife Grant T-32-P3, 18; the Arizona Game and Fish Department’s State Wildlife Grant (SWG) Program and Arizona Heritage Fund; and the U.S. Geological Survey Amphibian Research and Monitoring Initiative (ARMI).
Author information
Authors and Affiliations
Contributions
E. Muths, S. Oyler-McCance, M. Ryan, and B. Hossack conceived the study. J.T. Giermakowski, R. Harrow, S. Hedwall, I. Latella, R. Lovich, M. Ryan, S. Siefken, B. Sigafus, and B. Sullivancollected samples for the study. R. Cornman, S. Zimmerman, and S. Oyler-McCance analyzed the data. J. Fike coordinated sample collection and did library preparation for sequencing. E. Muths, S. Oyler-McCance, B. Sullivan, M. Ryan, J. Fike and R. Cornman wrote the initial draft. J.T. Giermakowski, R. Harrow, S. Hedwall, B. Hossack, I. Latella, R. Lovich, S. Siefken, B. Sigafus, and S. Zimmerman edited later drafts.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Oyler-McCance, S.J., Ryan, M.J., Sullivan, B.K. et al. Genetic connectivity in the Arizona toad (Anaxyrus microscaphus): implications for conservation of a stream dwelling amphibian in the arid Southwestern United States. Conserv Genet 25, 835–848 (2024). https://doi.org/10.1007/s10592-024-01606-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-024-01606-w