Abstract
The clouded apollo (Parnassius mnemosyne) used to have a wide distribution in Fennoscandia. Recent population declines have, however, led to regional extinctions and in Sweden it is currently one of the most endangered butterflies, confined to three geographically separated metapopulations: Blekinge, Roslagen and Västernorrland. Especially the Blekinge population has declined dramatically and few imagines have been observed during recent census efforts (< 10 in some localities). The clouded apollo is subject to a species action plan which includes both habitat restorations and captive breeding to produce individuals for release and reintroductions. Here, we apply whole-genome resequencing of clouded apollo individuals collected in the three natural populations and the captive population in Sweden and apply population genomic approaches to get a better understanding of the genetic structure and levels of genetic diversity in the species. We find that the clouded apollo populations in the different geographic regions have similar, but comparatively low levels of genetic diversity and we find evidence for significant genetic differentiation between the northernmost population and the populations in southern Sweden. Additional analysis, including previously available mitochondrial data, unveil that a bi-directional re-colonization of Fennoscandia after the latest glacial maximum most likely is the explanation for the considerable differentiation between some Swedish populations. Finally, we find evidence for population sub-structure in one of the Swedish populations. The results provide insights into the genetic consequences of population size declines and fragmentation in general and provide important information for direct conservation actions for the clouded apollo in Sweden in particular.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The clouded apollo (Parnassius mnemosyne) has a wide distribution that spans from central Asia to western Europe, but the species shows declining numbers and increasing fragmentation in many countries (Bolotov et al. 2013; Eliasson et al. 2005; Gratton et al. 2008). In Fennoscandia, the clouded apollo can currently be found in Finland, Sweden and Norway, but it has been extinct in Denmark since the early 1960:s (Eliasson et al. 2005). Within Sweden, the distribution range was previously more or less continuous from the far southern region, along the east coast, up to central Sweden (Artdatabanken 2020, 2021) and it was classified into two different subspecies (P. m. argiope in southernmost Sweden and P. m. romani in the north of the southernmost provinces) based on differences in size and pigmentation (Eliasson et al. 2005; Liljeblad 2021). Over the last decades, the distribution range has decreased and currently the clouded apollo is limited to three comparatively small and geographically isolated regions; Blekinge, Roslagen and Västernorrland (Fig 1) (Artdatabanken 2020, 2021). Besides retreating from large parts of the historical distribution, population sizes have also decreased dramatically—the clouded apollo is currently red-listed (classified as an endangered species) and since 2008 it has been subject to a national action plan initiated by the Swedish Environmental Protection Agency (SEPA, Naturvårdsverket) (Artdatabanken 2020; Franzén and Imby 2008). Recurrent efforts to estimate the census sizes within each geographic region have been conducted. In 1994–1995, clouded apollos were observed in 43 different localities within the three distribution areas and the numbers of individuals were estimated to be approximately 1500 in Roslagen and Västernorrland, respectively, and 150–200 in Blekinge (Artdatabanken 2021). Since, both the number of inhabited localities within the different geographic areas and the number of individuals has decreased further (Artdatabanken 2021; Eliasson 2021; Holst et al. 2021). The current status is that the situation for the species is critical in Blekinge—no egg laying females were observed 2020—while the situation seems to be more stable in Västernorrland (Artdatabanken 2021; Holst et al. 2021).
The most likely reason for the dramatic decline of the clouded apollo across the distribution range is continuous loss of suitable breeding habitat (Holst et al. 2021). The species prefers sheltered landscapes with a mosaic of moderately grazed pastures or mowed meadows, deciduous forest edges and shrubs, and access to larval host plants (Corydalis sp.) and a rich flora of flowering plants for nectar feeding imagines (Franzén and Imby 2008). As a consequence of the transition from small-scale agricultural practices to centralized land management, such habitats have been lost at a high rate due to locally heavier grazing pressure and overgrowth of small meadows that are no longer used as temporary pastures or for mowing/haymaking (Johansson et al. 2017). Efforts to restore preferred habitats in Roslagen and Blekinge have resulted in temporary breaks in the negative population trends, but it is likely that more regular and large-scale restoration initiatives will be needed to prevent further population fragmentation (Holst et al. 2021) and loss of genetic diversity as a consequence of genetic drift and inbreeding (e.g. Frankham et al. 2012; Sherpa et al. 2021).
Genetic tools to characterize and quantify potential population differentiation and levels of genetic diversity in the clouded apollo have previously been limited to a few microsatellites (Meglecz and Solignac 1998) and mitochondrial gene sequences (Gratton et al. 2008). Here, we use tissue samples from clouded apollos collected in the three geographically distinct Swedish natural populations and samples from a captive population at Nordens Ark and apply whole genome re-sequencing to generate polymorphism data. The data are used to quantify the levels of intra-population genetic diversity, levels of inter-population genetic differentiation and assess relatedness coefficients to characterize inbreeding in each population. Our results reveal comparatively low genetic diversity in all populations and significant genetic differentiation both between isolated populations and between subpopulations within Roslagen. We also find evidence for postglacial colonization of clouded apollo in Scandinavia from two directions.
Methods
Sampling
For the whole-genome re-sequencing, 10 individuals from each of the three natural populations and 10 individuals from the captive population at Nordens Ark were used. Note that the captive population was founded by individuals from Blekinge. All samples used for sequencing were collected between 2015 and 2020 and stored frozen in ethanol or dried, either as complete specimens or as pieces of legs or wings (Supplementary Table 1).
DNA extractions and sequencing
DNA-extractions were performed using a modified salt extraction protocol optimized for very small amounts of tissue. The protocol was optimized with trial extractions from other dried butterfly specimens already available at Uppsala University. The amount and quality of DNA extractions were assessed with Qubit and NanoDrop for a larger set of samples and 10 individuals with the highest yield from each population were sent to NGI/SciLifeLab Stockholm for library preparation and sequencing. For library preparation, the SMARTer ThruPLEX DNA-seq kit was used and the barcoded libraries were sequenced in multiplex on a single NovaSeq S4-300 flow cell with 2x151 bp read lengths. Before delivery of raw sequence data, NGI performed demultiplexing and basic quality control using FastQC (Andrews 2021). The sequencing yield varied between 47.68 and 106.60 million read pairs (14.30–31.98 Gb) per individual and no library had fewer than 92.5% of nucleotides with a Phred-score > 30 (Supplementary Table 2).
Identification of nuclear polymorphisms
The paired end reads for all 40 individuals were mapped to the repeat masked apollo butterfly (Parnassius apollo) reference genome (Podsiadlowski et al. 2021), using BWA aln (Li and Durbin 2010) as implemented in Stampy 1.0.31 (Lunter and Goodson 2011). The mapping failed for two individuals, one from Västernorrland and one from Nordens Ark (VN18 and NAB23), and those samples were excluded from further analyses. Single nucleotide polymorphisms (SNPs) were identified using BCFtools 1.13 (Li 2011) for all 38 samples together. In total, we called 93,064,863 SNPs. Since butterflies in the Parnassius genus have comparatively large genome sizes with a high fraction of transposable elements (TEs)—many of which have likely not been properly annotated and are therefore missing in previous TE-libraries used for repeat masking—there was a high risk for erroneously calling SNPs as a consequence of mapping errors in repetitive sequences. We therefore filtered out all SNPs that were not located in coding sequences (n = 28,006 genes) in the apollo genome (Podsiadlowski et al. 2021). To avoid biases introduced by potential paralogs, only SNPs within 1:1 single copy genes with complete open reading frames and without internal stop-codons were retained (n = 24,671).
Analysis of genetic diversity was performed for the three different codon positions, separately (8,413,530 1st-, 8,420,500 2nd-, and 8,413,565 3rd-codon positions), but for the analysis of genetic differentiation and population structure we restricted the analysis to 4-fold degenerate sites (n = 3,848,496)—i.e. positions where all point mutations are synonymous. The rationale behind this approach was to minimize the risk for biases introduced by sequencing errors, erroneous SNPs calling in regions where sequences were mapped incorrectly or direct effects of selection. The filtering reduced the number of SNPs used for analysis of genetic differentiation and population structure to 451,275. Since the sex-ratio of sequenced individuals was unknown, all analyses were based on autosomal markers and we only retained SNPs that were informative in all 38 samples (390,266 SNPs in 20,620 autosomal genes).
Population genetic analyses
The levels of genetic diversity were estimated for each respective population separately, using both the estimated probability of pair-wise differences between randomly selected chromosome pairs (θπ) and Watterson’s estimate based on segregating sites and corrected for sample size (θW). We also estimated potential deviations from the expected neutral allele frequency spectrum with Tajima’s D (TD) for each respective population. All intra-population estimates were calculated in non-overlapping 100 kb windows with at least 100 informative positions using the allele frequencies calculated using VCFtools v0.16 (Danecek et al. 2011) and our in-house developed scripts in python v3.9 (https://github.com/venta380/clouded_apollo_genomics). To visualize and characterize potential population structure, we applied both a standard Principal Component Analysis (PCA) implemented using SNPrelate v1.30.0 (Zheng et al. 2012) and Admixture (Alexander et al. 2009) as implemented in the R-package LEA 3.8.0 (Frichot et al. 2021) on the VCF file generated separately for autosomal 4-fold degenerate sites using VCFtools 0.16 (Danecek et al. 2011). A dendrogram based on genetic variation in 4D-sites for all 38 individuals and estimates of kinship coefficients between individuals within and between populations were performed using SNPrelate 1.30.0 in R/Bioconductor (Zheng et al. 2012).
Genome wide genetic differentiation (FST) and absolute divergence (DXY) between each population were estimated using the allele frequencies calculated with VCFtools 0.16 (Danecek et al. 2011). These allele frequencies were also used to calculate the total number of fixed, private and shared alleles in each pair-wise population comparison.
Mitochondrial DNA variation
To be able to compare the genetic relationships between Swedish and other European clouded apollo populations, we downloaded previously published mitochondrial DNA (mtDNA) sequences from selected individuals representing different regions of continental Europe, Finland and Åland (Gratton et al. 2008) (Supplementary Table 3). The sequences represent parts of the CO1 gene (931 bp). To get the orthologous sequences from the re-sequenced Swedish individuals, we first mapped all reads from each individual to the entire mtDNA genome sequence of the apollo (Podsiadlowski et al. 2021). Mapped sequences were then used in a second similarity search limited to the 931 bp of COI previously sequenced in multiple European clouded apollo individuals (Gratton et al. 2008). Mapped (orthologous) sequences were aligned with Muscle (Edgar 2004) and manually inspected for alignment errors. As a consequence of poor mapping success for the re-sequenced Swedish individuals in general, only a subset of samples could be used for subsequent analyses (5 from Nordens Ark, 6 from Blekinge, six from Västernorrland and 10 from Roslagen). The aligned COI sequences from the retained Swedish samples and the selection of previously published individuals were used to construct a haplotype network in PopArt (Leigh and Bryant 2015).
Results
Within population levels of genetic diversity
The estimated levels of intra-population, pair-wise genetic diversity (θπ) for autosomal 4D-positions were very similar between populations, ranging from 0.0049 (Nordens Ark) to 0.0052 (Västernorrland) (Table 1). The even levels of genetic diversity were also observed for the other site classes (Table 1). As expected, first and second codon positions had lower genetic diversity than third codon positions and 4D-sites in all populations (Table 1). Analogous patterns were observed for the estimator of genetic diversity based on segregating sites (θW), but the variation between populations was slightly larger with the highest diversity in Västernorrland and lowest diversity in Roslagen for all codon positions (Table 1). For each population, we also estimated Tajima’s D to get information about potential deviations from the expected allele frequency distribution under mutation-drift balance. All populations showed negative Tajima’s D for all codon positions but there was some variation between populations; Västernorrland and Blekinge had the most negative Tajima’s D (i.e. a larger than expected proportion of low frequency alleles) while the population in Roslagen had Tajima’s D close to zero (Table 1).
Genetic differentiation between clouded apollo populations in Sweden
As a first assessment of potential genetic differences between the populations, we categorized the SNPs identified in 4D-sites as fixed, private or shared for all pair-wise comparisons of populations. We found that all comparisons involving the population from Västernorrland had the highest ratio of fixed differences (1.5–2.2%) and lowest fraction of shared variants (36.4–39.2%) (Table 2). There were no fixed differences between Blekinge and Nordens Ark and the proportion of fixed differences between the population in Roslagen on the one hand and the populations in Blekinge and Nordens Ark, respectively, was low (0.1–0.3%) (Table 2).
To get quantitative estimates of the levels of genetic differentiation and divergence between the clouded apollo populations in Sweden, we estimated the fixation index (FST) and pair-wise genetic divergence (DXY) between all six population pairs. Again, the results revealed that all comparisons including the population in Västernorrland had considerably higher FST (≈ 0.1) and DXY (> 0.005) than any other population comparison (Table 3). The lowest level of differentiation and divergence was observed between Blekinge and Nordens Ark and the population in Roslagen was only slightly differentiated from the two populations in southern Sweden (Table 3).
Visualization of population structure in the clouded apollo in Sweden
To visualize the patterns of genetic structure in the clouded apollo in Sweden, we applied both PCA and Admixture analysis. The PCA verified a considerable level of population structure overall, but no apparent differentiation between the populations among Blekinge and Nordens Ark (Fig. 2).
The Admixture analysis was run with a fixed number of populations (K) from 2 to 5 and the results supported previous observations, with a considerable structure between populations. At K = 2, the population from Västernorrland was separated from the other three populations and at K = 3, the population in Roslagen came out as a discrete cluster (Supplementary Fig. 1). Estimated minimum cross-entropy scores suggested that the data was best explained at K = 3, i.e. that the clouded apollo in Sweden can be considered as consisting of three distinct populations (Supplementary Fig. 1).
As a complementary analysis, we also generated a dendrogram based on variation in 4D-sites. The dendrogram again verified that the individuals from Västernorrland form a monophyletic group with a long internal branch to the individuals from the other populations (Fig. 4). The results showed that the individuals from Roslagen formed a monophyletic group, but that individuals from Blekinge and Nordens Ark consist of a paraphyletic group. The dendrogram also revealed potential structure within the population in Roslagen, with two distinct clusters (Fig. 4). A deeper investigation of the Roslagen individuals revealed that the two clusters represent individuals from two different sampling locations in the region: Lötaholmen and Brudskäret (including the individual that originated from Roslagen but was kept at Nordens Ark, NAU37). To get a better understanding of the genetic differences between individuals from the two sampling sites in Roslagen, we performed both a PCA and an Admixture analysis based on individuals from Roslagen only. Both analyses supported intra-population structure in the Roslagen population with two distinct groups representing individuals from the two sampling sites, respectively (Supplementary Fig. 2). To get information about the levels of genetic variation within and between each respective sub-population in Roslagen, we also estimated genetic diversity, Tajima’s D and proportions of fixed, private and shared variants, and FST using the individuals from each sampling site, respectively. The estimated level of diversity was slightly lower in the population from Lötaholmen (θπ = 0.0046) than in the population from Brudskäret (θπ = 0.0053)—both estimates were however in the range observed for the other populations (Supplementary Table 4). Tajima’s D was positive for both sub-populations, but slightly higher for the population on Brudskäret (TD = 0.12) than for the population on Lötaholmen (TD = 0.056) (Supplementary Table 4). Most of the genetic variation was shared between sub-populations, but a large proportion (44%) of private alleles and some fixed differences (n = 78) were identified (Supplementary Table 5). The level of genetic differentiation between the two sampling sites in Roslagen was 0.057 ± 0.070 (Supplementary Fig. 3).
Analysis of the kinship coefficient and identity by descent (IBD)
To get information about the level of inbreeding within and between clouded apollo populations in Sweden, we estimated the kinship coefficient, both for individuals within and between populations. The data revealed that the highest kinship coefficient was observed in the population in Västernorrland, followed by Nordens Ark, Roslagen and Blekinge (Supplementary Fig. 4). As expected based on shared recent history, the inter-population kinship coefficients were highest for the Blekinge and Nordens Ark comparison. The lowest inter-population kinship coefficients were observed in comparisons including the individuals from Västernorrland and comparisons between Roslagen and the populations in southern Sweden showed intermediate values (Supplementary Fig. 4).
The analysis also again unveiled that alleles in individuals from Västernorrland have a lower level of IBD with alleles in individuals from the other populations, that the individuals from Blekinge and Nordens Ark can be treated as a single population, and that individuals from Roslagen are more differentiated from individuals in Västernorrland than from individuals in the population in southern Sweden (Supplementary Fig. 5).
Analysis of mitochondrial DNA
To get deeper insight into the genetic relationships between the individuals sampled in Sweden and other European populations, we used previously published sequence information from parts of the CO1-gene (Cytochrome C Oxidase Subunit 1) in a selected set of individuals representing populations within the distribution range in Europe (Gratton et al. 2008). We extracted the orthologous sequences from the Swedish individuals where the sequence could be identified with certainty. This was not possible for all Swedish samples and the analysis is therefore limited to a subset of our samples (Fig 3).
The results from the analysis of the mtDNA showed that individuals from Blekinge, Nordens Ark and Roslagen clustered with individuals collected in southern and central Europe and in the Åland archipelago in the Baltic sea, while individuals from Västernorrland clustered with samples from Finland and Lithuania (Fig. 4). Those two groups, previously named haplotype groups EN and W, respectively (Gratton et al. 2008), are predominantly characterized by two fixed differences in CO1. The analysis also supported the observation in the dendrogram analysis of 4D-sites, that the samples from Roslagen are subdivided into two clusters defined by a single mutation that probably occurred in the population at Brudskäret but that has not yet reached fixation in that population (4 of 5 of the individuals from Brudskäret that could be included in the analysis carry that mutation, Fig. 4).
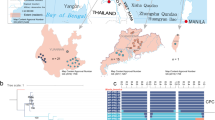
A haplotype network based on 931 bp of the mitochondrial CO1-gene. A subset of individuals from a previous study (Gratton et al. 2008) was used to get information about genetic relationships between individuals in different populations in Sweden and other populations in the distribution range in Europe. Roslagen L and Roslagen B correspond to the two sampling sites Lötaholmen and Brudskäret in the region, respectively.
Discussion
Genetic diversity within populations
All populations harbored approximately the same level of genetic diversity with median levels of θπ ≈ 0.5% for autosomal 4D-positions. This level of diversity is comparatively low for butterflies where θπ for 4D-positions has been shown to range between 0.5 and 4.1% (Mackintosh et al. 2019), but in line with observations in for example cabbage white (Pieris brassicae) and wood whites (Leptidea sp.) (Mackintosh et al. 2019), and not exceptionally low compared to many other organism groups (Romiguier et al. 2014). As expected based on the variation in intensity of natural selection between sites, the diversity was lower at 1st and 2nd codon positions. The estimates of genetic diversity based on numbers of segregating sites (θW) was marginally higher in all populations and also unveiled some variation in diversity between populations; highest in Västernorrland, followed by Blekinge/Nordens Ark and Roslagen. This indicates that the proportion of rare alleles is lower in the population in Roslagen, which is also supported by the higher Tajima’s D in the Roslagen population. The observed inter-population differences in θW (and, consequently Tajima’s D) indicates that the different populations have slightly different demographic histories. The overall negative Tajima’s D estimates are likely a consequence of the allele frequency spectrums still being affected by the rather recent re-colonization of Scandinavia after the last glacial maximum which led to founder effects—i.e. colonization of a few individuals and subsequent regional population expansions. These estimates also indicate that the rather recent population declines have not yet had a sufficient effect on rare variants to skew the allele frequency spectrum towards intermediate frequency alleles, but that the population in Roslagen might have had a more severe decline than the other populations, especially the population in Västernorrland. However, the observed sub-structure within the population in Roslagen means that different alleles can segregate at different frequencies in the two sub-populations, leading to a relative excess of intermediate frequency alleles when the populations are analyzed as a single unit. The follow-up analyses showed that both sub-populations had a marginal excess of intermediate frequency alleles which indicates an overall population decline in Roslagen, but it should be noted that the analysis was based on a very small sample set (n = 5 individuals from each sub-population, respectively) which might lead to larger uncertainties in allele frequency estimates.
Genetic structure in general
The quantitative assessments of genetic differences—both differentiation and divergence—between the clouded apollo populations in Sweden revealed a clear pattern of structure where the population in Västernorrland was significantly differentiated from the other populations. Both the proportion of fixed differences and the estimates of FST and DXY were highest in the comparisons including the population from Västernorrland. The populations in Blekinge (and Nordens Ark) and Roslagen had a larger proportion of shared alleles and significantly lower FST and DXY. These results hence point towards a historical barrier to genetic exchange between the populations in southern Sweden (Blekinge + Nordens Ark, Roslagen) and the population in central Sweden (Västernorrland). The population in Roslagen was also significantly differentiated from the other populations in Sweden, but less from the populations in Blekinge than from the population in Västernorrland. The population in Blekinge was not differentiated from the captive population at Nordens Ark, which is in line with expectations as the captive population was founded by individuals captured in Blekinge. In essence, the results from the estimated population genetic summary statistics are mirrored in both the PCA, the Admixture and the mtDNA analysis. It is noteworthy that the historical classification of the clouded apollo in Sweden into subspecies P. m. argiope and P. m. romani (Liljeblad 2021) has no support in the genetic analysis, since the individuals in the Roslagen population are obviously more closely related with individuals from southern Sweden than with individuals from Västernorrland.
The analysis of mtDNA also showed that the Roslagen and Blekinge populations belong to the same haplogroup, which is diagnostic for clouded apollo butterflies in central and western Europe and the Åland archipelago in the Baltic sea (’W’, Gratton et al. 2008). However, the analysis also revealed that 4 out of 5 individuals from the locality Brudskäret had a unique mtDNA haplotype characterized by a single point mutation. Based on previous data, we know that the most common haplotype in haplogroup W differs from the most common haplotype in group EN—the latter is diagnostic for clouded apollo individuals from northeastern Europe (Gratton et al. 2008)—by only two substitutions. We found that all individuals from Västernorrland harbored haplotype EN which is found in for example Finland and the Baltic states. This observation leads to the obvious conclusion that Scandinavia has been re-colonized from two different directions after the latest glacial period. The individuals that now exist in Blekinge (including Nordens Ark), Roslagen and the Åland archipelago likely have a common ancestry with individuals that colonized Sweden from the south (i.e. via Denmark), while individuals in Västernorrland are related to the ancestors that colonized Scandinavia from the southeast. This hypothesis has been proposed before, based on differences in mtDNA sequences between individuals from Åland and mainland Finland, but that study did not specifically include individuals sampled in Sweden (Gratton et al. 2008). The novel data generated here therefore gives additional information about the re-colonization of clouded apollos in Scandinavia after the latest ice age and provides novel information about the underlying demographic histories shaping genetic structure between clouded apollo populations within Sweden in particular.
Substructure in the Roslagen population
All analyses of population structure (PCA, Admixture, dendrogram based on autosomal 4D-sites and mtDNA) showed some differentiation between individuals sampled at Lötaholmen (Rådmansö, Stockholms län) and near Brudskäret (Östhammar, Uppsala län), respectively. Historically, the clouded apollo probably had a wider distribution in the region, in a mosaic of sub-populations (a metapopulation structure) along the coast of the Baltic sea, while the current distribution pattern is patchy with small sub-populations separated by considerable geographic distances (Hedin 2009; Hedin and Björklund 2008; Löf and Björklund 2019). The observed genetic differentiation between the sub-populations at Lötaholmen and Brudskäret indicate that there is limited gene-flow between these populations and that genetic drift has led to detectable differences in allele frequencies. The genetic diversity within the sub-populations in Roslagen was similar to the other populations in Sweden, but slightly lower in the Lötaholmen than in the Brudskäret population. This can indicate that the population on Lötaholmen has been isolated from other populations in the region and/or had a lower effective population size for a longer period of time. Tajima’s D estimates, however, point towards a more dramatic relative population size decline for the population at Brudskäret. It is important to keep in mind that the estimates of diversity and allele frequency distributions are based on a small sample set for the within sub-population analyses and the estimates should be considered relatively uncertain.
Analysis of kinship coefficients
We estimated kinship coefficients between individuals both within and between populations. Within populations, the highest degree of kinship was observed between the individuals in the populations in Västernorrland and Nordens Ark, repectively. The individuals in Roslagen had intermediate level kinship coefficients and individuals within Blekinge had the lowest level of kinship. It is expected that the individuals in the captive population at Nordens Ark should have higher kinship coefficients as the population was founded by a limited number of wild-caught specimens. The relatively high degree of kinship between individuals in Västernorrland indicates that the population might have been isolated for a longer time than the other natural populations in Sweden. It is reasonable to assume that the recolonization of Scandinavia can be characterized as a series of founder events and it is well established that the genetic variation generally decreases with distance from the center of a species’ distribution range—the so called center-periphery-hypothesis (Eckert et al. 2008; Pironon et al. 2017; Sherpa et al. 2021). Since the population in Västernorrland was most likely established by migrants from mainland Finland, it is straightforward to assume that the population in Västernorrland has been comparatively more isolated (less gene-flow with other populations) and this can have resulted in a higher level of inbreeding, and consequently higher intra-population kinship coefficients. The comparatively low level of kinship between individuals in Blekinge probably reflects a history of more genetic exchange with adjacent populations than what has occurred in Roslagen and Västernorrland. We cannot rule out, however, that some of the differences in kinship coefficients are explained by the sampling regime in each specific region. It is for example possible that more closely related individuals were sampled just by chance in Västernorrland than in Roslagen and Blekinge. However, the number of distinct sampling localities is highest in Västernorrland, which presumably indicates that the comparatively high kinship coefficients in the population reflect demography rather than the sampling.
The analysis of inter-population kinship coefficients again showed that the population in Västernorrland is genetically distinct and that the Roslagen population is more similar to the Blekinge/Nordens Ark population than to the population in Västernorrland—i.e. the kinship coefficients were higher for the comparison of individuals from Roslagen and Blekinge than for the comparison of Roslagen and Västernorrland. This analysis hence also questions the historical classification of subspecies in Sweden (Liljeblad 2021). As expected from the history of the captive population at Nordens Ark, the highest inter-population kinship coefficients were observed in the comparison between Blekinge and Nordens Ark.
Conservation implications
The clouded apollo is a sensitive species when it concerns climate changes and annual fluctuations in weather conditions during the breeding season. The reasons are the short life-span of imagines (only 3.8 days on average according to mark- recapture experiments in the population in Blekinge), the very specific habitat preferences (Artdatabanken 2021), the limited ability for dispersal (Kuussaari et al. 2015; Väisänen and Somerma 1985) and the potentially skewed sex ratio in imagines (Vlasanek et al. 2009). The rapid population declines across the distribution range, especially pronounced in some regions like Blekinge, can lead to regional extinction risks and immediate actions are needed to support local populations. Our analyses indicate that the best conservation plan in Sweden would be to take the significant differentiation between populations into account. The population in Västernorrland for example, should be treated as a distinct conservation unit. In the analysis of the mtDNA, we also observed that some of the individuals in Västernorrland—despite belonging to the same haplogroup—have unique mutations that separate them from the individuals in Finland and Lithuania. This indicates that the population in Västernorrland has limited gene-flow with mainland Finland as well. However, our results suggest translocations might be done between Roslagen and Blekinge. Such translocations can sometimes be successful despite the dispersal rate being very slow in clouded apollos. In southern Finland, for example, a long-term monitored translocation study found that suitable habitats within 10–200 meters from a release site were not colonized until 7 years after the translocation event and habitats within a 2 km range from the release site were not colonized until 13 years after the release (Kuussaari et al. 2015). However, the action was successful in the sense that both the total number of individuals and the number of sub-populations increased significantly after the translocation (Kuussaari et al. 2015). Thus, any conservation actions for the clouded apollos in Sweden should be carried out with appropriate time scales in mind. In each potential case, a translocation effort should be preceded by an assessment of the expected increase in genetic diversity and decreased risk for inbreeding on the one hand, and the risk for outbreeding depression on the other (Frankham et al. 2011). Finally, potential support actions involving translocation or rearing in captivity and releasing specimens at specific sites should crucially be combined with restoration of suitable habitat to ensure that the populations become established and interconnected in a metapopulation structure with potential for continuous gene-flow between sites. Our results indicate that translocations and captive rearing, mindful of data-supported conservation units, combined with habitat restoration, could be a potential strategy for conservation of the clouded apollo butterfly in Sweden.
References
Alexander DH, Novembre J, Lange K (2009) Fast model-based estimation of ancestry in unrelated individuals. Genome Research 19:1655–1664
Andrews, S. (2021). FastQC. Retrieved from https://www.bioinformatics.babraham.ac.uk/projects/fastqc/
Artdatabanken, S. (2020). Rödlistade arter i Sverige 2020. SLU Artdatabanken, Uppsala.
Artdatabanken, S. (2021). Artfakta: mnemosynefjäril (Parnassius mnemosyne). Retrieved from https://artfakta.se/naturvard/taxon/parnassius-mnemosyne-101510
Bolotov IN, Gofarov MY, Rykov AM, Frolov AA, Kogut YI (2013) Northern boundary of the range of the clouded apollo butterfly Parnassius mnemosyne (L.) (Papilionidae): climate influence or degradation of larval host plants? Nota Lepidoptera 36:19–33
Danecek P, Auton A, Abecasis G, Albers CA, Banks E, DePristo MA, Genomes Project Analysis (2011) The variant call format and VCFtools. Bioinformatics 27:2156–2158. https://doi.org/10.1093/bioinformatics/btr330
Eckert CG, Samis KE, Lougheed SC (2008) Genetic variation across species’ geographical ranges: the central-marginal hypothesis and beyond. Mol Ecol 17:1170–1188. https://doi.org/10.1111/j.1365-294X.2007.03659.x
Edgar RC (2004) MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 19:113
Eliasson CU, Ryrholm N, Holmer M, Jilg K, Gärdenfors U (2005) Nationalnyckeln till Sveriges flora och fauna. Fjärilar: Dagfjärilar. Hesperiidae - Nymphalidae. Laholm: Artdatabanken, SLU, Uppsala. Ruter
Eliasson K (2021) Mnemosyne i Sverige: en kvantifiering av artens etablering från1900-talet till dag. Retrieved from http://www.diva-portal.org/smash/record.jsf?pid=diva2%3A1597754&dswid=4830
Frankham R, Ballou JD, Briscoe DA (2012) Introduction to conservation genetics. Cambridge University Press, Cambridge
Frankham R, Ballou JD, Eldridge MD, Lacy RC, Ralls K, Dudash MR, Fenster CB (2011) Predicting the probability of outbreeding depression. Conserv Biol 25:465–475. https://doi.org/10.1111/j.1523-1739.2011.01662.x
Franzén, M., & Imby, L. (2008). Åtgärdsprogram för mnemosynefjäril 2008–2012. NVV rapport 5829. Retrieved from Bromma: https://www.naturvardsverket.se/978-91-620-5829-6
Frichot, E., Francois, O., & Gain, C. (2021). LEA: an R package for landscape and ecological association studies. Retrieved from https://bioconductor.org/packages/LEA/
Gratton P, Konopinski MK, Sbordoni V (2008) Pleistocene evolutionary history of the clouded apollo (Parnassius mnemosyne): genetic signatures of climate cycles and a “time-dependent” mitochondrial substitution rate. Mol Ecol 17:4248–4262. https://doi.org/10.1111/j.1365-294X.2008.03901.x
Hedin, E. (2009). Inventering av mnemosynefjäril (Parnassius mnemosyne) i Norrtälje kommun 2009. Retrieved from Norrtälje: https://www.norrtaljenaturcentrum.se/wp-content/uploads/2020/04/2009_8-Inventering-av-Mnemosynefj%C3%A4ril-2009.pdf
Hedin, E., & Björklund, J.-O. (2008). Mnemosynefjärilen i Uppland: historik och nuvarande utbredning. Retrieved from Stockholm: https://docplayer.se/12826603-Mnemosynefjarilen-i-uppland.html
Holst, I., Gylje Blank, S., & Svensson, M. (2021) Dagfjärilar som omfattas av åtgärdsprogram för hotade arter och naturtyper. Naturvårdsverket rapport 6982. Retrieved from Bromma: https://www.naturvardsverket.se/978-91-620-6982-7
Johansson V, Knape J, Franzén M (2017) Population dynamics and future persistence of the clouded Apollo butterfly in southern Scandinavia: the importance of low intensity grazing and creation of habitat patches. Biol Conserv 206:120–131
Kuussaari M, Heikkinen RK, Heliölä J, Luoto M, Mayer M, Rytteri S, von Bagh P (2015) Successful translocation of the threatened clouded apollo butterfly (Parnassius mnemosyne) and metapopulation establishment in southern Finland. Biol Conserv 190:51–59
Leigh JW, Bryant D (2015) POPART: full-feature software for haplotype network construction. Methods Ecol Evolut 6:1110–1116
Li H (2011) A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27:2987–2993
Li H, Durbin R (2010) Fast and accurate long-read alignment with Burrows-Wheeler transformation. Bioinformatics 26:589–595
Liljeblad, J. (2021). Dyntaxa. Svensk taxonomisk databas. Version 1.2. In (Vol. Version 1.2). SLU Artdatabanken.: GBIF.org.
Löf, A., & Björklund, J.-O. (2019) Miljöövervakning av mnemosynefjäril 2018 (Parnassius mnemosyne), Norrtälje kommun, Stockholms län. Retrieved from Stockholm: https://www.lansstyrelsen.se/download/18.3db3ed8a171ac1fbfcb235eb/1591090491095/Fakta%202019-06%20Mnemosynefj%C3%A4ril_rapport_2018%20slutlig.pdf
Lunter G, Goodson M (2011) Stampy: a statistical algorithm for sensitive and fast mapping of illumina sequence reads. Gen Res 21:936–939
Mackintosh A, Laetsch DR, Hayward A, Charlesworth B, Waterfall M, Vila R, Lohse K (2019) The determinants of genetic diversity in butterflies. Nat Commun 10:3466. https://doi.org/10.1038/s41467-019-11308-4
Meglecz E, Solignac M (1998) Microsatellite loci for Parnassius mnemosyne (Lepidoptera). Hereditas 128:179–180
Pironon S, Papuga G, Villellas J, Angert AL, Garcia MB, Thompson JD (2017) Geographic variation in genetic and demographic performance: new insights from an old biogeographical paradigm. Biol Rev 92:1877–1909. https://doi.org/10.1111/brv.12313
Podsiadlowski L, Tunström K, Espeland M, Wheat CW (2021) The genome assembly and annotation of the apollo butterfly Parnassius apollo, a flagship species for conservation biology. Gen Biol Evolut 13:122. https://doi.org/10.1093/gbe/evab122
Romiguier J, Gayral P, Ballenghien M, Bernard A, Cahais V, Chenuil A, Galtier N (2014) Comparative population genomics in animals uncovers the determinants of genetic diversity. Nature 515:261–263. https://doi.org/10.1038/nature13685
Sherpa S, Kebaïli C, Rioux D, Guéguen M, Renaud J, Després L, Andersen A (2021) Population decline at distribution margins: assessing extinction risk in the last glacial relictual but still functional metapopulation of a European butterfly. Divers Distrib 00:1–20. https://doi.org/10.1111/ddi.13460
Väisänen R, Somerma P (1985) The status of Parnassius mnemosyne (Lepidoptera, Papilionidae) in Finland. Notulae Entomologicae 65:109–118
Vlasanek P, Hauck D, Konvicka M (2009) Adult sex ratio in the Parnassius mnemosyne butterfly: effects of survival, migration, and weather. Israel J Ecol Evolut 55:233–252
Zheng X, Levine D, Shen J, Gogarten S, Laurie C, Weir B (2012) A high-performance computing toolset for relatedness and principal component analysis of SNP data. Bioinformatics 28:3326–3328. https://doi.org/10.1093/bioinformatics/bts606
Acknowledgements
This work was funded by Länsstyrelsen Blekinge (to N.B and J.H) and Nycanders Foundation, Uppsala University (to N.B.). The authors acknowledge support from the National Genomics Infrastructure in Stockholm funded by Science for Life Laboratory, the Knut and Alice Wallenberg Foundation and the Swedish Research Council, and SNIC/Uppsala Multidisciplinary Center for Advanced Computational Science for assistance with massively parallel sequencing and access to the UPPMAX computational infrastructure. The authors also thank Gunilla Engström for performing the DNA-extractions and Annika Lydänge and Jimmy Helgesson for organizing the samples used for sequencing.
Funding
Open access funding provided by Uppsala University.
Author information
Authors and Affiliations
Contributions
JH and NB: conceptualized and designed the study. All data analyses were performed by VM, VT and NB. The first draft of the manuscript was written by NB and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no relevant financial or non-financial interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Talla, V., Mrazek, V., Höglund, J. et al. Whole genome re-sequencing uncovers significant population structure and low genetic diversity in the endangered clouded apollo (Parnasssius mnemosyne) in Sweden. Conserv Genet 24, 305–314 (2023). https://doi.org/10.1007/s10592-023-01502-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-023-01502-9