Abstract
Reported effects of inbreeding vary among taxa and may depend on a number of factors, including what trait is measured, temporal variability, parental effects, or life history stage. To understand the effects of inbreeding during early life history stages, we estimated the effects of individual-level heterozygosity on hatching success and first year survival in a Swedish population of sand lizards (Lacerta agilis) over a period of almost a decade, using over 4000 eggs, 400 clutches, and over 3000 juveniles. Heterozygosity had a positive effect on hatching success, in standardized laboratory conditions, but no effect on first year survival. Also, both of these measures of viability varied across the years of the study, demonstrating the importance of temporal heterogeneity in pre and post-hatching conditions. Finally, we identified both paternal and maternal identity effects on hatching success. Thus, we show that selection on heterozygosity was not consistent across developmental life stages, emphasizing the need of considering a number of ontogenic stages, as well as potential parental and environmental effects, when studying the effects of heterozygosity on viability in natural populations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Inbreeding–mating between genetically similar individuals—is an important phenomenon in evolutionary and conservation biology (Darwin 1868; Charlesworth and Charlesworth 1987), as it occurs in a wide range of animal and plant populations, and may lead to inbreeding depression with potentially severe fitness consequences (Keller et al. 1994; Olsson et al. 1996a; Crnokrak and Roff 1999; Slate et al. 2000; Amos et al. 2001; Keller and Waller 2002; Huisman et al. 2016). Consanguineous matings result in a genome-wide increase in homozygosity, relative to matings between less related individuals. Such increase in homozygosity can affect fitness negatively through two proposed mechanisms, over-dominance or partial dominance. With over-dominance, inbreeding depression occurs due to the superiority of heterozygous individuals relative to homozygous individuals at fitness-related loci. Conversely, partial dominance can cause inbreeding depression when inbred individuals become homozygous for recessive or partially recessive deleterious alleles at fitness-related loci (Charlesworth and Charlesworth 1987). In evolutionary biology, inbreeding is studied due to its association with, for example, speciation (Lynch 1991), the evolution of mating systems (Pusey 1987) or the evolution of inbreeding avoidance mechanisms (Pusey and Wolf 1996). However, in conservation biology, considering genetic factors is important for the success of conservation efforts in small populations with a reduced genetic variation (Lacy 1997; Whiteley et al. 2015).
The magnitude of inbreeding depression can vary across an organism’s ontogeny (Hemmings et al. 2012), and reported relationships between inbreeding and fitness vary in the literature depending on the developmental stages of study organisms. A number of studies have found a significant relationship between inbreeding and hatching success (Keller and Waller 2002; Tregenza and Wedell 2002; Spottiswoode and Moller 2004; Hemmings et al. 2012; but see Michaelides et al. 2016). Other researchers have found no such effects of inbreeding on direct measures of offspring survival (Kempenaers et al. 1996; Kruuk et al. 2002) or on other traits that could indirectly affect survival, such as the incidence of malformations in larval tiger salamanders (Ambystoma tigrinum tigrinum) (Williams et al. 2008). Finally, somewhat surprisingly, a number of studies have found positive effects of inbreeding on offspring survival (Richardson et al. 2004; Weiser et al. 2016; Bichet et al. 2018). Additionally, it is important to recognize sources of error in calculating effects of inbreeding, which can change across life stages. For example, its effects may be underestimated or mistaken for parental infertility if inbred individuals die early during ontogeny (Hemmings et al. 2012).
The magnitude of inbreeding depression may also vary according to the trait measured or even the year of breeding, indicating potential interactions between genetic and environmental factors (Hedrick and Kalinowski 2000; Keller and Waller 2002). Furthermore, an offspring’s phenotype can be influenced by both maternal and paternal effects (Crean and Bonduriansky 2014). Thus, it is important to study the effects of inbreeding on viability over a wide genetic and ecological context, while considering many developmental stages, environmental variables, and the role of parental effects. This type of integrative framework is crucial for understanding the importance of inbreeding as a selection agent, in particular to improve the management of small or endangered populations (Madsen et al. 1999; Frankham 2005).
In the current study, we test the effects of inbreeding on mortality at two different life history stages, during incubation and embryonic development (‘hatching success’), and during the first year of life (‘first year survival’), using microsatellite-based heterozygosity data on a Swedish population of sand lizards (Lacerta agilis). We use data which was collected over a period of a decade, involving over 4000 eggs and 400 clutches, from 252 females. Sand lizards live for up to ca. 10 years, with a mean longevity of ca. 5 to 6 years (Olsson et al. 1996a). Males mature approximately at the age of two and females at three years of age, creating a generation overlap and a potential for consanguineous matings (Olsson et al. 1996a). Males are polygynous and female receptively asynchronous, with larger males dominating the number of copulations, hence skewing the operational sex ratio and decreasing the effective population size (Olsson et al. 1996a). Furthermore, the relatively high mortality rate in the first year post-hatching (Olsson et al. 1994) implies intense selection pressure on offspring viability. Thus, the ecology and life history of the sand lizard may be associated with extensive inbreeding, making it an ideal candidate for studying the effects of consanguineous matings on fitness during early ontogeny (Olsson et al. 1996a). In addition to studying the effects of inbreeding on hatching success and first year survival, we examine the potential influence of parental effects, juvenile sex, and breeding year, to account for temporal environmental heterogeneity.
Methods
Study system
Details of field work, husbandry, and laboratory techniques are described in previous work (Olsson 1994; Olsson and Shine 1997; Olsson et al. 2011). Therefore, they are only summarized here.
Sand lizards are small ground dwellers (up to 20 g) and have one of the largest distributions of any reptile species 8000 × 5000 km (Bischoff 1984). The main distribution is in central Europe. In Sweden, the distribution is fragmented and the species considered vulnerable. The genetic variability is low, compared to the populations in central Europe, and gene flow is limited between populations (Gullberg et al. 1999). The negative effects of matings between close kin is particularly pronounced in the Asketunnan population (our study population), shown by the incidence of malformations in inbred offspring (Olsson et al. 1996a). Malformed offspring have essentially no chance to survive in the wild (Olsson et al. 1996a), emphasizing the scale of the effect that continuing inbreeding may have on population dynamics at this site. Asketunnan is located on the Swedish West Coast (approximately N570 22′ E110 58″). In this population, females lay a single annual clutch of 5 to 15 eggs, depending on female body size (Olsson 1993). Approximately 1 week before egg laying, gravid females were captured by noose (with egg contours on their body sides indicating near oviposition), and brought to facilities at the University of Gothenburg, Sweden. They were marked by toe-clipping for identification (Olsson 1994), and kept individually in cages (40 × 50 × 60 cm) containing a flat rock, placed on wet sand for egg laying, and a 40-W spotlight at one end to allow thermoregulation. Eggs were collected within hours of egg laying and incubated at 25 °C, a temperature that optimizes hatching success and minimizes developmental asymmetries (Zakharov 1989). All clutches were incubated individually in separate boxes in as constant conditions as can be achieved, all in the same incubator. Most eggs hatched after approximately 40 days, and the juveniles were scored for hatching success (1 = hatched, 0 = died in egg). They were also measured snout to vent to the nearest 0.1 mm and weighed to the nearest 0.001 g, and marked by toe-clipping. Malformations, such as twisted spines, missing toes, and cranial deviations from symmetry, were screened by eye (scored as malformed or not malformed (Olsson et al. 1996a)), before release at random sites at the Asketunnan study site. Over a period of 10 years (between 1998 and 2007), 3627 eggs hatched, from a total of 4079 fertile eggs, yielding an overall hatching success of 89%. The eggs that failed to complete development were distinguished from non-fertilized eggs (93 eggs) by dissection (that did/did not identify blood vessels/embryo remnants), leaving 4079 fertile eggs available for analyses (Table 1). The released juveniles were recaptured the following year, or not (assumed dead). They were assessed during spring following their first winter, by systematic searches over the study site on all days that permitted lizard activity (May through mid-June) (Schwartz et al. 2011). Recaptured juveniles were given a score of 1, and those that were not recaptured a score of 0. The process of capture for egg laying and release, of females and juveniles, was repeated over a 10 year period, between 1998 and 2007 (Olsson et al. 2011). During this period, 252 different females were captured, and laid a total of 400 clutches. The number of females every year varied between 29 and 65, with some females being recaptured multiple times (Supplementary Fig. 1). In total, of 3627 laboratory-hatched juveniles, 3305 survived to be released. However, only 339 juveniles were recaptured, across all years, for an overall recapture rate of survivors of 10.2% (Table 1).
Hatchlings not recaptured were considered dead with a high degree of confidence, since juveniles disperse less than 135 m (Olsson et al. 1996a; Ryberg et al. 2004) and we monitored a 600 m perimeter around the study site. In previous work, no juveniles were found to leave Asketunnan through this buffer zone (Olsson et al. 1996a). A contributor to this result is the small home range of sand lizards (on average, ca. 10 × 10 m in females, and 30 × 30 m in males (Olsson et al. 1996a)). Furthermore, this relatively small population (≤ 500 adults) is concentrated along the rocky shoreline of a 400 m peninsula, thus limiting migration into and out of the area (Gullberg et al. 1998, 1999; Madsen et al. 2000).
Genotyping, parentage assignment and heterozygosity measurements
DNA was extracted from blood and tissue samples of 4534 adults, juveniles, and embryos that died during incubation. Parentage was assigned, using up to 21 microsatellite loci (with a minimum of 17 loci, see (Olsson et al. 2011) for a detailed description of methods). From 452 eggs that failed to hatch, we were only able to extract DNA for genotyping from 204 eggs. From the 3627 eggs that hatched, a total of 2601 juveniles were successfully assigned a father (Olsson et al. 2011) (Table 1). The genotyped microsatellites were then used to calculate individual heterozygosity, as a proxy for inbreeding, in the R package Rhh (Alho et al. 2010). Individual heterozygosity was estimated for 2983 observations, including juveniles and unhatched eggs. This was done using standardized heterozygosity (SH) (Coltman et al. 1999), with an average of 19.6 loci used per observation. SH represents the proportion of heterozygous typed loci divided by the mean heterozygosity of the typed loci. In addition, heterozygosity–heterozygosity correlations (HHC) were calculated in Rhh, to estimate the inbreeding signal at the 21 loci as a proxy for genome-wide inbreeding (Balloux et al. 2004). HHC was estimated by randomly dividing the genotyped loci into two groups, and calculating SH for each group 1000 times. The results from the two groups were used in correlations with each other, thus yielding a mean correlation, or heterozygosity–heterozygosity correlation HHC (Balloux et al. 2004). Evidence of inbreeding was further assessed by calculating the parameter g2 (David et al. 2007). g2 estimates identity disequilibrium (ID) in the form of correlations in heterozygosity or homozygosity across the genome (Weir and Cockerham 1973), which is fundamental for heterozygosity fitness correlations (HFC) (Szulkin et al. 2010). The value of g2 should be positive if there is variance in inbreeding, and thus potential for HFCs to arise in a population (Szulkin et al. 2010). Departure from 0 for g2 was assessed in RMES (Robust Multilocus Estimation of Selfing) based on 1000 iterations (David et al. 2007).
Statistical analyses
Generalized linear mixed models (GLIMMIX) with binomial error distributions, logit link functions, and crossed random effects (parental IDs appearing in a crossed design) were fitted in SAS (Statistical Analysis System) 9.4 to analyse the following effects:
Effects of parental identity on hatching success and first year survival
Two random intercept models were used, with maternal and paternal identities as categorical variables. The response variable was hatching success in the first model, and first year survival in the second.
Effect of individual-level heterozygosity on hatching success and first year survival, while accounting for year of study and juvenile sex
Two sets of models were fitted, one for hatching success and one for first year survival. First, year of study and standardized heterozygosity (SH) were sequentially added as explanatory variables (fixed effects) to the random intercept model with hatching success as the response variable. Year was added first, followed by SH. This sequential approach allowed us to test the effect of heterozygosity on hatching success, while accounting for year effects. Model fit was compared to that of the previous model each time a fixed effect was added, using a Log Likelihood ratio test (LL ratio test). The LL ratio represents the ratio of the − 2 Log Likelihoods of two models. Model fit difference was further assessed by comparing the Akaike information criterion (AIC) of the two models. The same protocol was used (LL ratio test, information criterion comparison, and sequential inclusion of fixed effects) in the second set of models, to assess the effect of SH on first year survival, starting with the random intercept model with first year survival as response variable. This time, juvenile sex was also added as a fixed effect, after year, to account for sex-specific survivorship. In this set of models, year was added to assess the effect of post-hatching conditions on first-year survival. However, it is important to mention that any year effects on hatching success would be most likely mediated through parental effects, because the laboratory and incubation conditions were identical among years. Also, juvenile sex was only added to the first year survival models, because the sex of the unhatched embryos was unknown. Finally, in a separate random intercept model, year and hatching date (as a proxy for release date) were included as random effects, with date nested under year, and first year survival as response variable. This was done to evaluate the influence of temporal environmental heterogeneity on first year survival at a finer scale. The juveniles were consistently released shortly after hatching. Therefore, hatching dates can be used as a reliable proxy for release dates.
Results
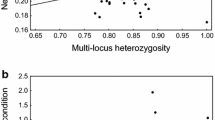
The mean SH value, for the whole data set, was 0.99, with a coefficient of variation of 22.22% (n = 3043; SD = 0.22) (Supplementary Fig. 2). The hatched eggs had a mean SH value of 0.99 (n = 2779; SD = 0.21), whereas the unhatched ones had a mean SH of 0.98 (n = 204; SD = 0.25) (Fig. 1). The mean SH values of the hatched and unhatched eggs did not differ significantly (mean ± s.e.m. = 0.98 ± 0.018 (unhatched) and 0.99 ± 0.004 (hatched); t = − 0.18; d.f. = 225.13; P = 0.86). The recaptured juveniles had a mean SH of 0.95 (n = 100; SD = 0.21), whereas those that were not recaptured had a mean SH of 0.99 (n = 2703; SD = 0.21) (Fig. 1). Again, these two mean values did not differ significantly (mean ± s.e.m. = 0.99 ± 0.004 (not recaptured) and 0.95 ± 0.021 (recaptured); t = 1.52; d.f. = 106.6; P = 0.13).
The boxes represent the range of standardized individual-level heterozygosity (SH) values for the observations used in the analyses. The ontogenic stage hatching refers to the hatched and unhatched eggs, status 1 and 0 respectively. Capture refers to the capture status of juveniles, 1 for recaptured and 0 for not recaptured. The symbols inside the boxes represent the mean values for each status, and the horizontal bars the medians. The means and medians are relatively close to each other for the two types of status (1 and 0), for both ontogenic stages (category)
The heterozygosity–heterozygosity correlation was positive and significant (mean = 0.1414; 95% confidence interval: 0.05559–0.19875), indicating the loci had sufficient information to provide estimates of inbreeding. The signature of inbreeding in the population was further confirmed by a positive, and statistically significant, value of g2 (n = 3786; s (g2) = 0.0025; P = 0.005).
In the following sections, we present the results of the generalized linear mixed models used to analyze the effect of individual-level heterozygosity on hatching success and first year survival, and the importance of parental identity, juvenile sex, and environmental effects during ontogeny.
Effects of parental identity on hatching success and first year survival
We found significant effects of paternal (n = 2557; \(\chi_{1}^{2}\) = 21.31; P < 0.0001; intraclass correlation coefficient (ICC) = 0.25; estimate = 20.137; 95% confidence limits 3.327–119.01) and maternal identity (n = 2557; \(\chi_{1}^{2}\) = 119.64; P < 0.0001; ICC = 0.70; estimate = 55.5626; 95% confidence limits 25.566–149.89) on hatching success. The estimates represent the variance in hatching success that is explained by parental effects. ICC is the proportion of the total variance in hatching success that is due to parental effects (correlations between juveniles belonging to the same parent). Conversely, first year survival was not significantly predicted by maternal identity (n = 2491; \(\chi_{1}^{2}\) = 1.09; P = 0.1484; ICC = 0.024; estimate = 0.084; 95% confidence limits 0-0.318), or by paternal identity (n = 2491; \(\chi_{1}^{2}\) = 2.69; P = 0.0506; ICC = 0.039; estimate = 0.138; 95% confidence limits 0-0.394), although the P value for the latter was close to statistical significance. Note, however, the confidence intervals that include 0.
Effect of individual-level heterozygosity on hatching success and first year survival, while controlling for year and sex effects
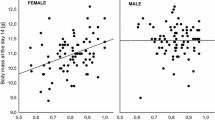
Individual level heterozygosity (SH) had a significant effect on hatching success (n = 2343; LL ratio test: \(\chi_{1}^{2}\) = 5.92, P = 0.015; odds ratio = 1.29), that did not vary across years (n = 2343; LL ratio test: \(\chi_{9}^{2}\) = 10.91, P = 0.2819) (Fig. 2, Table 2). Conversely, first year survival was not significantly predicted by SH (n = 2266; LL ratio test: \(\chi_{1}^{2}\) = 1.72, P = 0.1896; odds ratio = 0.92), that did not interact with year (n = 2266; LL ratio test: \(\chi_{9}^{2}\) = 8.12, P = 0.5221) (Fig. 3, Table 2). The probability of hatching, and of surviving the first year, varied significantly between years (n = 2557; LL ratio test: \(\chi_{9}^{2}\) = 70.74, P < 0.0001) (n = 2491; LL ratio test: \(\chi_{9}^{2}\) = 28.78, P = 0.0007) (Fig. 4, Table 2). However, at a finer scale, hatching date had no effect on first year survival (n = 3299; \(\chi_{1}^{2}\) = 1.07; P = 0.1507).Finally, juvenile sex did not significantly predict first year survival (n = 2477; LL ratio test: \(\chi_{2}^{2}\) = 2.26, P = 0.323; odds ratio = 0.81), or interact with SH (n = 2266; LL ratio test: \(\chi_{2}^{2}\) = 3.33, P = 0.1891) (Table 2).
Relationship between juvenile individual-level heterozygosity (JSH) and predicted first year survival probability, from a generalized linear mixed model with a binomial error distribution. An increase in individual-level heterozygosity is associated with a decrease in first year survival probability
The graph shows the hatching and first year survival probabilities for every year of the study period (lines). The bars represent the number of fertile eggs, and of released juveniles, for each year. Only oviposition and hatching data is available for the year 2007, since it was the last year of the study period
Discussion
Our aim was to test the effects of inbreeding at two key life history stages in the sand lizard population of Asketunnan, Sweden. We found a significant, positive, relationship between standardized individual-level heterozygosity (SH) and hatching success, confirmed by both the likelihood ratio test and the reduction in the Akaike information criterion (AIC) (Wang et al. 2011). This positive effect indicates that less inbred embryos experienced a higher hatching probability relative to more inbred embryos in our study population, a classic example of inbreeding depression. However, no such effect was found on first year survival.
The importance of inbreeding in driving the positive relationship between SH and hatching success, or significant heterozygosity fitness correlation (HFC), is supported by both a positive heterozygosity–heterozygosity correlation (HHC) and a positive g2 value. The mean SH value was 0.99, with a minimum of 0 (an individual homozygous for all typed loci) and a maximum of 1.65. Thus, the data set included individuals that varied substantially in their degree of inbreeding (coefficient of variation of 22.22%). This relatively large variance in inbreeding is likely to have underpinned the significant effect of SH on hatching success that we observed. Indeed, inbreeding-based theory predicts significant HFCs to arise under particular population structures, with a large variance in inbreeding coefficient combined with high statistical power (Szulkin et al. 2010; but see Chapman et al. 2009). However, according to meta-analysis, the effect sizes of HFCs are generally low, with individual multilocus heterozygosity explaining on average less than 1% of the variance in fitness in microsatellite studies (Chapman et al. 2009). Most (90%) of the incubated eggs hatched, and hence the overall hatching probability was high. The high mean hatching probability is likely due to the fact that the eggs were incubated in optimal laboratory conditions, in the absence of environmental extremes and predation. Furthermore, a relatively large number of dead embryos had low quality DNA, excluding them from the analyses. This left less than half of the dead embryos to be genotyped, thus further decreasing our potential to detect HFCs. Still, we found a substantial effect of heterozygosity on hatching probability (odds ratio 1.29), after controlling for parental effects, and thus observation dependency caused by the fact that a considerable number of juveniles shared the same parents. The odds ratio of 1.29 was estimated based on a 10% increase from the mean SH value in the population. In other words, a 10% increase in heterozygosity is associated with an increase in hatching odds by a factor of 1.29 (approximately 30%). However, as stated above, individual-level heterozygosity was estimated using SH, a standardised metric. Therefore, the odds ratios reported here cannot be used for comparisons among studies, as they relate to heterozygosity values which are relative to the average heterozygosity in the study population.
The deleterious effects of inbreeding on fitness are generally exacerbated under natural, and thus harsher, conditions (Keller and Waller 2002). Therefore, we expected inbreeding depression to be even more pronounced in the Asketunnan population, compared to the laboratory incubated eggs. It is, indeed, preferable to compare the relationship between individual heterozygosity and fitness across ontogeny using only natural settings. Our results, however, represent a useful indication of the effects of inbreeding on hatching success, emphasizing the relevance of this type of study in recovery plans for small populations with reduced genetic variation.
Interestingly, there was a non-significant effect of SH on first year survival, with an odds ratio of 0.92 (negative trend), a finding consistent with a number of studies in which diminishing effects of inbreeding on fitness across life history stages were reported (van Noordwijk and Scharloo 1981; Mainguy et al. 2009; Hemmings et al. 2012). Furthermore, no interaction between SH and juvenile sex, on first year survival, was observed. The lack of effect of heterozygosity on first year survival indicates a decrease in selection against inbreeding at this life stage. A strong episode of selection against inbreeding during early ontogeny can remove the most inbred individuals, leaving a subset of “higher quality” juveniles from the original cohort (Keller et al. 1994; Hemmings et al. 2012). However, we did not observe such a difference in individual genetic variation, the mean SH value of the unhatched eggs was not significantly different from the mean value of the hatched eggs. Nevertheless, one cannot exclude the fact that these values might differ in natural, and thus harsher, settings with environmental extremes and predation. The lack of difference in individual genetic variation that we observed demonstrate the importance of considering significant contextual variables, such as parental IDs, in the models, as they may conceal the effects of the fixed effect of interest (in this case individual-level heterozygosity) on survivorship. Most importantly, it indicates that the apparent lack of selection on heterozygosity during the first year post-hatching is not due to the presence of relatively non-inbred individuals at this ontogenic stage, relative to embryonic development. Instead, it is most likely due to the fact that the inbreeding load affected traits that are expressed during embryonic developments, and are either not expressed or expressed without detrimental effects during later ontogeny. This distinction is important, as it highlights the role of life-stage specific gene expression and regulation as determinants of the impact of inbreeding on viability, instead of solely the average level of heterozygosity in the population.
A number of ecological variables not included in the models, such as predation, may have played an important role, by levelling differences in survival probability between juveniles with different SH values. Therefore, one should generally be cautious in drawing definite conclusions with regards to the absence of effect of heterozygosity on first year survival. However, our large sample size (thousands of juveniles), in combination with the relatively narrow confidence interval for the odds ratio (0.824–1.039, including 1), allow us to infer with a certain degree of confidence that inbreeding did not have an impact on first year survival in the population. Importantly, the lack of effect of heterozygosity on first year survival stresses the importance of considering several life stages during conservation efforts, as the negative effect of inbreeding would not have been detected if we had not sampled the embryonic stage.
We found a relatively strong effect of paternal identity on hatching success (intraclass correlation (ICC) = 0.25), indicating that approximately one quarter of the total variance in hatching success is due to variation between paternal identities. Any direct benefits of males are difficult to envisage, since in sand lizards, they do not provide any resources but semen and genes to the female (Olsson and Madsen 2001a). The lack of transferred resources by males make genetic benefits, in the form of good genes (Moller and Alatalo 1999) and/or compatibility effects (Zeh and Zeh 1996, 1997), strong candidates to explain the observed male effects. Thus, the negative effect of homozygosity on hatching success that we observed, in combination with the significant paternal effect, suggests the potential existence of disassortative mating patterns with regards to genetic relatedness, if certain males are relatively more prone to choosing less related females.
Disassortative mating has been shown in previous work in this study population, with females preferring to associate with the odour of males more distantly related at the MHC (major histocompatibility complex) class I loci (Olsson et al. 2003). Also, it has been shown that associations in the wild are non-random with respect to MHC genotypes (Olsson et al. 2003). Work by other researchers have shown that MHC recognition may facilitate discrimination of kin from non-kin (Potts et al. 1994; but see Sherborne et al. 2007 and Sepil et al. 2015), and thus avoid the deleterious effects of consanguineous matings at MHC loci (Brown and Eklund 1994). So, if MHC class 1 genotypes act as a cue for kinship, the observed non-random associations in the wild (Olsson et al. 2003), and the paternal effects on hatching success that we observed, may result from inbreeding avoidance. It is also known that large and dominant males tend to associate with less related females at MHC loci (Olsson et al. 2003), suggesting that when a male has the opportunity to choose because of his higher social status, he may prefer more MHC-unrelated partners. Furthermore, we found no paternal effects on first year survival, a result which parallels the non-significant effects of SH at this life stage. This further supports the idea that certain males are capable of choosing less related females, and therefore experience an increase in fitness through an increased hatching potential, but not through juvenile survival. However, this idea is hypothetical, and it remains to be shown if individuals actively avoid inbreeding and/or seek MHC dissimilarity, a mating decision that potentially underlies the paternal effects observed in the present study. This can be done by contrasting the actual mating patterns, with regards to both relatedness and MHC similarity, against expectations under random mating, thus revealing the ultimate mechanisms that underlie the disassortative mating patterns observed in the population.
Maternal identity had an effect on hatching success (ICC = 0.70), which may be attributed to a variety of maternal effects, apart from genetic ones, such as differential investment into eggs with nutrients, antioxidants, or antibodies (Mousseau and Fox 1998). However, this relatively large effect disappeared after hatching. Similarly to the paternal effects mentioned above, the variance between mothers with regards to hatching probability may be, at least partly, explained by inbreeding avoidance. Unlike males in this population, females do not appear to exert pre-copulatory mate choice, mating with all courting males regardless of their degree of kinship (Olsson and Madssen 1995; Olsson et al. 1996a). Mate rejection by female sand lizards only appears to occur when females are not ready to mate (Olsson and Madssen 1995). Thus, if they exist, mechanisms of inbreeding avoidance in females are most likely restricted to the post-copulatory level.
The presence of post-copulatory mechanisms of mate discrimination in females is supported by the fact that offspring viability within clutches is correlated with the number of males with which the female mated, suggesting a bias of fertilization based on genetic compatibility (Olsson and Madsen 2001a). This was confirmed in another study in which males less related to a female, calculated as the number bands shared in DNA fingerprinting, sire more offspring per copulation, than do more related ones (Olsson et al. 1996b). So, this mechanism of mate discrimination, based on genetic similarity, may underlie the maternal effects that we observed. However, given the relatively large number of other sources of maternal effects, the relative importance of such mechanism remains to be shown.
In addition to the adult mechanisms of mate discrimination discussed above, proximate mechanisms of inbreeding avoidance may take place earlier in life. Male juveniles disperse over considerably longer distances than females (Olsson et al. 1996a). Therefore, inbreeding may also be avoided through sex-specific dispersal, although it cannot be rejected as being a pleiotropic effect for adult male mobility and mate acquisition (Olsson et al. 1996a, b). Regardless of the proximate mechanisms of kin discrimination that may be acting in the population, inbreeding avoidance should be considered as an ultimate factor affecting survivorship in small populations with low levels of genetic variation.
Year had a significant effect on first year survival. However, at a finer scale, hatching, and hence release, date did not predict first year survival. Considering the high mortality rates during a sand lizard’s first year of life (Olsson et al. 1994), the effect of year on first year survival suggests that certain years may have been more benign than others (Fig. 4) (Keller and Waller 2002), which agrees with Olsson and Madsen’s observations on a much smaller data set (Olsson and Madsen 2001b). More importantly, this illustrates the importance of long-term studies on inbreeding. Similarly, hatching success varied according to the year of study. Given the standardized laboratory setting in which the eggs were incubated, this effect was most likely mediated through an annual variation in maternal condition, again emphasizing the importance of maternal effects with regards to hatching success. Finally, year did not interact with SH on hatching success or on first year survival, showing that the influence of heterozygosity on viability remained constant throughout the study period.
In conclusion, in this long term study, we found a significant and negative effect of inbreeding on hatching success in the Asketunnan sand lizard population. However, we found no such effect on first year survival. This inconsistency in the strength of selection on individual heterozygosity highlights the complexity of the dynamics of inbreeding in natural populations, and the need to consider key ontogenic stages during conservation efforts, through, for example, genetic rescue (Whiteley et al. 2015). We also found that the probability of surviving the first year varied according to the year of study, stressing the importance of temporal heterogeneity in environmental conditions. Finally, we stress the importance of adopting an integrative approach when assessing the impact of inbreeding on early developmental stages, by considering the relative importance of parental effects and their interactions, as well as potential mechanisms of inbreeding avoidance. The field would benefit from studying adaptive parental interactions with regards to offspring viability in the future, to better understand the dynamics of inbreeding depression and its impact on populations at risk of extinction.
References
Alho JS, Välimäki K, Merilä J (2010) Rhh: an R extension for estimating multilocus heterozygosity and heterozygosity–heterozygosity correlation. Mol Ecol Resour 10:720–722
Amos W, Wilmer JW, Fullard K, Burg TM, Croxall JP, Bloch D, Coulson T (2001) The influence of parental relatedness on reproductive success. Proc Biol Sci 268:2021–2027
Balloux F, Amos W, Coulson T (2004) Does heterozygosity estimate inbreeding in real populations? Mol Ecol 13:3021–3031
Bichet C, Vedder O, Sauer-Gurth H, Becker PH, Wink M, Bouwhuis S (2018) Contrasting heterozygosity-fitness correlations across life in a long-lived seabird. Mol Ecol. https://doi.org/10.1111/mec.14979
Bischoff W (1984) Lacerta agilis (Linneaus 1758)—Zauneidechse. In: Böhme W (ed) Handbuch der reptilien und amphibien Europas 2/1. AULA-Verlag, Wiesbaden, pp 23–68
Brown JL, Eklund A (1994) Kin recognition and the major histocompatibility complex—an integrative review. Am Nat 143:435–461
Chapman JR, Nakagawa S, Coltman DW, Slate J, Sheldon BC (2009) A quantitative review of heterozygosity-fitness correlations in animal populations. Mol Ecol 18:2746–2765
Charlesworth D, Charlesworth B (1987) Inbreeding depression and its evolutionary consequences. Annu Rev Ecol Syst 18:237–268
Coltman DW, Pilkington JG, Smith JA, Pemberton JM (1999) Parasite-mediated selection against inbred Soay sheep in a free-living island population. Evolution 53:1259–1267
Crean AJ, Bonduriansky R (2014) What is a paternal effect? Trends Ecol Evol 29:554–559
Crnokrak P, Roff DA (1999) Inbreeding depression in the wild. Heredity 83:260–270
Darwin C (1868) The variation of animals and plants under domestication. John Murray, London
David P, Pujol B, Viard F, Castella V, Goudet J (2007) Reliable selfing rate estimates from imperfect population genetic data. Mol Ecol 16:2474–2487
Frankham R (2005) Genetics and extinction. Biol Conserv 126:131–140
Gullberg A, Olsson M, Tegelström H (1998) Colonization, genetic diversity, and evolution in the Swedish sand lizard, Lacerta agilis (Reptilia, Squamata). Biol J Linn Soc 65:257–277
Gullberg A, Olsson M, Tegelström H (1999) Evolution in populations of Swedish sand lizards: genetic differentiation and loss of variability revealed by multilocus DNA fingerprinting. J Evol Biol 12:17–26
Hedrick PW, Kalinowski ST (2000) Inbreeding depression in conservation biology. Annu Rev Ecol Syst 31:139–162
Hemmings NL, Slate J, Birkhead TR (2012) Inbreeding causes early death in a passerine bird. Nat Commun 3:863
Huisman J, Kruuk LE, Ellis PA, Clutton-Brock T, Pemberton JM (2016) Inbreeding depression across the lifespan in a wild mammal population. Proc Natl Acad Sci USA 113:3585–3590
Keller LF, Waller DM (2002) Inbreeding effects in wild populations. Trends Ecol Evol 17:230–241
Keller LF, Arcese P, Smith JNM, Hochachka WM, Stearns SC (1994) Selection against inbred song sparrows during a natural-population bottleneck. Nature 372:356–357
Kempenaers B, Adriaensen F, van Noordwijk AJ, Dhondt AA (1996) Genetic similarity, inbreeding and hatching failure in blue tits: are unhatched eggs infertile? Proc R Soc B 263:179–185
Kruuk LEB, Sheldon BC, Merila J (2002) Severe inbreeding depression in collared flycatchers (Ficedula albicollis). Proc R Soc B 269:1581–1589
Lacy RC (1997) Importance of genetic variation to the viability of mammalian populations. J Mamm 78:320–335
Lynch M (1991) The genetic interpretation of inbreeding depression and outbreeding depression. Evolution 45:622–629
Madsen T, Shine R, Olsson M, Wittzell H (1999) Conservation biology: restoration of an inbred adder population. Nature 402:34–35
Madsen T, Olsson M, Wittzell H, Stille B, Gullberg A, Shine R, Andersson S, Tegelstrom H (2000) Population size and genetic diversity in sand lizards (Lacerta agilis) and adders (Vipera berus). Biol Conserv 94:257–262
Mainguy J, Cote SD, Coltman DW (2009) Multilocus heterozygosity, parental relatedness and individual fitness components in a wild mountain goat, Oreamnos americanus population. Mol Ecol 18:2297–2306
Michaelides SN, While GM, Zajac N, Aubret F, Calsbeek B, Sacchi R, Zuffi MA, Uller T (2016) Loss of genetic diversity and increased embryonic mortality in non-native lizard populations. Mol Ecol 25:4113–4125
Moller AP, Alatalo RV (1999) Good-genes effects in sexual selection. Proc R Soc B 266:85–91
Mousseau TA, Fox CW (1998) The adaptive significance of maternal effects. Trends Ecol Evol 13:403–407
Olsson M (1993) Male preference for large females and assortative mating for body size in the sand lizard (Lacerta agilis). Behav Ecol Sociobiol 32:337–441
Olsson M (1994) Nuptial coloration in the sand lizard, Lacerta agilis: an intra-sexually selected cue to lighting ability. Anim Behav 48:607–613
Olsson M, Madsen T (1995) Female choice on male quantitative traits in lizards - why is it so rare. Behav Ecol Sociobiol 36:179–184
Olsson M, Madsen T (2001a) Promiscuity in sand lizards (Lacerta agilis) and adder snakes (Vipera berus): causes and consequences. J Hered 92:190–197
Olsson M, Madsen T (2001b) Between-year variation in determinants of offspring survival in the Sand Lizard, Lacerta agilis. Funct Ecol 15:443–450
Olsson M, Shine R (1997) The limits to reproductive output: offspring size versus number in the sand lizard (Lacerta agilis). Am Nat 149:179–188
Olsson M, Gullberg A, Tegelstrom H, Madsen T, Shine R (1994) Can female adders multiply. Nature 369:528
Olsson M, Gullberg A, Tegelström H (1996a) Malformed offspring, sibling matings, and selection against inbreeding in the sand lizard (Lacerta agilis). J Evol Biol 9:229–242
Olsson M, Shine R, Madsen T, Gullberg A, Tegelström H (1996b) Sperm selection by females. Nature 383:585
Olsson M, Madsen T, Nordby J, Wapstra E, Ujvari B, Wittsell H (2003) Major histocompatibility complex and mate choice in sand lizards. Proc R Soc B 270:S254–S256
Olsson M, Wapstra E, Schwartz T, Madsen T, Ujvari B, Uller T (2011) In hot pursuit: fluctuating mating system and sexual selection in sand lizards. Evolution 65:574–583
Potts WK, Manning CJ, Wakeland EK (1994) The role of infectious-disease, inbreeding and mating preferences in maintaining mhc genetic diversity—an experimental test. Philos T Roy Soc B 346:369–378
Pusey AE (1987) Sex-biased dispersal and inbreeding avoidance in birds and mammals. Trends Ecol Evol 2:295–299
Pusey A, Wolf M (1996) Inbreeding avoidance in animals. Trends Ecol Evol 11:201–206
Richardson DS, Komdeur J, Burke T (2004) Inbreeding in the seychelles warbler: environment-dependent maternal effects. Evolution 58:2037–2048
Ryberg K, Olsson M, Wapstra E, Madsen T, Anderholm S, Ujvari B (2004) Offspring-driven local dispersal in female sand lizards (Lacerta agilis). J Evol Biol 17:1215–1220
Schwartz TS, Perrin C, Wapstra E, Uller T, Olsson M (2011) Complex selection associated with Hox genes in a natural population of lizards. J Evol Biol 24:2520–2524
Sepil I, Radersma R, Santure AW, De Cauwer I, Slate J, Sheldon BC (2015) No evidence for mhc class I-based disassortative mating in a wild population of great tits. J Evol Biol 28:642–654
Sherborne AL, Thom MD, Paterson S, Jury F, Ollier WER, Stockley P, Beynon RJ, Hurst JL (2007) The genetic basis of inbreeding avoidance in house mice. Curr Biol 17:2061–2066
Slate J, Kruuk LEB, Marshall TC, Pemberton JM, Clutton-Brock TH (2000) Inbreeding depression influences lifetime breeding success in a wild population of red deer (Cervus elaphus). Proc R Soc B 267:1657–1662
Spottiswoode C, Moller AP (2004) Genetic similarity and hatching success in birds. Proc R Soc B 271:267–272
Szulkin M, Bierne N, David P (2010) Heterozygosity-fitness correlations: a time for reappraisal. Evolution 64:1202–1217
Tregenza T, Wedell N (2002) Polyandrous females avoid costs of inbreeding. Nature 415:71–73
van Noordwijk AJ, Scharloo W (1981) Inbreeding in an island population of the great tit. Evolution 35:674–688
Wang J, Xie H, Fisher JF (2011) Multilevel models: applications using SAS®. Walter De Gruyter, Berlin
Weir BS, Cockerham CC (1973) Mixed self and random mating at two loci. Genet Res 21:247–262
Weiser EL, Grueber CE, Kennedy ES, Jamieson IG (2016) Unexpected positive and negative effects of continuing inbreeding in one of the world’s most inbred wild animals. Evolution 70:154–166
Whiteley AR, Fitzpatrick SW, Funk WC, Tallmon DA (2015) Genetic rescue to the rescue. Trends Ecol Evol 30:42–49
Williams RN, Bos DH, Gopurenko D, Dewoody JA (2008) Amphibian malformations and inbreeding. Biol Letters 4:549–552
Zakharov VM (1989) Future prospects for population phenogenetics. Gordon and Breach, Amsterdam
Zeh JA, Zeh DW (1996) The Evolution of polyandry I: intragenomic conflict and genetic incompatibility. Proc R Soc B 263:1711–1717
Zeh JA, Zeh DW (1997) The evolution of polyandry II: post–copulatory defenses against genetic incompatibility. Proc R Soc B 264:69–75
Acknowledgements
We thank the Australian Research Council and the Swedish Science Council (MO, EW). We are also thankful to Dr Donal Blomqvist and Henrik Imberg for their valuable statistical advice.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
10592_2019_1180_MOESM1_ESM.pdf
Supplementary material 1 (PDF 9 kb) Figure. 1 (supplementary) Number of females captured every year, between 1998 and 2006. Note that certain females were recaptured multiple times. In total, 252 different females were captured, and the number varied between 29 and 65 during the study period. The last individuals were released in 2006, hence the lack of data for 2007. Figure. 2 (supplementary) Distribution of standardized individual-level heterozygosity (SH) values for hatched and unhatched embryos (n = 3043; Mean = 0.99; Coefficient of variation = 0.22; SD = 0.22; Maximum value = 1.65; minimum value = 0).
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Bererhi, B., Wapstra, E., Schwartz, T.S. et al. Inconsistent inbreeding effects during lizard ontogeny. Conserv Genet 20, 865–874 (2019). https://doi.org/10.1007/s10592-019-01180-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-019-01180-6