Abstract
CRISPR gene drive has recently been proposed as a promising technology for population management, including in conservation genetics. The technique would consist in releasing genetically engineered individuals that are designed to rapidly propagate a desired mutation or transgene into wild populations. Potential applications in conservation biology include the control of invasive pest populations that threaten biodiversity (eradication and suppression drives), or the introduction of beneficial mutations in endangered populations (rescue drives). The propagation of a gene drive is affected by different factors that depend on the drive construct (e.g. its fitness effect and timing of expression) or on the target species (e.g. its mating system and population structure). We review potential applications of the different types of gene drives for conservation. We examine the challenges posed by the evolution of resistance to gene drives and review the various molecular and environmental risks associated with gene drives (e.g. propagation to non target populations or species and unintended detrimental ecosystem impacts). We provide some guidelines for future gene drive research and discuss ethical, biosafety and regulation issues.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Endangered species and ecosystems can be protected using either direct interventions targeting the species of interest, or indirect interventions that focus on the surrounding species or environment. The potential use of synthetic biology technologies, such as gene drives, to meet these goals, has recently sparked interest (Piaggio et al. 2017), but also concerns (SynBioWatch 2016) among conservation biologists. Population management using gene drive is based on the release of genetically engineered individuals that are designed to propagate a desired mutation or transgene in natural populations (Fig. 1, Box 1). Depending on the fitness effect of a drive construct, we can distinguish three types of gene drives in conservation biology: eradication, suppression and rescue drives (Fig. 2). Compared to other typologies, for example whether or not the drive confers a new function to the organism (e.g. alteration or replacement drives; Gantz et al. 2015), we consider that this distinction into three categories is the most relevant for conservation purposes, as the fitness effect of a drive construct is an important parameter for its spread in a target population.
Comparison of Mendelian and gene drive inheritance in mice based on conversion rates from Grunwald et al. (2019). a A classical mutation is transmitted to 50% of the gametes of heterozygous individuals (in green; Mendelian inheritance). b A synthetic gene drive element, that targets the tyronase (Tyr) gene, is transmitted to 72% of the gametes of heterozygous individuals on average (in green; non-Mendelian inheritance). Gene drive transmission varies from 60 to 86% when comparing five different crosses of laboratory mice (Grunwald et al. 2019). A single pair of chromosomes is presented (black rectangle: wild chromosome, green rectangle: chromosome carrying a regular mutation or a gene drive). In this example, gene conversion takes place in the gonads. Original mouse picture by Donald Hobern-Flickr. (Color figure online)
Eradication and suppression drives are designed to extirpate or decrease the size of a population, respectively. They rely on the introduction of strongly or mildly deleterious mutations, respectively. These drives are primarily developed for their applications for human health and for the control of agricultural pests. They could also be applied for ecosystem management in conservation, to target invasive species that threaten biodiversity (Esvelt et al. 2014). Using gene drive for population management could have lower health, economic and environmental costs than traditional control methods (Harvey-Samuel et al. 2017).
Rescue drives, on the other hand, could be designed to save endangered populations by introducing beneficial mutations or removing deleterious ones (Fig. 2; Esvelt et al. 2014). Due to the non-Mendelian inheritance of gene drives, these mutations would spread more quickly in target populations than with natural selection only. Rescue drives could alleviate an important dilemma traditionally faced by conservation geneticists: should one introduce individuals from other regions that bring in useful genetic variability, or locally adapted individuals? Introducing individuals from a distant source population into an endangered population might inadvertently introduce deleterious alleles that could result in outbreeding depression or in the overall maladaptation of the population (Bucharova 2017). When only a single or a few loci with large fitness effects provide adaptation to a specific environmental factor, rescue drives would allow locus-specific assisted gene flow, by providing beneficial alleles for some adaptive traits, while maintaining alleles for other adaptive traits at high frequencies. Rescue drives could increase stress tolerance (e.g. using drought-tolerance genes in eastern white pine; Tang et al. 2007), or increase resistance to pathogens (e.g. using immunity genes conferring blight resistance in American chestnut; Kubisiak et al. 2013; Newhouse et al. 2014). Rescue drives could also be used in other contexts than conservation. In agriculture, they could for instance make honey bees and other important pollinators less susceptible to neonicotinoid insecticides. Regarding human health applications, rescue drives could help make bank voles more resistant to the tick-borne pathogen Borrelia afzelii, which is responsible for Lyme disease in humans (e.g. using Toll-like receptors; Tschirren et al. 2013).
Previous reviews on gene drives have focused either on the different types of drive systems (e.g. Gantz and Bier 2016; Champer et al. 2016; Harvey-Samuel et al. 2017; Marshall and Akbari 2018), on their applications for human health or for pest control in agriculture (e.g. Macias et al. 2017; Godfray et al. 2017; Scott et al. 2018; McFarlane et al. 2018) or on the challenges of their development in terms of identifying current knowledge gaps (Moro et al. 2018), biosafety (Benedict et al. 2018), regulation (Oye et al. 2014; Caplan et al. 2015; Meghani and Kuzma 2018) and ethics (Courtier-Orgogozo et al. 2017; Thompson 2018). Although a number of reviews presented some gene drive applications in conservation (Gould 2008; Esvelt et al. 2014; Thresher et al. 2014; Webber et al. 2015; Piaggio et al. 2017; Zentner and Wade 2017; Esvelt and Gemmell 2017; Moro et al. 2018; Min et al. 2018; Dearden et al. 2018; Phelps et al. 2019; Brossard et al. 2019), the fundamental differences between the risks associated with rescue drives and those associated with suppression and eradication drives have not been considered previously. In this paper, we fill this gap and review the use of gene drives for population management with a special emphasis on conservation biology. We focus on CRISPR-cas9-mediated gene drives (Box 1), a technology that can be applied to a wide variety of organisms and is more stable than alternative genome editing approaches (i.e. less prone to recombination events that result in non-functional enzymes; Champer et al. 2016). Unlike other drive systems (e.g. segregation distorters) however, such CRISPR-based gene drives present important molecular risks. In this review, we seek to provide conservation scientists and land managers with an in-depth understanding of CRISPR editing technology, so that they can better assess the risks and benefits associated with CRISPR gene drives. For readers interested in other drive systems, we recommend other publications (e.g. Gantz and Bier 2016; Champer et al. 2016; Harvey-Samuel et al. 2017; Marshall and Akbari 2018), including reviews on t-haplotype gene drives in invasive mice (Leitschuh et al. 2018; McFarlane et al. 2018).
Key features affecting gene drive propagation
Gene drive timing of expression
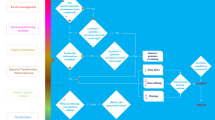
The timing of expression of the Cas9 endonuclease and gRNA (i.e., timing of conversion) is an important parameter for the successful propagation of a gene drive (Fig. 3; Deredec et al. 2008). For eradication and suppression drives, wild-type/drive heterozygotes have higher fitness than drive homozygotes, so that gene conversion late in life (in the gonads) favors drive spread more than early gene conversion (in the zygote) does (Fig. 3a). Conversely, early conversion (in the zygote) favors the spread of rescue drives (Fig. 3b). The timing of conversion can be adjusted experimentally by using promoters that drive expression at different stages, in the germline or in the early zygote (e.g. Champer et al. 2017; Hammond et al. 2018).
Conversion of wild-type allele into a gene drive allele can occur either (a) in the adult gonads (i.e. in its germline only) or (b) in the zygote. The selection coefficient, s, represents the fitness difference between wild-type homozygous versus drive homozygous individuals (s > 0, for eradication and suppression drives, s < 0 for rescue drives). When the fitness of wild-type/drive heterozygotes equals the fitness of wild-type homozygotes, the gene drive allele is completely recessive (dominance coefficient, h = 0), whereas when the fitness of wild-type/drive heterozygous equals the fitness of drive homozygous individuals, the gene drive allele is completely dominant (h = 1, as represented here). For eradication and suppression drives (s > 0), the fitness of heterozygous individuals equals or is higher than the fitness of homozygous individuals (whet ≥ whom). For rescue drives (s < 0), the fitness of homozygous individuals equals or is higher than the fitness of heterozygous individuals (whom ≥ whet). A single pair of chromosomes is presented (black rectangle: wild chromosome, green rectangle: chromosome carrying a gene drive). (Color figure online)
Gene drive genetic parameters
Theoretical studies have investigated the influence of different parameters on drive dynamics (e.g. Deredec et al. 2008; Unckless et al. 2015; Noble et al. 2018). Here we distinguish between deterministic models (that assume very large population sizes and ignore gene drift) and stochastic models (that take chance into account).
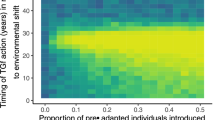
In a deterministic model, a gene drive allele can go to fixation, disappear, but also reach an intermediate equilibrium frequency (see Fig. 4, illustrating results from a model where gene conversion takes place in the gonads; see Supplementary Information for model details). For eradication or suppression drives, the final state depends on parameters such as the probability of successful gene conversion, the dominance coefficient (h), and the coefficient of selection (s) of the drive allele (Fig. 4). For some parameter combinations, the final state also depends on the introduction frequency of the drive (“WT or Drive” area in Fig. 4a and red curves in Fig. 4b, c). For rescue drives (s < 0), the drive allele is predicted to always fix eventually.
Effect of selection intensity (s) and dominance coefficient (h) on drive dynamics for suppression or eradication drives with conversion in the gonads. s and h are defined as in Fig. 3. a Parameter ranges of the different outcomes; “Drive fixation”: fixation of the drive allele, “Coex”: coexistence between drive and wild-type (WT) alleles, “WT”: fixation of the WT allele, “WT or Drive”: bistability, fixation of either the drive or the WT allele, depending on drive initial introduction frequency. Rescue drives (s < 0) always fix and are not represented. The conversion efficiency c is here set to 85%. b Dynamics of the frequency of the drive in the population, for four different sets of (s, h) parameters. The parameters corresponding to the line colors are shown with colored dots in a. The frequency of introduction of the drive allele is 0.1. c Same as (b), but with a higher frequency of introduction of the drive allele (0.6). Note the different outcome for the red curve (bistability conditions). See Deredec et al. (2008) and Supplementary Information for details. (Color figure online)
These results mostly hold true for stochastic models, but the fixation—or loss—of the drive allele is not always guaranteed whenever chance events (genetic drift) are taken into account. Stochasticity can indeed play an important role for small or fragmented populations. It has been confirmed that the release of a higher number of drive-carrying individuals increases the chance of its eventual fixation (Unckless et al. 2015).
Theoretical models have exclusively focused on fitness differences between gene drive and wild-type alleles during the diploid phase of the life cycle, ignoring potential differences also acting during the haploid phase (e.g. in the pollen for plants or in males of haplodiploid species such as invasive wasps). We expect such fitness differences to favor even further the spread of rescue drives, but to disfavor the spread of suppression and eradication drives.
Mating systems
Gene drives require sexual reproduction for their transmission. Mate limitation during the establishment phase of an invasive species is thought to favor species with mixed mating systems (i.e. species that can both outcross and self-fertilize or that can reproduce both sexually and asexually; Baker 1967; Pannell et al. 2015), compared to species that can only outcross or reproduce sexually. Gene drives are likely to be less efficient in invasive species with such mixed mating systems (Bull 2017). For example, a gene drive that targets a species with high rates of selfing and low inbreeding depression is unlikely to reach high frequencies (Noble et al. 2018). More generally, many of the important challenges in conservation genetics involve species that reproduce exclusively through selfing or asexual reproduction. For instance, despite the recent discovery of potential candidate genes to mitigate coral bleaching through locus-specific assisted gene flow (Jin et al. 2016), asexual reproduction of many coral species (e.g. Highsmith 1982) or of their endosymbionts (Andras et al. 2011) represents a major limitation to their management using gene drives. Similarly, gene drives cannot be used to modify asexually reproducing micro-organisms and increase their biodegradation of pollutants (Joutey et al. 2013) or to modify the asexual cells causing transmissible cancer in Tasmanian devils (Siddle et al. 2013).
Generation time
Although gene drives can invade populations in a few dozens of generations, this process could take several centuries in long-lived species. For instance, fixation of a gene drive within 20 generations in eastern white pine would take about 600 years, considering a generation time of 30 years (Nijensohn et al. 2005). As climate change occurs on a much shorter time scale (Pachauri et al. 2014), it is likely that rescue drives could not prevent extinction in this particular case.
Population structure in space and time
In spatially-structured populations, dispersal is another important parameter to consider to predict gene drive propagation. Population fragmentation with low dispersal rates between populations, as often observed in endangered species, would likely affect gene drive dynamics in a metapopulation. Very low dispersal rates could slow down the spread of a rescue drive in a metapopulation.
Population structure in time, corresponding to the presence of resting stages (e.g. seed bank in plants) might result in a constant inflow of wild-type individuals for years or decades (i.e. dispersal in time rather than space), which might reduce the spread of gene drives (NASEM 2016).
Density-dependence, age and social structure
Depending on the stage of the life cycle at which they occur, the effect of population density on mortality or reproduction could also affect gene drive dynamics (Godfray et al. 2017). While conversion rates can vary with age (e.g. as observed in Drosophila melanogaster; Preston et al. 2006), the influence of this age factor on drive propagation remains to be investigated. Finally, social structures that limit breeding to dominant pairs are also likely to affect gene drive spread (Moro et al. 2018), although their effect has never been thoroughly investigated theoretically.
Potential examples of applications in conservation
Population genetics studies have shown that new mutations are generally deleterious and that few mutations are advantageous (Eyre-Walker and Keightley 2007). Since the number of potential genetic targets is smaller for rescue drives than for suppression and eradication drives and since our knowledge regarding the genetic basis of adaptive traits remains limited, rescue drives represent a small fraction of potential applications of gene drives in conservation biology. We present a list of potential target species (Table 2), and provide details regarding four case studies (Fig. 2). Importantly, these illustrative case studies are hypothetical and are unlikely to be developed in the coming years; the reason for presenting them is merely to help better assess the different types of risks associated with gene drives.
Eradication of invasive black rats in New Zealand
Black rats were introduced to New Zealand following European colonization during the nineteenth century (Atkinson 1973). Their introduction resulted in the suppression of many endemic plants (through seed predation) and in the extinction or decline of several insect, snail, spider, reptile and bird species (Towns et al. 2006; Towns 2009).
Different population control methods can be used against rats, such as physical removal (e.g. trapping), pesticide treatments (e.g. anticoagulant toxicants), biological control (e.g. releasing cat predators) or female sterilization (e.g. contraception; NASEM 2016). In 2016, the New Zealand government financed a plan (Predator Free 2050) to eradicate black rats along with other invasive species by 2050 (Norton et al. 2016). Predator Free 2050, together with universities and non-profit organizations, is developing an international program on gene drive research against rodents (Genetic Biocontrol of Invasive Rodents program; Hall 2017). Thanks to the advances of CRISPR-cas9 genome editing in rat (Remy et al. 2017), gene drives currently developed in house mice (Grunwald et al. 2019) could be tested in rats in the future (Fig. 2a; Prowse et al. 2017).
Targeting rats in New Zealand using gene drives could have potential unintended consequences. Dispersal of gene drive rats to neighbouring countries would be an important international issue. Black rats can hybridize with the Asian rat, Rattus tanezumi (Chinen et al. 2005), which poses a risk of propagation to R. tanezumi native populations. In addition, removing rats (with any control method) could have some negative consequences for native species. For example, rats might have replaced native species originally responsible for dispersing seeds of native plants (Shiels and Drake 2011) or spores of native fungi (Vernes and McGrath 2009).
Protection of honeycreepers in Hawaii
Honeycreepers and other endemic birds in Hawaii have evolved in the absence of avian malaria and, consequently, are particularly susceptible to the invasive malaria parasite Plasmodium relictum (Lounibos 2002). The main vector of this parasite, the southern house mosquito, Culex quinquefasciatus, is invasive in Hawaii since the beginning of the nineteenth century (Lounibos 2002). An eradication drive targeting mosquito populations could protect endemic birds (Fig. 2a; NASEM 2016).
Potential methods include the disruption of mosquito genes that are required for female fertility (e.g. doublesex gene; Kyrou et al. 2018) or for the propagation of malaria parasites (e.g. a Culex gene equivalent to the FREP1 gene in A. gambiae; Dong et al. 2018). An alternative to eradication drives includes the introduction of cargo genes that code for antibodies preventing the reproduction and transmission of the parasites (Gantz et al. 2015). Note that strategies alternative to gene drives based on the sterilization of females with irradiation (sterile insect technique) or using the bacteria Wolbachia (incompatible insect technique) are currently being developed in different mosquito species (Lees et al. 2015; Ritchie et al. 2018).
Spreading Rht dwarfing alleles in invasive common ragweed
Common ragweed (Ambrosia artemisiifolia) is an annual plant, native to North America and invasive in South America, Europe, Africa, Asia and Australia (Smith et al. 2013). Ragweed produces different allelochemical metabolites and suppresses the growth of native plant species, hence reducing plant biodiversity (Smith et al. 2013). As ragweed causes important allergies in humans (Smith et al. 2013) and is considered as a weed in agriculture (Bassett and Crompton 1975), many countries and the European Union have launched eradication programs (Smith et al. 2013). However, control methods based on mechanical or herbicide treatments can have a negative impact on biodiversity (Alberternst et al. 2016).
In a recent perspective on gene drive applications in agriculture, Neve (2018) suggested developing suppression drives targeting homologues of Reduced height (Rht) genes in weeds. These genes are responsible for dwarfism in cultivated wheat and are also found in sunflowers (Ramos et al. 2013). Ragweed is related to sunflowers and suppression drives that target orthologous Rht genes could be developed to suppress this species. Height is an important trait for competition for light in ragweed (Coble et al. 1981), so that such a gene drive would reduce ragweed competitive ability (Fig. 2b). The evolution of selfing would not be an issue, as ragweed is an obligate outcrossing plant with separate sexes (Friedman and Barrett 2008). As ragweed is wind-pollinated, the decline of its population should not affect pollinator communities. However, ragweed populations are characterized by large seed banks (Brandes and Nitzsche 2006), which might impair the spread of suppression drives in this species.
Introducing MHC resistance alleles in endangered amphibian species
The chytrid fungus, Batrachochytrium dendrobatidis, has emerged as a global threat for up to 50% of amphibian species (Fisher et al. 2009). The fungus reproduces mostly asexually (Fisher et al. 2009), and therefore cannot be targeted with a gene drive. However, resistance to B. dendrobatidis infection varies both within and among amphibian species (Fu and Waldman 2017). Major histocompatibility complex (MHC) peptides play an important role in the innate immune system of vertebrates, by presenting antigens to lymphocytes. A specific MHC allele has been shown to increase the chance of survival of infected individuals in the lowland leopard frog Lithobates yavapaiensis (Savage and Zamudio 2011).
CRISPR-cas9 genome editing is advancing in clawed frogs, Xenopus spp. (e.g. Aslan et al. 2017), so the development of rescue drives using resistant MHC alleles could be considered for amphibian populations at risk (Fig. 2c; Esvelt et al. 2014). However, mounting an effective immune response against B. dendrobatidis might also carry immunity trade-offs (Fu and Waldman 2017). Although replacing an endogenous MHC gene by a resistant MHC allele would increase resistance to this fungus, it would also reduce allelic diversity at the MHC locus which could in turn increase population susceptibility to other pathogens. An alternative would be the insertion of a gene drive cassette with a cargo including a resistant MHC allele at a locus unlinked to the endogenous MHC locus. This strategy might preserve MHC variability but is likely to affect the stability of the gene drive cassette due to the risks of recombination with the endogenous MHC locus.
Issues associated with a lack of efficacy of gene drives
Evolution of resistance to gene drives
Resistance to gene drive can either occur at the molecular level when a chromosome is not recognized or cleaved by the Cas9 enzyme (Box 1c) or at the behavioral level when wild-type individuals avoid mating with gene drive individuals. To our knowledge, resistance studies have generally focused on molecular resistance, and behavioral resistance has never been investigated experimentally or theoretically. Molecular resistance can occur either through standing genetic variation (i.e. polymorphism at the target site) or through evolution by de novo mutations (Unckless et al. 2017).
Alleles that confer resistance to a gene drive through standing genetic variation are already present and segregate in the target population (Drury et al. 2017). Such resistance alleles could originate from mutations in the past, or be acquired through hybridization and introgression with a related drive-resistant species (as observed for anticoagulant resistance in house mouse; Song et al. 2011). Such resistance may not be a problem for rescue drives: alleles resistant to cleavage are expected to be more deleterious than the drive allele, and would therefore not prevent the spread of a drive-propagated beneficial mutation. For suppression and eradication drives, the risk of resistance via standing genetic variation can be reduced by targeting sequences that are functionally constrained and hence present low polymorphisms in natural populations (e.g. Kyrou et al. 2018). This would, however, increase the risks of gene drive propagation to non-target species (see below).
The evolution of de novo resistance represents an important risk, especially for eradication and suppression drives. When the Cas9-induced double-strand DNA breaks are not repaired by gene conversion, non-homologous end-joining (NHEJ) repair can result in insertions or deletions that make wild-type chromosomes resistant to further cleavage by the Cas9 endonuclease (Box 1c). A recent study in D. melanogaster shows that the probability of occurrence of such indel mutations in the germline could be several orders of magnitude higher in drive/wild-type heterozygotes compared to wild-type homozygotes (e.g. 20% vs. 10−8%; Sharp and Agrawal 2016; Champer et al. 2017). In addition, genetic variation in the NHEJ repair system both among and within target populations could select for increased resistance to gene drives (Champer et al. 2017, 2018b). Indel mutations conferring resistance are selected for when their fitness costs are lower than the fitness costs associated with gene drive alleles (Unckless et al. 2017). Studies suggest that, in plants, NHEJ repair predominates over homology directed repair (Gorbunova and Levy 1999; Li et al. 2013). The high occurrence of such indel mutations in target species where NHEJ predominates would impair the spread of suppression or eradication drives.
The emergence of resistance within a few generations is currently one of the main causes of failure in gene drive experimental evolution studies (Hammond et al. 2017; KaramiNejadRanjbar et al. 2018). Using several gRNAs that target multiple sites is predicted to decrease the rate of emergence of resistance alleles (Noble et al. 2017b). This strategy is similar to multi-drug therapy, whereby targeting multiple sites makes the evolution of resistance simultaneously at all sites less likely (Rex Consortium 2013). Two experimental studies found that targeting multiple sites decreases the appearance of alleles resistant to cleavage (Champer et al. 2018a; Oberhofer et al. 2018). However, the constructions differed and Oberhofer et al. (2018) found many instances of incomplete homology-directed repair where the CRISPR-cas9 cassette was only partially copied (e.g. without the cas9 gene). Individuals carrying partial copies of the cassette incur important fitness costs, which can prevent the spread of such gene drive constructs (Oberhofer et al. 2018).
Cas9 activity outside of the germline/zygote
As described above, there is an optimal timing for gene conversion (Fig. 3). When repaired by NHEJ events, cleavage of the wild-type allele outside of the optimal timing window could result in mosaic heterozygous individuals with low fitness. This issue has been studied only for suppression or eradication drives (Champer et al. 2018a; Oberhofer et al. 2018), but not for rescue drives.
Risks of unintended effects
Using gene drive in conservation biology could result in potential hazards at different scales, from molecular to ecosystem levels. Most of the molecular and ecosystem risks associated with population management using eradication and suppression drives are not specific to conservation and can also be expected in applications for human health and agriculture.
Molecular off-target mutations
To our knowledge, all gene drive experimental studies so far have focused on conversion at the target site, without testing for potential off-target effects outside of the targeted genomic region. In both heterozygous (wild-type/drive) or homozygous (drive/drive) individuals, the presence of the Cas9 endonuclease could induce double-strand breaks in genomic regions different from the target site. A random mutation in the gRNA could lead to the cleavage of off-target sites (i.e. gRNA “retargeting”; Scharenberg et al. 2016). A drive construct can therefore be considered as a mutagen, whose off-target effects will depend on the specificity of the gRNA and on its susceptibility to retargeting mutations.
Off-target double-strand breaks could be repaired by homology directed repair (Box 1b) or NHEJ (Box 1c). NHEJ repair could result in indel mutations with potential fitness costs (Barton and Zeng 2018). Homology-directed repair could result in the replacement of the chromosomal regions surrounding the off-target regions by corresponding regions in the homologous chromosomes, leading to a local loss of heterozygosity (Gorter de Vries et al. 2018). Furthermore, near-target mutations can also occur following the resection of DNA double-strand breaks and homology-directed repair of regions flanking the target site (Cho et al. 1998), which also results in loss of heterozygosity (Fig. 5). For example, frequent loss of heterozygosity over entire chromosome arms has been evidenced in yeast (ranging from 10 to 40% per meiosis; Gorter de Vries et al. 2018). If near- and off-target effects are frequent, they could globally increase the mutation load in target populations. They would represent an important risk of failure for rescue drives as beneficial effects could be overwhelmed by an increased inbreeding depression. In contrast, near- and off-target effects could accelerate population decline for eradication or suppression drives. Further studies are needed to determine the extent of these mutations and to model their impact on drive dynamics.
Risks of an extensive loss of heterozygosity. a Gene drive and wild-type chromosomes carry many different deleterious mutations, most of which are heterozygous. b Cleavage of the wild-type chromosome by the Cas9 and trimming (resection) of the DNA ends by the double-strand break DNA repair machinery. c Many deleterious mutations become homozygous following homology-directed repair, which can increase inbreeding depression. Red vertical rectangles indicate deleterious mutations. The sizes of the centromere, the target site, the cassette and the Cas9 protein are not to scale. (Color figure online)
Propagation to non-target populations
Several strategies could limit the spread of a gene drive from targeted populations to non-target populations. First, so-called “precision drives” (Esvelt et al. 2014) would target a gene or a sequence specific to a target population. A second strategy consists in first recoding an allele by propagating a gene drive with no fitness effect, and then releasing a second drive that would target the recoded allele only, and so forth with several successive drives (Esvelt et al. 2014). This approach would then limit the probability of spread of suppression and eradication drives to non-target populations. For rescue drives however, individuals carrying the final drive are selectively favored and could thus hybridize with non-target populations. Implementing this strategy would take a very long time, as each intermediate gene drive would need several generations to fix. To our knowledge, no theoretical model has investigated whether such precision drives could fail due to resistance through adaptive introgression with non-target populations (see above).
Propagation to non-target populations could also be avoided by the use of self-limiting drives, such as drives that only spread, at least theoretically, if introduced above a given frequency (e.g. a drive with the parameters of the red curve in Fig. 3, underdominance systems, or a combination of gene drive and underdominance systems; Dhole et al. 2018a). A strategy called “daisy-chain drive” (Noble et al. 2016) has been proposed to achieve self-limitation. This method involves a linear chain of unlinked genetic elements, such that gene conversion at locus i + 1 can only occur if a drive allele is present at locus i. Each of the lower elements confers some fitness cost, so that they are all sequentially lost from the population (Noble et al. 2016). So far, no laboratory report using daisy-chain drives has been published, but proof-of-concept experiments using nematodes have been proposed (Min et al. 2017). While temporally self-limited, daisy drives are not spatially self-limited, as they can easily spread to non-target populations (Dhole et al. 2018b).
Propagation to non-target populations is a key concern for the use of gene drive on islands. Islands are the primary biogeographical systems in which gene drives might be developed for conservation (e.g. 80% of the world’s islands now have invasive rodents; NASEM 2016). Dispersal may be rare, but a drive can spread with just one introduced individual. In addition, the deliberate unauthorized transport of gene drive individuals to non-target populations represents an important risk (Esvelt and Gemmell 2017). Eradication drives planned to be released to control invasive black rats and house mice in New Zealand could spread to the native range of these species (Leitschuh et al. 2018). Large-scale population genetics studies to estimate gene flow between New Zealand and other countries could help better estimate the risks of gene drive individuals escaping to other countries, and the risk of reinvasion of New Zealand following eradication.
The propagation of a transgene to non-target populations has been reported in genetically modified plants. For example, the dispersal of a transgene up to 3.8 km away from a test site has been observed following a field trial of glyphosate-resistant bentgrass in the USA (Reichman et al. 2006). The risks of transfer of a gene drive to non-target populations could be estimated using population genomic approaches, for instance by testing for potential gene flow between target and non-target populations and for the presence of the target sequence and flanking sequences of the gene drive cassette.
Propagation to non-target species
In addition to hybridization, DNA can be naturally transferred from one species to another through horizontal gene transfer. Such transfers are rarely detected, as most newly inserted DNA sequences are likely to be lost by genetic drift unless they confer strong fitness advantages (e.g. adaptive introgression of genes responsible for carotenoid biosynthesis in pea aphids; Moran and Jarvik 2010), or have a transmission advantage (self-replicating genetic elements).
Horizontal gene transfers can occur via parasites, pathogens or endosymbionts (e.g. viruses, bacteria, fungi and either sap-sucking insects in plants; Cho et al. 1998; or parasitoids in animals; Gilbert et al. 2010, 2014) and between extremely distantly related species (e.g. the BovB element moved at least 11 times between snakes, lizards, ruminants and marsupials; Ivancevic et al. 2017; and the Steamer element was transferred from molluscs to fish; Metzger et al. 2018). A natural transposable element, the P-element, has invaded D. melanogaster populations worldwide within 50 years, most likely following a single horizontal gene transfer event from an unrelated species, D. willistoni (Clark and Kidwell 1997). The P-element is now spreading in D. simulans populations (Hill et al. 2016). The transfer event might have occurred through hybridization of D. simulans with D. melanogaster (as some crosses between the two species can produce fertile hybrids; Davis et al. 1996), or horizontally (Kofler et al. 2015), or even maybe artificially (unintended escape of a few laboratory-raised D. simulans flies genetically engineered to carry a P-element), though the latter point is speculative.
Naturally occurring selfish endonuclease elements (so-called homing endonuclease genes; see Glossary) are in essence similar to gene drive constructs. The enzyme recognizes and cuts a specific target site on the homologous chromosome. Homology-directed repair converts the wild-type sequence into the homing endonuclease gene. Transfers of naturally occurring homing endonuclease genes have been documented between closely-related species (most likely through hybridization) and between distantly-related species (horizontal gene transfers). For example, the omega element has been transferred among different yeast species at least 15 independent times (Goddard and Burt 1999). Phylogenetic analyses in plants indicate that a class I intron homing endonuclease gene, that specifically targets the cox1 mitochondrial gene, has been transferred independently 70 times between 162 taxa belonging to 45 different families (Sanchez-Puerta et al. 2008). This element is also present in the genome of several species of fungi, green algae and liverworts, which suggests extensive horizontal gene transfers (Cho et al. 1998). However, this view may be biased as endonucleases that are easier to characterize are those that cut conserved sites that are shared among distantly related species, and hence more likely to be horizontally transferred. Whether the spread of gene drive cassettes in non-target species directly compares to the non-Mendelian propagation of natural endonuclease genes remains to be determined.
A target population fixed for an eradication drive will go extinct. For suppression or rescue drives that are fixed in a target population, there is no selective pressure to maintain a functional endonuclease, so that the CRISPR-cas9 cassette can eventually accumulate mutations (e.g. stop codons). These mutations have normal Mendelian inheritance (Fig. 1a) and can spread in target populations either through genetic drift (if they are neutral) or through natural selection (if they are beneficial, e.g. if the constitutive expression of Cas9 is costly). The risk of propagation to non-target species depends on the relative frequency of functional and non-functional gene drive cassettes and on the time before the non-functional CRISPR-cas9 cassette reaches fixation. The persistence time of gene drives is currently unknown. Whether it is the same order of magnitude as chemicals used for population management (several decades for many persistent organic pollutants such as DDT; Jones and De Voogt 1999) remains to be investigated.
Consequences for ecosystems
Removing invasive species might have unanticipated negative impacts on ecosystems through indirect effects on food webs (Zavaleta et al. 2001). Eradicating an invasive population might lead to the subsequent reinvasion by the same species or a different species with the same ecological niche. Other potential indirect effects depend on the position of the invasive species in the food chain. Invasive prey species can be heavily consumed by predators so that their sudden removal might result in increased predation on endemic species (Courchamp et al. 2003). For example, poisoning of black rats in a New Zealand forest resulted in invasive stoats (Mustela erminea), one of the main rat predators, switching their diet to native birds and bird eggs (Murphy and Bradfield 1992). Conversely, removing an invasive predator or an invasive herbivore might cascade down the food chain. For example, the eradication of feral goats and pigs on Sarigan islands (a US territory in the northwestern Pacific) led to the release of a previously undetected invasive vine (Operculina ventricosa) that subsequently covered most of the native forest and surrounding grassland (Kessler 2002). When two invasive species compete for the same niche (e.g. rats and mice), targeting only one competing species can result in an increase of the population of the other (Caut et al. 2007). Invasive species can also create new habitats or provide a food source for native species. For example, the worldwide invasion of the American brine shrimp Artemia franciscana has led to the extinction of many native Artemia parthenogenetica populations in Southern France (Rode et al. 2013). Contrary to A. parthenogenetica, A. franciscana is present throughout the winter in the area and represents a food source for native birds, including the greater flamingo (Phoenicopterus roseus). Eradicating the invasive A. franciscana might negatively affect native bird communities. More generally, eradicating an invasive species can move the ecosystem further away from its equilibrium without returning to its pre-invasion state, sometimes even making the system more susceptible to new invasions (David et al. 2017).
All of the risks listed above are not specific to population management using gene drives. As the pace of population suppression or eradication is likely to differ between gene drives and other control strategies, theoretical models could help anticipate and mitigate potential negative effects (David et al. 2013). For instance, the release of gene drive individuals might transiently increase population size with potentially long-lasting ecological consequences (David et al. 2013). Finally, rescue drives could destabilize food webs, for example by turning an endangered species into an invasive one.
Risk of failure of countermeasures to stop an ongoing drive
The reversibility (or not) of the modification is a key issue for the genetic modification of wild organisms. A first straightforward method to stop an ongoing gene drive is to release drive-resistant individuals that carry a functional copy of the targeted gene without the recognition sequence (Box 1c; Vella et al. 2017). This approach is expected to be effective for eradication drives, which impose strong fitness costs, but not for rescue drives or suppression drives imposing mild fitness costs (see above).
A second method consists in stopping the spread of a gene drive using a so-called gene drive brake (hereafter “brake”; Wu et al. 2016). Depending on whether the brake allele includes the cas9 gene, one can distinguish “immunizing reversal drives” and “reversal drives”. The former are used to replace both the initial drive and wild-type alleles with a second drive immune to the initial drive (Esvelt et al. 2014). Immunizing drives include the cas9 gene and have two different gRNAs that target the wild-type sequence and the sequence of the initial gene drive (Esvelt et al. 2014). The latter do not possess the cas9 gene and only target the sequence of the initial gene drive (Gantz and Bier 2016; Wu et al. 2016). In gene drive/brake heterozygotes, the gRNA binds with Cas9 to disrupt the functional copy of the cas9 gene (Fig. 6). In wild-type/brake heterozygotes, the brake has a regular Mendelian inheritance. A laboratory experiment in D. melanogaster showed that a reversal brake can inactivate a gene drive with a high efficiency (> 90%; Wu et al. 2016). Both immunizing reversal drives and reversal drives can in theory include a functional copy of the gene(s) disrupted by a prior suppression or an eradication drive, and thus have a fitness similar to that of the wild-type allele (Esvelt et al. 2014).
Transformation of a chromosome carrying a CRISPR-cas9 cassette into a chromosome carrying a reversal brake. a The reversal brake includes a gRNA recognizing a sequence of the cas9 gene on the gene drive cassette. b The Cas9 endonuclease produced by the gene drive allele can bind to the gRNA produced by the brake allele to recognize and cleave the cas9 gene. c The double-strand break is repaired by homology-directed repair using the brake chromosome as a template, resulting in the conversion of the gene drive allele into a brake allele
Brakes are not a perfect cure against drives: the presence of a drive allele in a population during the time between brake release and drive elimination can have long-lasting effects on the recovered populations, including inbreeding depression due to a temporary decrease in population size or to the presence of off-target mutations. Moreover, countermeasures against rescue drives are likely to fail, as drive alleles have a higher fitness than wild-type alleles.
A recent theoretical study shows that the release of brake-carrying individuals can lead to the fixation of the brake allele, the restoration of the wild-type allele or oscillations around a polymorphic equilibrium where both wild-type, gene drive and brake alleles are maintained through time (Vella et al. 2017). The polymorphic equilibrium is observed when the brake allele has a lower fitness than the wild-type allele. Overall, immunizing reversal drives are better at stopping an ongoing gene drive than reversal drives, as they target both the wild-type and drive alleles (Vella et al. 2017). However, once the drive is eliminated, the population still contains a copy of the cas9 gene and the continued presence of the Cas9 protein can increase the risks of potential negative off-target mutations (Gantz and Bier 2016; see above). Populations fixed for a reversal drive are also genetically modified, as they express the gRNA directed against the cas9 gene, but they do not carry a cas9 gene. Finally, the probability of stochastic elimination of an ongoing gene drive decreases with the cost of the brake allele (Vella et al. 2017). Calvez et al. (2018) studied the spatial spread of both drive and brake alleles. The brake allele can catch up with the drive allele and remove it from the population if the fitness reduction due to the drive allele is strong enough. When a drive has milder negative effects on fitness, it cannot be stopped and keeps spreading spatially.
Guidelines for gene drive usage in conservation
Informed decision-making
Due to their potential low costs and large scale of action compared to other biocontrol methods, gene drives have been considered by some authors as multi-purpose silver bullets in conservation biology, agriculture and public health (Esvelt et al. 2014). Because their implementation could have far-reaching unintended consequences and trigger irremediable modification of the natural environment, other authors (e.g. Webber et al. 2015) pointed that gene drives also pose strong conservation threats. The long history of successes and failures in classical biological control can help making several recommendations for gene drive research. In the USA, the National Academies have issued guidelines for gene drive research (NASEM 2016). At the international level, decision 14/19 of the United Nations Convention on Biological Diversity (CBD; https://www.cbd.int/doc/decisions/cop-14/cop-14-dec-19-en.pdf) highlights the need of a case-by-case risk assessment to minimize potential adverse effects and the importance of obtaining the informed consent of local communities that could be impacted. The assessment of biodiversity conservation and synthetic biology under International Union for Conservation of Nature (IUCN) resolution 6.086 (https://portals.iucn.org/library/node/46503) should also be published soon after the publication of this article (http://www.iucn.org/synbio).
Whether gene drives should be added to the conservation toolkit to protect endangered species or ecosystems depends on their added value relative to alternative strategies. The field of synthetic biology is moving fast and conservation geneticists might be unaware of the most recent alternative strategies (Phelps et al. 2019), or of strategies that are becoming less cost-prohibitive (Kandul et al. 2019). Previous experience with failed classical biocontrol strategies can also provide valuable information regarding the relevance of using gene drives as a last resort solution. Gould (2008), the NASEM (2016) and Moro et al. (2018) provide comprehensive recommendations to fill existing knowledge gaps and reduce the risks of implementing gene drives.
Overall, three types of information about the target and non-target species are required before implementing a gene drive strategy:
-
Genetic and technical information needed includes how to breed and conduct controlled experiments in the target species. Gene drive research also requires the availability of genome editing technology in the focal species or a related species, and the availability of an annotated reference genome to identify potential targets and design gRNAs that are specific of these loci (Moro et al. 2018).
-
Ecological information needed includes behavioral and demographic data (e.g. spatio-temporal variation in size; Moro et al. 2018), and a good understanding of the mating system and of gene flow between populations (e.g. quantifying dispersal ability as well as anthropogenic dispersal; Webber et al. 2015). Spatially explicit theoretical models can help predict gene drive dynamics.
-
Ecological and evolutionary data on potential non-target species includes quantification of gene flow between target and non-target species (hybridization or horizontal gene transfer), checking for the presence of potential target sites in non-target species, and appropriate modeling of food web structure to forecast long-term ecosystem impacts (Moro et al. 2018).
Biosafety and gene drive research
The unintentional release of gene drive individuals in the environment can represent an important risk for the safety of humans and their environment. Best practice guidelines have been proposed by various groups of experts (NASEM 2016; Lunshof and Birnbaum 2017; Krishnan and Gillum 2017; van der Vlugt et al. 2018). Gene drive strains should be managed using an appropriate combination of confinement strategies to mitigate these risks (Akbari et al. 2015):
-
Ecological confinement, by conducting gene drive research in countries where the target species is not present and cannot establish in the wild;
-
Physical containment, by using physical barriers (e.g. nets, secured lab facilities, etc.) or animal anesthesia;
-
Reproductive confinement, by using lab strains that cannot reproduce with wild individuals (e.g. Drosophila strain with chromosomal rearrangements; Akbari et al. 2015);
-
Molecular confinement, by using split gene drives with cas9 gene and gRNA on different chromosomes, or gene drive targeting an artificial sequence (DiCarlo et al. 2015; Champer et al. 2019);
-
Molecular identification, by tagging gene drive strains with specific phenotypic markers with dominant expression (Benedict et al. 2018).
There are currently no guidelines for the transport of gene drive strains, and some researchers have suggested that they should not be distributed to other laboratories (Akbari et al. 2015). For split gene drives, strains carrying the gRNA could be kept and distributed separately from the strains carrying the cas9 gene. The safety of gene drive research projects should be assessed by independent experts (e.g. institutional biosafety committees; Benedict et al. 2018). Funding agencies should ensure that appropriate guidelines are followed and enforced when necessary. Finally, we believe that a broad national and international consensus is required before carrying on deliberate release in controlled field trials, provided the high risks of propagation to wild populations.
Ethical and regulatory issues
Besides identifying possible risks, regulating gene drives requires ethical principles considering both human social values and non-human environmental values (NASEM 2016). Altering an organism or the environment poses ethical questions and can result in important risks for humans and ecosystems (Lunshof and Birnbaum 2017). Independent ethical committees are needed to help shaping the goals and justifications of gene drive research projects.
Scientists should be socially responsible for informing lawmakers and engaging with the “various publics that will use, be affected by, take an interest in, benefit from or be at risk from gene drives” (Thompson 2018). Such engagement is key so that stakeholders and local communities can make informed decisions, considering both the benefits and risks associated with gene drives as well as potential alternatives to the genetic engineering of wild populations.
Given the high risks of propagation of gene drive individuals across borders, there is a pressing need to build a strong international regulatory framework. As genetically modified organisms (hereafter GMOs) containing foreign pieces of DNA, gene drives are subject to GMO national and international regulation and their provisions. At the international level, GMOs are regulated under the 2003 Cartagena Protocol on Biosafety, a supplement to the Convention on Biodiversity (ratified by 167 nations with the exception of the United States of America and Canada; CBD 2003) and under two directives of the European Union on GMO legislation. National agencies have also issued more specific recommendations for the safe use of gene drives (e.g., Germany, ZKBS 2016, USA; NASEM 2016; Australia; AAS 2017; France; HCB 2017; the Netherlands; RIVM 2018). However, as the technology is evolving rapidly, some of the international and national GMO regulatory frameworks need to be adapted to the specificities of gene drive organisms (Oye et al. 2014; van der Vlugt et al. 2018). In 2016, 160 civil society organizations called for a global moratorium on the development and release of the gene drive technology (ETC Group 2016).
Gene drive organisms can be seen as an efficient technology for population control but also as potential bioweapons (Gurwitz 2014). The recent $100-million program including gene drive research projects (“Safe Genes program”) funded by the American Defense Advanced Research Programs Agency might contribute to these concerns (Reeves et al. 2018). The debate about a potential use of gene drive technology requires the transparency of gene drive research programs (including their funding sources and an appropriate risk assessment) and a broad engagement of evolutionary biologists with the public (Oye et al. 2014; Meghani and Kuzma 2018; Kofler et al. 2018).
Conclusions
Potential applications of gene drive in conservation include the extirpation of invasive pest populations that threaten biodiversity and the introduction of beneficial mutations in endangered populations. We highlighted the peculiarities associated with rescue drives compared to suppression and eradication drives. Rescue drives are likely to have different dynamics (e.g. no risk of resistance evolution, but no known countermeasure to recover the wild-type population). Overall, evolutionary and conservation geneticists can help better assess environmental risks associated with gene drives using both experimental (primarily in the lab) and theoretical approaches. Conservation geneticists could identify candidate genes for gene drives, estimate gene flow between target and non-target populations or species using population genomics approaches, and develop custom demographic models for different drive scenarios. Finally, conservation ecologists could help design appropriate gene drive management policies by quantifying interaction networks, such as food web diversity, structure and functioning. We believe that it is essential that conservation geneticists and land managers develop an expertise on gene drive technologies to engage in the current debate regarding their potential applications. This engagement should help stakeholders, policymakers and the local communities make informed decisions regarding the use and regulation of gene drives.
References
AAS (2017) Synthetic gene drives in Australia: implications of emerging technologies. Australian Academy of Science. https://www.science.org.au/support/analysis/reports/synthetic-gene-drives-australia-implications-emerging-technologies
Akbari OS, Bellen HJ, Bier E et al (2015) Safeguarding gene drive experiments in the laboratory. Science 349:927–929
Alberternst B, Nawrath S, Starfinger U (2016) Biodiversity impacts of common ragweed. Julius-Kühn-Archiv. https://doi.org/10.5073/jka.2016.455.45
Andras JP, Kirk NL, Drew Harvell C (2011) Range-wide population genetic structure of Symbiodinium associated with the Caribbean Sea fan coral, Gorgonia ventalina. Mol Ecol 20:2525–2542
Aslan Y, Tadjuidje E, Zorn AM, Cha S-W (2017) High-efficiency non-mosaic CRISPR-mediated knock-in and indel mutation in F0 Xenopus. Development 144:2852–2858
Atkinson UAE (1973) Spread of the ship rat (Rattus r. rattus L.) III New Zealand. J R Soc N Z 3:457–472
Baker HG (1967) Support for Baker’s law—as a rule. Evolution 21:853–856
Barton HJ, Zeng K (2018) New methods for inferring the distribution of fitness effects for INDELs and SNPs. Mol Biol Evol 35:1536–1546
Bassett IJ, Crompton CW (1975) The biology of Canadian weeds: 11. Ambrosia artemisiifolia L. and A. psilostachya DC. Can J Plant Sci 55:463–476. https://doi.org/10.4141/cjps75-072
Benedict MQ, Burt A, Capurro ML et al (2018) Recommendations for laboratory containment and management of gene drive systems in arthropods. Vector-Borne Zoonotic Dis 18:2–13. https://doi.org/10.1089/vbz.2017.2121
Brandes D, Nitzsche J (2006) Biology, introduction, dispersal, and distribution of common ragweed (Ambrosia artemisiifolia L.) with special regard to Germany. Nachrichtenblatt-Dtsch Pflanzenschutzdienstes Braunschw 58:286–291
Brossard D, Belluck P, Gould F, Wirz CD (2019) Promises and perils of gene drives: navigating the communication of complex, post-normal science. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.1805874115
Bucharova A (2017) Assisted migration within species range ignores biotic interactions and lacks evidence: missing evidence for assisted migration. Restor Ecol 25:14–18. https://doi.org/10.1111/rec.12457
Bull JJ (2017) Lethal gene drive selects inbreeding. Evol Med Public Health 1:1–16. https://doi.org/10.1093/emph/eow030
Burt A, Trivers R (2006) Genes in conflict: the biology of selfish genetic elements. Harvard University Press, Cambridge
Calvez V, Débarre F, Girardin L (2018) Catch me if you can: a spatial model for a brake-driven gene drive reversal. ArXiv Prepr ArXiv181206641
Caplan AL, Parent B, Shen M, Plunkett C (2015) No time to waste—the ethical challenges created by CRISPR: CRISPR/Cas, being an efficient, simple, and cheap technology to edit the genome of any organism, raises many ethical and regulatory issues beyond the use to manipulate human germ line cells. EMBO Rep 16:1421–1426. https://doi.org/10.15252/embr.201541337
Caut S, Casanovas JG, Virgos E et al (2007) Rats dying for mice: modelling the competitor release effect. Austral Ecol 32:858–868
CBD (2003) Cartagena protocol on biosafety to the Convention on Biological Diversity. CBD, Rio de Janeiro
Champer J, Buchman A, Akbari OS (2016) Cheating evolution: engineering gene drives to manipulate the fate of wild populations. Nat Rev Genet 17:146–159. https://doi.org/10.1038/nrg.2015.34
Champer J, Reeves R, Oh SY et al (2017) Novel CRISPR/Cas9 gene drive constructs reveal insights into mechanisms of resistance allele formation and drive efficiency in genetically diverse populations. PLOS Genet 13:e1006796. https://doi.org/10.1371/journal.pgen.1006796
Champer J, Liu J, Oh SY et al (2018a) Reducing resistance allele formation in CRISPR gene drive. Proc Natl Acad Sci USA 115:5522–5527. https://doi.org/10.1073/pnas.1720354115
Champer J, Wen Z, Luthra A et al (2018b) Multiple loci of small effect confer wide variability in efficiency and resistance rate of CRISPR gene drive. bioRxiv. https://doi.org/10.1101/447615
Champer J, Chung J, Lee YL et al (2019) Molecular safeguarding of CRISPR gene drive experiments. eLife 8:e41439. https://doi.org/10.7554/eLife.41439
Chan Y-S, Naujoks DA, Huen DS, Russell S (2011) Insect population control by homing endonuclease-based gene drive: an evaluation in Drosophila melanogaster. Genetics 188(1):33–44
Chinen AA, Suzuki H, Aplin KP et al (2005) Preliminary genetic characterization of two lineages of black rats (Rattus rattus sensu lato) in Japan, with evidence for introgression at several localities. Genes Genet Syst 80:367–375
Cho Y, Qiu Y-L, Kuhlman P, Palmer JD (1998) Explosive invasion of plant mitochondria by a group I intron. Proc Natl Acad Sci USA 95:14244–14249
Clark JB, Kidwell MG (1997) A phylogenetic perspective on P transposable element evolution in Drosophila. Proc Natl Acad Sci USA 94:11428–11433
Coble HD, Williams FM, Ritter RL (1981) Common ragweed (Ambrosia artemisiifolia) interference in soybeans (Glycine max). Weed Sci 29:339–342
Courchamp F, Chapuis J-L, Pascal M (2003) Mammal invaders on islands: impact, control and control impact. Biol Rev 78:347–383
Courtier-Orgogozo V, Morizot B, Boëte C (2017) Agricultural pest control with CRISPR-based gene drive: time for public debate: should we use gene drive for pest control? EMBO Rep 18:878–880. https://doi.org/10.15252/embr.201744205
David AS, Kaser JM, Morey AC et al (2013) Release of genetically engineered insects: a framework to identify potential ecological effects. Ecol Evol 3:4000–4015. https://doi.org/10.1002/ece3.737
David P, Thébault E, Anneville O et al (2017) Impacts of invasive species on food webs. In: Bohan DA, Dumbrell AJ, Massol F (eds) Advances in ecological research. Elsevier, Amsterdam, pp 1–60
Davis AW, Roote J, Morley T et al (1996) Rescue of hybrid sterility in crosses between D. melanogaster and D. simulans. Nature 380:157
Dearden PK, Gemmell NJ, Mercier OR et al (2018) The potential for the use of gene drives for pest control in New Zealand: a perspective. J R Soc N Z 48:225–244. https://doi.org/10.1080/03036758.2017.1385030
Deredec A, Burt A, Godfray HCJ (2008) The population genetics of using homing endonuclease genes in vector and pest management. Genetics 179:2013–2026. https://doi.org/10.1534/genetics.108.089037
Dhole S, Loyd AL, Gould F (2018a) Tethered homing gene drives: a new design for spatially restricted population replacement and suppression. bioRxiv. https://doi.org/10.1101/457564
Dhole S, Vella MR, Lloyd AL, Gould F (2018b) Invasion and migration of spatially self-limiting gene drives: a comparative analysis. Evol Appl 11:794–808. https://doi.org/10.1111/eva.12583
DiCarlo JE, Chavez A, Dietz SL et al (2015) Safeguarding CRISPR-Cas9 gene drives in yeast. Nat Biotechnol 33:1250–1255. https://doi.org/10.1038/nbt.3412
Dong Y, Simões ML, Marois E, Dimopoulos G (2018) CRISPR/Cas9 -mediated gene knockout of Anopheles gambiae FREP1 suppresses malaria parasite infection. PLOS Pathog 14:e1006898. https://doi.org/10.1371/journal.ppat.1006898
Drury DW, Dapper AL, Siniard DJ et al (2017) CRISPR/Cas9 gene drives in genetically variable and nonrandomly mating wild populations. Sci Adv 3(5):e1601910
Esvelt KM, Gemmell NJ (2017) Conservation demands safe gene drive. PLOS Biol 15:e2003850. https://doi.org/10.1371/journal.pbio.2003850
Esvelt KM, Smidler AL, Catteruccia F, Church GM (2014) Emerging technology: concerning RNA-guided gene drives for the alteration of wild populations. Elife 3:e03401
ETC Group (2016) Common call for a global moratorium on genetically-engineered gene drives. Action Group on Erosion, Technology and Concentration. http://www.synbiowatch.org/gene-drives/gene-drives-moratorium/
Eyre-Walker A, Keightley PD (2007) The distribution of fitness effects of new mutations. Nat Rev Genet 8:610
Fisher MC, Garner TWJ, Walker SF (2009) Global Emergence of Batrachochytrium dendrobatidis and Amphibian Chytridiomycosis in space, time, and host. Annu Rev Microbiol 63:291–310. https://doi.org/10.1146/annurev.micro.091208.073435
Friedman J, Barrett SC (2008) High outcrossing in the annual colonizing species Ambrosia artemisiifolia (Asteraceae). Ann Bot 101:1303–1309
Fu M, Waldman B (2017) Major histocompatibility complex variation and the evolution of resistance to amphibian chytridiomycosis. Immunogenetics 69:529–536. https://doi.org/10.1007/s00251-017-1008-4
Gandhi S, Haeussler M, Razy-Krajka F et al (2017) Evaluation and rational design of guide RNAs for efficient CRISPR/Cas9-mediated mutagenesis in Ciona. Dev Biol 425:8–20
Gantz VM, Bier E (2016) The dawn of active genetics. BioEssays 38:50–63. https://doi.org/10.1002/bies.201500102
Gantz VM, Jasinskiene N, Tatarenkova O et al (2015) Highly efficient Cas9-mediated gene drive for population modification of the malaria vector mosquito Anopheles stephensi. Proc Natl Acad Sci USA 112:E6736–E6743. https://doi.org/10.1073/pnas.1521077112
Gilbert C, Schaack S, Pace IIJK et al (2010) A role for host–parasite interactions in the horizontal transfer of transposons across phyla. Nature 464:1347
Gilbert C, Chateigner A, Ernenwein L et al (2014) Population genomics supports baculoviruses as vectors of horizontal transfer of insect transposons. Nat Commun 5:3348
Goddard MR, Burt A (1999) Recurrent invasion and extinction of a selfish gene. Proc Nat Acad Sci 96(24):13880–13885
Godfray HCJ, North A, Burt A (2017) How driving endonuclease genes can be used to combat pests and disease vectors. BMC Biol. https://doi.org/10.1186/s12915-017-0420-4
Gorbunova V, Levy AA (1999) How plants make ends meet: DNA double-strand break repair. Trends Plant Sci 4:263–269
Gorter de Vries AR, Couwenberg LGF, van den Broek M et al (2018) Allele-specific genome editing using CRISPR-Cas9 is associated with loss of heterozygosity in diploid yeast. Nucleic Acids Res. https://doi.org/10.1093/nar/gky1216
Gould F (2008) Broadening the application of evolutionarily based genetic pest management. Evol Int J Org Evol 62:500–510
Grunwald HA, Gantz VM, Poplawski G et al (2019) Super-Mendelian inheritance mediated by CRISPR–Cas9 in the female mouse germline. Nature. https://doi.org/10.1038/s41586-019-0875-2
Gurwitz D (2014) Gene drives raise dual-use concerns. Science 345:1010–1010
Hall SS (2017) Could genetic engineering save the Galápagos? Sci Am 317:48–57
Hammond AM, Kyrou K, Bruttini M et al (2017) The creation and selection of mutations resistant to a gene drive over multiple generations in the malaria mosquito. PLOS Genet 13:e1007039. https://doi.org/10.1371/journal.pgen.1007039
Hammond AM, Kyrou K, Gribble M et al (2018) Improved CRISPR-based suppression gene drives mitigate resistance and impose a large reproductive load on laboratory-contained mosquito populations. bioRxiv. https://doi.org/10.1101/360339
Harvey-Samuel T, Ant T, Alphey L (2017) Towards the genetic control of invasive species. Biol Invasions 19:1683–1703. https://doi.org/10.1007/s10530-017-1384-6
HCB (2017) Avis relatif à l’utilisation de moustiques génétiquement modifiés dans le cadre de la lutte antivectorielle. Haut Conseil des Biotechnologies. http://www.hautconseildesbiotechnologies.fr/fr/actualite/publication-lavis-sur-moustiques-gm
Highsmith RC (1982) Reproduction by fragmentation in corals. Mar Ecol Prog Ser Oldendorf 7:207–226
Hill T, Schlötterer C, Betancourt AJ (2016) Hybrid dysgenesis in Drosophila simulans associated with a rapid invasion of the P-element. PLOS Genet 12:e1005920. https://doi.org/10.1371/journal.pgen.1005920
Ivancevic A, Kortschak D, Bertozzi T, Adelson D (2017) Re-evaluating inheritance in genome evolution: widespread transfer of LINEs between species. bioRxiv. https://doi.org/10.1101/106914
Jin YK, Lundgren P, Lutz A et al (2016) Genetic markers for antioxidant capacity in a reef-building coral. Sci Adv 2:e1500842
Jinek M, Chylinski K, Fonfara I et al (2012) A programmable dual-RNA—guided DNA endonuclease in adaptive bacterial immunity. Science 337(6069):816–821
Jones KC, De Voogt P (1999) Persistent organic pollutants (POPs): state of the science. Environ Pollut 100:209–221
Joutey NT, Bahafid W, Sayel H, El Ghachtouli N (2013) Biodegradation: involved microorganisms and genetically engineered microorganisms. In: Chamy R, Rosenkranz F (eds) Biodegradation-life of science. InTech, London, pp 289–320
Kandul NP, Liu J, Wu SL et al (2019) Transforming insect population control with precision guided sterile males with demonstration in flies. Nat Commun 10:84
KaramiNejadRanjbar M, Eckermann KN, Ahmed HM et al (2018) Consequences of resistance evolution in a Cas9-based sex-conversion suppression gene drive for insect pest management. Proc Natl Acad Sci USA 115(24):6189–6194
Kessler CC (2002) Eradication of feral goats and pigs and consequences for other biota on Sarigan Island, Commonwealth of the Northern Mariana Islands. In: Veitch CR, Clout MN (eds) Turning the tide: the eradication of invasive species. WorldConservation Union (IUCN), Species Survival Commission, InvasiveSpecies Specialist Group. IUCN, Gland, Switzerland, pp. 132–140
Kofler R, Hill T, Nolte V et al (2015) The recent invasion of natural Drosophila simulans populations by the P-element. Proc Natl Acad Sci USA 112(21):6659–6663
Kofler N, Collins JP, Kuzma J et al (2018) Editing nature: Local roots of global governance. Science 362:527. https://doi.org/10.1126/science.aat4612
Krishnan P, Gillum D (2017) Gene drive 101: a basic guidance resource for biosafety professionals. Appl Biosaf 22:181–184. https://doi.org/10.1177/1535676017731318
Kubisiak TL, Nelson CD, Staton ME et al (2013) A transcriptome-based genetic map of Chinese chestnut (Castanea mollissima) and identification of regions of segmental homology with peach (Prunus persica). Tree Genet Genomes 9:557–571
Kyrou K, Hammond AM, Galizi R et al (2018) A CRISPR–Cas9 gene drive targeting doublesex causes complete population suppression in caged Anopheles gambiae mosquitoes. Nat Biotechnol. https://doi.org/10.1038/nbt.4245
Lees RS, Gilles JR, Hendrichs J et al (2015) Back to the future: the sterile insect technique against mosquito disease vectors. Curr Opin Insect Sci 10:156–162
Leitschuh CM, Kanavy D, Backus GA et al (2018) Developing gene drive technologies to eradicate invasive rodents from islands. J Responsib Innov 5:S121–S138. https://doi.org/10.1080/23299460.2017.1365232
Li J-F, Norville JE, Aach J et al (2013) Multiplex and homologous recombination–mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat Biotechnol 31:688
Lounibos LP (2002) Invasions by insect vectors of human disease. Annu Rev Entomol 47:233–266
Lunshof JE, Birnbaum A (2017) Adaptive risk management of gene drive experiments: biosafety, biosecurity, and ethics. Appl Biosaf 22:97–103
Macias V, Ohm J, Rasgon J (2017) Gene drive for mosquito control: where did it come from and where are we headed? Int J Environ Res Public Health 14:1006. https://doi.org/10.3390/ijerph14091006
Marshall JM, Akbari OS (2018) Can CRISPR-based gene drive be confined in the wild? A question for molecular and population biology. ACS Chem Biol 13:424–430. https://doi.org/10.1021/acschembio.7b00923
McFarlane GR, Whitelaw CBA, Lillico SG (2018) CRISPR-based gene drives for pest control. Trends Biotechnol 36:130–133. https://doi.org/10.1016/j.tibtech.2017.10.001
Meghani Z, Kuzma J (2018) Regulating animals with gene drive systems: lessons from the regulatory assessment of a genetically engineered mosquito. J Responsible Innov 5:S203–S222. https://doi.org/10.1080/23299460.2017.1407912
Metzger MJ, Paynter AN, Siddall ME, Goff SP (2018) Horizontal transfer of retrotransposons between bivalves and other aquatic species of multiple phyla. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.1717227115
Min J, Noble C, Najjar D, Esvelt KM (2017) Daisyfield gene drive systems harness repeated genomic elements as a generational clock to limit spread. bioRxiv. https://doi.org/10.1101/104877
Min J, Smidler AL, Najjar D, Esvelt KM (2018) Harnessing gene drive. J Responsible Innov 5:S40–S65. https://doi.org/10.1080/23299460.2017.1415586
Moran NA, Jarvik T (2010) Lateral transfer of genes from fungi underlies carotenoid production in aphids. science 328:624–627
Moro D, Byrne M, Kennedy M et al (2018) Identifying knowledge gaps for gene drive research to control invasive animal species: The next CRISPR step. Glob Ecol Conserv 13:e00363
Murphy E, Bradfield P (1992) Change in diet of stoats following poisoning of rats in a New Zealand forest. N Z J Ecol 16:137–140
NASEM (2016) Gene drives on the horizon: advancing science, navigating uncertainty, and aligning research with public values. National Academies Press, Washington, D.C.
Neve P (2018) Gene drive systems: do they have a place in agricultural weed management?: Gene drive and weed management. Pest Manag Sci. https://doi.org/10.1002/ps.5137
Newhouse AE, Polin-McGuigan LD, Baier KA et al (2014) Transgenic American chestnuts show enhanced blight resistance and transmit the trait to T1 progeny. Plant Sci 228:88–97
Nijensohn SE, Schaberg PG, Hawley GJ, DeHayes DH (2005) Genetic subpopulation structuring and its implications in a mature eastern white pine stand. Can J For Res 35:1041–1052
Noble C, Min J, Olejarz J et al (2016) Daisy-chain gene drives for the alteration of local populations. bioRxiv. https://doi.org/10.1101/057307
Noble C, Olejarz J, Esvelt KM et al (2017) Evolutionary dynamics of CRISPR gene drives. Sci Adv 8:e1601964
Noble C, Adlam B, Church GM et al (2018) Current CRISPR gene drive systems are likely to be highly invasive in wild populations. eLife 7:e33423
Norton DA, Young LM, Byrom AE et al (2016) How do we restore New Zealand’s biological heritage by 2050? Ecol Manag Restor 17:170–179
Oberhofer G, Ivy T, Hay BA (2018) Behavior of homing endonuclease gene drives targeting genes required for viability or female fertility with multiplexed guide RNAs. Proc Natl Acad Sci USA 115:E9343. https://doi.org/10.1073/pnas.1805278115
Oye KA, Esvelt K, Appleton E et al (2014) Regulating gene drives. Science 345:626–628. https://doi.org/10.1126/science.1254287
Pachauri RK, Allen MR, Barros VR et al (2014) Climate change 2014: synthesis report. Contribution of Working Groups I, II and III to the fifth assessment report of the Intergovernmental Panel on Climate Change. IPCC
Pannell JR, Auld JR, Brandvain Y et al (2015) The scope of Baker’s law. New Phytol 208:656–667
Phelps MP, Seeb LW, Seeb JE (2019) Transforming ecology and conservation biology through genome editing. Conserv Biol. https://doi.org/10.1111/cobi.13292
Piaggio AJ, Segelbacher G, Seddon PJ et al (2017) Is it time for synthetic biodiversity conservation? Trends Ecol Evol 32:97–107. https://doi.org/10.1016/j.tree.2016.10.016
Preston CR, Flores C, Engels WR (2006) Age-dependent usage of double-strand-break repair pathways. Curr Biol 16:2009–2015
Prowse TAA, Cassey P, Ross JV et al (2017) Dodging silver bullets: good CRISPR gene-drive design is critical for eradicating exotic vertebrates. Proc R Soc B Biol Sci 284:20170799. https://doi.org/10.1098/rspb.2017.0799
Ramos ML, Altieri E, Bulos M, Sala CA (2013) Phenotypic characterization, genetic mapping and candidate gene analysis of a source conferring reduced plant height in sunflower. Theor Appl Genet 126:251–263. https://doi.org/10.1007/s00122-012-1978-4
Reeves RG, Voeneky S, Caetano-Anollés D et al (2018) Agricultural research, or a new bioweapon system? Science 362:35–37
Reichman JR, Watrud LS, Lee EH et al (2006) Establishment of transgenic herbicide-resistant creeping bentgrass (Agrostis stolonifera L.) in nonagronomic habitats. Mol Ecol 15(13):4243–4255
Remy S, Chenouard V, Tesson L et al (2017) Generation of gene-edited rats by delivery of CRISPR/Cas9 protein and donor DNA into intact zygotes using electroporation. Sci Rep 7:16554
Rex Consortium (2013) Heterogeneity of selection and the evolution of resistance. Trends Ecol Evol 28:110–118
Ritchie SA, van den Hurk AF, Smout MJ et al (2018) Mission accomplished? We need a guide to the ‘post release’world of Wolbachia for Aedes-borne disease control. Trends Parasitol 34(3):217–226
RIVM (2018) Risk assessment method for activities involving organisms with a gene drive under contained use. RIVM Report
Rode NO, Lievens EJP, Segard A et al (2013) Cryptic microsporidian parasites differentially affect invasive and native Artemia spp. Int J Parasitol 43:795–803. https://doi.org/10.1016/j.ijpara.2013.04.009
Sanchez-Puerta MV, Cho Y, Mower JP, Alverson AJ, Palmer JD (2008) Frequent, phylogenetically local horizontal transfer of the cox1 group I Intron in flowering plant mitochondria. Mol Biol Evol 25(8):1762–1777
Savage AE, Zamudio KR (2011) MHC genotypes associate with resistance to a frog-killing fungus. Proc Natl Acad Sci USA 108(40):16705–16710
Scharenberg AM, Stoddard BL, Monnat RJ, Nolan A (2016) Retargeting: an unrecognized consideration in endonuclease-based gene drive biology. bioRxiv. https://doi.org/10.1101/089946
Scott MJ, Gould F, Lorenzen M et al (2018) Agricultural production: assessment of the potential use of Cas9-mediated gene drive systems for agricultural pest control. J Responsible Innov 5:S98–S120. https://doi.org/10.1080/23299460.2017.1410343
Sharp NP, Agrawal AF (2016) Low genetic quality alters key dimensions of the mutational spectrum. PLoS Biol 14:e1002419
Shiels AB, Drake DR (2011) Are introduced rats (Rattus rattus) both seed predators and dispersers in Hawaii? Biol Invasions 13:883–894
Siddle HV, Kreiss A, Tovar C et al (2013) Reversible epigenetic down-regulation of MHC molecules by devil facial tumour disease illustrates immune escape by a contagious cancer. Proc Natl Acad Sci USA 110:5103–5108. https://doi.org/10.1073/pnas.1219920110
Smith M, Cecchi L, Skjøth CA et al (2013) Common ragweed: a threat to environmental health in Europe. Environ Int 61:115–126. https://doi.org/10.1016/j.envint.2013.08.005
Song Y, Endepols S, Klemann N et al (2011) Adaptive introgression of anticoagulant rodent poison resistance by hybridization between old world mice. Curr Biol 21:1296–1301. https://doi.org/10.1016/j.cub.2011.06.043
SynBioWatch (2016) A call for conservation with a conscience: no place for gene drives in conservation. http://www.synbiowatch.org/wp-content/uploads/2016/09/letter_vs_genedrives.pdf
Tang W, Newton RJ, Li C, Charles TM (2007) Enhanced stress tolerance in transgenic pine expressing the pepper CaPF1 gene is associated with the polyamine biosynthesis. Plant Cell Rep 26:115–124
Thompson PB (2018) The roles of ethics in gene drive research and governance. J Responsible Innov 5:S159–S179. https://doi.org/10.1080/23299460.2017.1415587
Thresher RE, Hayes K, Bax NJ et al (2014) Genetic control of invasive fish: technological options and its role in integrated pest management. Biol Invasions 16:1201–1216. https://doi.org/10.1007/s10530-013-0477-0
Towns DR (2009) Eradications as reverse invasions: lessons from Pacific rat (Rattus exulans) removals on New Zealand islands. Biol Invasions 11:1719–1733
Towns DR, Atkinson IA, Daugherty CH (2006) Have the harmful effects of introduced rats on islands been exaggerated? Biol Invasions 8:863–891
Tschirren B, Andersson M, Scherman K et al (2013) Polymorphisms at the innate immune receptor TLR2 are associated with Borrelia infection in a wild rodent population. Proc R Soc Lond B Biol Sci 280:20130364
Unckless RL, Messer PW, Connallon T, Clark AG (2015) Modeling the manipulation of natural populations by the mutagenic chain reaction. Genetics 201:425–431. https://doi.org/10.1534/genetics.115.177592
Unckless RL, Clark AG, Messer PW (2017) Evolution of resistance against CRISPR/Cas9 gene drive. Genetics 205:827–841. https://doi.org/10.1534/genetics.116.197285
van der Vlugt CJB, Brown DD, Lehmann K et al (2018) A framework for the risk assessment and management of gene drive technology in contained use. Appl Biosaf 23:25–31. https://doi.org/10.1177/1535676018755117
Vella MR, Gunning CE, Lloyd AL, Gould F (2017) Evaluating strategies for reversing CRISPR-Cas9 gene drives. Sci Rep https://doi.org/10.1038/s41598-017-10633-2
Vernes K, McGrath K (2009) Are introduced black rats (Rattus rattus) a functional replacement for mycophagous native rodents in fragmented forests? Fungal Ecol 2:145–148. https://doi.org/10.1016/j.funeco.2009.03.001
Webber BL, Raghu S, Edwards OR (2015) Opinion: Is CRISPR-based gene drive a biocontrol silver bullet or global conservation threat?: Fig. 1. Proc Natl Acad Sci USA 112:10565–10567. https://doi.org/10.1073/pnas.1514258112
Wu B, Luo L, Gao XJ (2016) Cas9-triggered chain ablation of cas9 as a gene drive brake. Nat Biotechnol 34:137–138. https://doi.org/10.1038/nbt.3444
Zavaleta ES, Hobbs RJ, Mooney HA (2001) Viewing invasive species removal in a whole-ecosystem context. Trends Ecol Evol 16:454–459
Zentner GE, Wade MJ (2017) The promise and peril of CRISPR gene drives: genetic variation and inbreeding may impede the propagation of gene drives based on the CRISPR genome editing technology. BioEssays 39:1700109. https://doi.org/10.1002/bies.201700109
ZKBS (2016) Position statement of the ZKBS on the classification of genetic engineering operations for the production and use of higher organisms using recombinant gene drive systems. Zentrale Kommission für die Biologische Sicherheit (Central Committee of Biological Safety)
Acknowledgements
We thank Sean Hoban for encouragements to prepare this review. We thank four anonymous reviewers, Sean Hoban, Ruth Hufbauer, Eric Marois and Renaud Vitalis for their comments that help improve previous versions of the manuscript. We also thank Clément Rabourdin for helping with artworks. Credits of pictures used with the permission of Ngā Manu Nature Images (rat) and under CC BY-NC-SA 2.0 (mosquito), CC BY-NC 4.0 (ragweed), and CC BY 2.0 (mouse and frog) licenses appear in the captions of Figs. 1 and 2. Funding for this project was provided by the CeMEB LabEx/University of Montpellier (ANR-10-LABX-04-01) to NOR, ANR-14-ACHN-0003 to FD, INRA to AE and DB, INRA-SPE (SUZUCTICID project) to NOR and AE and European Research Council (FP7/2007–2013 Grant Agreement No. 337579) to VCO.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Rode, N.O., Estoup, A., Bourguet, D. et al. Population management using gene drive: molecular design, models of spread dynamics and assessment of ecological risks. Conserv Genet 20, 671–690 (2019). https://doi.org/10.1007/s10592-019-01165-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-019-01165-5