Abstract
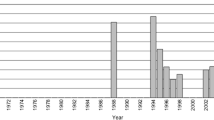
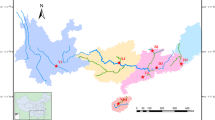
Japanese eel (Anguilla japonica) is an important food source in East Asia whose population has dramatically declined since the 1970s. Despite past analysis with DNA sequencing, microsatellite and isozyme methods, management decisions remain hampered by contradictory findings. For example, it remains unresolved whether Japanese eels are a single panmictic population or whether they harbor significant substructure. Accurate assessment of population genetic substructure, both spatial and temporal, is essential for determining the relevant number of distinct management units appropriate for this species. In the present study, we assayed genetic variation genome-wide using Restriction Site Associated DNA Sequencing (RAD-seq) technology to analyze the population genetic structure of Japanese eels. For analysis of temporal isolation, five “cohort” samples were collected yearly from 2005 to 2009 in the Yangtze River Estuary. For analysis of spatial structure, five “arrival wave” samples were collected in China in 2009, and two arrival wave samples were collected in Japan in 2001. In each cohort of each arrival wave, five individuals were collected for a total of 55 eels sampled. In total, 214,210 loci were identified from these individuals, 106,652 of which satisfied quality checks and were retained for further analysis. There was relatively little population differentiation between arrival waves and cohorts collected either at different locations during the same year (Fst = 0.077) or at the same location collected over subsequent years (Fst = 0.082), and locations displayed no consistent isolation-by-distance.






Similar content being viewed by others
References
Arai T, Kotake A, Lokman PM, Miller MJ, Tsukamoto K (2004) Evidence of different habitat use by New Zealand freshwater eels Anguilla australis and A. dieffenbachii, as revealed by otolith microchemistry. Mar Ecol Prog Ser 266:213–225. https://doi.org/10.3354/meps266213
Avise JC, Helfman GS, Saunders NC, Hales LS (1986) Mitochondrial DNA differentiation in North Atlantic eels: population genetic consequences of an unusual life history pattern. Proc Natl Acad Sci USA 83:4350–4354. https://doi.org/10.1073/pnas.83.12.4350
Baird NA, Etter PD, Atwood TS, Currey MC, Shiver AL, Lewis ZA, Selker EU, Cresko WA, Johnson EA (2008) Rapid SNP discovery and genetic mapping using sequenced RAD markers. PLoS ONE 3:1–7. https://doi.org/10.1371/journal.pone.0003376
Baltazar-Soares M, Biastoch A, Harrod C, Hanel R, Marohn L, Prigge E, Evans D, Bodles K, Behrens E, Böning CW, Eizaguirre C (2014) Recruitment collapse and population structure of the european eel shaped by local ocean current dynamics. Curr Biol 24:104–108. https://doi.org/10.1016/j.cub.2013.11.031
Catchen J, Hohenlohe PA, Bassham S, Amores A, Cresko WA (2013) Stacks: an analysis tool set for population genomics. Mol Ecol 22:3124–3140. https://doi.org/10.1111/mec.12354
Chan I, Chan D, Lee S, Tsukamoto K (1997) Genetic variability of the Japanese eel Anguilla japonica (Temminck & Schlegel) related to latitude. Ecol Freshw Fish 6:45–49. https://doi.org/10.1111/j.1600-0633.1997.tb00141.x
Chang K, Han Y, Tzeng W (2007) Population genetic structure among intra-annual arrival waves of the Japanese Eel. Zool Stud 46:583–590
Edmands S, Moberg PE, Burton RS (1996) Allozyme and mitochondrial DNA evidence of population subdivision in the purple sea urchin Strongylocentrotus purpuratus. Mar Biol 126:443–450. https://doi.org/10.1007/BF00354626
Gong X, Ren S, Cui Z, Yue L (2014) Genetic evidence for panmixia of Japanese eel (Anguilla japonica) populations in China. Genet Mol Res 13:768–781. https://doi.org/10.4238/2014.January.31.3
Han Y, Sun Y, Liao Y, Tzeng WN (2008) Temporal analysis of population genetic composition in the overexploited Japanese eel Anguilla japonica. Mar Biol 155:613–621. https://doi.org/10.1007/s00227-008-1057-1
Han Y, Hung C, Liao Y, Tzeng W (2010a) Population genetic structure of the Japanese eel Anguilla japonica: panmixia at spatial and temporal scales. Mar Ecol Prog Ser 401:221–232. https://doi.org/10.3354/meps08422
Han Y, Iizuka Y, Tzeng W (2010b) Does variable habitat usage by the Japanese eel lead to population genetic differentiation? Zool Stud 49:392–397
Hedgecock D (1994) Does variance in reproductive success limit effective population sizes of marine organisms? Genet Evol Aquat Org 122–134
Hellberg ME, Burton RS, Neigel JE, Palumbi SR (2002) Genetic assesment of connectivity among marine populations. Bull Mar Sci 70:273–290
Helyar SJ, Hemmer-Hansen J, Bekkevold D, Taylor MI, Ogden R, Limborg MT, Cariani A, Maes GE, Diopere E, Carvalho GR, NielEE (2011) Application of SNPs for population genetics of nonmodel organisms: New opportunities and challenges. Mol Ecol Resour 11:123–136. https://doi.org/10.1111/j.1755-0998.2010.02943.x
Hendry AP, Day T (2005) Population structure attributable to reproductive time: Isolation by time and adaptation by time. Mol Ecol 14:901–916. https://doi.org/10.1111/j.1365-294X.2005.02480.x
Henkel CV, Dirks RP, de Wijze DL, Minegishi Y, Aoyama J, Turner B, Knudsen BJ, Bundgaard M, Hvam K, Boetzer M, Pirovano W, Weltzien F, Dufour S, Tsukamoto K, Spaink HP, van den Thillart GE (2012) First draft genome sequence of the Japanese eel, Anguilla japonica. Gene 511:195–201. https://doi.org/10.1016/j.gene.2012.09.064
Ishikawa S, Aoyama J, Tsukamoto K, Nishida M (2001) Population structure of the Japanese eel Anguilla japonica as examined by mitochondrial DNA sequencing. Fish Sci 67:246–253
Johnson MS, Black R (1982) Chaotic genetic patchiness in an intertidal limpet, Siphonaria sp. Mar Biol 70:157–164. https://doi.org/10.1007/BF00397680
Kettle AJ, Haines K (2006) How does the European eel (Anguilla anguilla) retain its population structure during its larval migration across the North Atlantic Ocean? Can J Fish Aquat Sci 63:90–106. https://doi.org/10.1139/f05-198
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10:R25. https://doi.org/10.1186/gb-2009-10-3-r25
Maes GE, Pujolar JM, Hellemans B, Volckaert FAM (2006) Evidence for isolation by time in the European eel (Anguilla anguilla L.). Mol Ecol 15:2095–2107. https://doi.org/10.1111/j.1365-294X.2006.02925.x
Mantel N (1967) The detection of disease clutersing and a generalized regression approach. Cancer Res 27:209–220
Miller MR, Dunham JP, Amores A, Cresko WA, Johnson EA (2007) Rapid and cost-effective polymorphism identification and genotyping using restriction site associated DNA (RAD) markers. Genome Res 17:240–248. https://doi.org/10.1101/gr.5681207
Moberg PE, Burton RS (2000) Genetic heterogeneity among adult and recruit red sea urchins, Strongylocentrotus franciscanus. Mar Biol 136:773–784. https://doi.org/10.1007/s002270000281
Paschou P, Ziv E, Burchard EG, Burchard FG, Choudhry S, Rodriguez-Cintron W, Mahoney MW, Drineas P (2007) PCA-correlated SNPs for structure identification in worldwide human populations. PLoS Genet 3:1672–1686. https://doi.org/10.1371/journal.pgen.0030160
Pujolar JM, Maes GE, Volckaert FAM (2007) Genetic and morphometric heterogeneity among recruits of the European eel, Anguilla anguilla. Mar Ecol Prog Ser 81:297–308. https://doi.org/10.3354/meps307209
Pujolar J, Daniele B, Andrello M, Capoccioni F, Ciccotti E, Leo GAD, Zane L (2011) Genetic patchiness in European eel adults evidenced by molecular genetics and population dynamics modelling. Mol Biol Evol 58:198–206. https://doi.org/10.1016/j.ympev.2010.11.019
Pujolar JM, Jacobsen MW, Frydenberg J, Als TD, Larsen PE, Maes GE, Zane L, Jian JB, Cheng L, Hansen MM (2013) A resource of genome-wide single-nucleotide polymorphisms generated by RAD tag sequencing in the critically endangered European eel. Mol Ecol Resour 13:706–714. https://doi.org/10.1111/1755-0998.12117
Pujolar JM, Jacobsen MW, Als TD, Frydenberg J, Magnussen E, Jónsson B, Jiang X, Cheng L, Bekkevold D, Maes GE, Bernatchez L, Hansen MM (2014) Assessing patterns of hybridization between North Atlantic eels using diagnostic single-nucleotide polymorphisms. Heredity 112:627–637. https://doi.org/10.1038/hdy.2013.145
Pujolar1 JM, Maes1 GE, Volckaert1 FAM (2006) Genetic patchiness among recruits in the European eel Anguilla anguilla. Mar Ecol Prog Ser 307:209–217
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, Maller J, Sklar P, de Bakker PIW, Daly MJ, Sham PC (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575. https://doi.org/10.1086/519795
Ragauskas A, Butkauskas D (2013) The formation of the population genetic structure of the European eel Anguilla anguilla (L.): a short review. Ekologija 59:143–154
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145:1219–1228
Sambrook J, Russell DW (2006) Purification of nucleic acids by extraction with phenol:chloroform. CSH Protoc. https://doi.org/10.1101/pdb.prot4455
Sang T, Chang H, Chen C, Hui C (1994) Population structure of the Japanese eel, Anguilla japonica. Mol Biol Evol 11:250–260
Tseng M, Tzeng W, Lee S (2006) Population genetic structure of the Japanese eel Anguilla japonica in the northwest Pacific Ocean: evidence of non-panmictic populations. Mar Ecol Prog Ser 308:221–230. https://doi.org/10.3354/meps308221
Tseng M, Tzeng W, Lee S (2008) Genetic differentiation of the Japanese Eel. Am Fish Soc Symp 58:59–69
Tsukamoto K (1992) Discovery of the spawning area for Japanese eel. Nature 356:789–791. https://doi.org/10.1038/356789a0
Tsukamoto K, Arai T (2001) Facultative catadromy of the eel Anguilla japonica between freshwater and seawater habitats. Mar Ecol Prog Ser 220:265–276. https://doi.org/10.3354/meps220265
Tsukamoto K, Umezawa A (1990) Early life history and oceanic migration of the eel Anguilla japonica. La mer 188–198
Tzeng W, Shiao J, Iizuka Y (2002) Use of otolith Sr:Ca ratios to study the riverine migratory behaviors of Japanese eel Anguilla japonica. Mar Ecol Prog Ser 245:213–221. https://doi.org/10.3354/meps245213
Weir BS (1996) Genetic data analysis II: methods for discrete population genetic data. Sinauer Assoc Sunderl MA 150–156. https://doi.org/10.1136/jmg.29.3.216
Willing EM, Dreyer C, van Oosterhout C (2012) Estimates of genetic differentiation measured by Fst do not necessarily require large sample sizes when using many SNP markers. PLoS ONE 7:1–7. https://doi.org/10.1371/journal.pone.0042649
Wirth T, Bernatchez L (2001) Genetic evidence against panmixia in the European eel. Nature 409:1037–1040. https://doi.org/10.1038/35059079
Acknowledgements
We sincerely thank Dr. Minhui Wang for assistance with data analysis. Research supported by the Cornell China Faculty Development Program,the National Natural Science Foundation of China (31201995), the China-ASEAN Maritime Cooperation Fund from Shanghai University, the Shanghai Municipal Agricultural Commission (No. 2013 2–2), the Shanghai Municipal Science and Technology Commission of Chongming (Grant No. 13231203504) and the Open Foundation of Engineering Research Centre of Modern Industrial Technology for Eels (No. RE201501), Ministry of Education. ERD is supported by NIH F32 DK109595.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gong, X., Davenport, E.R., Wang, D. et al. Lack of spatial and temporal genetic structure of Japanese eel (Anguilla japonica) populations. Conserv Genet 20, 467–475 (2019). https://doi.org/10.1007/s10592-019-01146-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-019-01146-8