Abstract
Understanding genetic structure and diversity underpins the management of isolated populations. Small populations confined to islands may require effective genetic management for population persistence due to inbreeding and reduced genetic diversity. Endemic to the offshore islands of mainland Mauritius, the Bojer’s skink (Gongylomorphus bojerii) has previously been managed as two genetic units due to divergence between populations to the north and south-east. In 2009, a few individuals were discovered on the south-eastern island Ile de la Passe (IDLP), an island within its former range where the species was believed to have gone extinct. This island was later supplemented with translocations from other south-eastern islands, but individual geographic origin and the genetic consequences of these translocations remains unknown. Demographic population history and translocation events were reconstructed using a suite of microsatellite markers and the mitochondrial DNA cytochrome b gene, assessing two northern and three south-eastern populations to infer the geographic origin of the IDLP individuals, and to assess the population’s genetic diversity and structure. Although IDLP showed significant differentiation from all other populations, all analyses indicated a south-eastern origin, possibly founded by a single gravid female. Subsequent translocations have so far failed to ameliorate IDLP’s genetic diversity. In addition, admixture analysis was used to track translocation and gene-flow in IDLP. We recommend using additional translocations from other south-eastern islands to further genetically support IDLP. Our study highlights the need to clarify cryptic population structure and utilise post-translocation genetic monitoring among similarly managed populations, particularly endangered island populations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Anthropogenic activity can alter the genetic structure of populations by creating new barriers to dispersal or eliminating obstacles previously preventing gene flow, leading to admixture (Crispo et al. 2011; Helmus et al. 2014). Translocations within a species’ former range have been advocated for maintaining genetic diversity within existing populations and for establishing new populations as buffers against localised extinctions (Griffith et al. 1989; Madsen et al. 1999; Tuberville et al. 2005). Understanding such levels of genetic diversity and population structure within and among isolated populations is essential for ensuring appropriate conservation and management decisions (Frankham et al. 2017). For example, admixture can be beneficial for populations; increasing genetic diversity, masking deleterious mutations and producing new genotypes, potentially leading to transgressive segregation and heterosis (Crispo et al. 2011; Verhoeven et al. 2011). Admixture into small, inbred populations may prevent inbreeding, thereby reducing the likelihood of extinction (Madsen et al. 1999; Hogg et al. 2006; Johnson et al. 2010; Heber et al. 2013; Bateson et al. 2014). Additionally, admixture has the potential to supplement recently introduced populations, or help populations survive in an environment where native genomes are not pre-adapted (Verhoeven et al. 2011).
In contrast, the mixing of locally adapted, isolated populations can have adverse consequences with the loss of locally adapted allele combinations, breakdown of co-adapted gene complexes or the reintroduction of genetic load which had previously been purged (Frankham 2015). Populations can suffer from hybrid breakdown, genomic extinction and outbreeding depression (Rhymer and Simberloff 1996; Crispo et al. 2011; Verhoeven et al. 2011). In such cases, conservation managers predominantly choose to prevent the mixing of genetically distinct types (Moritz 1999; Allendorf et al. 2010; Weeks et al. 2011); instead populations of the same genetic provenance are considered as more appropriate stock when conducting translocations (IUCN 2013). Nevertheless, mixed populations may result when previous translocations have been poorly documented and/or populations have not been thoroughly characterised (Latch et al. 2006). Similarly, accidental translocations within a species’ range can be difficult to distinguish from natural migrations and may remain undetected, resulting in admixture and introgression (Michaelides et al. 2013) and impacting long-term population viability and future population management. The genetic risks associated with mixing populations should be carefully weighed against the benefits, therefore when planning and evaluating translocation programs, careful consideration of genetic structure has become a necessity (IUCN 2013).
Genetic analyses reveal patterns of diversity and structure within and among populations, allowing the prediction and identification of admixture pre- and post-translocation. In the case of accidental and poorly documented translocations, previous studies have attempted to identify the origin of the introduced population, for example when differentiating natural range expansions and translocations in the red fox (Vulpes vulpes; Statham et al. 2012), in characterising the subspecies structure of the cryptically distributed wild turkey (Meleagris gallopavo; Latch et al. 2006), and identifying source populations for colonisation events in desert bighorn sheep (Ovis canadensis nelson; Epps et al. 2010). Similarly, genetics, when used in conjunction with demographic, behavioural and ecological data, can be particularly valuable in post-translocation monitoring to evaluate the success of a program. In order to improve population viability, translocations should aim to minimise inbreeding, and maximise genetic variation and effective population size (IUCN 2013).
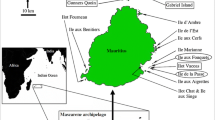
Investigating translocation history using genetic data is of particular importance for the future management of the Critically Endangered Bojer’s skink (Gongylomorphus bojerii) (Cole and Payne 2015) in order to monitor the outcome of reintroductions and augmentations of insular island populations. Bojer’s skinks are endemic to Mauritius and are restricted to a number of small offshore islands (Fig. 1), now only found to the north and south-east of the mainland (Jones 1993; Cole and Payne 2015). This is due to the arrival of humans on the Mascarenes in the seventeenth century and the subsequent appearance of commensal invasive species (Tonge 1990; Jones 1993). The species still faces a substantial threat of extinction and relies on active conservation management. Current efforts focus on translocating individuals among islands, augmenting existing populations, and reintroducing skinks onto neighbouring islands that were once part of their historical range. Bojer’s skinks are currently treated as two separate Management Units (MUs; northern and south-eastern) based on mitochondrial DNA (mtDNA) haplogroup divergence which differs by ~ 1.7% (Austin et al. 2009). The northern offshore islands feature one haplogroup; whereas the other haplogroup was described from Ilot Vacoas (IV) in the south-east (Austin et al. 2009). Subsequently, two additional populations were founded from IV individuals on the south-eastern islands Ile de la Passe (IDLP) and Ile aux Fouquets (IAF) where Bojer’s skinks existed until the 1930s and 1970s, respectively (Cole et al. 2009; Fig. 1). Ten individuals of each sex were translocated from IV to IAF in January 2007, and again in 2008 and 2010 (Cole et al. 2009; Michaelides et al. 2015). In 2009, a small population was found to have re-established on the small island of IDLP (approx. 200 m × 200 m), seemingly independent of human intervention, and were later supplemented with 16 individuals from IV and 20 individuals from IAF in 2013 (Cole et al. 2013). The discovered skink population on IDLP could represent a relic of the historic population that survived the presence of invasive musk shrews, although this is unlikely given the rapid extirpation of a considerably larger skink population by musk shrews elsewhere (Cole and Payne 2015). Furthermore, IDLP is topographically simplistic with sparse and degraded habitat, such that the highly active and inquisitive diurnal skink would have been detected during extensive herpetofaunal surveys of the small islet from the 1960s to 2008 (Vinson and Vinson 1969; Tonge 1990; Jones 1993; Freeman 2003; Cole 2005; Cole et al. 2009). The origin of the individuals occupying the island in 2009 is therefore unknown and there are currently no data on mtDNA diversity on IDLP, an issue of particular relevance considering the impacts of tourism on the offshore islets of Mauritius and the biosecurity risks that pertain with regular movement of people and boats (Cole et al. 2013).
Distribution of Bojer’s skinks on Mauritian offshore islands. The number of individuals translocated during the re-establishment and supplementation of populations on Ile de la Passe and Ile aux Fouquets are given and the direction of movement is depicted by arrows. Flat Island was included in this study but has since been eradicated in 2011. (Color figure online)
Evaluating the management actions implemented so far and future management options for Bojer’s skinks, particularly in terms of translocation, requires an accurate assessment of the genetic status of the island populations. We used molecular tools to analyse the genetic diversity and differentiation among Bojer’s skink insular populations. In particular, we aimed to identify and quantify any admixture between the 2009 individuals and subsequent deliberate translocations to IDLP, and assess the population’s genetic diversity, following the addition of individuals into this small, genetically uncharacterised recipient population.
Materials and methods
A total of 254 Bojer’s skinks were sampled across five Mauritian island populations between 2008 and 2016 (Table S1). These included the remaining northern and south-eastern populations on Round Island (RI) and IV, re-introduced south-eastern populations on IAF and IDLP, and the recently extirpated northern population from Flat Island (FI). Tissue samples were obtained with permission of the Government of Mauritius following ethical review by the Durrell Wildlife Conservation Trust by removing the tail tip (< 1 cm) during routine monitoring of populations by the Mauritius Reptile Recovery Programme. All specimens were caught by hand during three capture periods on IAF and IDLP, and one capture period on all other islands. All individuals were photographed and identified by their dorsal pattern to ensure no skink was sampled twice. Tail tips were stored in 99% ethanol and frozen prior to DNA extraction. Samples were imported into the United Kingdom under the conditions of a general permit IMP/GEN/2014/06 issued to the Durrell Wildlife Conservation Trust (Jersey, Channel Islands). DNA was extracted from tail tips following a HotSHOT extraction protocol (Truett et al. 2000). Briefly, 0.5 cm of tissue was added to 100 µL lysis solution (25 mM NaOH, 0.2 mM Na2 EDTA, pH 12.0) and incubated at 95 °C for 4–5 h. After cooling, the equivalent volume of neutralising solution (40 mM Tris–HCL, pH 5.0) was added, before DNA samples were stored at − 20 °C.
Microsatellite genotyping
A suite of polymorphic microsatellite markers were used for genotyping (Table S2), all of which were previously developed for Gongylomorphus skinks (Austin, unpublished data; Dawson and Freeman, unpublished data; Freeman 2003). For all downstream statistical analyses we used 14 microsatellites, except in the case of admixture and relatedness analyses, and to assess private alleles. We performed these analyses using nine loci because admixture and relatedness were assessed among south-eastern populations only, and the majority of samples from these populations were genotyped at nine loci. For all PCRs, forward and reverse primers (0.2 µM each) were multiplexed (four multiplex reactions given in Table S2) and combined with 0.1 µL of 20 mg/mL Bovine Serum Albumin (BSA), 5 µL of QIAGEN Multiplex PCR MasterMix and 1 µL (50–100 ng/µL) of extracted DNA, in a final volume of 10 µL. For multiplexes GongA and GongB, the cycling conditions were modified from Dawson and Freeman (unpublished data): 95 °C for 15 min, 40 cycles of 94 °C for 60 s, 60 °C for 45 s, 72 °C for 60 s followed with 72 °C for 30 min, and for multiplexes GBC and GBD, conditions were modified from Michaelides et al. (2015): 95 °C for 15 min, 17 touchdown cycles of 94 °C for 30 s, 60 °C for 30 s and 72 °C for 30 s, with the annealing temperature (60 °C) decreasing by 0.7 °C each cycle, followed with 30 cycles of 94 °C for 30 s, 48 °C for 30 s and 72 °C for 45 s, and 72 °C for 20 min (Table S2). PCRs were performed using a GeneAmp™ 9700 thermal cycler (Applied Biosystems). PCR fragments were analysed at MRC Protein Phosphorylation and Ubiquitylation (PPU) DNA Sequencing and Services, Dundee. Allele sizes were assigned using GeneMarker v.2.7.0 (SoftGenetics). Individuals missing > 20% data following re-amplification were excluded from further analyses. Data were checked for evidence of null alleles, large allele dropout or stutter peaks using MICROCHECKER v.2.2.3 (Van Oosterhout et al. 2004), and deviations from Hardy–Weinberg equilibrium (HWE) and linkage disequilibrium (LD) using GENEPOP v.4.6 (Raymond and Rousset 1995; Rousset 2008).
Mitochondrial DNA
A subset of 51 individuals including all islands were used as representatives for the mitochondrial analysis (Tables S1, S3). A fragment of the mitochondrial cytochrome b gene (~ 700 bp) was amplified using primers CBH-3 and L14841, following conditions described in Austin et al. (2009), with an initial activation step of 95 °C for 15 min. PCR was performed in 15 µL, including 5 µL of QIAGEN Multiplex PCR MasterMix, 0.1 µL of 20 mg/mL BSA, 0.33 µM of each primer and 50–100 ng/µL DNA. Forward and reverse sequences were aligned and trimmed to 642 bp using Geneious R.10.2.3 (Kearse et al. 2012) and all consensus sequences were aligned in ClustalX (Thompson et al. 1997).
Statistical analyses
Microsatellites
Indices of microsatellite genetic diversity were calculated using MSA v.4.05 (Dieringer and Schlötterer 2003), including the number of alleles (A), allelic richness (AR), observed (HO) and expected (HE) heterozygosity. Inbreeding coefficient (FIS) was estimated with FSTAT v.2.9.3.2 (Goudet 2001). The above statistics were calculated for IDLP based on temporal sampling in order to address the effects of translocations. Significant differences between populations were assessed using an ANOVA, followed by a Tukey’s HSD test in R v.3.3.2 (R Core Team 2016). The number of alleles unique to one or more populations were calculated across the nine loci successfully genotyped in at least 65% of samples from each population, and visualised as a Venn diagram. Pairwise FST values were estimated using FreeNA (Chapuis and Estoup 2007). As null alleles can lead to biased estimates of genetic differentiation (Dabrowski et al. 2014) and our dataset showed evidence of null alleles, we also used the ENA method of Chapuis and Estoup (2007) with 5000 bootstrap iterations to estimate unbiased FST. All analyses were performed twice, accounting for the presence and the absence of null alleles.
Bayesian clustering was used to infer the degree of genetic structure among populations based on multilocus microsatellite data using STRUCTURE v.2.3.4 (Pritchard et al. 2000). An admixture model was used with no location prior and allele frequencies were assumed to be correlated. One million MCMC iterations were run, following a burn-in period of 100,000 to determine the optimal number of clusters (K). The K-values considered ranged from 1 to 7, and 10 independent simulations were performed per value. Structure Harvester Web v.0.6.94 (Earl and vonHoldt 2012) was used to select the most appropriate value of K based on the posterior probability of the data for the given value of K (LnPr(X|K)) and Delta K (Evanno et al. 2005).
The likelihood of each individual being assigned to a given cluster or hybrid class was estimated using STRUCTURE and NewHybrids v.1.1 beta (Anderson and Thompson 2002), respectively. These analyses were limited to the south-eastern clusters (IAF/IV and IDLP). NewHybrids was used to assign individuals into six classes based on the proportion of their genetic material originating from each reference population (Table 1): ‘pure’ IDLP, ‘pure’ IAF/IV, first generation hybrids (F1), second generation hybrids (F2) and back-crossed hybrids (IAF/IV_Bx or IDLP_Bx). The Jeffreys-like prior was used due to the low allele frequencies observed in the populations compared (Anderson and Thompson 2002). We used a burn-in period of 100,000 steps and 1,000,000 MCMC iterations. The threshold value (Tq) defines the probability above which an individual can be assigned. HybridLab v.1.0 (Nielsen et al. 2006) was used to generate simulated datasets and identify the impact of changing Tq on efficiency, accuracy and overall performance of assignment (see Vähä and Primmer 2006). For assessing the performance of NewHybrids, all hybrid classes were averaged, to give a parameter summarising F1, F2 and backcrossed classes. From the 60 most ‘pure’ individuals of IDLP and IAF/IV, 100 genotypes were generated for each of the six hybrid classes (n = 600) and repeated 10 times. Each dataset was then analysed in STRUCTURE and NewHybrids under the same conditions as the empirical dataset, while accuracy, efficiency and overall performance were assessed using Tq values from 0.70 to 0.95. If an assignment probability for a given cluster or class surpassed Tq, the individual was assigned to that cluster or class, however, if all assignment probabilities for a given individual failed to surpass Tq for all clusters or classes, the individual was termed ‘admixed’ in STRUCTURE and ‘unassigned’ in NewHybrids.
To assess temporal changes in relatedness among individuals on IDLP, relatedness and likelihood-based approaches were taken. First, Queller and Goodnight’s relatedness (Queller and Goodnight 1989) was calculated using GenAlEx v.6.503 (Peakall and Smouse 2012). Pairwise relatedness (r) was estimated from observed allele frequencies and was used to categorise each dyad into first order (r ≥ 0.50), second order (0.25 ≤ r < 0.50), third order relatives (0.125 ≤ r < 0.25) or unrelated (r < 0.125). We also estimated the mean relatedness for each temporal sample. Secondly, sibships were inferred using COLONY v.2.0.6.1 (Wang and Santure 2009). This full-likelihood analysis was performed with no information on parental genotypes, assuming both male and female polygamy and without a sibship size prior distribution. The non-inbreeding model was selected as the low FIS value indicated no inbreeding in the IDLP population. Allele frequencies were estimated from the dataset. Five replicated runs of long length and very high precision were performed for each sample. Individuals were classified as full siblings, half siblings and unrelated pairs within the best configuration that maximises the probability of all relationships. These analyses were limited to nine loci because individuals sampled in 2009 and 2014, i.e. two of three sampling years analysed, were not genotyped at the five additional loci.
Msvar v.1.3 (Beaumont 1999; Storz and Beaumont 2002) was used to investigate signatures of recent demographic change. Each cluster identified in STRUCTURE was tested under three demographic scenarios performed with different prior distributions (Table S4): firstly the same prior for current (N0) and ancestral (N1) effective population sizes, i.e. a stable population; secondly a larger prior for N0 compared to N1, i.e. an expansion; and finally a larger prior for N1 compared to N0, i.e. a bottleneck. We parameterised the model using a generation time of 3 years (Cole and Payne 2015) and an average mutation rate of 10−4 for vertebrate microsatellites (Brohede et al. 2002; Bulut et al. 2009; Orozco-terWengel et al. 2013). Each Markov chain consisted of 67,770 simulations, of which the first 20% was disregarded to allow for burn-in. Convergence between the three independent chains per cluster was assessed with Brooks, Gelman and Rubin’s diagnostics (Brooks and Gelman 1998) using the boa package (Smith 2007) implemented in R v.3.3.2 (R Core Team 2016). Parameter medians (N0, N1 and t) and highest probability density limits (Alpha level = 0.1) were also assessed using boa.
Mitochondrial DNA
The mtDNA sequences were combined with 15 previously published haplotypes for this species (Table S5; Austin et al. 2009) and were aligned and trimmed in MEGA v.7 (Tamura et al. 2007). The aligned sequences were used to construct a haplotype network with 95% confidence limits, using the program TCS (Clement et al. 2000), which was superimposed onto a map to qualitatively investigate the geographic distribution of the haplotypes. Diversity indices such as haplotype diversity (Nei and Tajima 1981) and nucleotide diversity, π (Nei 1987) were calculated for the whole sample using DNAsp v.6.10.03 (Rozas et al. 2017). Alignment gaps and missing data were excluded from all calculations. In addition, the neutrality test of Tajima (Tajima 1989a, b, 1993) was performed to determine whether the pattern of sequence polymorphism in populations conformed to a neutral model and to test for a demographic population expansion or contraction. We also performed the Fu and Li’s F test (Fu 1997) to test the neutrality of the data. Both these tests were conducted in DNAsp v.6.10.03 (Rozas et al. 2017).
Results
Null alleles were detected in Gong2, GB06 and GB14. However, correcting for them in the dataset had no qualitative change to the FST results (ENA method) or downstream analyses, therefore results are presented for all 14 loci. While two loci (Gong2 and GB06) were identified as deviating significantly from HWE after Bonferroni correction (p < 7.14 × 10−4), this occurred in only one of five populations and therefore was most likely due to random variation. Allelic richness (ANOVA; F = 30.47, p < 0.001) and expected heterozygosity (ANOVA; F = 31.09, p < 0.001) differed significantly among populations (Table 2), showed significantly higher diversity in northern populations, and showed significantly lower heterozygosity in IDLP than all other islands (Tukey’s HSD test, p < 0.05). Significant FIS (p < 0.05) indicated some inbreeding in IAF (Table 2). Pairwise FST values indicated high and significant differentiation among populations (FST = 0.02–0.36, p < 0.01; Table 3), particularly when the north was compared with the south-east (FST = 0.24, p < 0.01). IDLP showed high differentiation from the other south-eastern populations (FST = 0.23–0.24, p < 0.01), although these values were lower than when IDLP was compared to northern populations.
STRUCTURE analysis identified the highest ΔK at K = 3, however the largest log-likelihood was K = 4 (Figs. S1, S2), with the fourth cluster featuring FI individuals, with scattered contribution from all other islands, thus K = 3 was considered more biologically meaningful. K = 3 separated the populations by geographic location, with IDLP identified as distinct from both the northern and south-eastern islands, which were also distinct from one another (Fig. 2).
Population clustering predicted by STRUCTURE at K = 3. Vertical bars represent individual genotypes, divided proportionately into coloured assignment probabilities to each cluster. White bars separate populations: RI Round Island, FI Flat Island, IV Ilot Vacoas, IAF Ile aux Fouquets, IDLP Ile de la Passe. Samples from IDLP are further split by year sampled. (Color figure online)
The simulated dataset was analysed in both STRUCTURE and NewHybrids to test efficiency, accuracy and overall performance of both programmes under a range of threshold values (0.70–0.95; Figs. S3, S4). In STRUCTURE, overall performance peaked at Tq = 0.85, reaching 79.9%. This Tq maximised both accuracy (89.0–97.4%), and efficiency (89.8–100%) for purely IAF/IV or IDLP individuals, and hybrids, within the simulated datasets. In NewHybrids, the overall performance continuously decreased as Tq increased. Accuracy for all groups, at all thresholds, did not fall below 95.7%, however the efficiency of identifying pure IDLP individuals ranged between 51.7 and 75.0%. This decreasing trend in efficiency resulted in a decreasing trend in overall performance, outweighing the impact of an increase in accuracy of 5%. When IDLP was removed, overall performance was highest at Tq = 0.85 at 95.8%. To avoid sacrificing accuracy for all groups for the efficiency of IDLP alone, a Tq value of 0.85 was used.
Of the 180 south-eastern individuals analysed, STRUCTURE identified 76 individuals of IAF/IV origin and 89 of IDLP origin (Table 4). The remaining 15 individuals, featuring intermediate q values (0.16–0.84), were assigned as admixed (Table 4). NewHybrids identified 73 pure IAF/IV individuals, 92 pure IDLP individuals, 5 F1 hybrids, and did not assign 10 individuals (Table 4). No F2 or backcrossed individuals were identified (Table 4). The temporal analysis of IDLP using STRUCTURE suggested an increase in admixed individuals from 11.1 to 13.1% between 2009 and 2016, and NewHybrids identified no hybrids in 2009 and 2014, but 5 (2.8%) F1 individuals in 2016 (Table 4). The majority of admixed individuals were identified from IDLP rather than IAF or IV, and these were predominantly F1 individuals (Fig. 3).
Assignments of all south-eastern Bojer’s skink samples (n = 180), as pure or admixed. a Plot of membership coefficients (q) of each individual belonging to both genetic clusters using STRUCTURE (K = 2). Tq = 0.85, as indicated by the dashed black lines at q = 0.85 and 0.15. b Plot of posterior probabilities (Q) of each individual belonging to six classes as identified using NewHybrids (Tq = 0.85). Each individual is represented by a vertical bar, split by colour to represent the likelihood of an individual belonging to one of the six categories: pure IAF/IV, pure IDLP, F1, F2, backcrossed to IDLP and backcrossed to IAF/IV. White bars separate populations: IV Ilot Vacoas, IAF Ile aux Fouquets, IDLP Ile de la Passe. Samples from IDLP are further split by year sampled. (Color figure online)
The percentage of relationships on IDLP categorised as Queller and Goodnight first and second order relatives was lowest in 2009, increased in 2014 and decreased again in 2016 (Table 5). The number of third order relatives increased from 2009 to 2016. Among 36 pairwise comparisons between individuals sampled in 2009, 11 (30.56%) were related. While among 435 comparisons from 2014 to 1830 comparisons from 2016, 212 (48.74%) and 766 (41.86%) pairs were related, respectively. Mean pairwise relatedness was − 0.119 (± 0.556), − 0.015 (± 0.750) and − 0.012 (± 0.524) in 2009, 2014 and 2016, respectively. From the optimal configuration estimated by COLONY, the percentage of dyads inferred as full siblings was highest in 2009 (5.56%), decreased in 2014 (1.15%) and increased slightly in 2016 (1.69%), whilst the percentage of dyads inferred as half siblings and overall sibships consistently decreased from 2009 to 2016: 38.89–19.56% and 44.44–21.26%, respectively (Table 6).
Msvar results suggested all inferred clusters experienced a reduction in their effective population size in the recent past (Table 7), with limited overlap in the posterior distributions for N0 and N1 (Fig. 4a–c). The effective sizes of the northern populations, IAF/IV and IDLP were inferred to have been reduced approximately 60-fold, 250-fold, and 15,000-fold, during bottlenecks estimated at occurring 80, 400 and 30 years ago respectively. Within clusters, each demographic model resulted in similar posterior distributions for the three tested parameters (N0, N1 and t; Fig. 4a–c).
Posterior distribution of parameters produced by Msvar. N0 = current effective population size, N1 = the ancestral effective population size, and T = the time since bottleneck in years. Each figure corresponds to a cluster identified by STRUCTURE; a IAF/IV, b North c IDLP. Solid lines represent the posterior distribution of parameters during three independent runs; a population expansion (green), a population bottleneck (blue), or a stable population (red). Dotted lines represent prior distributions. (Color figure online)
We obtained 615 bp of aligned mtDNA sequences from 66 Bojer’s skink samples, representing all seven extant populations in Mauritius. Nine haplotypes were identified, for which TCS connected alleles separated by up to six mutational steps (95% confidence limits). Two major haplogroups (A and B, Fig. 5) predominantly distinguished northern from south-eastern samples, however two samples from IV classified as haplotype B01, and one sample from IAF classified as haplotype B04, grouped with northern samples (Haplogroup A). Haplogroup A featured seven haplotypes with up to six mutational steps between them. In contrast, Haplogroup B featured only two haplotypes (of which one is extinct) separated by only two mutational steps. Seven of the nine haplotypes were found on just one island (Fig. 6). Only haplotype B01 was shared between the northern (RI, FI and Pigeon House Island) and south-eastern islands (IV). There were no northern haplotypes represented in IDLP, despite it having the largest sample size. The south-eastern islands showed low haplotype diversity with 40 of the 44 samples classified as haplotype B08 and this haplotype was the only haplotype present on all three islands. Haplotype diversity was highest on IV, lower on IAF and lower still on IDLP, with haplotype B08 becoming increasingly prominent. There were three haplotypes sampled from IAF; two were unique but were sampled from extinct populations (B04 and B09), and one was an extant haplotype from the most recent IAF samples (B08). When all 66 sequences were considered as one population, including the sequences downloaded from GenBank, the overall haplotype diversity was estimated at 0.60 and the nucleotide diversity was 1.09%. Both these values corresponded to values reported for other reptiles (Dutton et al. 1999; Rato et al. 2010). The Tajima D test for neutrality (0.88; Tajima 1989a, b, 1993) was not statistically significant (p > 0.10), suggesting that the cytochrome b gene is neutral. The positive value for the Tajima D test showed an overall signature of population contraction (Tajima 1989b), corresponding to the Msvar results. Likewise, the Fu and Li’s F test (0.81; Fu 1997) rejected the null hypothesis of neutrality (p < 0.10) and this test also suggested the presence of a population bottleneck.
TCS network based on 615 bp of the mitochondrial cytochrome b gene. The area of each circle is proportional to the number of samples, black circles represent unobserved haplotypes and the lines between each circle (coloured or black) represent a single nucleotide polymorphism. TCS only connects alleles with a 95% confidence limit, i.e. six steps. Extinct haplotypes are taken from Austin et al. (2009). (Color figure online)
The geographical distribution of mitochondrial haplotypes derived from this study and previously published sequences (Austin et al. 2009). Haplotypes are designated by colour and pie chart sizes are proportional to the number of individuals sampled from each island. Extinct haplotypes taken from Austin et al. (2009) are denoted by an asterisk. (Color figure online)
Discussion
Nuclear and mtDNA analyses detected three and two highly differentiated clusters among populations of Bojer’s skinks, respectively. Although divergent in allele and haplotype frequencies, the 2009 individuals from IDLP possessed the same allelic complement as the other south-eastern islands (Fig. S5), allowing us to infer a southern origin. Admixture could be observed among IDLP founders and translocated individuals, demonstrating successful integration. However, the translocations still need to be augmented because genetic diversity is lower in IDLP relative to its source population(s), likely caused by founder effect and genetic drift. These findings enable more informed strategies to ensure the long-term population viability of Bojer’s skinks on IDLP.
Similar to previous studies (Freeman 2003; Michaelides et al. 2015), nuclear DNA shows significantly lower genetic diversity in south-eastern compared to northern populations. Populations with low carrying capacities, and those subjected to demographic bottlenecks are expected to experience reductions in genetic variation due to genetic drift (MacArthur and Wilson 1963; England et al. 2003). As such, we would expect to find lower diversity among the smaller south-eastern islands of Mauritius (0.01–0.03 km2), compared to those in the north (2.19–2.53 km2). During sourcing for translocations, adequate levels of genetic variation must be retained in source populations and achieved in recipient populations to reduce the risk of inbreeding depression and loss of adaptive potential (IUCN 2013). The low levels of genetic variation exhibited by IDLP are therefore a concern for population viability, particularly when compared to its source populations of IV and IAF. Sampling years were not regularly spaced and therefore do not show all variation within the trend, complicating inference of the result of translocations on temporal changes in diversity for IDLP. Samples from IV represent genetic diversity in 2014, 1 year after translocation to IDLP (2013). When comparing our values to Michaelides et al. (2015), i.e. pre-translocation (HO = 0.62 ± 0.18, HE = 0.68 ± 0.13, A = 6.53 ± 3.24), the genetic diversity of this source population seems to have been unaffected by the translocation.
Bayesian clustering recapitulated the genetic differentiation between northern and south-eastern populations, corroborating previous microsatellite (Michaelides et al. 2015) and mtDNA (Austin et al. 2009) analyses, providing further support for the subdivision of Bojer’s skinks into two management units. High differentiation is also supported by pairwise FST values, little mitochondrial haplotype sharing and a large number of alleles unique to the northern populations (Fig. S5). Differentiation of northern and south-eastern genotypes is to be expected due to the absence of natural migration between these regions, and similar structure is commonly shown in island reptile populations (e.g. Ciofi and Bruford 1999; Ciofi et al. 2002; MacAvoy et al. 2007; Jordan and Snell 2008; Hurston et al. 2009). STRUCTURE analysis also showed differentiation between IDLP and the other south-eastern islands, assigning the population to a distinct cluster. Although highly differentiated, individuals from IDLP share more of their alleles with other south-eastern compared to northern populations, as indicated by pairwise FST values and the shared alleles Venn diagram (Fig. S5). Additionally, differentiation between IDLP and IAF/IV in nuclear DNA and the absence thereof in mitochondrial cytochrome b suggests the small re-established population in 2009 was likely derived from neighbouring south-eastern populations but has since suffered from founder effect and genetic drift. It has been documented in many instances that translocated populations can rapidly show a high degree of differentiation from their source, including the re-introduction of sea otters (Enhydra lutris; Larson et al. 2002), relocation of north American wapiti (Cervus elaphus; Polziehn et al. 2000), translocations of the Seychelles warbler (Acrocephalus sechellensis; Wright et al. 2014) and the bridled nailtail wallaby (Onychogalea fraenata; Sigg 2006). Although our data show translocations from IV to IAF were successful, supporting those from Michaelides et al. (2015), translocations to IDLP appear to have so far failed in creating a back-up population of similar viability.
The accidental introduction of the musk shrew (Suncus murinus) is considered responsible for the extirpation of Bojer’s skink populations in Mauritius, and this invasive species was eradicated from IDLP in 2000 as part of a program to reintroduce native reptile species to their former ranges (Varnham et al. 2002). Ile da la Passe is approximately 0.02 km2 in size and extensive surveys on the island until 2004 found no skinks present. While it is possible for small lizards to raft between islands even further apart than those south-east of Mauritius (Carranza et al. 2000; Calsbeek and Smith 2003), strong currents around these islands were expected to limit dispersal to IDLP. It is therefore more likely that an individual(s), possibly a single gravid female, was accidentally translocated to IDLP between 2004 and 2009 via touristic activities on the islands. Individuals found re-established on IDLP in 2009 show evidence that genetic differentiation from other populations had already accumulated before the deliberate translocations, probably only one or two generations after the population was founded.
Admixture was identified on IDLP using both STRUCTURE and NewHybrids in the 2016 sample, following translocations. NewHybrids showed an increase from 0 to 8.2% of samples identified as F1 admixed individuals just a single generation after translocation. Kennington et al. (2012) recorded a shift from one genetically distinct cluster to another within 8 generations following translocation, but differences in the proportion of individuals belonging to that cluster could be observed after just a single generation. The low levels of admixture on IDLP in 2016 is likely due to the short time period since translocations rather than a lack of breeding between genetically distinct individuals. Significant divergence of populations within a species, as seen on IDLP, provides an unusual scenario with which to evaluate the success of translocations by monitoring admixture, as illustrated during the supplementation of wild common hamster populations (Cricetus cricetus; Le Haye et al. 2017) and the management of the peregrine falcon (Falco peregrinus; Jacobsen et al. 2008). Admixture analyses can function as an indicator of population viability, for example after domestication of the honey bee (Apis mellifera; Harpur et al. 2012), and reintroductions of the Perdido Key beach mouse (Peromyscus polionotus trissyllepsis; Greene et al. 2017). Biebach and Keller (2012) identified the degree of admixture within a founding group to be a better predictor of final genetic variation than the number of founders in Alpine ibex (Capra ibex) populations, enabling efficient management strategies to be implemented. Assessing both the rate of admixture and the change in genetic diversity enables a greater insight into the impacts of the translocation and, in turn, enables improved subsequent management.
The terrestrial fauna on the islands of Mauritius remain at risk from invasive predators and competitors, and other human-induced impacts. Since sampling, the invasion of FI by the musk shrew has completely eradicated the largest population of Bojer’s skink (Cole and Payne 2015). All populations analysed in this study showed signatures of recent historical bottlenecks. Although the 95% HPDs are broad, the inferred bottleneck of IAF/IV approximately 400 years ago coincides well with the earliest documented arrival of humans and their commensal species. These bottlenecks have produced a small effective population size across the islands, particularly in IDLP where the current effective size is inferred to be less than unity (N0 = 0.5, 95% HPD = 0–34). The rate of genetic drift and the accumulation of inbreeding and loss of adaptive potential are ultimately linked to population size (Kalinowski and Waples 2002). Despite small effective population sizes, significant FIS values only indicated inbreeding in the IAF population. Conversely, relatedness among individuals on IDLP increased between 2009 and 2014, which is to be expected if the population was founded by few individuals and samples in 2014 were only taken 0.3 generations after the translocation. In 2016, one generation after the translocations, the overall percentage of individuals that were related decreased, demonstrating translocations have begun to alleviate some of the problems the population experienced before.
The mtDNA haplotype network suggests a general distinction between northern and south-eastern populations. Only haplotype B01 was shared between RI (north) and IV (south-east), supporting previous mitochondrial evidence (Austin et al. 2009) that there was once historical gene flow between the north and south-east which may have been possible due to transient land bridges that formed between the islands and the mainland during the last glacial maxima (Nichols and Freeman 2004). Post-glacial migration may have continued by skinks rafting from the islands to the mainland on debris, however this would have been at a low level and would have ceased around the early 1900s when Bojer’s skinks became extinct on the mainland (Freeman 2003; Austin et al. 2009; Cole et al. 2009). The star-like structure of Haplogroup A qualitatively suggests a population that may have undergone a demographic expansion followed by an extinction event, removing many alleles, as indicated by Austin et al. (2009). It is possible that some shared haplotypes between the north and south-east were among those that went extinct. However, while many of these results may represent genuine genetic processes and results of the translocation, it is difficult to rule out the role of sampling effort missing certain intermediate haplotypes. The mitochondrial data supports the successive loss of genetic diversity from IV to IAF to IDLP which is also represented in the microsatellite data. Haplotype B08 became increasingly prominent from IV to IAF to IDLP and was the only extant haplotype sampled on IDLP and IAF. This is most likely as a result of founder effect and genetic drift during translocation. Haplotype B01 is not present on either IDLP or IAF suggesting that translocated individuals did not represent the full genetic diversity of the population of origin.
In characterising the genetic structure among island populations of Bojer’s skink, we were able to infer with some confidence the origin of individuals on IDLP. While the observed patterns of diversity and structure could also result if a small population had always survived inconspicuously on IDLP, extensive surveys and monitoring on the offshore islands of Mauritius suggest this is not the case. Thus, while the IDLP population still shows signs of divergence at nuclear DNA and low diversity 3 years after translocations, managers now have the option to use future translocations from other south-eastern islands as a form of genetic rescue. However, monitoring admixture and diversity will be necessary to determine the long-term success of the translocations already implemented and inform future management strategies. To maintain unique northern and south-eastern diversity, we recommend the continuation of two management units in conservation planning. Populations of Bojer’s skink would also benefit from strict biosecurity regulations and continual removal of invasive species where possible. While this study focused on small island populations of Bojer’s skink, the same methodology can be applied to almost any species, enabling informed management strategies concerning the translocation of wildlife. Genetic data should be routinely used pre- and post-translocation alongside other success measures to determine the long-term viability of a translocated population.
References
Allendorf FW, Hohenlohe PA, Luikart G (2010) Genomics and the future of conservation genetics. Nat Rev Genet 11:697–709
Anderson EC, Thompson EA (2002) A model-based method for identifying species hybrids using multilocus genetic data. Genetics 160:1217–1229
Austin JJ, Arnold EN, Jones CG (2009) Interrelationships and history of the slit-eared skinks (Gongylomorphus, Scincidae) of the Mascarene Islands, based on mitochondrial DNA and nuclear gene sequences. Zootaxa 2153:55–68
Bateson ZW, Dunn PO, Hull SD, Henschen AE, Johnson JA, Whittingham LA (2014) Genetic restoration of a threatened population of greater prairie-chickens. Biol Conserv 174:12–19
Beaumont MA (1999) Detecting population expansion and decline using microsatellites. Genetics 153:2013–2029
Biebach I, Keller LF (2012) Genetic variation depends more on admixture than number of founders in reintroduced Alpine ibex populations. Biol Conserv 147:197–203
Brohede J, Primmer CR, Moller A, Ellegren H (2002) Heterogeneity in the rate and pattern of germline mutation at individual microsatellite loci. Nucleic Acids Res 30:1997–2003
Brooks SP, Gelman A (1998) General methods for monitoring convergence of iterative simulations. J Comput Graph Stat 7:434–455
Bulut Z, McCormick CR, Gopurenko D, Williams RN, Bos DH, DeWoody JA (2009) Microsatellite mutation rates in the eastern tiger salamander (Ambystoma tigrinum tigrinum) differ 10-fold across loci. Genetica 136:501–504
Calsbeek R, Smith TB (2003) Ocean currents mediate evolution in island lizards. Nature 426:552–555
Carranza S, Arnold EN, Mateo JA, Lopez-Jurado LF (2000) Long-distance colonization and radiation in gekkonid lizards, Tarentola (Reptilia: Gekkonidae), revealed by mitochondrial DNA sequences. Proc R Soc Lond B 267:637–649
Chapuis M, Estoup A (2007) Microsatellite null alleles and estimation of population differentiation. Mol Biol Evol 24:621–631
Ciofi C, Bruford MW (1999) Genetic structure and gene flow among Komodo dragon populations inferred by microsatellite loci analysis. Mol Ecol 8:17–30
Ciofi C, Milinkovitch MC, Gibbs JP, Caccone A, Powell JR (2002) Microsatellite analysis of genetic divergence among populations of giant Galápagos tortoises. Mol Ecol 11:2265–2283
Clement M, Posada D, Crandall K (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1660
Cole N, Payne C (2015) Gongylomorphus bojerii [Online] The IUCN red list of threatened species. http://www.iucnredlist.org/details/biblio/62251/0. Accessed 6 Oct 2017
Cole N (2005) The ecological impact of the invasive house gecko Hemidactylus frenatus upon endemic Mauritian geckos. PhD, University of Bristol, UK
Cole N, Jones C, Buckland S, Jhumka Z, Mootoocurpen R, Tatayah V, Bachraz V, Nundlaul V, Roopa P, Seewajee P (2009) The reintroduction of endangered Mauritian reptiles Mauritian Wildlife Foundation. Unpublished report, Vacoas, Mauritius
Cole N, Goder M, Vencatasamy D, Mootoocurpen R, Havery S, Gamble F, Nundlaul V (2013) Restoration of island ecosystems in Mauritius: the Mauritius Reptile Recovery Programme Annual Report 2013. Durrell Wildlife Conservation Trust, Jersey
Crispo E, Moore JS, Lee-Yaw JA, Gray SM, Haller BC (2011) Broken barriers: human-induced changes to gene flow and introgression in animals. Bioessays 33:508–518
Dabrowski MJ, Pilot M, Kruczyk M, Żmihorski M, Umer HM, Gliwicz J (2014) Reliability assessment of null allele detection: inconsistencies between and within different methods. Mol Ecol Resour 14:361–373
Dieringer D, Schlötterer C (2003) Microsatellite analyser (MSA): a platform independent analysis tool for large microsatellite data sets. Mol Ecol Notes 3:167–169
Dutton PH, Bowen BW, Owens DW, Barragan A, Davis SK (1999) Global phylogeography of the leatherback turtle (Dermochelys coriacea). J Zool 248:397–409
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
England P, Osler G, Woodworth L, Montgomery M, Briscoe D, Frankham R (2003) Effects of intense versus diffuse population bottlenecks on microsatellite genetic diversity and evolutionary potential. Conserv Genet 4:595–604
Epps CW, Wehausen JD, Palsboll PJ, McCullough DR (2010) Using genetic tools to track desert bighorn sheep colonizations. J Wildl Manag 74:522–531
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Frankham R (2015) Genetic rescue of small inbred populations: meta-analysis reveals large and consistent benefits of gene flow. Mol Ecol 24:2610–2618
Frankham R, Ballou JD, Ralls K, Eldridge M, Dudash MR, Fenster CB, Lacy RC, Sunnucks P (2017) Genetic management of fragmented animal and plant populations. Oxford University Press, Oxford
Freeman KL (2003) Ecology and conservation genetics of the Gongylomorphus genus in Mauritius. PhD, Queen Mary University, London
Fu YX (1997) Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147:915–925
Goudet J (2001) FSTAT, a program to estimate and test gene diversities and fixation indices (version 2.9.3). http://www2.unil.ch/popgen/softwares/fstat.htm
Greene DU, Gore JA, Austin JD (2017) Reintroduction of captive-born beach mice: the importance of demographic and genetic monitoring. J Mammal 98:513–522
Griffith B, Scott JM, Carpenter JW, Reed C (1989) Translocation as a species conservation tool: status and strategy. Science 245:477–480
Harpur BA, Minaei S, Kent CF, Zayed A (2012) Management increases genetic diversity of honey bees viaadmixture. Mol Ecol 21:4414–4421
Heber S, Varsani A, Kuhn S, Girg A, Kempenaers B, Briskie J (2013) The genetic rescue of two bottlenecked South Island robin populations using translocations of inbred donors. Proc R Soc B 280:20122228
Helmus MR, Mahler DL, Losos JB (2014) Island biogeography of the Anthropocene. Nature 513:543–546
Hogg JT, Forbes SH, Steele BM, Luikart G (2006) Genetic rescue of an insular population of large mammals. Proc R Soc B 273:1491–1499
Hurston H, Voith L, Bonanno J, Foufopoulos J, Pafilis P, Valakos E, Anthony N (2009) Effects of fragmentation on genetic diversity in island populations of the Aegean wall lizard Podarcis erhardii (Lacertidae, Reptilia). Mol Phylogenet Evol 52:395–405
IUCN (2013) Guidelines for reintroductions and other conservation translocations. IUCN/SSC, Gland
Johnson WE, Onorato DP, Roelke ME, Land ED, Cunningham M, Belden RC, Mcbride R, Jansen D, Lotz M, Shindle D, Howard J, Wildt DE, Penfold LM, Hostetler JA, Oli MK, O’Brien SJ (2010) Genetic restoration of the Florida panther. Science 329:1641–1645
Jones CG (1993) The ecology and conservation of Mauritian skinks. Proc R Soc Arts Sci Maurit 5:71–95
Jordan MA, Snell HL (2008) Historical fragmentation of islands and genetic drift in populations of Galápagos lava lizards (Microlophus albemarlensis complex). Mol Ecol 17:1224–1237
Kalinowski ST, Waples RS (2002) Relationship of effective to census size in fluctuating populations. Conserv Biol 16:129–136
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Mentjies P, Drummond A (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649
Kennington WJ, Hevroy TH, Johnson MS (2012) Long-term genetic monitoring reveals contrasting changes in the genetic composition of newly established populations of the intertidal snail Bembicium vittatum. Mol Ecol 21:3489–3500
Larson S, Jameson R, Bodkin J, Staedler M, Bentzen P (2002) Microsatellite DNA and mitochondrial DNA variation in remnant and translocated sea otter (Enhydra lutris) populations. J Mammal 83:893–906
Latch EK, Applegate RD, Rhodes OE Jr (2006) Genetic composition of wild turkeys in Kansas following decades of translocations. J Wildl Manag 70:1698–1703
Le Haye MJJ, Reiners TE, Raedts R, Verbist V, Koelewign HP (2017) Genetic monitoring to evaluate reintroduction attempts of a highly endangered rodent. Conserv Genet 18:877–892
MacArthur RH, Wilson EO (1963) Equilibrium theory of insular zoogeography. Evolution 17:373–387
MacAvoy E, McGibbon L, Sainsbury J, Lawrence H, Wilson C, Daugherty C, Chambers G (2007) Genetic variation in island populations of tuatara (Sphenodon spp.) inferred from microsatellite markers. Conserv Genet 8:305–318
Madsen T, Shine R, Olsson M, Wittzell H (1999) Conservation biology: restoration of an inbred adder population. Nature 402:34
Michaelides S, While GM, Bell C, Uller T (2013) Human introductions create opportunities for intra-specific hybridization in an alien lizard. Biol Invasions 15:1101–1112
Michaelides S, Cole N, Funk SM (2015) Translocation retains genetic diversity of a threatened endemic reptile in Mauritius. Conserv Genet 16:661–672
Moritz C (1999) Conservation units and translocations: strategies for conserving evolutionary processes. Hereditas 130:217–228
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Nei M, Tajima F (1981) DNA polymorphism detectable by restriction endonucleases. Genetics 97:145–163
Nichols RA, Freeman KLM (2004) Using molecular markers with high mutation rates to obtain estimates of relative population size and to distinguish the effects of gene flow and mutation: a demonstration using data from endemic Mauritian skinks. Mol Ecol 13:775–787
Nielsen EEG, Arvebach L, Kotlicki P (2006) HYBRIDLAB (version 1.0): a program for generating simulated hybrids from population samples. Mol Ecol Notes 6:971–971
Orozco-terWengel P, Andreone F, Louis E, Vences M (2013) Mitochondrial introgressive hybridization following a demographic expansion in the tomato frogs of Madagascar, genus Dyscophus. Mol Ecol 22:6074–6090
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetics software for teaching and research-an update. Bioinformatics 28:2537–2539
Polziehn RO, Hamr J, Mallory FF, Strobeck C (2000) Microsatellite analysis of North American wapiti (Cervus elaphus) populations. Mol Ecol 9:1561–1576
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Queller DC, Goodnight KF (1989) Estimating relatedness using genetic markers. Evolution 43:258–275
R Core Team (2016) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/
Rato C, Carranza S, Perera A, Carretero MA, Harris DJ (2010) Conflicting patterns of nucleotide diversity between mtDNA and nDNA in the Moorish gecko, Tarentola mauritanica. Mol Phylogenet Evol 56:962–971
Raymond M, Rousset F (1995) GENEPOP (Version 1.2): a population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Rhymer JM, Simberloff D (1996) Extinction by hybridization and introgression. Annu Rev Ecol Evol Syst 27:83–109
Rousset F (2008) Genepop’007: a complete re-implementation of the genepop software for Windows and Linux. Mol Ecol Resour 8:103–106
Rozas J, Ferrer-Mata A, Sánchez-DelBarrio JC, Guirao-Rico S, Librado P, Ramos-Onsins SE, Sánchez-Gracia A (2017) DnaSP v6: DNA sequence polymorphism analysis of large datasets. Mol Biol Evol 34:3299–3302
Sigg DP (2006) Reduced genetic diversity and significant genetic differentiation after translocation: comparison of the remnant and translocated populations of bridled nailtail wallabies (Onychogalea fraenata). Conserv Genet 7:577–589
Smith BJ (2007) boa: an R package for MCMC output convergence assessment and posterior inference. J Stat Softw 21:1–37
Statham MJ, Sacks BN, Aubry KB, Perrine JD, Wisely SM (2012) The origin of recently established red fox populations in the United States: translocations or natural range expansions? J Mammal 93:52–65
Storz JF, Beaumont MA (2002) Testing for genetic evidence of population expansion and contraction: an empirical analysis of microsatellite DNA variation using a hierarchical Bayesian model. Evolution 56:154–166
Tajima F (1989a) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Tajima F (1989b) The effect of change in population size on DNA polymorphism. Genetics 123:597–601
Tajima F (1993) Measurement of DNA polymorphism. In: Takahata N, Clark GA (eds) Mechanisms of molecular evolution. Introduction to molecular paleopopulation biology. Japan Scientific Societies Press, Sinaur Associates, Inc., Sunderland, pp 37–59
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thompson JD, Gibson TJ, Plewniak TF, Jeanmougin F, Higgins DG (1997) The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 24:4876–4882
Tonge S (1990) The past, present and future of the herpetofauna of Mauritius. Bull Chic Herpetol Soc 25:220–226
Truett GE, Heeger P, Mynatt RL, Truett AA, Walker JA, Warman ML (2000) Preparation of PCR-quality mouse genomic DNA with hot sodium hydroxide and Tris (HotSHOT). Biotechniques 29:52–54
Tuberville TD, Clark EE, Buhlmann KA, Gibbons JW (2005) Translocation as a conservation tool: site fidelity and movement of repatriated gopher tortoises (Gopherus polyphemus). Anim Conserv 8:349–358
Vähä JP, Primmer CR (2006) Efficiency of model-based Bayesian methods for detecting hybrid individuals under different hybridization scenarios and with different numbers of loci. Mol Ecol 15:63–72
Van Oosterhout C, Hutchinson WF, Wills DP, Shipley P (2004) Micro-checker: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Varnham KJ, Roy SS, Seymour A, Mauremootoo JR, Jones CG, Harris S (2002) Eradicating Indian musk shrews (Suncus murinus, Soricidae) from Mauritian offshore islands. In: Veitch CR, Clout MN (eds) Turning the tide: the eradication of invasive species. Invasive Species Specialist Group, Species Survival Commission, World Conservation Union, Gland, pp 342–349
Verhoeven KJF, Macle M, Wolfe LM, Biere A (2011) Population admixture, biological invasions and the balance between local adaptation and inbreeding depression. Proc R Soc B 278:2–8
Vinson J, Vinson JM (1969) The saurian fauna of the Mascarene Islands. Mauritius Inst Bull 6:203–320
Wang J, Santure AW (2009) Parentage and sibship inference from multilocus genotype data under polygamy. Genetics 181:1579–1594
Weeks AR, Sgro CM, Young AG, Frankham R, Mitchell NJ, Miller KA, Byrne M, Coates DJ, Eldridge MDB, Sunnucks P, Breed MF, James EA, Hoffmann AA (2011) Assessing the benefits and risks of translocations in changing environments: a genetic perspective. Evol Appl 4:709–725
Wright DJ, Spurgin LG, Collar NJ, Komdeur J, Burke T, Richardson DS (2014) The impact of translocations on neutral and functional genetic diversity within and among populations of the Seychelles warbler. Mol Ecol 23:2165–2177
Acknowledgements
The authors gratefully acknowledge the Mauritius Reptile Recovery Programme (Durrell Wildlife Conservation Trust, Mauritian Wildlife Foundation and National Parks and Conservation Service) for assistance with the fieldwork in this study, and the Ministry of Agro Industry, Government of Mauritius for permission to export samples for this research. We thank Josie Jackson, Helen Cadwallader and Mafalda Costa for help in laboratory and statistical analyses. Funding for this research was provided by Durrell Wildlife and Conservation Trust and Cardiff University.
Author information
Authors and Affiliations
Contributions
NC, RY and MWB conceived the study. NC and RM did the field work. SJDP, JH-M, ZM and ECS carried out the laboratory work and analysed the data, supervised by IMR, SJDP and ECS drafted the manuscript, with contribution from ZM. All authors read and approved the final manuscript.
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
du Plessis, S.J., Howard-McCombe, J., Melvin, Z.E. et al. Genetic diversity and cryptic population re-establishment: management implications for the Bojer’s skink (Gongylomorphus bojerii). Conserv Genet 20, 137–152 (2019). https://doi.org/10.1007/s10592-018-1119-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-018-1119-y