Abstract
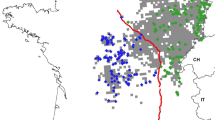
The European wildcat (Felis silvestris silvestris) underwent a severe decline across Europe in the early twentieth century. Remaining populations are often very small and isolated, though there are indications that wildcat populations are currently expanding their range. However, linear landscape elements such as rivers and roads are thought to present barriers to dispersal, inhibiting gene flow and, thus, affecting the recolonization process. In this study, we investigated the fine-scale genetic structure of wildcats in the Upper Rhine Valley. We specifically analysed wildcats on both sides of the Rhine River by genotyping 55 individual wildcats, using 20 microsatellite loci. Genetic differentiation was weak and positive spatial autocorrelation was found up to a distance of 10 km (females: 5 km, males: 10 km) indicating substantial gene flow among sampling sites. High levels of gene flow, even across the Rhine River, indicated that the water body itself does not necessarily have a strong barrier effect, which is in contrast to other studies. Our findings could best be explained by the populations’ history, a local extinction east of the River Rhine and a current ongoing population expansion. Our study highlights that potential barriers, such as rivers, may have different effects in different local wildcat populations and that the history of the populations is important to interpret genetic results. As many wildcats still occur in isolated and patchy forest fragments, maintaining connectivity between populations is crucial to ensure their viability in the long term.



Similar content being viewed by others
References
Balkenhol N, Waits LP (2009) Molecular road ecology: exploring the potential of genetics for investigating transportation impacts on wildlife. Mol Ecol 18:4151–4164. doi:10.1111/j.1365-294X.2009.04322.x
Birlenbach K, Klar N (2009) Aktionsplan zum Schutz der Europäischen Wildkatze in Deutschland. Naturschutz Landsch 41:325–332
Biró Z, Szemethy L, Heltai M (2004) Home range sizes of wildcats (Felis silvestris) and feral domestic cats (Felis silvestris f. catus) in a hilly region of Hungary. Mamm Biol—Z Säugetierkd 69:302–310. doi:10.1078/1616-5047-00149
Blanco JC, Cortés Y, Virgós E (2005) Wolf response to two kinds of barriers in an agricultural habitat in Spain. Can J Zool 83:312–323
Brooks TM, Mittermeier RA, Mittermeier CG et al (2002) Habitat loss and extinction in the hotspots of biodiversity. Conserv Biol 16:909–923
Chen C, Durand E, Forbes F, François O (2007) Bayesian clustering algorithms ascertaining spatial population structure: A new computer program and a comparison study. Mol Ecol Notes 7:747–756
Cozzi G, Broekhuis F (2013) Comparison of the effects of artificial and natural barriers on large African carnivores: implications for interspecific relationships and connectivity. J Anim Ecol 82:707–715
Devictor V, Julliard R, Jiguet F (2008) Distribution of specialist and generalist species along spatial gradients of habitat disturbance and fragmentation. Oikos. doi:10.1111/j.2008.0030-1299.16215.x
Earl DA, VonHoldt BM (2011) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the evanno method. Conserv Genet Resour 4:359–361. doi:10.1007/s12686-011-9548-7
Eckert I (2003) DNA-Analysen zum genetischen Status der Wildkatze (Felis silvestris) in Deutschland. 1–101
Eckert I, Suchentrunk F, Markov G, Hartl GB (2010) Genetic diversity and integrity of German wildcat (Felis silvestris) populations as revealed by microsatellites, allozymes, and mitochondrial DNA sequences. Mamm Biol—Z Säugetierkd 75:160–174. doi:10.1016/j.mambio.2009.07.005
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. doi:10.1111/j.1365-294X.2005.02553.x
Excoffier L, Laval G, Schneider S (2005) Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Goossens B, Waits LP, Taberlet P (1998) Plucked hair samples as a source of DNA: reliability of dinucleotide microsatellite genotyping. Mol Ecol 7:1237–1241
Greenwood PJ (1980) Mating systems, philopatry and dispersal in birds and mammals. Anim Behav 28:1140–1162
Harestad AS, Bunnell FL (1979) Home range and body weight—a reevaluation. Ecology 60:389–402
Hartmann SA, Steyer K, Kraus RHS et al (2013) Potential barriers to gene flow in the endangered European wildcat (Felis silvestris). Conserv Genet 14:413–426. doi:10.1007/s10592-013-0468-9
Hepenstrick D, Thiel D, Holderegger R, Gugerli F (2012) Genetic discontinuities in roe deer (Capreolus capreolus) coincide with fenced transportation infrastructure. Basic Appl Ecol 13:631–638. doi:10.1016/j.baae.2012.08.009
Herdtfelder M, Strein M, Suchant R (2007) Wildkatzen am Kaiserstuhl. Naturschutz Landsch 10:320–328
Herrmann M, Vogel C (2005) Wildkatze Felis silvestris sivestris Schreber, 1777. In: Braun M, Dieterlen F (eds) Die Säugetiere Baden-Württembergs, vol 2. Eugen Ulmer, Stuttgart, pp 363–376
Hertwig ST, Schweizer M, Stepanow S et al (2009) Regionally high rates of hybridization and introgression in German wildcat populations (Felis silvestris, Carnivora, Felidae). J Zool Syst Evol Res 47:283–297
Holderegger R, Di Giulio M (2010) The genetic effects of roads: a review of empirical evidence. Basic Appl Ecol 11:522–531. doi:10.1016/j.baae.2010.06.006
Jakobsson M, Rosenberg N (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806. doi:10.1093/bioinformatics/btm233
Jombart T (2008) Adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24(11):1403–1405
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genetics 11(1):1
Klar N, Herrmann M, Kramer-Schadt S (2009) Effects and mitigation of road impacts on individual movement behavior of wildcats. J Wildl Manag 73:631–638. doi:10.2193/2007-574
McKelvey KS, Schwartz MK (2005) Dropout: a program to identify problem loci and samples for noninvasive genetic samples in a capture-mark-recapture framework. Mol Ecol Notes 5:716–718. doi:10.1111/j.1471-8286.2005.01038.x
Menotti-Raymond M, David VA, Lyons LA et al (1999) A genetic linkage map of microsatellites in the domestic cat (Felis catus). Genomics 57:9–23. doi:10.1006/geno.1999.5743
Monterroso P, Brito JC, Ferreras P, Alves PC (2009) Spatial ecology of the European wildcat in a Mediterranean ecosystem: dealing with small radio-tracking datasets in species conservation. J Zool 279:27–35. doi:10.1111/j.1469-7998.2009.00585.x
Müller U, Strein M, Suchant R (2003) Wildtierkorridore in Baden-Württemberg. Berichte Freiburger forstliche Forschung 48:46
Navidi W, Arnheim N, Waterman MS (1992) A multiple-tubes approach for accurate genotyping of very small DNA samples by using PCR: statistical considerations. Am J Hum Genet 50:347–359
Noss RF, Quigley HB, Hornocker MG et al (1996) Conservation biology and carnivore conservation in the Rocky mountains. Conserv Biol 10:949–963
Nowell K, Jackson P (1996) Wild cats. status survey and conservation action plan. IUCN/SSC Cat Specialist Group, Switzerland, pp 1–383
Nussberger B, Greminger MP, Grossen C et al (2013) Development of SNP markers identifying European wildcats, domestic cats, and their admixed progeny. Mol Ecol Resour 13:447–460. doi:10.1111/1755-0998.12075
Oliveira R, Godinho R, Randi E et al (2007) Molecular analysis of hybridisation between wild and domestic cats (Felis silvestris) in Portugal: implications for conservation. Conserv Genet 9:1–11. doi:10.1007/s10592-007-9297-z
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in excel. Population genetics software for teaching and research. Mol Ecol Notes 6:288–295
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in excel. Population genetic software for teaching and research–an update. Bioinformatics 28:2537–2539. doi:10.1093/bioinformatics/bts460
Piechoki R (1990) Die Wildkatze - Felis silvestris., Neue Brehm BüchereiA Ziemsen, Wittenberg Lutherstadt
Pierpaoli M, Biro ZS, Herrmann M et al (2003) Genetic distinction of wildcat (Felis silvestris) populations in Europe, and hybridization with domestic cats in Hungary. Mol Ecol 12:2585–2598. doi:10.1046/j.1365-294X.2003.01939.x
Piry S, Luikart G, Cornuet JM (1999) BOTTLENECK: a computer program for detecting recent reductions in the effective population size using allele frequency data. J Hered 90:502–503
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Raimer F (1988) Die Wildkatze in Hessen und Niedersachsen - Biotop, Umwelt, Verbreitung, Bestandsentwicklung, Gefährdung, Schutz
R Development Core Team (2013) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.r-project.org/
Revilla E, Wiegand T, Palomares F et al (2004) Effects of matrix heterogeneity on animal dispersal: from individual behavior to metapopulation-level parameters. Am Nat 164:E130–E153. doi:10.1086/424767
Rosenberg NA (2003) Distruct: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138. doi:10.1046/j.1471-8286.2003.00566.x
Say L, Devillard S, Léger F et al (2012) Distribution and spatial genetic structure of European wildcat in France. Anim Conserv 15:18–27. doi:10.1111/j.1469-1795.2011.00478.x
Stahl P, Artois M (1991) Status and conservation of the wildcat in Europe and around the Mediterranean rim. Council of Europe, Strasbourg
Stahl P, Artois M, Aubert MFA (1988) Organisation spatiale et déplacement des chats forestiers adultes (Felis silvestris Schreber 1777) en Lorraine. Rev d’Ecologie, Terre Vie 43:113–132
Steyer K, Simon O, Kraus RHS et al (2012) Hair trapping with valerian-treated lure sticks as a tool for genetic wildcat monitoring in low-density habitats. Eur J Wildl Res 59:39–46. doi:10.1007/s10344-012-0644-0
Streif S, Kraft S, Veith S et al (2012) Monitoring and research of the European wildcat (Felis silvestris) in Baden-Württemberg. Säugetierkundliche Informationen 8:411–416
Taberlet P, Waits L, Luikart G (1999) Noninvasive genetic sampling: look before you leap. Trends Ecol Evol 14:323–327
Thiel C (2004) Streifgebiete und Schwerpunkte der Raumnutzung von Felis silvestris silvestris (Schreber 1777) in der Nordeifel. 1–195
Thompson DJ, Jenks JA (2010) Dispersal movements of subadult cougars from the Black Hills: the notions of range expansion and recolonization. Ecosphere. doi:10.1890/ES10-00028.1
Vallinoto M, Araripe J, Rego PS et al (2006) Tocantins river as an effective barrier to gene flow in Saguinus niger populations. Genet Mol Biol 29:215–219
Wilcove DS, McLellan CH, Dobson AP (1986) Habitat fragmentation in the temperate zone. In: Soule ME (ed) Conservation biology: the Science of Scarcity and Diversity. Sinauer Associates, Sunderland, pp 237–256
Woodroffe R, Ginsberg J (1998) Edge effects and the extinction of populations inside protected areas. Science 280:2126–2128
Wright S (1943) Isolation by distance. Genetics 28(2):114
Acknowledgments
We are grateful to the numerous persons who helped to conduct this research, in particular, our collegues at the Forest Research Institute Baden Wuerttemberg, Stéphanie Kraft, Sarah Veith, Mara Sandrini and Rudi Suchant for their support and constructive discussions. Sampled wildcat material and expertise was provided by François Léger (Office National de la Chasse et de la Faune Sauvage), Beatrice Nussberger (Institute of Evolutionary Biology and Environmental Studies, University of Zurich), Mathias Herrmann (ÖKO-LOG), Kathrin Steyer (Conservation Genetics, Senckenberg Research Institution and Natural History Museum Frankfurt), the Zentrum für Fisch- und Wildtiermedizin (Vetsuisse-Faculty, University of Bern) and the numerous people who regularly controlled our lure sticks. We acknowledge the funding for this study from the Landesjagdabgabe Baden Wuerttemberg. Stephanie Witczak helped revising the English of the present paper. We thank the two anonymous referees for providing constructive comments and help in improving the contents of this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Würstlin, S., Segelbacher, G., Streif, S. et al. Crossing the Rhine: a potential barrier to wildcat (Felis silvestris silvestris) movement?. Conserv Genet 17, 1435–1444 (2016). https://doi.org/10.1007/s10592-016-0874-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-016-0874-x