Abstract
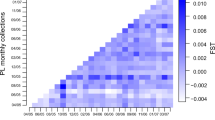
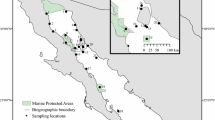
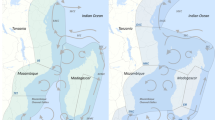
Marine protected areas (MPAs) are an important ecosystem-based management approach to help improve the sustainability of the spiny lobster fishery (Panulirus argus), but information is lacking concerning levels of lobster population connectivity among MPAs. Given their prolonged (~6 months) pelagic larval duration, population connectivity must be considered in any spatial management plan for P. argus. We used genetic techniques to uncover spatial patterns of connectivity among MPAs along the Mesoamerican Barrier Reef (MBRS) of Central America. We hypothesized that connectivity would be greater and genetic differentiation diminished among lobster populations within MPAs in the southern MBRS, which is dominated by a retentive oceanographic environment, as compared to MPAs in the more advective environment further north. We found that levels of connectivity are high among spiny lobster populations residing in MPAs in Central America, although overall F ST was low (F ST = 0.00013) but significant (P = 0.037). MPAs in the northern MBRS contained significantly more individuals that were genetically determined outliers or migrants than southern MPAs (P = 0.008, R 2 = 0.61), which may have contributed to the higher levels of genetic differentiation observed in northern MPAs. Direct genetic testing of larvae and adults will be required to confirm this hypothesis. The high level of connectivity among MPAs provides additional evidence of the importance of international cooperation in the management of Caribbean lobster fisheries. However, uncertainty regarding the ecological and physical drivers of genetic differentiation in Northern MPAs implies that managers should hedge against uncertainty.






Similar content being viewed by others
References
Acosta C, Robertson D (2003) Comparative spatial ecology of fished spiny lobsters Panulirus argus and an unfished congener P. guttatus in an isolated marine reserve at Glover’s Reef atoll, Belize. Coral Reefs 22:1–9
Akima H (1996) Algorithm 761; scattered-data surface fitting that has the accuracy of a cubic polynomial. ACM Trans Math Softw 22:362–371
Alberto F (2009) MsatAllele_1.0: an R package to visualize the binning of microsatellite alleles. J Hered 100:394–397
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B 57:289–300
Bird CE, Stephen AK, Smouse PE, Toonen RJ (2011) Detecting and measuring genetic differentiation. In: Phylogeography and population genetics in Crustacea, vol. 19. pp. 31–35.
Blouin MS, Parsons M, Lacaille V, Lotz S (1996) Use of microsatellite loci to classify individuals by relatedness. Mol Ecol 5:393–401
Botsford LW, Brumbaugh DR, Grimes C, Kellner JB, Largier J, O’Farrell MR, Ralston S, Soulanille E, Wespestad V (2008) Connectivity, sustainability, and yield: bridging the gap between conventional fisheries management and marine protected areas. Rev Fish Biol Fish 19:69–95
Botsford LW, White JW, Coffroth MA, Paris CB, Planes S, Shearer TL, Thorrold SR, Jones GP (2009) Connectivity and resilience of coral reef metapopulations in marine protected areas: matching empirical efforts to predictive needs. Coral Reefs 28:327–337
Briones-Fourzán P, Candela J, Lozano-Álvarez E (2008) Postlarval settlement of the spiny lobster Panulirus argus along the Caribbean coast of Mexico: patterns, influence of physical factors, and possible sources of origin. Limnol Oceanogr 53(3):970–985
Butler MJ IV, Herrnkind WF (1997) A test of recruitment limitation and the potential or artificial enhancement of spiny lobster populations in Florida. Can J Fish Aquat Sci 54:452–463
Butler MJ IV, Paris CB, Goldstein JS, Matsuda H, Cowen RK (2011) Behavior constrains the dispersal of long-lived spiny lobster larvae. Mar Ecol Prog Ser 422:223–237
Carvajal-Rodríguez A, de Uña-Alvarez J, Rolán-Alvarez E (2009) A new multitest correction (SGoF) that increases its statistical power when increasing the number of tests. BMC Bioinform 10:209
Chavez EA (2009) Socio-economic assessment for the management of the Caribbean spiny lobster. Proc Gulf Carib Fish Inst 60:193–196
Chittaro PM, Hogan JD (2012) Patterns of connectivity among populations of a coral reef fish. Coral Reefs 32:341–354
Cho L (2005) Marine protected areas: a tool for integrated coastal management in Belize. Ocean Coast Manag 48:932–947
Christie MR, Johnson DW, Stallings CD, Hixon MA (2010) Self-recruitment and sweepstakes reproduction amid extensive gene flow in a coral-reef fish. Mol Ecol 19:1042–1057
Cowen R, Gawarkiewicz G, Pineda J, Thorrold S, Werner F (2007) Population connectivity in marine systems: an overview. Oceanography 20:14–21
Cruz R, Bertelsen RD (2008) The spiny lobster (Panulirus argus) in the wider Caribbean: a review of life cycle dynamics and implications for responsible fisheries management. Proc Gulf Carib Fish Inst 59:433–446
Diniz FM, Maclean N, Paterson IG, Bentzen P (2004) Polymorphic tetranucleotide microsatellite markers in the Caribbean spiny lobster, Panulirus argus. Mol Ecol Notes 4:327–329
Fanning L, Mahon R, McConney P (2011) Towards marine ecosystem-based management in the wider Caribbean. Amsterdam University Press, Amsterdam
Food and Agriculture Organization (2006) Fifth regional workshop on the assessment and management of the Caribbean spiny lobster. ftp://ftpfao.org/docrep/fao/010/a1518b/a1518b00.pdf
Gaines SD, White C, Carr MH, Palumbi SR (2010) Marine reserves special feature: designing marine reserve networks for both conservation and fisheries management. Proc Natl Acad Sci 107:18286–18293
Galpern P, Manseau M, Hettinga P, Smith K, Wilson P (2012) Allelematch: an R package for identifying unique multilocus genotypes where genotyping error and missing data may be present. Mol Ecol Resour 12:771–778
Goldstein JS, Matsuda H, Takenouchi T, Butler MJ IV (2008) A description of the complete development of larval Caribbean spiny lobster Panulirus argus (LATREILLE, 1804) in culture. J Crustac Biol 28:306–327
Goudet J (2005) hierfstat, a package for r to compute and test hierarchical F-statistics. Mol Ecol Notes 5:184–186
Hedgecock D (1994) Does variance in reproductive success limit effective population size of marine organisms? In: Beaumont AR (ed) Genetics and evolution of aquatic organims. Chapman and Hall, London, pp 122–135
Hedgecock D, Barber P, Edmands S (2007) Genetic approaches to measuring connectivity. Oceanography 20:70–79
Hellberg ME (2009) Gene flow and isolation among populations of marine animals. Annu Rev Ecol Evol Syst 40:291–310
Hogan JD, Thiessen RJ, Sale PF, Heath DD (2011) Local retention, dispersal and fluctuating connectivity among populations of a coral reef fish. Oecologia 168:61–71
Iacchei M, Ben-Horin T, Selkoe KA, Bird CE, García-Rodríguez FJ, Toonen RJ (2013) Combined analyses of kinship and FST suggest potential drivers of chaotic genetic patchiness in high gene-flow populations. Mol Ecol 22:3476–3494
Jombart T (2008) adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24:1403–1405
Jones GP, Almany GR, Russ GR, Sale PF, Steneck RS, Oppen MJH, Willis BL (2009) Larval retention and connectivity among populations of corals and reef fishes: history, advances and challenges. Coral Reefs 28:307–325
Jost L (2008) GST and its relatives do not measure differentiation. Mol Ecol 17:4015–4026
Kough AS, Paris CB, Butler MJ IV (2013) Larval connectivity and the international management of fisheries. PLoS One 8:e64970
Kraemer P, Gerlach G (2013) R Package “Demerelate.” cran.r-project.org. pp 1–33
Kramer PA, Kramer PR (2002) Ecoregional Conservation Planning for the Mesoamerican Caribbean Reef. World Wildlife Fund
Lipcius RN, Eggleston DB, Schreiber SJ, Seitz RD, Shen J, Sisson M, Stockhausen WT, Wang HV (2008) Importance of metapopulation connectivity to restocking and restoration of marine species. Rev Fish Sci 16:101–110
Loiselle BA, Sork VL, Nason J, Graham C (1995) Spatial genetic structure of a tropical understory shrub, Psychotria officinalis (Rubiaceae). Am J Bot 82:1420–1425
Marko PB, Hart MW (2011) The complex analytical landscape of gene flow inference. Trends Ecol Evol 26:448–456
Maxwell KE, Matthews TR, Bertelsen RD, Derby CD (2013) Age and size structure of Caribbean spiny lobster, Panulirus argus, in a no-take marine reserve in the Florida Keys, USA. Fish Res 144:84–90
Meirmans PG, van Tienderen PH (2004) genotype and genodive: two programs for the analysis of genetic diversity of asexual organisms. Mol Ecol Notes 4:792–794
Meirmans PG (2006) Using the AMOVA framework to estimate a standardized genetic differentiation measure. Evolution 60:2399–2402
Meirmans PG, Hedrick PW (2011) Assessing population structure: F(ST) and related measures. Mol Ecol Resour 11:5–18
Menzies RA, Kerrigan JM (1979) Implications of spiny lobster recruitment patterns in the Caribbean—a biochemical genetic approach. Proc Gulf Carib Fish Inst 31:164–178
Naro-Maciel E, Reid B, Holmes KE et al (2011) Mitochondrial DNA sequence variation in spiny lobsters: population expansion, panmixia, and divergence. Mar Biol 158:2027–2041
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
OSPESCA Regulation OSP-02-09. Ordenamiento Regional de la Pesquería de la Langosta del Caribe (Panulirus argus). http://www.sica.int/busqueda/busqueda_archivo.aspx?Archivo=regl_65694_1_26012012.pdf
Puebla O, Bermingham E, McMillan WO (2012) On the spatial scale of dispersal in coral reef fishes. Mol Ecol 21:5675–5688
Queller DC, Goodnight KF (1989) Estimating relatedness using genetic markers. Evolution 43:258–275
Raymond M, Rousset F (1995) GENEPOP (Version 1.2): Population Genetics Software for Exact Tests and Ecumenicism. J Hered 3:248–249
R Core Team (2013) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org/
Rousset F (2008) genepop’007: a complete re-implementation of the genepop software for Windows and Linux. Mol Ecol Resour 8:103–106
Sale PF, Cowen RK, Danilowicz BS, Jones GP, Kritzer JP, Lindeman KC, Planes S, Polunin NV, Russ GR, Sadovy YJ (2005) Critical science gaps impede use of no-take fishery reserves. Trends Ecol Evol 20:74–80
Schunter C, Pascual M, Garza JC et al (2014) Kinship analyses identify fish dispersal events on a temperate coastline. Proc R Soc B 281:20140556
Selkoe KA, Gaines SD, Caselle JE, Warner RR (2006) Current shifts and kin aggregation explain genetic patchiness in fish recruits. Ecology 87:3082–3094
Silberman JD, Sarver SK, Walsh PJ (1994) Mitochondrial DNA variation and population structure in the spiny lobster Panulirus argus. Mar Biol 120:601–608
Strasburg JL, Rieseberg LH (2010) How robust are “isolation with migration” analyses to violations of the IM model? A simulation study. Mol Biol Evol 27:297–310
Tringali MD, Seyoum S, Schmitt SL (2008) Ten di- and trinucleotide microsatellite loci in the Caribbean spiny lobster, Panulirus argus, for studies of regional population connectivity. Mol Ecol Resour 8:650–652
Truelove N (2014) The conservation genetics of ecologically and commercially important coral reef species. Ph.D. thesis, University of Manchester, UK, pp 1–254
van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) micro-checker: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Wang Y, Hey J (2010) Estimating divergence parameters with small samples from a large number of loci. Genetics 184:363–379
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Acknowledgments
We are grateful for the logistical support provided by the Belize Fisheries Department biologists and rangers and staff at Glover’s Reef Marine Reserve managed by the Wildlife Conservation Society. We would particularly like to thank James Azueta and Isaias Majil at the Belize Fisheries Department. Without their help and hard work this research project would not have been possible. Numerous individuals helped to collect samples throughout Central America. We would also like to thank the Comisión Nacional de Áreas Naturales Protegidas in Mexico and particularly María del Carmen García Rivas for her assistance at Banco Chinchorro. We thank the following individuals for help with sample collection and logistics: Dr. Alfonso Aguilar (Universidad Autónoma de Yucatán Mexico); Miguel Alamilla, Kira Forman, Luis Novelo, Elias Cantun, Samuel Novelo, Martinez, Merve, Shakera Arnold, Ali, Aldo, and Islop, Wilfredo Pott and Barbi Gentle (Belize Fisheries) Friederike Clever (University of Manchester); Alex Tilley, Danny Wesby, Janet Gibson, and Sarah Pacyna (Wildlife Conservation Society); and Alicia Medina (WWF). This research was supported by funding for a PhD fellowship for NKT from the Sustainable Consumption Institute and Faculty of Life Sciences at the University of Manchester, and by a grant (OCE-0928930) from the US National Science Foundation to MJB and DCB. This is manuscript contribution number 966 of the Smithsonian Marine Station at Fort Pierce, Florida.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Truelove, N.K., Griffiths, S., Ley-Cooper, K. et al. Genetic evidence from the spiny lobster fishery supports international cooperation among Central American marine protected areas. Conserv Genet 16, 347–358 (2015). https://doi.org/10.1007/s10592-014-0662-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-014-0662-4