Abstract
Metastasis to bone is the leading cause of death from prostate cancer. Interaction between tumor cells and bone cells can promote progression and influence tumor phenotype. It is known that prostate cancer cells support osteoclast differentiation, and degradation of bone matrix by osteoclasts releases growth factors stimulating tumor cell proliferation and invasion. In the present study osteolytic (PC-3) and osteoblastic (LNCaP-19) castration-resistant prostate cancer (CRPC) cells were co-cultured with mature osteoclasts or their precursor cells (RAW 264.7) to characterize direct effects of mature osteoclasts on CRPC cells. Osteoclasts increased proliferation and decrease apoptosis of CRPC cells as assessed with flow cytometry. RNA sequencing revealed that osteolytic CRPC cells were more responsive to osteoclast stimulation regarding gene expression, but the overall induced expression patterns were similar between the prostate cancer cell lines. Genes related to DNA repair were upregulated by osteoclasts, while genes related to endoplasmic reticulum stress-induced apoptosis and cholesterol synthesis were downregulated. The results of this study shows that osteoclasts directly influence CRPC cells, increasing proliferation, decreasing apoptosis, and affecting gene expression pathways that can affect sensitivity to DNA damage and endoplasmic reticulum function. This suggests targeting of osteoclasts to be a possible way to affect efficacy of other drugs by combination regimens in treating prostate cancer metastases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Prostate cancer metastasis to bone is the major cause for morbidity and mortality of prostate cancer [1]. Prostate cancer cells form metastatic lesions characterized by increased bone formation [2, 3]. This contrasts to other cancer forms where skeletal metastasis is common, such as lung and breast cancer, where osteolytic lesions dominate [4]. However, in prostate cancer the resulting osteogenic phenotype, where bone marrow is suppressed by newly formed osteoid, still includes osteolytic areas, and ongoing osteolysis is a prerequisite for the increased osteogenic process [2, 5].
Osteoclasts and osteoblasts regulate their reciprocal action, leading to the balance of continuous formation and degradation of bone maintaining the bone structure intact. Osteoblasts secrete RANKL, which binds to its receptor on osteoclast precursors initiating the differentiation into mature osteoclasts, a process regulated by M-CSF as a rate limiting factor. Multinucleated osteoclasts sit on the bone surface releasing release hydrogen ions, collagenase, cathepsin K and other hydrolytic enzymes. The acidic environment increases solubility of bone mineral and the enzymes digest and degrade collagen and other organic components of the decalcified bone matrix which releases growth factors previously bound to the bone matrix [6].
Metastatic prostate cancer is treated with androgen deprivation therapy (ADT), most often gonadotropin-releasing hormone (GnRH) agonists, to block testosterone synthesis and inhibit activation of the androgen receptor (AR), a mechanism critical for survival and proliferation of prostate cancer cells. In the majority of patients, this decreases tumor burden, and prolongs time to progression, although eventually castration-resistant prostate cancer (CRPC) relapses. However, ADT also causes a decrease in bone mineral density since androgens [7], especially by their conversion into estrogens, are important for the maintenance of bone. Estrogen inhibits bone resorption by osteoclasts by shifting the balance of osteoprotegerin (OPG)/ RANKL secretion from osteoblasts as well as limiting the number of tumor necrosis factor-producing T-cells [8,9,10,11,12]. Degradation of bone by osteoclasts in osteoporosis leads to release of several growth factors. This promotes proliferation of resident prostate cancer cells, in a pattern described as a vicious cycle, since the tumor cells in turn, via secretion of parathyroid hormone related protein, promote osteoblasts to secrete RANKL increasing osteoclastic activity [13, 14].
Inhibition of osteoclast differentiation and activation, by blocking of RANKL signaling or RANK expression has been shown to decrease osteolytic prostate cancer growth in bone in experimental in vivo settings [15,16,17]. In humans, therapy inducing osteoclast apoptosis [18] and RANKL-inhibitors aiming to inhibit osteoclast differentiation and activation is frequently used among prostate cancer patients in order to increase bone strength and decrease bone fractures. Both are used with approximately the same efficacy [19]. Denosumab, an inhibitory RANKL antibody, was demonstrated to increase bone metastasis-free survival [20] and to delay skeletal related events in patients with CRPC [21]. However, no survival benefit for prostate cancer patients could be determined [19, 20]. Another possible treatment option for prostate cancer patients, besides different forms of androgen inhibition, is the PARP-inhibitors targeting cells with impaired DNA repair mechanism [22].
The indirect stimulatory role of osteoclasts on metastatic growth by degradation of bone has been acknowledged in prostate cancer, despite the osteogenic phenotype of the metastases. Also, several studies demonstrate the promoting effect of prostate cancer cells on osteoclastogenesis [23, 24]. Osteoblasts influence CRPC cells’ expression of osteogenic genes [25]. Furthermore, osteogenic CRPC cells responded to osteoblast stimulation with increased proliferation and osteogenic properties [25], as well as expression of genes encoding steroidogenic enzymes [26]. It has also been reported that bone-derived stroma cells can influence AR signaling in CRPC cells [27]. A recent study shows that Wnt-activation of RAW 264.7 derived osteoclasts promoted migration and invasion of breast cancer cells, an effect that could be inhibited by activation of the Wnt pathway in the osteoclasts [28]. In addition, we have recently reported that osteoclasts can influence steroidogenesis in prostate cancer cells [29]. However, the influence of osteoclasts on prostate cancer cell proliferation and gene expression has previously not been extensively investigated.
In the present study, we analyze the effects of osteoclasts on proliferation and survival of CRPC cells and use RNA sequencing to illustrate the gene expression changes in CRPC cells induced by osteoclasts, together with a focused study on genes related to bone metabolism. This will define any direct influence of osteoclasts on CPRC progression as bone metastases.
Material and methods
Cell culture
The osteogenic castration-resistant cell line LNCaP-19 is developed from LNCaP and has been characterized previously [25, 30, 31]. Osteolytic CRPC PC-3 cells were obtained from the European Collection of Cell Cultures ECCC (Wiltshire, UK). LNCaP-19 and PC-3 cells were maintained in RPMI 1640 medium supplemented with glucose, sodium pyruvate medium and 10% charcoal–dextran stripped serum ((CSS), Invitrogen) or 10% fetal bovine serum ((FBS), Invitrogen). Murine RAW 264.7 cells, osteoclast precursors, were obtained from ATCC (ATCC TIB-71) (Rockville MD) and maintained in DMEM (ATCC® 30–2002™) with 10% FBS. All cultures were supplemented with 1% penicillin/streptomycin and confirmed mycoplasma free.
Formation of osteoclasts
RAW 264.7 macrophages were transformed into osteoclasts by the stimulation of soluble RANKL (EMD Millipore GF091). The RAW 264.7 cells were seeded with 5 × 103 cells/cm2 concentration with 10 ng/ml of sRANKL in DMEM (ATCC 30–2002) with 10% FBS in six well plates. Medium (including sRANKL) was changed every 72 h for 6 days. At day 7, cells were washed with PBS and fixed with 4% paraformaldehyde for 15 min. Mature osteoclasts were defined by positivity for tartrate-resistant acid phosphatase (TRAP) positivity using the Acid Phosphatase, Leukocyte (TRAP) Kit (Sigma-Aldrich 387A-1KT).
Co-culture of osteoclasts and prostate cancer cells
Co-culture with osteoclasts and LNCaP-19 or PC-3 was performed using the Transwell permeable support (24 mm insert, 0.4 µm pore size, polyester membrane) in 6 well plates (Corning 3450, Kennebunk, ME, USA). Pre-incubation of Transwell inserts with the culture medium for LNCaP or PC-3 cells, as specified above, was performed for 24 h before seeding of 3 × 105 cells/insert of PC-3 or LNCaP-19 cells on the membrane. After 36 h, the inserts with PC cells were moved into plates with mature osteoclast (after the formation procedure described above) in the lower compartments. RAW 264.7 cells were seeded 5 × 103 cells/cm2 in control wells at the start of osteoclast maturation. Both osteoclasts and RAW 264.7 cells were washed with PBS, and LNCaP-19 or PC-3 culture medium (without sRANKL) was added prior to start of the co-culture. Co-culturing was performed for 48 h after which cells were harvested for later analysis. All culture experiments were repeated at least three times.
UV-treatment
UV-C irradiation is able to induce highly efficiently DNA damage [32]. LNCaP-19 or PC-3 cells were cultured in transwells for 36 h as specified in co-culture section. The culture medium was removed before the UV-C treatment, cells were irradiated for 1 min, and then the medium was added immediately after the treatment. The transwells with UV treated PC cells were moved into plates with mature osteoclast or untreated RAW 264.7 cells as controls in the lower compartments. After 48 h of co-culturing, PC cells were harvested for DNA damage analysis with Western blotting using phosphorylated histone protein γ-H2AX as a marker for DNA strand breaks.
Flow cytometry
For cell apoptosis assay, after 48 h co-culturing, the cells were collected and washed in ice-cold PBS. PE Annexin V Apoptosis Detection kit (BD Pharmingen™, USA) was used to assess cell apoptosis according to the manufacturer’s instructions. 7-AAD was used to stain dead cells. The cells were incubated with 5 µl of PE Annexin V and 5 µl 7-AAD for 15 min at RT (25 °C) in the dark. Unstained cells, cells stained with PE Annexin V (no 7-AAD) and cells stained with 7-AAD (no PE Annexin V) were used to set up compensation and quadrants.
For the cell proliferation assay, EdU Staining Proliferation Kit (iFluor 488) (Abcam, UK) was used to analyze the proliferation of cells after 48 h co-culturing. EdU solution was added and incubated together with the cells for 4 h under optimal growth conditions. Both cell apoptosis and cell proliferation were analyzed with a flow cytometer (BD Accuri™ C6, BD Biosciences).
RNA isolation and RNA sequencing
Total RNA was isolated using RNeasy Mini Plus kit (Qiagen, USA) according to the manufacturer’s instructions. RNA concentration was measured on a NanoDrop (Thermo Fisher, USA). RNA samples from LNCaP-19 cells and PC-3 cells co-cultured with mature osteoclasts or control cells (RAW 264.7 cells) were quantified and checked for contamination using (Dropsense96, Trinean). After normalization for equal input (500 ng in 5 ul), samples were spiked with ERCC RNA Spike-In Control Mix (ThermoFisher). Libraries were prepared using QuantSeq 3′ mRNA-Seq (FWD) Library Prep Kit for Illumina (Lexogen) including individual indexing for multiplex sequencing. Finalized libraries were quality checked with capillary electrophoresis (Fragment Analyzer) and quantified with Picogreen (ThermoFisher). The libraries were normalized and pooled before NextSeq500 sequencing (Illumina).
Protein preparation and Western blotting
The cells were lysed with RIPA Lysis and Extraction Buffer (Invitrogen™, USA) supplemented with PhosSTOP™ phosphatase inhibitor and cOmplete™ protease inhibitor cocktail (Roche Diagnosis; DE). The protein concentration was determined using a Coomassie Plus (Bradford) Assay Kit (Pierce™, USA) according to the manufacturer’s instructions. Protein lysates were separated by NuPAGE™ 4–12%, Bis–Tris gel and then transferred to PVDF membranes (Invitrogen™, USA). Membranes were blocked in Pierce™ Clear Milk Blocking Buffer (Pierce Biotechnology; USA) and incubated with primary antibodies (Supplementary X) at 4 °C overnight. Then the membranes were washed with PBS-T and incubated with secondary antibodies for 1 h at RT (Antibodies are specified in Supplementary Table 1). Amersham™ ECL Select™ western blotting detection reagent (GE Healthcare; UK) was used for visualizing and detection following manufacturer’s instruction.
Data handling and statistics
The pipeline Human (GRCh38) Lexogen QuantSeq 2.6.1 was used for the sequence data analysis. After mapping the reads against the ENSEMBL reference genome, we merged the reads count files using a custom bash script. ENSEMBL identifiers were translated into HGNC gene names using a custom R script that retrieved the corresponding gene names from the BioMart database. Differential gene expression was performed using the R-package edgeR [33]. Multiple testing correction is performed using Benjamini–Hochberg false discovery rate (FDR) method. A gene is considered differentially expressed if FDR < 5% and the log-transformed fold change (log2(FC)) > 1 (up-regulated) or log2(FC) < − 1 (down-regulated). All non-protein coding RNAs were removed prior to analysis. Lists of significantly affected genes were assessed for gene ontology (GO) enrichment using PANTHER Overrepresentation Test (released 2020-07-28) and the PANTHER Enrichment test annotated with GO Ontology database https://doi.org/10.5281/zonodo.4081749 (released 2020-10-09) [34]. The annotation Data sets used was GO biological process complete and Reactome Pathways. The analysis was performed using the Fisher’s Exact test with Bonferroni correction for multiple testing. Only GO terms with a Bonferroni adjusted P-value < 0.05 were considered. QIAGEN IPA (QIAGEN Inc., https://digitalinsights.qiagen.com/IPA) software was used for further analysis of upstream regulators and for illustration of affected pathways in the significant genes. Differences in gene expression in the osteogenesis PCR array were statistically evaluated using Student’s t-test. These calculations were performed in IBM SPSS Statistics 26. Flow cytometry results were handled in the BD Accuri TM C6 analysis software and statistically evaluated with unpaired t-test with Welch’s correction using the GraphPad Prism 9.0.0.
Results
Osteoclasts diminish apoptosis and promote proliferation in CRPC cells
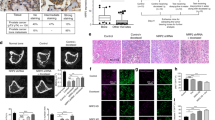
Osteolysis is known to stimulate proliferation of metastatic cells by release of growth factors bound to bone tissue. To investigate any direct effect of mature osteoclasts on PC cell proliferation or apoptosis, a flow cytometry assay was used on PC cells after co-culturing with mature osteoblasts or control RAW cells. The apoptosis assay using Annexin V and 7-AAD showed a decrease of end stage apoptotic cells in LNCaP-19 co-cultured with osteoclasts compared to the control condition, while in PC-3 cells, the decrease in apoptosis was not statistically significant (Fig. 1a–c). Furthermore, mature osteoclasts promoted proliferation in both LNCaP-19 and PC-3 cells using an EdU incorporation assay (Fig. 1d–f).
Effects of osteoclasts on prostate cancer cell apoptosis and proliferation a Representative images of the flow cytometry of PC-3 and LNCaP-19 cells labelled with 7AAD and PE Annexin V after 48 h co-culture with mature osteoclasts or control cells (unstimulated RAW 264.7 cells). b, c Effects on osteoclasts on apoptosis (7AAD+ and Annexin V+) in PC-3 (b) and LNCaP-19 (c). Bars represent mean ± SEM. d Representative images of the flow cytometry of PC-3 and LNCaP-19 cells labelled with EdU after 48 h co-culture with osteoclasts or control cells. e, f Effects on osteoclasts on proliferation (EdU+) in PC-3 (e) and LNCaP-19 (f)
The majority of osteoclast-induced expression changes are upregulation
To further investigate the influence of osteoclasts on CRPC cells, analysis of the gene expression changes induced by the co-culture was performed with RNA-sequencing. A principal component analysis of the RNA sequencing data showed that the two cell lines were well separated and samples from co-cultures with mature osteoclasts and the macrophage control cells were separated within the individual clusters for the cell lines (Fig. S1).
The sequencing generated read counts across 56,324 genes. After filtering out genes with low counts (< 2 cpm in the majority of samples) that was reduced to 13,511. Of these, 6226 (46%) were significantly affected by osteoclast stimulation (FDR < 0.05). Although non-protein coding types of RNA have been shown to influence osteoclasts and bone metabolism [35, 36], the result in the present study is focused on the protein coding genes.
In total, the expression of 3566 protein coding genes was significantly affected by co-culture with mature osteoclasts compared to precursor cells (FDR < 0.05, −1 > log2(FC) > 1). Gene expression in PC-3 was generally more responsive to osteoclast stimulation with 2756 affected genes, compared to 1479 in LNCaP-19. Of the 3566 genes, approximately 20% (669 genes) responded with a change in the same direction in both cell lines. In general, both cell lines responded with more increased gene expression than decreased (in total 2100 upregulated and 1466 downregulated) (Fig. S1).
Gene ontology and pathway analysis
GO overrepresentation analyses revealed that osteoclast stimulation of LNCaP-19 and PC-3 cells affect largely the same groups of genes in both cell lines. GO terms that appear as overrepresented for both PC-3 and LNCaP-19 were cell cycle, chromosome organization, DNA repair, DNA replication, tRNA processing/metabolic process, and macromolecule methylation/modification. Underrepresented in both was the G-protein-coupled receptor signaling pathway (Table 1a, b). Adding the fold change of the altered genes into the analysis, the PANTHER enrichment test revealed differences between the cell lines’ response to osteoclasts. This test compares the average fold change of genes within an identified GO term with the average fold change of the whole data set. PC-3, due to its larger amount of affected genes, presented with a broader range of enriched GO terms; the only positively enriched GO terms were associated with RNA modification, while the list of negatively enriched GO terms was topped by regulation of apoptosis, regulation of cell differentiation, and regulation of cell population proliferation. In LNCaP-19, only four GO terms were identified with the PANTHER enrichment test. Positively enriched were double-strand break repair via homologous recombination and DNA-templated DNA replication, while response to endoplasmic reticulum stress and regulation of epithelial cell differentiation displayed a more downregulated profile compared to the whole data set (Table 2a, b).
Using the Reactome Pathway option built into the PANTHER tool, enriched pathways were identified in the cell lines after co-culture with osteoclasts. In LNCaP-19, only two pathways were identified (Table 2b). In PC-3 however, some distinct pathways not clearly overlapping with the enriched GO terms appear. The most significantly affected pathway was activation of gene expression by SREBF, which regulates cholesterol biosynthesis (Table 2a). A parallel Core Analysis of fold change in significantly altered genes was carried out in QIAGEN IPA to identify affected pathways in an untargeted fashion for each cell line. As with the previous analyses, pathways related to cancer (cell cycle, DNA replication and cell death) were identified by IPA as being the most affected by co-culture with osteoclasts, especially in LNCaP-19. In PC-3, a partly different picture emerges including the SREBP2 pathway (Graphical illustration in Supplementary Fig. S3). IPA was then used to generate illustrative pathways using a gene list for genes coupled to SREBF2 (Supplementary Fig. S4). All 18 genes in the pathway were significantly altered when PC3 cells were co-cultured with osteoclasts, whereas only 9 of the 18 were significantly altered in LNCaP-19 cells (Supplementary Table 2).
The most significantly affected genes
Studying the most significantly affected genes (based on lowest false discovery rate), it appears that the upregulated profile in PC-3 is the most specific response to osteoclast secretory products. Of the 20 most significantly upregulated genes in PC-3, 14 were not affected in LNCaP-19. In comparison, 16 of the top 20 upregulated genes in LNCaP-19 were also upregulated in PC-3. No such difference was observed among the top 20 downregulated genes, where most of the genes were affected in both cell lines. Eight genes (DDIT3, DDIT4, GDF15, GADD45B, ATF3, SAT1, BBC3, and INSIG1) appear on the top 20 lists for both cell lines, indicating a strikingly similar pattern for the most downregulated genes in response to osteoclast stimulation (Supplementary Table 3). In line with this data, a DDIT3 centered network cluster was identified in the QIAGEN IPA analysis in both cell lines limiting the analysis to genes with fold change > 3 (Supplementary Fig. S5).
In line with the GO and pathway analyses, many of the most upregulated genes, especially in LNCaP-19, were related to regulation of RNA. Five genes for heterogeneous nuclear ribonucleoproteins (HNRNPM, HNRNPP2, HNRNPD, HNRNPAB, and HNRNPDL) which complex with heterogeneous nuclear RNAs were found, together with additionally at least three genes involved in RNA splicing (SFPQ, SRSF3, U2AF2). The homogenous pattern regarding the most downregulated genes in both cell lines clearly matched the GOs and pathways related to ER-induced stress response, where BBC3, DDIT3, ATF3, ATF4, and CHAC1 all co-operate in apoptosis induced by ER-stress (Supplementary Table 3).
Upstream regulators
The QIAGEN IPA software was used to identify possible upstream regulators for the changes induced by osteoclasts in PC-3 and LNCaP-19 (Table 3a, b). This module in IPA uses identified pathways integrated with previously reported associations in the literature to identify putative upstream regulators of the observed effects. In PC-3, the transcriptional regulators NUPR1, HNF4A, and p53 could possibly mediate the effects on apoptosis and DNA repair. In LNCaP-19, ATF4 was identified as a possible upstream inducer of the ER stress related effects, while the inhibited action of EIF2AK3 could contribute to the regulation of expression of many types of genes. Suggested as a common upstream regulator for PC-3 and LNCaP-19 was the mTOR inhibitor 2-(4-amino-1-isopropyl-1H-pyrazolo[3,4- d]pyrimidin-3-yl)-1H-indol-5-ol (TORKinib or PP242), a mammalian target of rapamycin (mTOR) inhibitor, whose action was predicted by the IPA software to be inhibited.
Downregulation of proteins in cholesterol synthesis pathway
Cholesterol is an important component of cell membranes as well as the precursor for steroid synthesis. The RNA-sequencing data indicated that osteoclast simulation downregulates SREBP2 regulated genes, of which several are enzymes involved in cholesterol synthesis, such as 3-Hydroxy-3-Methylglutaryl-CoA Synthase 1 (HMGCS1), Farnesyl-Diphosphate Farnesyltransferase 1 (FDFT1), and Farnesyl Diphosphate Synthase (FDPS) [37]. To verify the effects of osteoclasts on this pathway, selected proteins were investigated by Western blotting. Among the selected genes, SRBEF2 itself and its co-transcription factor SP-1, as well as squalene epoxidase (SQLE), which is one of the rate-limiting enzymes in this pathway, were downregulated after osteoclast stimulation in both PC-3 and LNCaP-19. Conversely, FDFT1, the first specific enzyme in cholesterol biosynthesis was upregulated in PC-3 (Fig. 2a).
Protein analysis of selected genes a Western blotting of selected proteins in the SREBP2/cholesterol pathway, and b Western blotting of selected proteins in the ER stress induced apoptosis pathway in PC-3 and LNCaP-19 cells after co-culture with mature osteoclasts or control cells (unstimulated RAW 264.7 cells). GAPDH, β-actin and tubulin were used as loading controls
Downregulation of proteins related to ER-induced stress response
To investigate whether the osteoclasts-induced downregulation of the proteins related to the ER-stress induced apoptosis was translated into changes also on the protein level, Western blotting analysis was performed. Among the selected genes, DDIT3 which induces cell cycle arrest and apoptosis in response to ER stress [38] was downregulated by co-culture with osteoclasts at the protein level in both cell lines, while the transcription factor ATF3 was downregulated only in LNCaP-19 (Fig. 2b).
DNA damage after UV exposure of CRPC cells is affected by osteoclasts
To investigate whether the induction of expression of genes related to DNA repair could be translated into the protein level, Western blotting analysis was performed on selected genes in this GO group. Two of the analyzed proteins, BRCA1 and PALB2 was increased in PC-3, while the effect was not seen as clearly in LNCaP-19 after co-culture with osteoclasts compared to the control condition. The expression level of BRCA2 was not affected by osteoclasts on the protein level (Fig. 3a). To investigate whether this altered expression of genes related to DNA repair by osteoclasts influences the accumulation of DNA breaks, we exposed LNCaP-19 and PC-3 cells to UV-light and analyzed the amount of DNA breaks after 48 h of co-culture. In both cell lines, co-culture with osteoclasts results in less DNA breaks, reflected by the lower levels of accumulated γ-H2AX compared to cells co-cultured with precursor RAW 264.7 cells (Fig. 3b).
Protein analysis of DNA repair pathway a Western blotting of selected proteins in the DNA repair pathway in PC-3 and LNCaP-19 after co-culture with mature osteoclasts or control cells (unstimulated RAW 264.7 cells), and b accumulation of γ-H2AX as a marker for DNA breaks in UV-exposed PC-3 and LNCaP-19 cells after co-culture with mature osteoclasts or control cells (unstimulated RAW 264.7 cells). β-actin was used as loading control
Genes related to bone metabolism
To specifically study the effects of osteoclasts on genes expression related to bone metabolism and enable comparisons with the effects of osteoblasts previously described [25], an osteogenesis directed PCR panel was used. Similar to the RNA sequencing results, PC-3 was more sensitive to osteoclast simulation compared to LNCaP-19, with 12 genes statistically changed (2FC 0.05) compared to only one (TGFB2) in LNCaP-19 (Table 4). These changes indicate a pattern where osteoclasts induce PC-3 cells to express genes for secreted factors that promote osteogenesis (BMP6, VEGFA, TGFB2) and inhibit osteoclast differentiation and activation (CSF2), while decreasing expression of CSF3, which stimulates osteoclastogenesis and induces apoptosis in osteoblasts [39].
The increased expression of CSF2 in PC-3 cells after co-culture with osteoclasts, indicating an inhibiting effect on osteoclasts, was confirmed at the protein level, while CSF3 levels were unchanged. In addition, osteoclasts decreased both osteoblast cadherin (CDH11) and N-cadherin (CDH2) indicating a decreased capacity of attachment of the PC-3 cells to other cell types in the bone microenvironment. No altered expression of secreted factors promoting osteogenesis could be confirmed on the protein level (Fig. 4a, b).
Protein analysis selected genes involved in osteogenesis a Growth factors regulating bone metabolism and tumor growth, and b proteins involved in osteoclast differentiation and cell adhesion in PC-3 and LNCaP-19 cells after co-culture with mature osteoclasts or control cells (unstimulated RAW 264.7 cells). β-actin was used as loading control
Discussion
Interaction with bone cells is important for prostate cancer cells to grow as metastases in the skeleton. The present study demonstrates direct influence of osteoclasts on prostate cancer cell proliferation and gene expression. In particular, genes regulating DNA repair, RNA modifications, and ER stress-induced apoptosis were affected. Further, osteoclasts influence osteolytic PC cells to a gene expression pattern characteristic for osteogenesis, thus stabilizing bone metabolism.
Osteoclasts have previously been attributed an important role in prostate cancer progression in bone, by degrading bone tissue and releasing tumor-promoting factors in the so-called vicious cycle. It is also known that prostate cancer cells directly promote differentiation of both osteoclasts and osteoblasts from their precursors [5, 40, 41]. However, a direct role of osteoclasts in regulating prostate cancer function has previously not been established. The present study suggests that mature osteoclasts, in comparison to macrophage precursor cells, increased proliferation and decreased apoptosis in prostate cancer cells. The effect was larger for osteoblastic prostate cancer cells compared to osteolytic and more dedifferentiated prostate cancer cells, in line with the effect of osteoblast stimuli in a previous study [25]. This indicates that these osteoblastic cells are more susceptible to bone cell stimulation regulating proliferation, compared to the osteolytic prostate cancer cells, which display overall more dedifferentiated features such as decreased AR expression. It may also indicate that osteoblastic prostate cancer cells are more sensitive to therapies aiming to inhibit proliferation, while cytotoxic strategies targeting stably proliferating cells may be more efficient in metastases dominated by osteolysis. Further, the phenotypical characteristics of bone remodeling has been linked to molecular phenotypes with impact on prognosis [42, 43].
In contrast to the proliferative response, osteolytic PC-3 cells displayed larger differences in overall gene expression after osteoclast stimulation compared to LNCaP-19. More genes were affected and consequently more GO terms describing their roles were identified. In general, few of classical tumor metastasis associated genes or pathways, such as those related to epithelial-mesenchymal transition, invasion or angiogenesis, were affected by co-culture with osteoclasts. Since the control cells in this setting are their macrophage precursor cells this might not be surprising. It is known that tumor infiltrating macrophages affect the progression of tumors and metastases [44], and supposedly their progeny, the osteoclasts, retain that ability, supporting a beneficial function of inhibiting the differentiation of or directly targeting this cell population in tumors. Despite the close relation between macrophages and osteoclasts, a large number of genes and functions were affected specifically by mature osteoclasts in the present study.
Of special interest in relation to cancer therapy are effects on DNA repair mechanisms. In both PC-3 and LNCaP-19, genes related to DNA-repair were overrepresented among affected genes. Several of the DNA repair genes upregulated by osteoclasts overlaps with the homologous recombination repair pathway (BRCA1, BRCA2, PALB2, BRIP1, and RAD51C), of which two (BRCA1 and PALB2) were shown to be affected also on the protein level in the present study. The decreased DNA damage, as demonstrated by lower γ-H2AX levels, in osteoclast-stimulated CRPC cells after exposure to UV-light supports that osteoclasts influence the response to DNA damage in CRPC cells. Deletion or mutation in DNA repair genes are suggested to indicate sensitivity to PARP-inhibitors, [45] as evaluated in the AMPLITUDE trial for prostate cancer (NCT04497844). However, the possible impact on sensitivity of different drugs needs to be investigated further, especially since a connection between sensitivity to PARP-inhibitors and AR signaling has been established [46].
Interaction between tumor cells and the bone microenvironment is necessary to convey the different phenotypes of the metastases defined by different prostate cancer cell types. Osteolytic prostate cancer cells promote osteoclast differentiation and activity while osteoblastic prostate cancer cells stimulate proliferation and differentiation of osteoblasts [5]. In addition, osteoblastic PC cells display osteomimetic properties such as expression of osteoblast specific markers and mineralization of bone matrix in vitro, features that were increased by stimulation by osteoblasts. In the present study, osteolytic PC cells were more sensitive than osteoblastic PC cells to osteoclast stimulation regarding gene expression related to bone metabolism. The parallel, i.e. that osteoblastic PC cells responded more to osteoblasts, was previously described, indicating a phenotype dependent gene regulation by bone cells [25]. However, only a few of the observed changes in the present study could be verified on the protein level, indicating that other regulatory mechanisms than RNA transcription, possibly provided by other parts of the bone microenvironment not present in the in vitro system, are important for the final phenotypic changes. Colony stimulating factor 2 (CSF2), which inhibits osteoclast differentiation [47], was still upregulated in PC-3 cells after osteoclast stimulation when assessed on the protein level, indicating a possible regulatory mechanism balancing the osteolytic progression. Together with the altered CSF2, the two cell adhesion proteins, osteoblast-cadherin (CDH11) and N-cadherin (CDH2) were downregulated, suggesting that osteoclasts decrease the tumor cell ability to adhere to mesenchymal cells in the microenvironment.
One pathway identified as significantly downregulated in PC-3 in this study was regulation by sterol regulatory element-binding protein 2 (SREBP2) including several enzymes involved in cholesterol synthesis. Cholesterol is a vital component in cell membranes and a precursor for steroid synthesis, including androgens. In prostate cancer, high cholesterol and cholesterol biosynthesis have been associated with for example progression [48], metastasis [49], and poor treatment response to taxanes [50]. Together with the downregulation of the cholesterol synthesis pathway by ostecoclasts in the present study, this would indicate a suppressive role of osteoclasts on aggressive features associated with cholesterol. However, a detectable effect on cholesterol levels needs to be confirmed in further studies.
Osteoclasts decreased gene expression related to ER stress induced apoptosis in both cell lines, identified both from the enriched GO terms and the list of the most affected genes. RNA expression for several genes, such as ATF3, ATF4, DDIT3, TRIB3, and CHAC1, involved in ER stress induced apoptosis was downregulated after co-culture with osteoclasts compared to the control cells, and both ATF3 and DDIT3 were confirmed as downregulated on the protein level. ER stress results in accumulation of un-folded proteins, which elicit either rescuing programs or activated apoptosis [51, 52]. The lower expression of genes associated with ER stress induced apoptosis may indicate that this mechanism for apoptosis induction was specifically decreased by osteoclasts. It may also suggest that osteoclasts decrease the sensitivity to drugs inducing cell death due to unfolded protein responses, highlighting an area for further studies regarding a possible protective role of osteoclasts for tumor cells in the bone microenvironment.
A limitation of the present study is the complexity of the co-culture system, where, in the osteoclast-stimulated situation, precursor cells are intermixed with the differentiated osteoclasts. To which extent the matured osteoclasts influence the remaining precursor cells, and how that affect their interaction with the tumor cells, as compared to the control situation, is not known. Thus, a contaminating effect by the remaining precursor cells in the osteoclast stimulation of tumor cells cannot be ruled out. Further studies are needed to identify the specific function of pure osteoclasts. Another limitation is the absence of other bone cells in the model system, thus the interpretations from the study must be further evaluated in the full complexity of the bone microenvironment. The present study does not identify the mechanism(s) by which osteoclasts act on tumor cells, an information that would be of great interest in terms of future targeting of this interaction. The putative upstream regulators suggested by the bioinformatic analysis may indicate certain targets, such as mTOR, for future studies in this regard.
In conclusion, the present study suggests that osteoclasts directly influence the growth and function of prostate cancer cells. In addition, osteoclasts affect gene expression pathways involved in DNA repair and apoptosis, indicating that they can influence the sensitivity to cytotoxic drugs. This broadens the putative role of therapeutic osteoclast inhibition to include combination therapies.
Data availability
The datasets generated in and analyzed in the current study are available from the corresponding author on reasonable request.
Abbreviations
- CRPC:
-
Castration-resistant prostate cancer
- ADT:
-
Androgen deprivation therapy
- RANKL:
-
Receptor activator of nuclear factor kappa-Β ligand
- FDR:
-
False discovery rate
- ER:
-
Endoplasmic reticulum
References
Thobe MN, Clark RJ, Bainer RO, Prasad SM, Rinker-Schaeffer CW (2011) From prostate to bone: key players in prostate cancer bone metastasis. Cancers (Basel) 3(1):478–493. https://doi.org/10.3390/cancers3010478
Roudier MP, True LD, Higano CS, Vesselle H, Ellis W, Lange P et al (2003) Phenotypic heterogeneity of end-stage prostate carcinoma metastatic to bone. Hum Pathol 34(7):646–653. https://doi.org/10.1016/s0046-8177(03)00190-4
Coleman RE, Croucher PI, Padhani AR, Clezardin P, Chow E, Fallon M et al (2020) Bone metastases. Nat Rev Dis Primers 6(1):83. https://doi.org/10.1038/s41572-020-00216-3
Mundy GR (2002) Metastasis to bone: causes, consequences and therapeutic opportunities. Nat Rev Cancer 2(8):584–593. https://doi.org/10.1038/nrc867
Keller ET, Brown J (2004) Prostate cancer bone metastases promote both osteolytic and osteoblastic activity. J Cell Biochem 91(4):718–729. https://doi.org/10.1002/jcb.10662
Stenbeck G (2002) Formation and function of the ruffled border in osteoclasts. Semin Cell Dev Biol 13(4):285–292. https://doi.org/10.1016/s1084952102000587
Sountoulides P, Rountos T (2013) Adverse effects of androgen deprivation therapy for prostate cancer: prevention and management. ISRN Urol 2013:240108. https://doi.org/10.1155/2013/240108
Eghbali-Fatourechi G, Khosla S, Sanyal A, Boyle WJ, Lacey DL, Riggs BL (2003) Role of RANK ligand in mediating increased bone resorption in early postmenopausal women. J Clin Invest 111(8):1221–1230. https://doi.org/10.1172/JCI17215
Hofbauer LC, Khosla S (1999) Androgen effects on bone metabolism: recent progress and controversies. Eur J Endocrinol 140(4):271–286. https://doi.org/10.1530/eje.0.1400271
Hofbauer LC, Schoppet M (2004) Clinical implications of the osteoprotegerin/RANKL/RANK system for bone and vascular diseases. JAMA 292(4):490–495. https://doi.org/10.1001/jama.292.4.490
Modder UI, Roforth MM, Hoey K, McCready LK, Peterson JM, Monroe DG et al (2011) Effects of estrogen on osteoprogenitor cells and cytokines/bone-regulatory factors in postmenopausal women. Bone 49(2):202–207. https://doi.org/10.1016/j.bone.2011.04.015
Roggia C, Gao Y, Cenci S, Weitzmann MN, Toraldo G, Isaia G et al (2001) Up-regulation of TNF-producing T cells in the bone marrow: a key mechanism by which estrogen deficiency induces bone loss in vivo. Proc Natl Acad Sci USA 98(24):13960–13965. https://doi.org/10.1073/pnas.251534698
Clines GA, Guise TA (2005) Hypercalcaemia of malignancy and basic research on mechanisms responsible for osteolytic and osteoblastic metastasis to bone. Endocr Relat Cancer 12(3):549–583. https://doi.org/10.1677/erc.1.00543
Vessella RL, Corey E (2006) Targeting factors involved in bone remodeling as treatment strategies in prostate cancer bone metastasis. Clin Cancer Res 12(20 Pt 2):6285s–6290s. https://doi.org/10.1158/1078-0432.CCR-06-0813
Takayama K, Inoue T, Narita S, Maita S, Huang M, Numakura K et al (2017) Inhibition of the RANK/RANKL signaling with osteoprotegerin prevents castration-induced acceleration of bone metastasis in castration-insensitive prostate cancer. Cancer Lett 397:103–110. https://doi.org/10.1016/j.canlet.2017.03.034
Virk MS, Alaee F, Petrigliano FA, Sugiyama O, Chatziioannou AF, Stout D et al (2011) Combined inhibition of the BMP pathway and the RANK-RANKL axis in a mixed lytic/blastic prostate cancer lesion. Bone 48(3):578–587. https://doi.org/10.1016/j.bone.2010.11.003
Zheng Y, Basel D, Chow SO, Fong-Yee C, Kim S, Buttgereit F et al (2014) Targeting IL-6 and RANKL signaling inhibits prostate cancer growth in bone. Clin Exp Metastasis 31(8):921–933. https://doi.org/10.1007/s10585-014-9680-3
Oades GM, Coxon J, Colston KW (2002) The potential role of bisphosphonates in prostate cancer. Prostate Cancer Prostatic Dis 5(4):264–272. https://doi.org/10.1038/sj.pcan.4500607
Jakob T, Tesfamariam YM, Macherey S, Kuhr K, Adams A, Monsef I et al (2020) Bisphosphonates or RANK-ligand-inhibitors for men with prostate cancer and bone metastases: a network meta-analysis. Cochrane Database Syst Rev 12:CD013020. https://doi.org/10.1002/14651858.CD013020.pub2
Smith MR, Saad F, Coleman R, Shore N, Fizazi K, Tombal B et al (2012) Denosumab and bone-metastasis-free survival in men with castration-resistant prostate cancer: results of a phase 3, randomised, placebo-controlled trial. Lancet 379(9810):39–46. https://doi.org/10.1016/S0140-6736(11)61226-9
Fizazi K, Carducci M, Smith M, Damiao R, Brown J, Karsh L et al (2011) Denosumab versus zoledronic acid for treatment of bone metastases in men with castration-resistant prostate cancer: a randomised, double-blind study. Lancet 377(9768):813–822. https://doi.org/10.1016/S0140-6736(10)62344-6
Flippot R, Patrikidou A, Aldea M, Colomba E, Lavaud P, Albiges L et al (2022) PARP inhibition, a new therapeutic avenue in patients with prostate cancer. Drugs. https://doi.org/10.1007/s40265-022-01703-5
Mori K, Le Goff B, Charrier C, Battaglia S, Heymann D, Redini F (2007) DU145 human prostate cancer cells express functional receptor activator of NFkappaB: new insights in the prostate cancer bone metastasis process. Bone 40(4):981–990. https://doi.org/10.1016/j.bone.2006.11.006
Zhang J, Dai J, Qi Y, Lin DL, Smith P, Strayhorn C et al (2001) Osteoprotegerin inhibits prostate cancer-induced osteoclastogenesis and prevents prostate tumor growth in the bone. J Clin Invest 107(10):1235–1244. https://doi.org/10.1172/JCI11685
Hagberg Thulin M, Jennbacken K, Damber JE, Welen K (2014) Osteoblasts stimulate the osteogenic and metastatic progression of castration-resistant prostate cancer in a novel model for in vitro and in vivo studies. Clin Exp Metastasis 31(3):269–283. https://doi.org/10.1007/s10585-013-9626-1
Hagberg Thulin M, Nilsson ME, Thulin P, Ceraline J, Ohlsson C, Damber JE et al (2016) Osteoblasts promote castration-resistant prostate cancer by altering intratumoral steroidogenesis. Mol Cell Endocrinol 422:182–191. https://doi.org/10.1016/j.mce.2015.11.013
Villagran MA, Gutierrez-Castro FA, Pantoja DF, Alarcon JC, Farina MA, Amigo RF et al (2015) Bone stroma-derived cells change coregulators recruitment to androgen receptor and decrease cell proliferation in androgen-sensitive and castration-resistant prostate cancer cells. Biochem Biophys Res Commun 467(4):1039–1045. https://doi.org/10.1016/j.bbrc.2015.10.009
Li KX, Sun X, Li BY, Yokota H (2021) Conversion of osteoclasts into bone-protective, tumor-suppressing cells. Cancers (Basel) 13(22):5593. https://doi.org/10.3390/cancers13225593
Huang J, Hagberg Thulin M, Damber JE, Welen K (2021) The roles of RUNX2 and osteoclasts in regulating expression of steroidogenic enzymes in castration-resistant prostate cancer cells. Mol Cell Endocrinol 535:111380. https://doi.org/10.1016/j.mce.2021.111380
Gustavsson H, Welen K, Damber JE (2005) Transition of an androgen-dependent human prostate cancer cell line into an androgen-independent subline is associated with increased angiogenesis. Prostate 62(4):364–373. https://doi.org/10.1002/pros.20145
Jennbacken K, Gustavsson H, Welen K, Vallbo C, Damber JE (2006) Prostate cancer progression into androgen independency is associated with alterations in cell adhesion and invasivity. Prostate 66(15):1631–1640. https://doi.org/10.1002/pros.20469
Cadet J, Anselmino C, Douki T, Voituriez L (1992) Photochemistry of nucleic acids in cells. J Photochem Photobiol B 15(4):277–298. https://doi.org/10.1016/1011-1344(92)85135-h
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26(1):139–140. https://doi.org/10.1093/bioinformatics/btp616
Mi H, Muruganujan A, Ebert D, Huang X, Thomas PD (2019) PANTHER version 14: more genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Res 47(D1):D419–D426. https://doi.org/10.1093/nar/gky1038
Patil S, Dang K, Zhao X, Gao Y, Qian A (2020) Role of LncRNAs and CircRNAs in bone metabolism and osteoporosis. Front Genet 11:584118. https://doi.org/10.3389/fgene.2020.584118
Weivoda MM, Lee SK, Monroe DG (2021) miRNAs in osteoclast biology. Bone 143:115757. https://doi.org/10.1016/j.bone.2020.115757
Sato R (2010) Sterol metabolism and SREBP activation. Arch Biochem Biophys 501(2):177–181. https://doi.org/10.1016/j.abb.2010.06.004
Yamaguchi H, Wang HG (2004) CHOP is involved in endoplasmic reticulum stress-induced apoptosis by enhancing DR5 expression in human carcinoma cells. J Biol Chem 279(44):45495–45502. https://doi.org/10.1074/jbc.M406933200
Christopher MJ, Link DC (2008) Granulocyte colony-stimulating factor induces osteoblast apoptosis and inhibits osteoblast differentiation. J Bone Miner Res 23(11):1765–1774. https://doi.org/10.1359/jbmr.080612
Jin JK, Dayyani F, Gallick GE (2011) Steps in prostate cancer progression that lead to bone metastasis. Int J Cancer 128(11):2545–2561. https://doi.org/10.1002/ijc.26024
Logothetis CJ, Lin SH (2005) Osteoblasts in prostate cancer metastasis to bone. Nat Rev Cancer 5(1):21–28. https://doi.org/10.1038/nrc1528
Nordstrand A, Bovinder Ylitalo E, Thysell E, Jernberg E, Crnalic S, Widmark A et al (2018) Bone cell activity in clinical prostate cancer bone metastasis and its inverse relation to tumor cell androgen receptor activity. Int J Mol Sci 19(4):1223. https://doi.org/10.3390/ijms19041223
Thysell E, Köhn L, Semenas J, Järemo H, Freyhult E, Lundholm M, et al. (2021) Verification of the transcriptomic-based prostate cancer metastasis subtypes MetA-C, their clinical and biological relevance, and their independent prognostic value to genomic drivers. Manuscript.
Qian BZ, Pollard JW (2010) Macrophage diversity enhances tumor progression and metastasis. Cell 141(1):39–51. https://doi.org/10.1016/j.cell.2010.03.014
Ceccaldi R, Sarangi P, D’Andrea AD (2016) The Fanconi anaemia pathway: new players and new functions. Nat Rev Mol Cell Biol 17(6):337–349. https://doi.org/10.1038/nrm.2016.48
Asim M, Tarish F, Zecchini HI, Sanjiv K, Gelali E, Massie CE et al (2017) Synthetic lethality between androgen receptor signalling and the PARP pathway in prostate cancer. Nat Commun 8(1):374. https://doi.org/10.1038/s41467-017-00393-y
Atanga E, Dolder S, Dauwalder T, Wetterwald A, Hofstetter W (2011) TNFalpha inhibits the development of osteoclasts through osteoblast-derived GM-CSF. Bone 49(5):1090–1100. https://doi.org/10.1016/j.bone.2011.08.003
Wang X, Sun B, Wei L, Jian X, Shan K, He Q et al (2022) Cholesterol and saturated fatty acids synergistically promote the malignant progression of prostate cancer. Neoplasia 24(2):86–97. https://doi.org/10.1016/j.neo.2021.11.004
Xu Z, Huang L, Dai T, Pei X, Xia L, Zeng G et al (2021) SQLE mediates metabolic reprogramming to promote LN metastasis in castration-resistant prostate cancer. Onco Targets Ther 14:4285–4295. https://doi.org/10.2147/OTT.S315813
Marin-Aguilera M, Pereira MV, Jimenez N, Reig O, Cuartero A, Victoria I et al (2021) Glutamine and cholesterol plasma levels and clinical outcomes of patients with metastatic castration-resistant prostate cancer treated with taxanes. Cancers (Basel) 13(19):4960. https://doi.org/10.3390/cancers13194960
Gorman AM, Healy SJ, Jager R, Samali A (2012) Stress management at the ER: regulators of ER stress-induced apoptosis. Pharmacol Ther 134(3):306–316. https://doi.org/10.1016/j.pharmthera.2012.02.003
Hetz C (2012) The unfolded protein response: controlling cell fate decisions under ER stress and beyond. Nat Rev Mol Cell Biol 13(2):89–102. https://doi.org/10.1038/nrm3270
Acknowledgements
The authors want to acknowledge the help with the performance of the RNA sequencing by TATAA Biocenter, Gothenburg, Sweden, the research group of Jonas Nilsson, Sahlgrenska Center for Cancer Research, Department of Surgery, Institute for Clinical Sciences, University of Gothenburg, Sweden for help with setting up the flow cytometry analysis, Manuel Luna, Institute of Biomedicine, University of Gothenburg for help with preliminary interpretation of sequencing data. This work was supported by the Swedish Cancer Society [grant numbers CAN2017/478, CAN 2017/380, and 20 1055]; the Swedish state under the agreement between the Swedish government and the county councils, the ALF-agreement [grant number ALFGBG-138351], the Swedish Prostate Cancer Federation, Knut and Alice Wallenberg Foundation, and the research foundations of A. Gabrielsson [grant number FB19-107], and U and K-E. Winberg.
Funding
Open access funding provided by University of Gothenburg.
Author information
Authors and Affiliations
Contributions
JH: Performed all experiments, as well as initial data curation of sequencing data, interpretations of results, and drafting the manuscript. EF: Performed the statistical analysis on the RNA sequencing data. RB: Performed the QIAGEN IPA Analysis, AJ: Bioinformatic analysis, interpretation of results, JED: Conceptualization, of the study and interpretation of results. KW: Supervision, conceptualization, interpretation of the results, finalizing the manuscript. All authors participated in editing the manuscript and agree to the final version.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no potential conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, J., Freyhult, E., Buckland, R. et al. Osteoclasts directly influence castration-resistant prostate cancer cells. Clin Exp Metastasis 39, 801–814 (2022). https://doi.org/10.1007/s10585-022-10179-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10585-022-10179-2