Abstract
Background
Angiotensin-converting enzyme (ACE) inhibitors reduce clinical symptoms and improve outcome in patients with hypertension, heart failure, and stable coronary artery disease (CAD) and are among the most frequently used drugs in these patient groups. For hypertension, treatment is guided by the level of blood pressure. In the secondary prevention setting, there are no means of guiding therapy. Prior attempts to target ACE-inhibitors to those patients that are most likely to benefit have not been successful, mainly due to the consistency in the treatment effect in clinical subgroups. Still, for prolonged prophylactic treatment with ACE-inhibitors it would be best to target treatment to only those patients most likely to benefit, which would considerably lower the number needed to treat and increase cost-effectiveness. A new approach for such “tailored-therapy” may be to integrate information on the genetic variation between patients. Until now, pharmacogenetic research of the efficacy of ACE-inhibitor therapy in CAD patients is still in a preliminary stage.
Methods
The PERindopril GENEtic association study (PERGENE) is a substudy of the EUROPA trial, a randomized double-blind placebo-controlled multicentre clinical trial which demonstrated a beneficial effect of the ACE-inhibitor perindopril in reducing cardiovascular morbidity and mortality in 12.218 patients with stable coronary artery disease (mean follow-up 4.2 years). Blood tubes were received from patients at the beginning of the EUROPA trial and buffy coats were stored at −40°C at the central core laboratory. Candidate genes were selected in the renin–angiotensin-system and bradykinin pathways. Polymorphisms were selected based on haplotype tagging principles using the HapMap genome project, Seattle and other up-to-date genetic database platforms to comprehensively cover all common genetic variation within the genes. Selection also took into consideration the functionality of SNP’s, location within the gene (promoter) and existing relevant literature. The main outcome measure of PERGENE is the effect of genetic factors on the treatment benefit with ACE-inhibitors. The size of this pharmacogenetic substudy allows detection with a statistical power of 98% to detect a difference in hazard ratios (treatment effect) of 20% between genotypes with minor allele frequency of 0.20 (two-sided alpha 0.05).
Conclusion
The PERGENE study is a large cardiovascular pharmacogenetic study aimed to assess the feasibility of pharmacogenetic profiling of the treatment effect of ACE-inhibitor use with the perspective to individualize treatment in patients with stable coronary artery disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The efficacy of ACE-inhibitors to improve outcome has been demonstrated by several large clinical trials in patients with cardiovascular disease. These include post-myocardial infarction (MI) patients, patients with asymptomatic left ventricular systolic dysfunction, heart failure or a history of cerebrovascular disease, and stable CAD patients with preserved left ventricular function [1–7]. Currently, the use of ACE inhibitors is recommended in guidelines on the management of hypertension, stable CAD, MI, heart failure, and in the prevention of the progression of renal insufficiency in diabetes mellitus related kidney disease [8–10]. The ACE-inhibitor perindopril has been extensively studied in several large, controlled trials in a variety of patient groups with different etiologies [7, 11–14]. Of these, the EUROPA trial is noteworthy, as it is the only secondary prevention study with perindopril in a stable CAD population. ACE inhibitor research has also led to a better understanding of pathophysiological processes and maladaptive responses in the renin–angiotensin aldosterone system (RAAS). In particular, several sub-studies of EUROPA (including PERFECT, PERSPECTIVE and PERTINENT), have established that ACE inhibitors have additional effects beyond the blood pressure reduction alone such as the improvement of endothelial function, improvement of the neuro-humoral balance, and reduction of unfavorable remodeling of the coronary arteries [15–18].
For hypertension, treatment is guided by the level of blood pressure. In the secondary prevention setting, there are no means by which we can guide therapy. Many patients need to be treated, and absolute benefits are modest. It is not yet possible to predict in advance which patients are to benefit most from treatment. Prior attempts to target ACE-inhibitors to those patients who are most likely to benefit have not been successful, mainly because of the consistency in the treatment effect in clinically relevant subgroups based on simple clinical characteristics [19–21]. Treating only those patients who are most likely to benefit would considerably lower the number needed to treat and increase cost-effectiveness. A new approach to “tailored-therapy” concerns cardiovascular pharmacogenetics which examine the genetic determinants of patients’ responses to cardiovascular drugs; in other words, understanding why some drugs work better for some patients than others and why some patients are more likely to experience serious side-effects than others. Many (if not all) aspects of human physiology have genetic determinants and could therefore be subject to pharmacogenetic studies.
Several factors may be expected to play a dominant role in determining the response of a patient to therapy. In particular, in the case of ACE inhibition, genetic factors within the RAS pathway are likely to affect its pharmacodynamics and clinical efficacy. Genetic factors causing differences in drug absorption and metabolic clearance are also relevant. However, until now there are no strong leads to explore this pharmaco-genetically since there are no metabolic (CYP) genes linked specifically to ACE-inhibitors, although this might also be a relevant pathway to investigate in future research.
In previous studies, several genetic polymorphisms in RAAS genes have been associated with high blood pressure levels or an increased cardiovascular risk [22, 23]. Nearly all these studies focused at two polymorphisms, the ACE I/D polymorphism and the M235T polymorphism in the angiotensinogen gene. Because of limited power, due to limited study sample size, results have been inconsistent and the underlying questions not answered adequately. With regard to interaction between genetic factors and treatment response, the results are scarce. No prior studies have been performed yet with ACE-inhibitors at a large scale neither in a randomized setting nor in stable CAD (one of the major indications of ACE-inhibitors). Another important limitation of prior research in cardiovascular pharmacogenetics is the investigation of one or two polymorphisms within only one gene, thereby ignoring the well-documented feedback mechanisms within the RAAS and the fact that there are two angiotensin II receptors (AT1 and AT2), which have counteracting effects also in humans. We suggest that a more comprehensive coverage of genetic variation in multiple RAAS genes is needed, by using a correct haplotype approach. This is achieved by using the latest information on genetic variation and linkage disequilibrium patterns in the selection of haplotype-tagging SNP’s.
The PERGENE study aims at assessing the feasibility of pharmacogenetic profiling of ACE-inhibitor therapy in patients with stable CAD. We hypothesized that genetic polymorphism in the RAAS and kininogen–kallikrein–bradykinin pathways may influence the treatment effect of ACE-inhibitors in patients with stable CAD. The PERGENE study is unique in the field of pharmacogenetic studies because of the large sample size, a randomized and placebo-controlled design, and the availability of extensive and accurate phenotypic data. Also, the extensive selection of tagging SNP’s in multiple genes in both pathways ensures a new and comprehensive coverage of common genetic variation in the candidate genes. A detection of heterogeneity in the treatment benefit according to genetic determinants may lead to significant advances in tailored-therapy and personalized medicine (Appendix).
2 Methods
2.1 Study population and design
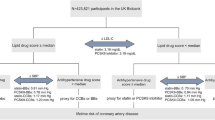
The PERGENE study is a substudy of the EUROPA-trial that will investigate genetic determinants of the treatment effect of ACE-inhibition in all subjects (Fig. 1). The study design of the EUROPA trial has been described in detail elsewhere [24]. In short, the EUROPA-trial was a randomized, double-blind, placebo-controlled clinical trial, with 12,218 patients who were randomized after a 4-week run-in period. Mean follow-up was 4.2 years. The study recruited men and women aged ≥18 years without clinical evidence of heart failure and with evidence of coronary artery disease documented by either previous MI, percutaneous or surgical coronary revascularization or angiographic evidence of ≥70% narrowing of ≥1 major coronary artery. Men were also recruited if they had a history of chest pain and a positive exercise test or regional wall motion abnormalities during stress echocardiography or nuclear scintigraphy or with transient perfusion defects during scintigraphy perfusion imaging.
The EUROPA study comprised a run-in period of 2 weeks during which patients received perindopril 4 mg/day, followed by 2 weeks during which patients received perindopril 8 mg/day provided that the 4 mg/day of perindopril was well tolerated. At the end of the run-in period, a double-blind treatment period of at least 36 months started during which patients received either perindopril 8 mg/day or placebo. Patients continued in the study until the last patient included completed the follow-up period. Following randomization, patients were seen at 3, 6 and 12 months and thereafter at six monthly intervals. Written informed consent was obtained from all patients for performing genetic association analyses.
2.2 Outcome measures
The main outcome measure of PERGENE is to assess (1) the effect of RAAS and bradykinin polymorphisms on the risk reduction of the primary endpoint of EUROPA (cardiovascular mortality, non-fatal MI, or successful resuscitated cardiac arrest) by perindopril. Other outcome measures are: (2) the effect of RAAS- and bradykinin polymorphisms on the amount of blood pressure reduction by perindopril treatment during the run-in period (in which all patient were treated with perindopril), (3) the effect of RAAS- and bradykinin polymorphisms on blood pressure, (4) the effect of RAAS- and bradykinin polymorphisms on incident cardiovascular risk during 4 years follow-up, and (5) the effect of RAAS- and bradykinin polymorphisms in relation to intolerance of ACE-inhibitors (Table 1). Other relevant hypotheses can be investigated in the future.
3 Data collection
3.1 Large scale blood sample collection (Fig. 1)
A logistic procedure and a high-quality program for the creation of a DNA bio-bank was established within the EUROPA trial [25]. Our group defined a successful protocol for large-scale blood samples collection in a large multicentre clinical trial. Blood sample were sent to a central laboratory (TNO Leiden, The Netherlands), registered, labeled and erythrocytes were lyzed. Buffy coats were frozen at −40°C in three aliquots. A total of 10,497 blood tubes of participants of the EUROPA-trial were received at the central core laboratory for storage. The average time between blood collection and processing was 10.1 (SD 8.4) days. The effect of transport time in the EUROPA trial was validated and shown to have no major effect on the DNA quality and quantity in a selection of 61 blood samples representing a wide range of transport times and countries. In all cases, sufficient amounts of DNA could be isolated and obtained from all samples; there was no relation with the region of origin, or with varying environmental temperatures, and the amount of DNA isolated. The quality and quantity of the DNA was high and well suited for performing genetic association studies [25].
3.2 DNA isolation
DNA was isolated from the stored white blood cells at the Genetic Laboratory of the department of Internal Medicine at the Erasmus MC using an automated isolation process (Hamilton liquid handler coupled with Magnetic separator for automated DNA extraction). The isolated DNA was stored in matrix 2D tubes and normalized and reformatted, using a pipetting robot, and dispensed into 384-well PCR plates using a Caliper Sciclone ALH3000 pipetting robot (Caliper LS, Mountain View, CA),
3.3 Candidate genes
The candidate genes to be studied are located in the RAAS and bradykinin pathways as presented in Fig. 2. ACE-inhibitors inhibit the angiotensin-converting enzyme, which is a central component of the RAAS and genes in this pathway are therefore highly relevant to be studied with respect to treatment effect of ACE-inhibitors. Candidate genes are ACE, angiotensinogen (AGT) and the angiotensin II receptor type 1 and type 2 gene (AGTR1, AGTR2). Also, several new and relatively unexplored genes in this pathway are of interest to investigate pharmaco-genetically, such as the renin (REN) and aldosterone synthase genes (CYP11b2). Moreover, as ACE cleaves bradykinin into inactive peptides, ACE-inhibitors increase bradykinin, which, amongst others, results in anti-remodeling, anti-atherosclerotic and anti-thrombotic effects, improves endothelial function and is also a strong vasodilator. As such, bradykinin counteracts the effect of angiotensin II in many ways. This pathway has not been explored yet with respect to pharmacogenetics, but may be very interesting, especially since the blood pressure effect and clinical effect of ACE-inhibitors are likely to be caused by bradykinin [26]. Relevant genes in this pathway are kallikrein (KLK), kininogen (KNG), and bradykinin-receptors (BDKRB 1 and 2), and the resultant endothelial nitric oxide synthase genes (eNOS3). This list of candidate genes ensures a comprehensive coverage of relevant genes in the RAAS and bradykinin pathways, which have not been investigated to this extent in prior studies.
Selected candidate genes in the renin–angiotensin–aldosteron-system and bradykinin pathways. The arrows with a plus mark correspond to a stimulatory effect and arrows with a minus mark to a inhibitory effect. To the left and right is a list of candidate genes attached at each corresponding level within the RAAS and bradykinin pathways. AGT angiotensinogen, REN renin, ACE angiotensin-converting enzyme, AGTR1 angiotensin II receptor type 1, AGTR2 angiotensin II receptor type 2, CYP11B aldosteron synthase, KLK kallikrein, KNG kininogen, BDKRB1 bradykinin receptor type 1, BDKRB2 bradykinin receptor type 2, eNOS3 nitric oxide synthase
3.4 Selection of single nucleotide polymorphisms (SNPs)
To cover the genetic variation in these candidate genes more comprehensively, the list of selected polymorphisms in the candidate genes were selected using the latest genetic information from available databases (such as dbSNP: http://www.ncbi.nlm.nih.gov/SNP and Celera http://www.celeradiscoverysystem.com) and our own research (WAVE dHPLC; sequencing). The final selection will also take into account the haplotype block structure of these SNPs by using HapMap data (HapMap Release 23a/Phase II Mar 08/on NCBI B36 assembly/ DbSNP b126), and SeattleSNP databases (see: http://hapmap.jst.go.jp/index.html and http://pga.gs.washington.edu), as well as Ensemble and OMIM. Tagging SNP’s are representative SNP’s in a region of the gene with high linkage disequilibrium, which makes it possible to identify genetic variation without genotyping every SNP within the gene and reach a high coverage of common variation within the gene with a limited number of tagging SNP’s. Linkage disequilibrium, as provided by the HapMap project, is a measure of the non-random association between polymorphisms at different loci and it describes a situation in which some combinations of alleles or genetic markers occur with a higher or lower frequency in a population than would be expected from a random formation of haplotypes from alleles based on their frequencies. We selected tagging SNP’s are within the candidate genes using Caucasian subjects as reference population since in the EUROPA-trial more than 99% of the patients were of Caucasian origin. In Haploview, we will use a cut-off of minor allele frequency of 5% and haplotype frequency of 5% with r 2 of 0.80 to select the haplotype tagging SNP’s within the candidate genes and aim to achieve more than 90% coverage of common genetic variation. The selected area of the gene always should contain about 2 kb at the 5′ and 3′ ends of the gene to ensure maximum coverage of functionally relevant genetic areas. In selecting these haplotype-tagging SNP’s, we prefer to use functional SNP’s or SNP’s located in regulatory or promoter regions of the gene. In addition, we will add several relevant SNP’s to our list based on prior literature.
3.5 Genotyping
We will use a high-throughput genotyping facility including a Caliper Sciclone ALH 3000 pipetting robot (including a TwisterII, and integrated plate sealer, plate reader (OD260/280)), and polymerase-chain-reaction (PCR) machine (ABI 9700, 2 × 384)), an ABI7900HT Taqman (running 2 ng gDNA in 2 μL reactions). The most commonly used genotyping techniques are Taqman (for one to ten SNP’s) and Sequenom (for five to 40 SNP’s).
3.6 Allelic discrimination with Taqman
Genotypes will be determined in genomic DNA with the Taqman allelic discrimination assay (Applied Biosystems, Foster City, CA, USA). Assay-by-Design service (http://www.appliedbio-systems.com) will be used to set up a TaqMan allelic discrimination assay for the selected SNP’s; primer designs are readily available at ABI. The PCR mixture includes 1–2 ng genomic DNA in a 2-μl volume and the following reagents: probes (200 nM), primers (0.9 μM), 2× Taqman PCR master mix (AB gene, or ABI). Reagents are dispensed in a 384-well plate using the Deerac Equator NS808 (Deerac Fluidics, Dublin, Ireland). PCR cycling reactions will be performed in 384-well PCR plates in an ABI 9700 PCR system (Applied Biosystems Inc., Foster City, CA). These consist of initial denaturation for 15 min at 95°C and 40 cycles with denaturation for 15 s at 95°C and annealing and extension for 60 s at 60°C. Allele-specific fluorescence was then analyzed on an ABI Prism 7900HT Sequence Detection System with SDS v 2.1 (Applied Biosystems) and results analyzed by the ABI TaqMan 7900HT using the sequence detection system 2.22 software (Applied Biosystems). To confirm the accuracy of the genotyping, 5% randomly selected samples and duplicates will be included and genotyped additionally in the same procedure.
3.7 Mass-spectrometric genotyping
Polymerase-chain-reaction (PCR) assays and extension primers for these SNP’s are designed with the use of MassARRAY software, version 3.0 (Sequenom). The MassARRAY Designer software can automatically design both PCR and MassEXTEND primers for multiplexed assays. MassEXTEND is a primer extension process designed to detect sequence differences at the single nucleotide level. The primer is extended, dependent upon the template sequence, resulting in an allele-specific difference in mass between extension products. This mass difference allows the data analysis software to differentiate between SNP alleles. PCR and extension reactions will be performed according to the manufacturer's instructions, and extension product sizes determined by mass spectrometry (Sequenom). The iPLEX Gold assay uses a single termination mix and universal reaction conditions for all SNP’s. The SpectroCHIP arrays are placed into the MALDI-TOF mass spectrometer and the mass correlating genotype is determined in real time. Duplicate test samples (control plates, 5%) and six water samples per plate (PCR-negative controls), of which the technician is unaware, are included in each 96-well plate. The rate of concordant results between duplicate samples will be checked.
Quality control for the genotyping will further involve testing for Hardy–Weinberg equilibrium and repeated laboratory analyses in a random group of samples. To ensure high-quality output of SNP, eg. if the call rate or clustering was difficult, the SNP can be tested with alternative approaches (e.g. Taqman reaction mixtures) or on the other genotyping platform (Sequenom) likewise.
4 Data analysis
4.1 Statistical analysis and statistical power
In the EUROPA study, 9% of patients (n = 1091) had a major cardiovascular event during follow-up (cardiovascular death, MI, cardiac arrest: 603 (10%) placebo and 488 (8%), 20% relative risk reduction (95% CI 9–29, p = 0.0003). A secondary endpoint (total mortality, non-fatal MI, hospital admission for UAP, cardiac arrest) occurred in 16% of patients (n = 1,947). Event rates will be compared between treatment groups and the treatment effect will be compared between genotype strata (for each gene). Gene–drug interaction for relative risk reductions will be assessed using Cox proportional hazards regression analysis. Hardy–Weinberg equilibrium of each polymorphism will be tested using Chi-square analysis. All analyses will be based on intention to treat. A p-value of 0.05 or less will be deemed significant.
4.2 Haplotype analysis
Haplotypes will be inferred with use of the program Haplo.Stats and R. Haplotype alleles present in the patient population were inferred by means of the haplo.em function of the program Haplo Stats (http://cran.r-project.org/src/contrib/Descriptions/haplo.stats.html), which computes maximum likelihood estimates of haplotype probabilities.
4.3 Sample size considerations
The large study size and large number of events (n > 1,000) will provide sufficient power for the detection of interaction effects. The size of this pharmacogenetic substudy allows detection with a power of 98% to detect a difference in hazard ratios (treatment effect for the primary endpoint) of 20% between genotypes with minor allele frequency of 0.20, based upon ten-thousand patients (two-sided alpha 0.05). For other genotype distributions, power will be less but for most comparisons still above 80%. For a minor allele frequency of 0.10, statistical power is 88% to detect a difference in treatment effect of 20% with a two-sided alpha 0.05 (Fig. 3).
5 Discussion
In European countries and in other countries worldwide, millions of patients have coronary artery disease, and are at risk of (recurrent) events, particularly cardiovascular death and myocardial infarction. The ACE-inhibitors are among one of the most frequently prescribed drugs for secondary prevention in patients with stable coronary artery disease. Still, in the secondary prevention setting, there are no means of guiding therapy to those patients who are most likely to benefit. Prior attempts to target ACE-inhibitors have not been successful, largely due to because of the consistency in the treatment effect in clinical subgroups [19–21].
In the EUROPA study, treatment with perindopril in 50 patients for a period of 4 years (200 patient years) was required to prevent one major cardiovascular event. The efficiency, and the cost-effectiveness of such prolonged prophylactic treatment would be significantly enhanced if patients who do benefit and those who do not benefit from an ACE-inhibitor could be distinguished prior to the start of treatment. For this purpose, it is important to study possible heterogeneities in the treatment effect to assess whether a variation in the treatment effect does exist which then can be used to guide ACE-inhibitor therapy to the patients most likely to benefit of treatment.
There are several arguments that this variation in treatment effect may indeed exist. For instance, the activity of ACE and angiotensinogen (plasma levels) have been shown to vary widely between patients, also in the response to an ACE-inhibitor [17]. Also, a large inter-individual variability in blood pressure response to ACE inhibition is well documented. Whether the treatment effect on outcome reduction varies in a similar way is unknown. Until now subgroup analysis based on clinical characteristics has resulted in consistent treatment effects [19–21]. Still, relative and absolute risk reductions with ACE inhibitor therapy vary. A larger risk reduction is seen in heart failure patients as compared to coronary artery disease patients [19].
Therefore it is likely that the treatment effect of ACE-inhibitor use on outcome (in CAD patients) will also differ, some patients will benefit others will not. The integration of genetic information, which is highly specific for each individual patient, can be a new way to identify a true heterogeneity in the treatment effect of ACE-inhibitors. A pharmacogenetic profile related to drug response can be used to target ACE-inhibitors to those patients most likely to benefit of treatment.
It has been suggested that the response to drug therapy may be influenced by genetic polymorphisms in different ways. Firstly, variations within genes of the RAAS and related systems may influence the disease process (atherosclerosis) and inherent differences in accessibility to therapeutic agents such as ACE-inhibitors. Secondly, pharmacodynamics may be affected by polymorphisms in the genes of all proteins involved in the RAAS and related systems, including receptors and signal transduction molecules. Thirdly, variations in drug absorption and metabolic clearance may cause inter-individual variation in pharmacokinetics.
Genetic polymorphisms in the ACE and AGT genes have been shown to influence plasma levels of these enzymes, mainly M235T and T174M in AGT and ACE I/D in ACE [27]. For example with the M235T polymorphism, the TT-allele results in higher levels of angiotensinogen.
Regarding intermediate endpoints, a series of relatively small studies (34 to 345 patients) reported that polymorphisms in the ACE, angiotensinogen and AGTR1 and AGTR2 genes modulated the effects of ACE-inhibitors [28]. Such interactions were shown for specific alleles in relation with blood pressure reduction, regression of left ventricular hypertrophy, diastolic cardiac function and re-stenosis after percutaneous coronary intervention.
Regarding outcomes, pharmacogenetic data is very scarce. The PROGRESS study, which included 5,688 patients with history of stroke, did not show an association of the ACE I/D polymorphism and ACE-inhibition on risk reduction of cerebrovascular events [29]. The inconsistency in these studies could be explained by the fact that in virtually all association studies only one RAAS gene was taken into consideration, thereby ignoring the well-documented feedback mechanisms within the RAAS [30]. For instance, the elevated angiotensinogen levels which are found in Thr235 homozygotes are accompanied by reduced renin levels, so that the angiotensin I-generating capacity returns to normal in those with high angiotensinogen concentrations. In addition, the two angiotensin II receptors (AT1 and AT2) that exist in humans have counteracting effects [31, 32]. Thus, future studies should preferably investigate more than one RAAS gene (and ideally all genes: renin, angiotensinogen, ACE, AGTR1, AGTR2, aldosterone synthase).
The DNA samples collected in the EUROPA study offer a unique opportunity to investigate the relations between polymorphisms in genes of the RAAS with the treatment benefit of an ACE inhibitor on cardiovascular events in a sufficiently large population and in a randomized double-blind setting. As mentioned, the available studies of this subject have been of small size, not randomized and therefore, reported relationships may have been due to chance findings. Furthermore, the majority of studies so far included only one polymorphism or one gene of the RAAS.
In contrast, the PERGENE study uses a haplotype tagging selection procedure to comprehensively cover all common genetic variations (>90%) in the relevant genes within the RAAS and bradykinin pathways. We will use the latest information from HapMap Genome project, SEATTLE and other up to date genetic information platforms as well as sophisticated software packages as Haplostats and R for these haplotype analyses.
Haplotypes are a combination of alleles at different markers along the same chromosome that are inherited as a unit (linkage disequilibrium-patterns). The determination of haplotypes is essential for understanding genetic variation and the inheritance of complex diseases. An analysis based on haplotypes is advantageous over an analysis based on individual SNPs, especially in the presence of multiple susceptibility alleles, and when linkage disequilibria between SNPs are weak. With a single SNP approach, associations may be missed when the causal SNP is not in linkage disequilibrium with the single analyzed SNP. It is more informative to simultaneously analyze multiple markers in a region of interest that identifies genetic variants underlying various human traits; Also, these markers should be selected based on tagging principles and linkage disequilibrium. By combining information from multiple SNP’s in the RAAS and bradykinin pathway genes, a more efficient and comprehensive in-depth analysis of common genetic variation in relation to ACE-inhibitor therapy is performed, which is more likely to unravel any important pharmacogenetic associations.
In summary, this project is unique because of its size, design (randomized-setting), accurate phenotypic data, complete coverage of two pathways (RAAS and bradykinin), but also because of the extensive and comprehensive SNP selection procedure which involves multiple SNP in multiple genes of both pathways with integrating information on the haplotype structure of RAAS and bradykinin genes.
Until now, attempts to target therapy using simple clinical patient characteristics have been insufficient to guide ACE-inhibition therapy and it is not yet possible to say in advance who to treat or not [19–21]. New and improved approaches that integrate more patient-specific characteristics are needed to target ACE-inhibitor therapy. We will investigate whether specific genetic polymorphisms in RAAS genes modify the treatment effect of ACE-inhibitor therapy. Our aim is to develop a pharmacogenetic profile associated with the benefit of ACE inhibitor therapy in patients with stable coronary artery disease.
If it is possible to construct a pharmacogenetic profile related to treatment benefit, this could lead to a significant reduction of the number of patients needed to treat. It should be realized in this regard that the absolute treatment effects are only modest (about 2% absolute risk reduction) in stable CAD patients.
In the EUROPA trial, 200 patient years of treatment with perindopril 8 mg was necessary to prevent one event in the primary endpoint (number needed to treat 50 for 4 years) [7, 33]. A pharmaco-genetic profile related to the benefit of perindopril may enable the selection of those patients advance of treatment. Likewise, targeting therapy to only those patients that are to benefit will considerably increase the cost-effectiveness of treatment. Until now, cardiovascular pharmacogenetic research is still in a premature stage but has the potential to enhance personalized medicine and tailored-therapy in cardiovascular medicine.
References
Pfeffer MA, Braunwald E, Moye LA, SAVE Investigators, et al. Effect of captopril on mortality and morbidity in patients with left ventricular dysfunction and myocardial infarction: results of the survival and ventricular enlargement trial. N Engl J Med 1992;327:669–77.
The SOLVD investigators. Effect of enalapril on survival in patients with reduced left ventricular ejection fractions and congestive heart failure. N Engl J Med 1991;325:293–302.
The Acute Infarction Ramipril Efficacy (AIRE) study investigators. Effect of ramipril on mortality and morbidity of survivors of acute myocardial infarction with clinical evidence of heart failure. Lancet 1993;342:821–8.
Torp-Pedersen C, Kober L, TRACE Study Group. Effect of ACE inhibitor trandolapril on life expectancy of patients with reduced left-ventricular function after acute myocardial infarction. Lancet 1999;354:9–12.
The Heart Outcomes Prevention Evaluation study investigators. Effects of an angiotensin-converting enzyme inhibitor, ramipril, on cardiovascular events in high-risk patients. N Engl J Med 2000;342:145–53.
The PEACE Trial Investigators. Angiotensin-converting-enzyme inhibition in stable coronary artery disease. N Engl J Med 2004;351:2058–68.
The European Trial on Reduction of Cardiac Events with Perindopril in Patients with Stable Coronary Artery Disease investigators. Efficacy of perindopril in reduction of cardiovascular events in stable coronary artery disease: randomized, double-blind, placebo controlled, multicentre trial (the EUROPA study). Lancet 2003;362:782–8.
The Task Force on the Management of Stable Angina Pectoris of the European Society of Cardiology. Guidelines on the management of stable angina pectoris. Eur Heart J 2006;27:1341–81.
Wood D, DeBacker G, Faergam O, Graham I, Mancia G, Pyörälä K, and the task force. Prevention of coronary disease in clinical practice: recommendation of the second Joint Task Force of European and other societies on Coronary Prevention. Atherosclerosis 1998;140:99–270.
Gibbons RJ, Abrams J, Chatterjee K, et al. ACC/AHA 2002 guideline update for the management of patients with chronic stable angina—summary article: a report of the American College of Cardiology/American Heart Association Task Force on practice guidelines (Committee on the Management of Patients With Chronic Stable Angina). J Am Coll Cardiol 2003;41:159–68.
Dahlöf B, Sever PS, Östergren J. Prevention of cardiovascular events with an antihypertensive regimen of amlodipine adding perindopril as required versus atenolol adding bendroflumethiazide as required, in the Anglo-Scandinavian Cardiac Outcomes Trial-Blood Pressure Lowering Arm (ASCOT-BPLA): a multicentre randomized controlled trial. Lancet 2005;366:895–906.
PROGRESS Collaborative Group. Randomised trial of a perindopril based blood pressure-lowering regimen among 6105 individuals with previous stroke or transient ischemic attack. Lancet. 2001;200(358):1033–41.
The PREAMI Investigators. Effects of angiotensin-converting enzyme inhibition with perindopril on left ventricular remodeling and clinical outcome. Results of the Randomized Perindopril and Remodeling in Elderly with Acute Myocardial Infarction (PREAMI) Study. Arch Intern Med 2006;166:659–66.
ADVANCE Collaborative Group. Effects of a fixed combination of perindopril and indapamide on macrovascular and microvascular outcomes in patients with type 2 diabetes mellitus (the ADVANCE trial): a randomized controlled trial. Lancet 2007;370:829–40.
Bots ML, Remme WJ, Lüscher TF, et al. ACE inhibition and endothelial function: main findings of PERFECT, a sub-study of the EUROPA trial. Cardiovasc Drugs Ther 2007;21:269–79.
Rodriguez-Granillo GA, Vos J, De Feyter PJ. Long-term effect of perindopril on coronary atherosclerosis progression (from the PERindopril’s Prospective Effect on Coronary aTherosclerosis by Angiography and IntraVascular Ultrasound Evaluation [PERSPECTIVE] Study). Am J Cardiol 2007;100:159–63.
Ceconi C, Fox KM, Ferrari R, et al. ACE inhibition with perindopril and endothelial function. Results of a substudy of the EUROPA study: PERTINENT. Cardiovasc Res 2007;73:237–46.
Remme WJ, Deckers JW, Fox KM, Ferrari R, Bertrand M, Simoons ML. for the EUROPA investigators. Secondary prevention of coronary disease with ACE inhibition—does blood pressure reduction with perindopril explain the benefits in EUROPA? Cardiovasc Drugs Ther 2009;23 (this issue).
Dagenais GR, Pogue J, Fox K, Simoons ML, Yusuf S. Angiotensin-converting-enzyme inhibitors in stable vascular disease without left ventricular systolic dysfunction or heart failure: a combined analysis of three trials. Lancet 2006;368:581–8.
Deckers JW, Goedhart DM, Boersma E, et al. Treatment benefit by perindopril in patients with stable coronary artery disease at different levels of risk. Eur Heart J 2006;27:796–801.
Brugts JJ, Boersma E, Chonchol M, et al. The cardioprotective effects of the angiotensin-converting enzyme inhibitor perindopril in patients with stable coronary artery disease are not modified by mild to moderate renal insufficiency: insights from the EUROPA trial. J Am Coll Cardiol 2007;50:2148–55.
Brugts JJ, de Maat M, den Uil CA, et al. Pharmacogenetics of ACE inhibition in stable coronary artery disease: steps towards tailored drug therapy. Curr Opin Cardiol 2008;23:296–301.
Brugts JJ, den Uil CA, de Maat M, Danser AHJ. The renin–angiotensin–aldosteron system: approaches to guide angiotensin-converting enzyme inhibition in patients with coronary artery disease. Cardiology 2008;112:303–12.
Gomma AH, Fox KM, for the EUROPA investigators. The EUROPA trial: design, baseline demography and status of the substudies. Cardiovasc Drugs Ther 2001;15:169–79.
Nederhand RJ, Droog S, Kluft C, Simoons ML, de Maat MP. Logistics and quality control for DNA sampling in large multicenter studies. J Thromb Haemost 2003;1:987–91.
Gainer JV, Morrow JD, Loveland A, King DJ, Brown NJ. Effect of bradykinin-receptor blockade on the response to angiotensin-converting-enzyme inhibitor in normotensive and hypertensive subjects. N Engl J Med 1998;339:1285–92.
Danser AHJ, Schunkert H. Renin–angiotensin system gene polymorphisms: potential mechanisms for their association with cardiovascular diseases. Eur J Pharmacol 2000;410:303–16.
Schelleman H, Stricker BH, De Boer A, et al. Drug–gene interactions between genetic polymorphisms and antihypertensive therapy. Drugs 2004;64:1801–16.
Harrap SB, Tzourio C, Cambien F. The ACE gene I/D polymorphism is not associated with the blood pressure and cardiovascular benefits of ACE inhibition. Hypertension 2003;42:297.
Danser AH, Derkx FH, Hense HW, Jeunemaitre X, Riegger GA, Schunkert H. Angiotensinogen (M235T) and angiotensin converting enzyme (I/D) polymorphisms in association with plasma renin and prorenin levels. J Hypertens 1998;16:1879–83.
Deinum J, van Gool JM, Kofflard MJ, ten Cate FJ, Danser AH. Angiotensin II type 2 receptors and cardiac hypertrophy in women with hypertrophic cardiomyopathy. Hypertension 2001;38:1278–81.
Batenburg WW, Garrelds IM, Bernasconi CC, et al. Angiotensin II type 2 receptor-mediated vasodilation in human coronary microarteries. Circulation 2004;109:2296–301.
Briggs A, Mihaylova B, Sculpher M, EUROPA-trial investigators, et al. Cost effectiveness of perindopril in reducing cardiovascular events in patients with stable coronary artery disease using data from the EUROPA study. Heart 2007;93:1081–6.
Acknowledgements
The PERGENE study is supported by a grant from the Netherlands Heart Foundation (NHS2005B219). Dr. Brugts is supported by a grant from the Netherlands Heart Foundation (NHS2005B219) and a grant from the Netherlands Organization for Health Research and Development (ZonMW).
Conflict of interest
None.
Financial disclosure
None.
The EUROPA trial was supported by Servier, Paris, France. For the current pharmacogenetic analysis of PERGENE, the funding source had no participating role in any form.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Consortia
Corresponding author
Appendix
Appendix
1.1 Participating PERGENE center
The Netherlands: Erasmus MC Thoraxcenter, Department of Cardiology, Rotterdam.
1.2 PERGENE investigators
Brugts JJ, de Maat MPM, Boersma E, Witteman JCM, van Duijn C, Uitterlinden AG, Danser AHJ, Simoons ML.
We gratefully acknowledge the EUROPA-investigators.
1.3 EUROPA Executive Committee
Bertrand M, Ferrari R, Fox K, Remme WJ, Simoons M.L.
1.4 EUROPA Steering Committee
Aldershville J, Denmark; Hildebrandt P, Bassand JP, France; Cokkinos D, Greece; Toutouzas P, Greece, Eha J, Estonia; Erhardt L, Sweden; Erikssen J, Noway; Grybauskas R, Lithuania; Kalnins U, Latvia; Karsch K, Sechtem U, Germany; Keltai M, Hungary; Klein W, Austria; Luescher T, Switzerland; Mulcahy D, Ireland; Nieminen M, Finland; Oto A, Ozsaruhan O, Turkey; Paulus W, Belgium; Providencia L, Portugal; Riecansky I, Slovakia; RuzylloW, Poland; Santini U, Tavazzi L, Italy; Soler-Soler J, Spain;WidimskyP, Czech Republic.
1.5 EUROPA Safety Committee
Julian D, Murray, Dargie H, United-Kingdom; Kobler W, Germany;
1.6 EUROPA End Point Validation Committee
Duprez D, Belgium; Steg G, France; Thygesen K, Denmark;
1.7 EUROPA investigators
Austria, Drexel H, Gombotz G, Klein W, Stoeckl G, Belgium, Duprez D, Heyndrickx GH, Legrand V, Materne P, Van Mieghem W, Czech Republic, Bocek P, Branny M, Cech M, Charouzek J, Drazka J, Fabik L, Florian J, Francek L, Groch L, Havranek P, Herman A, Hradec J, Jansky P, Jirmar R, Jokl I, Krejcova H, Kvasnak M, Maratka T, Marcinek G, Moravcova J, Nedbal P, Peterka K, Povolny J, Ass. Rosolova H, Semrad B, Sochor K, Spacek R, Spinar J, Stipal R, Stuchlik K, Sulda M, Ulman J, Vaclavicek A, Vojtisek P, Denmark, Bjerregard Andersen H, Dorff B, Hildebrandt P, Kristensen K, Madsen JK, Markenvard J, Meibom J, Norgaard A, Scheibel M, Estonia, Eha J, Leht A, Teesalu R, Vahulaa V, Finland, Itkonen A, Juvonen J, Karmakoski J, Kilkki E, Koskela E, Kotila MJ, Melin J, Nieminen MS, Savola R, Terho T, Voipio Pulkki LM, France, Apffel F, Attali P, Baron B, Bassand JP, Berthier Y, Bertrand M, Dambrine P, Danchin N, Decoulx E, Deshayes P, Fouche R, Genest M, Godard S, Guillot JP, Hanania G, Lelguen C, Leroy F, Mansourati J, Mery D, Michel, Quiret JC, Raynaud P, Rondepierre D, Roynard JL, Van Belle E, Veyrat A, Germany, Gaudron P, Karsch KR, Lauer B, Rettig Stˇsrmer G, Riessen R, RutschW, Sechtem U, Sigel HA, Simon R, Stork S, Von Schacky C, Winkelmann BR, Greece, Christakos S, Cokkinos D, Feggos S, Geleris P, Georgiadis S, Gialafos J, Goudevenos I, Kardara D, Kardaras F, Karidis C, Kelesides C, Kyriakidis M, Koliopoulos N, Kremastinos D, Liberi S, Manolis AN, Pyrgakis V, Papasteriadis E, Papazoglou N, Skoufas P, Stamatelopoulos S, Stambola S, Stavridis A, Syribeis S, Vardas P, Vassiliadis I, Voudris V, Zacharoulis A, Zobolos S, Zouras C, Hungary, Berenyi I, Bocsa Z, Csendes E, Edes I, Gelesz E, Janosi A, Kalo E, Karpati P, Kornel S, Pap I, Pinter I, Polak G, Reiber I, Rusznak M, Simon A, Tarjan J, Tihanyi L, Timar S, Toth K, Veress G, Ireland, Barton J, Crean P, Daly K, Kearney P, Meany TB, Mulcahy D, Quigley P, Italy, Azzolini P, Barone G, Barsotti A, Bellone E, Borghetti A, Branzi A, Brunelli C, Capponi E, Capucci A, Casaccia M, Casali G, Cecchetti E, Ceci V, Celegon L, Chimini C, Colombo A, Corsini G, Cucchini F, Dalla Volta S, De Luca I, De Servi S, Delise P, Di Donato M, Di Giacomo U, Di Pasquale G, Fiorentini C, Gaddi O, Giannetto M, Giannuzzi P, Giordano A, Giovannini E, Iacono A, Inama G, Ippoliti G, Leghissa R, Lorusso R, Marzilli M, Minutiello L, Moretti G, Mosele GM, Pasotti C, Pettinati G, Pezzano A, Polimeni MR, Portaluppi F, Proto C, Riva S, Sanguinetti M, Santini M, Severi S, Sinagra G, Tantalo L, Tavazzi L, Vajola SF, Vincenzi M, Volterrani M, Zavatteri G, Zogno M, Latvia, Gailiss E, Gersamija A, Kalnins U, Ozolina MA, Skards J, Lithuania, Baubiniene A, Berukstis E, Grigoniene L, Grybauskas, Kibarskis A, Kirkutis A, Marcinkus R, Milvidaite I, Vasiliauskas D, Netherlands, Aalders JCA, Bruggeling WAJ, Bucx JJJ, De Feyter PJ, De Leeuw MJ, De Waard DEP, De Weerd GJ, De Zwaan C, Dijkgraaf R, Droste HT, Freericks MP, Hagoort Kok AW, Jap WTJ, Jochemsen GM, Kiemeney K, Kuijer PJP, Mannaerts HFJ, Piek JJ, Saelman JPM, Simoons ML, Slob FD, Smits WCG, Spierenburg HAM, Suttorp MJ, Tan TB, Van Beek GJ, Van Daele MERM, Van Den Merkhof LFM, Van Den Toren EW, Van Hessen MWJ, Van Langeveld RAM, Van Loo LWH, Van Nierop PR, Van Rey FJW, Van Straalen MJ, Vos J, Werner HA, Westendorp JJC, Zwiers G, Norway, Erikssen J, Poland, Achremczyk P, Adamus J, Baska J, Bolinska Soltysiak H, Bubinski R, Ceremuzynski L, Cieslinski A, Dariusz D, Deptulski T, Drewla P, Drozdowski P, Dubiel JS, Dudek D, Galewicz M, Ghlebus K, Halawa B, Jakubowska Majnigier M, Janion M, Jaworska K, Kaszewska I, Kleinrok A, Kornacewicz Jach Z, Krawczyk W, Krynicki R, Krzciuk M, Krzeminska Pakula M, Kuch J, Kuzniar J, Legutko J, Liszewska Pfejfer D, Loboz Grudzien K, Musial W, Opolski G, Pasyk S, Piwowarska W, Pulkowski G, Radziszewski B, Romanski B, Ruzyllo W, Rynkiewicz A, Skura M, Slowinski S, Smielak Korombel W, Targonski R, Templin W, Tendera M, Tracz W, Trusz Gluza M, Turek P, Tyminska Sedek K, Wodniecki J, Zalewski M, Zinka E, Portugal, Carrageta M, Coelho Gil J, Ferreira R, Leitao Marques A, Santos Andrade CM, Seabra Gomes R, Slovakia, Bada V, Belicova M, Buksarova E, Dukat A, Gonsorcik J, Jonas P, Kamensky G, Karvaj M, Micko K, Mikes Z, Palinsky M, Pella D, Renker B, Riecansky I, Sefara P, Sojka G, Sulej P, Szakacs M, Szentivanyi M, Spain, Aguirre Salcedo JM, Alonso Orcajo N, Ancillo Garcia P, Auge Sanpera JM, Ayuela Azcarate J, Bardaji Mayor JL, Bertomeu Martinez V, Blanco Coronado JL, Bros Caimari R, Bruguera Cortada J, CaparrosValderrama J, DeArmas Trujillo D, Del Rio Ligorit A, Espinosa Caliani JS, Fernandez Aviles F, Garcia Guerrero JJ, Garcia Lopez D, Gonzalez Cocina E, Guallar Urena C, Jodar Lorente L, Lopez Garcia ArandaV, MacayaDeMiguel C, Maroto Montero J, Martinez Romero P, Navarro Lopez F, Noriega Peiro F, Olague De Ros J, Orellana Mas J, Paz Bermejo MA, Placer Peralta LJ, Rodriguez Padial L, Salvador Sanz A, Segui Bonnin J, Simarro Martin E, Sobrino Daza J, Valles Belsue F, Sweden, Ekdahl S, Erhardt L, Forslund L, Ohlin H, Persson S, Switzerland, Pieper M, Moccetti T, Turkey, Acartˇsrk E, Guzelsoy D, Oto A, TMZsaruhan O, Ozturk M, SansoyV, Turkoglu C, Yˇsksel H, United Kingdom, Adgey AAJ, Ahsan A, Al Khafaji M, Ball SG, Birkhead J, Boon N, Bowker T, Brack M, Bridges A, Buchalter M, Buchalter B, Calder B, Cooke RA, Corr L, Cowell R, Curzen NP, Davidson C, Davies E, Davies J, De Belder M, Dhiya L, Doig JC, Findlay IN, Fox K, Francis CM, Glancy JM, Glen S, GreenwoodTW, Groves P, Hall AS, Hamilton G, Hillman R, Holdright D, HubbardW, Travill C, Hutton I, Ilsley C, Innes M, James M, Jennings K, Jones CJH, Joy M, Keeling P, Kooner J, Lawson C, Levy RD, Lip G, Lorimer AR, Marshall H, Mclachlan B, Montgomery H, Morley C, Murdoch DL, Muthusamy R, Oakley GD, PennyW, Pohl JEF, Purvis J, Pye M, Ramsdale D, Reid D, Roberts DH, Rowlands D, Rozkovec A, Saltissi S, Schofield PM, Scott M, Shapiro LM, Silke B, Stephens J, Sutherland S, Swan JW, Shakespeare C, Tildesley G, Travill C, Watson RDS, Wilkinson P.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Brugts, J.J., de Maat, M.P.M., Boersma, E. et al. The Rationale and Design of the Perindopril Genetic Association Study (PERGENE): A Pharmacogenetic Analysis of Angiotensin-Converting Enzyme Inhibitor Therapy in Patients with Stable Coronary Artery Disease. Cardiovasc Drugs Ther 23, 171–181 (2009). https://doi.org/10.1007/s10557-008-6156-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10557-008-6156-1