Abstract
Purpose
Artificial intelligence (AI) for reading breast screening mammograms could potentially replace (some) human-reading and improve screening effectiveness. This systematic review aims to identify and quantify the types of AI errors to better understand the consequences of implementing this technology.
Methods
Electronic databases were searched for external validation studies of the accuracy of AI algorithms in real-world screening mammograms. Descriptive synthesis was performed on error types and frequency. False negative proportions (FNP) and false positive proportions (FPP) were pooled within AI positivity thresholds using random-effects meta-analysis.
Results
Seven retrospective studies (447,676 examinations; published 2019–2022) met inclusion criteria. Five studies reported AI error as false negatives or false positives. Pooled FPP decreased incrementally with increasing positivity threshold (71.83% [95% CI 69.67, 73.90] at Transpara 3 to 10.77% [95% CI 8.34, 13.79] at Transpara 9). Pooled FNP increased incrementally from 0.02% [95% CI 0.01, 0.03] (Transpara 3) to 0.12% [95% CI 0.06, 0.26] (Transpara 9), consistent with a trade-off with FPP. Heterogeneity within thresholds reflected algorithm version and completeness of the reference standard. Other forms of AI error were reported rarely (location error and technical error in one study each).
Conclusion
AI errors are largely interpreted in the framework of test accuracy. FP and FN errors show expected variability not only by positivity threshold, but also by algorithm version and study quality. Reporting of other forms of AI errors is sparse, despite their potential implications for adoption of the technology. Considering broader types of AI error would add nuance to reporting that can inform inferences about AI’s utility.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Artificial intelligence (AI), a rapidly evolving field of data science in which computer algorithms are developed to perform complex tasks, has been applied to screening mammography for the early detection of breast cancer with the aim of improving outcomes for screening participants [1]. AI has the potential to identify cancers in mammograms that are not perceptible to human readers, thereby potentially increasing the sensitivity of screening and improving outcomes for women through initiation of treatment for early-stage disease. Other proposed benefits of AI include fewer false positive findings that lead to anxiety and unnecessary investigations, and workforce efficiencies for screening programmes that may translate to lower programme costs and improvements in the screening experience for women. Such benefits assume that AI performs at least as accurately as human readers in detecting breast cancer, and research has therefore focussed on evaluating the comparative accuracy of algorithms and human readers. However, there is recognition that even when algorithms exhibit high performance in selected research datasets, AI errors in cancer detection (false positives, FP; false negatives, FN) may be greater when algorithms are applied in “real-world” settings or transferred between populations [2]. Furthermore, technological updates can produce subtle changes to medical images which may not be obvious to humans but can alter the AI’s output [3]. Such errors may be difficult to detect and explain by humans [4] and may strongly influence decision making by human readers (automation bias) [5]. Given the theoretical ease for AI algorithms to be scaled up and applied to large populations, unpredictable or unexpected errors may lead to harmful consequences.
Beyond the potential for FP or FN cancer findings, the concept of AI “error” in automated mammography interpretation has not been clearly delineated. Other types of error may include a (true positive) cancer detected in the wrong location, or technical errors that result in the algorithm failing to process images or generate a result. Earlier systematic reviews presented AI error as FP and FN, which is consistent with the focus on test accuracy in the literature [6,7,8]. However, imaging or lesion features associated with these FP and FN were not elaborated, and other potential forms of error were not reported. At present, it is unclear what forms of AI error are reported in the literature, as well as the frequency and lesion or imaging features of these AI errors.
In this study, we aim to identify the range of outcomes that have been reported as AI errors; quantify the frequency AI errors; and to describe the study, imaging, or lesion features associated with AI errors in practice. To meet this objective, we performed a systematic literature review of external validation studies of AI algorithms for independent mammographic interpretation using real-world screening data.
Materials and methods
This systematic review was performed and reported in accordance with Preferred Reporting Items for Systematic Reviews and Meta-Analyses of Diagnostic Test Accuracy (PRISMA-DTA) statement [9]. The review protocol was registered in the International Prospective Register of Systematic Reviews (PROPSERO) (CRD42022340331).
Information sources and literature search
A literature search was conducted without language restrictions for diagnostic accuracy studies published from 1 January 2010 to 11 July 2022. To capture contemporary AI algorithms, the search was limited from January 2010, coinciding with technical and hardware developments that facilitated efficient processing of machine learning [10]. Databases searched include MEDLINE, EMBASE, SCOPUS and a pre-print database, ArXiv. We reviewed reference lists of relevant systematic reviews to identify the additional studies. Details of the search strategy are listed in Online Resource 1.
Study selection
One reviewer (A.Z.) independently screened titles and abstracts and subsequent full-text articles against eligibility criteria (Online Resource 2). A second reviewer (M.L.M.) independently screened a 25% sample of titles and abstracts and the final set of included full-text studies for quality assurance.
Eligible study designs were external validation studies performed in population breast cancer screening settings where the AI algorithm acted as an independent reader (defined as a standalone system for replacement of radiologist reading, or as a pre-screen to triage whether the mammogram requires further review by a radiologist). Where studies included both conventional mammography and tomosynthesis, data on mammograms only were included.
Studies were excluded if more than 5% of included mammograms were incomplete; AI was used as a prediction tool (e.g. cancer subtypes, lesion characteristics or risk) or to assist radiologist reading (meaning the read was not solely from the AI algorithm); or AI was implemented for other imaging types (e.g. magnetic resonance imaging or ultrasound). Studies were excluded if outcomes did not include AI errors.
Data extraction and risk of bias assessment
Two reviewers (A.Z. and N.N.) independently extracted data on a pre-designed standardised collection form. Data that were systematically extracted included study design and setting, population characteristics, commercial availability, frequency and characteristics of pre-specified AI errors [false positives (FP), false negatives (FN), location error, technical error] and reference standard. Other forms of AI error were extracted when reported. FP was defined as incorrect presence of a suspicious finding (in cases where no cancer is found). FN was defined as cancer not detected by AI but detected by radiologist(s) or found at follow-up. Location error was defined as correct diagnosis of cancer, but the region of interest indicated by AI does not correspond with cancer location. Technical error was defined as failure of AI to process and interpret the mammogram or output a finding.
From studies reporting AI accuracy, we extracted raw values to derive 2 × 2 tables cross-classifying the AI result (positive or negative) and the reference standard finding (cancer present or absent). From these values, we calculated false positive proportions (FPP) and false negative proportions (FNP) per study (FP or FN divided by total number of examinations). When studies reported data at comparable positivity thresholds (including multiple possible thresholds per study), we extracted data and calculated FNP and FPP, and sensitivity and specificity estimates at those thresholds. Only common positivity thresholds across studies were reported.
Two reviewers (A.Z. and M.L.M.) independently assessed methodological quality using the Quality Assessment of Diagnostic Accuracy Studies (QUADAS-2) tool modified to the review question and QUADAS-AI preliminary extension [11] (Online Resource 3). Risk of bias of individual studies was assessed under four domains including (i) patient selection, (ii) index test, (iii) reference standard and (iv) flow and timing. The first three domains were assessed in terms of concerns regarding applicability. Reference standards were recorded to assess comparability across studies. All discrepancies were resolved by discussion and consensus.
Data synthesis
Narrative synthesis was conducted because of methodological variations between studies. FPP, FNP, location, technical and other errors were tabulated. FNP and FPP estimates and their 95% Wald confidence intervals (CIs) were plotted in a forest plot. Estimates were pooled within each positivity threshold using inverse variance random-effects meta-analysis with the restricted maximum likelihood estimator [12, 13]. Tests for subgroup differences between thresholds were not calculated because data in each subgroup were not independent.
Sensitivity and specificity of single and consensus readers were plotted against AI positivity thresholds (when reported) in receiver operating characteristic (ROC) space to complement FPP and FNP estimates.
Analyses were undertaken using the metafor package and visual summaries were generated using the ggplot2 package [14, 15] in R version 4.2.2 (R Project for Statistical Computing in Vienna, Austria).
Results
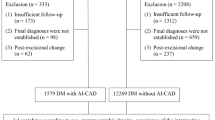
After deduplication, 1760 unique results were screened, of which 73 potentially eligible full-text articles were assessed. Seven studies were included in this review [16,17,18,19,20,21,22]. Figure 1 summarises the screening and eligibility process and documents reasons for exclusion.
Study characteristics of included studies
Characteristics of the seven included studies, comprising 447,676 examinations, are presented in Table 1. One study reported AI error as the location of false markings on the mammogram [20]. The remaining six studies reported AI sensitivity and specificity [16,17,18,19, 21, 22], and five of those reported AI errors as FNR and FPR according to the positivity threshold applied [16,17,18,19, 22]. One study reported additional error information as AI technical failures [19].
Two studies evaluated datasets on real-world screening populations from Sweden, and one each evaluated screening populations from Denmark, Norway, Germany, Spain, and the United States (US). Evaluation datasets were sourced from screening programmes [18, 19, 21], sub-cohorts of randomised controlled trials [17, 22] and specialist cancer centres [16, 20]. Screening mammograms all had 4 views (2 views per breast). Mammogram units were Siemens Mammomat [17,18,19] or Hologic Selenia [20, 22], and one study reported the use of both [16]. All were retrospective cohort studies with consecutive screens. Years of enrolment ranged from 2008 to 2018. Study-level mean age of the women ranged between 53 and 60 years.
For six studies reporting on FPR and FNR and/or sensitivity and specificity, two studies used a reference standard of screen-detected cancers only [16, 17], and four included interval cancers (in addition to screen-detected cancers) with follow-up of either 12 months [21] or 24 months [18, 19, 22]. Cancer prevalence in studies with screen-detected cancers only ranged between 0.64 and 0.71%, whereas this was from 0.71 to 1.22% for studies that included screen-detected and interval cancers. The reference standard in an additional study reporting cancer location was clinical review of established biopsied cancer location (all screen-detected; ≥ 2 year follow-up confirmed no interval cancers). An AI cancer marking was considered correct if its location intersected with the geometric centre of ground-truth (radiologists’) region of interest (ROI) [20].
Four studies using AI for triage reported “high”- or “moderate”-risk cases would be reviewed by radiologists [16, 17, 19, 22]. “Low-risk” cases (i.e. case deemed as low suspicion of cancer) would have no human-reading or reading by a single radiologist. AI performance was compared to radiologists (single or consensus reading) with 3 to 15 years of experience [16, 19, 22]. Three studies used standalone AI to evaluate its accuracy compared to either double reading (1–20 + years’ experience) [18, 21] or computer-aided detection (CAD) to reduce false positive markings on mammograms [20].
Table 2 summarises the risk of bias and applicability concerns of included studies. Overall, four studies had high risk of bias or applicability concerns in at least one of the four domains [16, 17, 20, 21]. Two studies had high risk of bias and applicability concerns for the reference standard [16, 17] and three studies had unclear risk of bias for patient selection [17, 20, 22]. Four studies had either high or unclear risk of bias in flow and timing [16, 17, 20, 21].
Characteristics of AI tools and positivity thresholds
Of the six studies reporting FP and FN errors, five evaluated different versions (V1.4.0 to V1.7.0) of a commercially available algorithm (Transpara, Screen Point Medical) [16,17,18,19, 22] and one used an ensemble model [21] (Online Resource 4). An additional study reporting location errors used a prototype AI-CAD system [20].
In studies applying AI for triage, two used a single prospective threshold [17, 22], one used multiple prospective thresholds [19] and one study used multiple retrospective thresholds [16]. The most commonly reported thresholds were Transpara Score 5 [17, 19] or 7 [16, 22], where 10 equates to the highest suspicion of cancer on a scale of 0–10.
In studies using standalone AI, one used a single prospective threshold (Transpara Score 9) in addition to retrospective thresholds [18] and one used a retrospective single threshold to match radiologists’ specificity [21]. The study that assessed the location of any AI markings on the mammogram did not specify an AI positivity threshold [20].
Reported AI errors and associated factors
Table 3 presents the frequency and features associated with reported AI error.
Six studies reported sensitivity and specificity [16,17,18,19, 21, 22], five of which also reported AI error as false negatives and false positives according to comparable positivity thresholds [16,17,18,19, 22]. One study reported on AI error as technical failure which was defined as failure of the AI to process mammograms[19]. One study referred to location error as an AI cancer marking that is incorrectly highlighted on a mammogram (i.e. false positive marks) [20].
False positive proportion (FPP) and false negative proportion (FNP)
Pooled FPP decreased incrementally with increasing Transpara threshold (71.83% [95% CI 69.67, 73.90] at Transpara 3 to 10.77% [95% CI 8.34, 13.79] at Transpara 9). The most commonly reported prospective triage thresholds were Transpara 5 (pooled FPP 46.86% [95% CI 39.33, 54.53]) and Transpara 7 (pooled FPP 29.86% [95% CI 26.59, 33.35) (Fig. 2).
The pooled FNP increased incrementally from 0.02% [95% CI 0.01, 0.03] at Transpara 3 to 0.12% [95% CI 0.06, 0.26] (Transpara 9), consistent with a trade-off with FPP.
There was heterogeneity within Transpara thresholds, reflecting study-level differences in Transpara version and reference standard (Fig. 2, Online Resource 5). For studies using later versions of Transpara (V1.6.0 and V1.7.0), FPP was lower (and FNP higher) within each threshold when the reference standard included screen-detected and interval cancers [16], compared with studies including only screen-detected cancers in the reference standard [18, 19, 22]. One study that evaluated an earlier Transpara version (V1.4.0) included only screen-detected cancers in the reference standard [17]; it reported lower FPP and higher FNP relative to a study using the same reference standard and a later Transpara version [16].
Table 3 describes the lesion or imaging features associated with AI missed cancers (i.e. FN). Three studies reported on the lesion or imaging characteristics associated with FN, each at different Transpara scores (i.e. 5, 7, 9). Two studies reported that the majority of FN were invasive cancers (88–100%) [17, 18]. A majority of FN (53–100%) were interval cancers [18, 22]. One study reported FN cancers generally had a radiographic appearance of spiculated mass and were in Breast Imaging Reporting and Data System Density C and D breasts [17]. Two studies reported that median tumour size for cancers missed by AI ranged from ≤ 7 to 25 mm [17, 18]. When compared to AI detected cancers, the majority of these (77–84%) were also invasive cancers [17, 18].
Sensitivity and specificity
In studies that compared AI performance to radiologists, two reported on single reading [16, 21] and four [16, 18, 21, 22] reported on consensus reading. Transpara was the AI tool used in five of these studies (the other used an ensemble system [21]). As expected, we observed a trade-off with higher specificity and lower sensitivity as the Transpara positivity threshold increased. Regardless of single or consensus reading, the radiologists’ specificity remained consistent relative to the varied specificity and sensitivity at different AI positivity thresholds (Fig. 3). The range in sensitivity of a single reader is comparable to Transpara Score 7 (0.83–0.92) or 9 (0.77–0.88).
Other types of error
Two studies reported other forms of AI errors. One study reported that no technical failures were encountered in which the AI model failed to process mammograms [19]. A second study investigated the location of AI false marking on the mammogram [20]. Location error was weakly associated with radiographic features including calcification or masses. Eight of the 18 location errors were ultimately confirmed as benign lesions. No other forms of AI error were identified from included studies.
Discussion
This systematic review of externally validated AI algorithms for cancer detection in breast screening identified relatively few studies that report AI errors. Four types of AI errors were identified, the most commonly reported being false positive and false negative findings, which is consistent with a focus on diagnostic accuracy in studies of AI in breast cancer screening [16,17,18,19, 22]. Previous systematic reviews have assessed the diagnostic accuracy of AI in external validation studies; however, none have reported on AI error in detail [6,7,8]. This review is a novel attempt to report findings and imaging features associated with AI errors and identify other types of error. Technical and location errors were reported relatively infrequently and inconsistently, despite their importance in establishing the utility of AI in population breast cancer screening practice.
The findings highlight factors relating to algorithm, study, and imaging characteristics that may plausibly influence the FPP and FNP of AI in the breast screening context. The exploration of multiple AI positivity thresholds showed the expected trade-off between FPP and FNP, with progressively lower FPP (and higher FNP) as the positivity threshold increased. However, there was considerable heterogeneity of FPP and FNP within thresholds. Between-studies comparisons suggested that the frequency of these errors varied according to the version of the AI algorithm. Lower FNP was observed with more recent (v1.6.0 and 1.7.0) compared with earlier (v1.4.0) versions of Transpara, suggesting that improvements in AI over time have resulted in a lower likelihood of errors leading to missed cancers. However, a corresponding increase in FPR was also found, indicating that technical improvements to enhance AI sensitivity have the potential to result in increased recall. Studies that have integrated AI into the screen-reading workflow as an adjunct to radiologist reading have used recent Transpara versions[18, 23]; absent comparison with earlier versions, it is unclear if these observed differences in AI error rates may have translated to increased cancer detection and recall over time.
The comparisons also highlight the importance of appropriately defining the reference standard to classify FP and FN results. Studies that included both screen-detected and interval cancers reported lower FPP for AI compared with studies including screen-detected cancers only, logically reflecting the limitation of the latter design in validating true positive AI results that are deemed negative by radiologists [6]. “Missed” interval cancers by AI also contributed to higher FNP in such studies. Incompleteness of interval cancer ascertainment has been identified as source of potential bias in studies of AI [6, 8], with empirical studies showing an inflation of overall accuracy [2]. Studies investigating AI errors should, at minimum, include all interval cancers (ideally through registry linkage to minimise the potential for bias) [8]; extended follow-up should also be considered [8, 24], acknowledging the desirability of aligning follow-up with screening intervals which may differ between settings.
The above suggestion that studies of AI for mammography screening should use sufficient follow-up to include interval cancers is reinforced in the study from Larsen et al. [18] which used cancer registry linkage to ascertain interval cancers—it showed more of the cancers missed by the AI were interval (than screen-detected) cancers. It also showed that the AI was more likely to miss a smaller tumour than a larger tumour on the mammogram, evidenced in a median tumour diameter of cancers missed by AI ranging between 7 and 25 mm, whereas those correctly detected by the AI ranged between 9 and 30 mm.
Technical AI errors—in which the algorithm fails to generate output—may have important implications for implementation of AI in screening programmes. Such failures require remediation in the workflow, and systematic failures have the potential to have disproportionate impacts on different sub-populations [3]. Location errors—where AI identifies abnormality in an incorrect location of the breast—have potentially adverse clinical consequences for women and may erode radiologists’ confidence in AI findings. However, it should be noted that at present, there is no gold standard for defining these ‘location-specific’ errors which require clinical (imaging) review and subjective judgement, and this is an area worthy of further exploration. The one study identified in this review referred to location error as an AI cancer marking that is incorrectly highlighted on a mammogram and used a retrospective clinical review process [20]. Breast imaging fellowship-trained radiologists generated a region of interest to establish the location of the biopsied lesion, and AI markings were considered to be correct if the geometric centre lay within the region of interest.
Despite the importance of understanding the frequency and nature of both technical and location errors that occur when AI reads mammography, this review found that they were reported infrequently. In the case of technical errors, these were reported in only one instance to confirm the absence of such errors [21]. Additional emphasis on enumerating and describing such AI errors would be a valuable complement to the current research focus on accuracy (including FP and FN errors), enabling better understanding of the potential impact of AI on screening workflow and clinical outcomes. Improving knowledge on these issues is highly relevant for potential implementation of AI in breast screening practice, noting that women have high expectations that AI will improve mammography screening accuracy and outcomes [25].
This review did have limitations. Firstly, it focused on AI as standalone reader, not as an aid to reader interpretation. This may have limited the search strategy to exclude studies that are more likely to report location errors. However, these errors have been reported mostly in reader studies using cancer-enriched datasets and may not be generalisable to population breast cancer screening settings [7]. Furthermore, the review interprets between-study comparisons to explain heterogeneity in error estimates. Although the observed differences in FPP and FNP are in the expected directions, there are likely to be clinical and methodological differences between studies beyond those considered in our analyses. Within-study comparisons would provide stronger evidence from which to draw inferences. Where possible, authors should be encouraged to facilitate such comparisons by reporting FP and FN errors for screen-detected and interval cancers separately, and multiple follow-up times for ascertaining interval cancers [2].
Conclusions
Current evidence on AI algorithms in breast cancer screening demonstrates that false positives and false negatives are the predominantly reported forms of AI errors, which is consistent with the focus on diagnostic accuracy in the literature. Further reporting of other types of errors, including technical errors, could provide a better understanding of AI’s utility in breast screening practice. Further studies on AI errors using real-world data could also allow future systematic reviews to explore plausible factors (e.g. clinical or radiological characteristics) associated with errors that are generalisable to real populations. This could complement co-existing AI accuracy research, to ensure the safe integration of AI into future screening practice.
Data availability
Data sharing is not applicable to this article as no datasets were generated or analysed during the current study.
References
Marinovich ML, Wylie E, Lotter W, Pearce A, Carter SM, Lund H et al (2022) Artificial intelligence (AI) to enhance breast cancer screening: protocol for population-based cohort study of cancer detection. BMJ Open 12(1):e054005. https://doi.org/10.1136/bmjopen-2021-054005
Marinovich ML, Wylie E, Lotter W, Lund H, Waddell A, Madeley C et al (2023) Artificial intelligence (AI) for breast cancer screening: BreastScreen population-based cohort study of cancer detection. EBioMedicine 90:104498. https://doi.org/10.1016/j.ebiom.2023.104498
Oakden-Rayner L, Dunnmon J, Carneiro G, Ré C (2020) Hidden stratification causes clinically meaningful failures in machine learning for medical imaging. Proc ACM Conf Health Inference Learn 2020(2020):151–159. https://doi.org/10.1145/3368555.3384468
Finlayson SG, Bowers JD, Ito J, Zittrain JL, Beam AL, Kohane IS (2019) Adversarial attacks on medical machine learning. Science 363(6433):1287–1289
Dratsch T, Chen X, Rezazade Mehrizi M, Kloeckner R, Mähringer-Kunz A, Püsken M et al (2023) Automation bias in mammography: the impact of artificial intelligence BI-RADS suggestions on reader performance. Radiology 307(4):e222176. https://doi.org/10.1148/radiol.222176
Anderson AW, Marinovich ML, Houssami N, Lowry KP, Elmore JG, Buist DSM et al (2022) Independent external validation of artificial intelligence algorithms for automated interpretation of screening mammography: a systematic review. J Am College Radiol 19(21):259–73. https://doi.org/10.1016/j.jacr.2021.11.008
Houssami N, Kirkpatrick-Jones G, Noguchi N, Lee CI (2019) Artificial Intelligence (AI) for the early detection of breast cancer: a scoping review to assess AI’s potential in breast screening practice. Expert Rev Med Devices 16(5):351–362. https://doi.org/10.1080/17434440.2019.1610387
Freeman K, Geppert J, Stinton C, Todkill D, Johnson S, Clarke A et al (2021) Use of artificial intelligence for image analysis in breast cancer screening programmes: systematic review of test accuracy. BMJ 374:n1872. https://doi.org/10.1136/bmj.n1872
McInnes MD, Moher D, Thombs BD, McGrath TA, Bossuyt PM, Clifford T et al (2018) Preferred reporting items for a systematic review and meta-analysis of diagnostic test accuracy studies: the PRISMA-DTA statement. JAMA 319(4):388–396. https://doi.org/10.1001/jama.2017.19163
Lee JH, Shin J, Realff MJ (2018) Machine learning: overview of the recent progresses and implications for the process systems engineering field. Comput Chem Eng 114:111–121. https://doi.org/10.1016/j.compchemeng.2017.10.008
Sounderajah V, Ashrafian H, Rose S, Shah NH, Ghassemi M, Golub R et al (2021) A quality assessment tool for artificial intelligence-centered diagnostic test accuracy studies: QUADAS-AI. Nat Med 27(10):1663–1665. https://doi.org/10.1038/s41591-021-01517-0
Viechtbauer W (2005) Bias and efficiency of meta-analytic variance estimators in the random-effects model. J Educ Behav Stat 30(3):261–293. https://doi.org/10.3102/10769986030003261
Raudenbush SW (2009) Analyzing effect sizes: random-effects models. The handbook of research synthesis and meta-analysis, 2nd edn. Russell Sage Foundation, New York, pp 295–315
Viechtbauer W (2010) Conducting meta-analyses in R with the metafor package. J Stat Softw 36(3):1–48. https://doi.org/10.18637/jss.v036.i03
Wickham H, editor An implementation of the grammar of graphics in R: ggplot. Book of Abstracts; 2006.
Balta C, Rodriguez-Ruiz A, Mieskes C, Karssemeijer N, Heywang-Köbrunner SH. Going from double to single reading for screening exams labeled as likely normal by AI: what is the impact? Proceedings of SPIE; 2020. https://doi.org/10.1117/12.2564179
Lang K, Dustler M, Dahlblom V, Akesson A, Andersson I, Zackrisson S (2021) Identifying normal mammograms in a large screening population using artificial intelligence. Eur Radiol 31(3):1687–1692. https://doi.org/10.1007/s00330-020-07165-1
Larsen M, Aglen CF, Lee CI, Hoff SR, Lund-Hanssen H, Lang K et al (2022) Artificial intelligence evaluation of 122969 mammography examinations from a population-based screening program. Radiology 303:212381. https://doi.org/10.1148/radiol.212381
Lauritzen AD, Rodriguez-Ruiz A, von Euler-Chelpin MC, Lynge E, Vejborg I, Nielsen M et al (2022) An artificial intelligence-based mammography screening protocol for breast cancer: outcome and radiologist workload. Radiology 304:210948. https://doi.org/10.1148/radiol.210948
Mayo RC, Kent D, Sen LC, Kapoor M, Leung JWT, Watanabe AT (2019) Reduction of false-positive markings on mammograms: a retrospective comparison study using an artificial intelligence-based CAD. J Digit Imaging 32(4):618–624. https://doi.org/10.1007/s10278-018-0168-6
Schaffter T, Buist DSM, Lee CI, Nikulin Y, Ribli D, Guan Y et al (2020) Evaluation of combined artificial intelligence and radiologist assessment to interpret screening mammograms. JAMA Netw 3(3):e200265. https://doi.org/10.1001/jamanetworkopen.2020.0265
Raya-Povedano JL, Romero-Martin S, Elias-Cabot E, Gubern-Merida A, Rodriguez-Ruiz A, Alvarez-Benito M (2021) AI-based strategies to reduce workload in breast cancer screening with mammography and tomosynthesis: a retrospective evaluation. Radiology 300(1):57–65. https://doi.org/10.1148/radiol.2021203555
Larsen M, Aglen CF, Hoff SR, Lund-Hanssen H, Hofvind S (2022) Possible strategies for use of artificial intelligence in screen-reading of mammograms, based on retrospective data from 122,969 screening examinations. Eur Radiol 32(12):8238–8246. https://doi.org/10.1007/s00330-022-08909-x
Lee CI, Houssami N, Elmore JG, Buist DSM (2020) Pathways to breast cancer screening artificial intelligence algorithm validation. Breast 52:146–149
Lennox-Chhugani N, Chen Y, Pearson V, Trzcinski B, James J (2021) Women’s attitudes to the use of AI image readers: a case study from a national breast screening programme. BMJ Health Care Inform 28(1):e100293. https://doi.org/10.1136/bmjhci-2020-100293
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions. Aileen Zeng was a recipient of The Daffodil Centre Postgraduate Research Scholarship. Dr Luke Marinovich is funded by a National Breast Cancer Foundation Investigator Initiated Research Scheme grant (2023/IIRS0028). Prof Nehamat Houssami is funded through a National Breast Cancer Foundation Chair in Breast Cancer Prevention grant (EC-21-001) and a National Health and Medical Research Council Investigator Leader grant (1194410). Dr Brooke Nickel is supported by a National Health and Medical Research Council (NHMRC) Emerging Leader Research Fellowship (1194108).
Author information
Authors and Affiliations
Contributions
Conceptualization: Nehmat Houssami, Aileen Zeng and Luke Marinovich; Literature Search: Aileen Zeng; Data Extraction: Aileen Zeng, Naomi Noguchi and Luke Marinovich; Data Analysis: Aileen Zeng and Luke Marinovich; Writing – original draft preparation: Aileen Zeng and Luke Marinovich; Writing – review and editing: Nehmat Houssami, Luke Marinovich, Naomi Noguchi and Brooke Nickel.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Ethics approval
This a systematic review and does not require ethics approval.
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zeng, A., Houssami, N., Noguchi, N. et al. Frequency and characteristics of errors by artificial intelligence (AI) in reading screening mammography: a systematic review. Breast Cancer Res Treat 207, 1–13 (2024). https://doi.org/10.1007/s10549-024-07353-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-024-07353-3