Abstract
Purpose
The progression of ductal carcinoma in situ (DCIS) to invasive breast carcinoma (IBC) in humans is highly variable. To better understand the relationship between them, we performed a multi-omic characterization of co-occurring DCIS and IBC lesions in a cohort of individuals.
Methods
Formalin-fixed paraffin-embedded tissue samples from 50 patients with co-occurring DCIS and IBC lesions were subjected to DNA-seq and whole transcriptome RNA-seq. Paired DCIS and IBC multi-omics profiles were then interrogated for DNA mutations, gene expression profiles and pathway analysis.
Results
Most small variants and copy number variations were shared between co-occurring DCIS and IBC lesions, with IBC exhibiting on average a higher degree of additional mutations. However, 36% of co-occurring lesions shared no common mutations and 49% shared no common copy number variations. The most frequent genomic variants in both DCIS and IBC were PIK3CA, TP53, KMT2C, MAP3K1, GATA3 and SF3B1, with KMT2C being more frequent in DCIS and TP53 and MAP3K1 more frequent in IBC, though the numbers are too small for definitive conclusions. The most frequent copy number variations were seen in MCL1, CKSB1 and ERBB2. ERBB2 changes were not seen in IBC unless present in the corresponding DCIS. Transcriptional profiles were highly distinct between DCIS and IBC, with DCIS exhibiting upregulation of immune-related signatures, while IBC showed significant overexpression in genes and pathways associated with cell division and proliferation. Interestingly, DCIS and IBC exhibited significant differential expression of different components of extracellular matrix (ECM) formation and regulation, with DCIS showing overexpression of ECM-membrane interaction components while IBC showed upregulation of genes associated with fibronectin and invadopodia.
Conclusion
While most co-occurring DCIS and IBC were mutationally similar and suggestive of a common clonal progenitor, transcriptionally the lesions are highly distinct, with IBC expressing key pathways that facilitate invasion and proliferation. These results are suggestive of additional levels of regulation, epigenetic or other, that facilitate the acquisition of invasive properties during tumor evolution.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The diagnosis of ductal carcinoma in situ (DCIS) has increased significantly with the introduction of screening mammography and now represents 20% of newly diagnosed breast cancer in the United States. DCIS may be found either with or without simultaneous invasive disease. Invasive ductal cancer presents with synchronous DCIS approximately 60% of the time [1]. Additionally, invasive breast carcinoma (IBC) may develop either without preexisting DCIS or temporally distant from earlier pure DCIS. While isolated DCIS has an excellent prognosis, patients with DCIS have an increased risk of invasive disease and breast cancer death, particularly at younger ages [2,3,4,5,6]. Studies have suggested that DCIS can evolve into invasive disease but the frequency of this process and whether or not DCIS and invasive disease may represent independent processes in the same breast are unclear [7].
A variety of studies have shown that genomic copy number variations both genome-wide and at specific loci are a prognostic factor for progression and/or recurrence for DCIS [8,9,10,11]. Moreover, gene expression studies have shown significant heterogeneity in DCIS expression profiles, with hallmarks of the major gene expression subtypes for IBC often already evident in DCIS lesions [12,13,14,15,16]. There also appears to be a complex interplay between DCIS progression and the tumor immune microenvironment (TIME), with tumor infiltrating lymphocytes (TIL) largely associated with higher-grade DCIS lesions but largely transitioning to an immune-excluded environment in IBC [17,18,19]. A variety of studies have also examined the genomics and transcriptomics of DCIS and IBC in co-occurring and non-co-occurring lesions to implicate a variety of genes potentially associated with progression [1, 20,21,22,23]. However, there remain gaps in our understanding of the key molecular processes associated with DCIS transformation and invasion. As such, we performed a multi-omic investigation of genomic and transcriptomic similarities and differences in a cohort of patients who presented with simultaneous DCIS and invasive breast cancer in the same quadrant of the same breast.
Methods
Patient recruitment
Patients were identified from the Swedish Cancer Institute (SCI) Breast Cancer Database starting sequentially with 2018 backwards to identify candidates with simultaneous DCIS and invasive ductal cancer. Individual pathologic review was performed from the cohort of patients who had had modified radical mastectomy to select 50 cases for analysis of normal breast, DCIS, and invasive disease.
Molecular analysis
Formalin-fixed paraffin-embedded (FFPE) tissue blocks were retrieved from pathology archives. Ten 5 µm sections were prepared on microscopy slides for next-generation sequencing (NGS) analysis. DNA and RNA extraction, library preparation and sequencing were performed by Tempus Labs, Inc. (Chicago, Ill) using the Tempus xT workflow (DNA-seq of 648 genes at 500 × coverage), as previously described [24, 25]. DNA-seq alignment, mapping and variant calling were performed using the Tempus xT informatics pipeline. Clinically significant mutations identified by this assay include germline and/or somatic single nucleotide variants, insertions/deletions, and copy number variations, as well as structural rearrangements in a subset of 22 genes. Additionally, whole transcriptome RNA-seq was performed using the KAPA RNA HyperPrep kit. Tempus uses total RNA into library prep and exome hybrid capture to select the library. While the kits used across the two sample batches were the same, there was a change in adapters used for the second batch of RNA-seq, resulting in a difference in library strandedness. Both DNA-seq and bulk RNA-seq were performed in two batches.
Informatics
Raw reads were demultiplexed using bcl2fastq v2.20 into FASTQ format. FASTQ files were aligned to GRCh38 GENCODE Human release 39 using STAR v2.7-10a. STAR run parameters were adapted from the ENCODE RNA-Seq pipeline for gene count quantification [26]. Strandedness between batches of sequencing was identified using the checkstrandedness function from how_are_we_stranded_here to ensure the correct read counts column was pulled for downstream analyses [27]. The counts were preprocessed by removing genes with less than an average of 5 counts across all samples. Differential gene expression analysis was performed on pairwise samples for patient cases who had both a DCIS and IBC sample sequenced using DESeq2 [28] with visuals created with ComplexHeatmap and EnhancedVolcano [29,30,31].
Two methods of pathway enrichment analysis were utilized. First, a pre-ranked gene list was generated from sorting the DESeq2 contrast results by log fold change for Gene Set Enrichment Analysis (GSEA) against MSigDB collections C2, C3 transcription factor targets, and C5 biological processes [32]. Gene sets were visualized using the EnrichmentMap app in Cytoscape [33]. Significantly enriched pathways were assessed at an FDR corrected p-value ≤ 0.1 with node color indicating the GSEA assigned normalized enrichment score (NES). Secondly, the DAVID functional annotation tool was used on the lists of differentially expressed genes at q-value ≤ 0.001 focusing on GO biological processes, KEGG, and Reactome pathway databases [34, 35]. Reactome Pathway Browser was utilized by inputting the lists of differentially expressed genes enriched for each side of the contrast separately to visualize pathway specific gene expression [36].
Samples excluded from downstream RNA analyses were due to a patient only having a single sample, RNA quantity not sufficient, failed sequencing quality control metrics, or failed DNA-seq data which was the case for a single patient for DNA-seq and RNA-seq integrative analyses. Whole transcriptome similarity between samples was assessed by calculating Euclidean distances on the variance stabilizing transformed DESeq2 object and the heatmap of sample-to-sample distances was created via pheatmap. Statistical tests comparing the mean Euclidean distance for patient’s pairwise samples whose DCIS and IBC shared clinically significant variants versus those that did not was conducted via Student’s T-test.
Results
Cohort characteristics
We selected a series of thirty-nine cases of co-occurring DCIS and IBC in the same quadrant of the same breast with adequate tissue for both DCIS and IBC for DNA-seq. (Thirty-one of these cases also had adequate tissue RNA for RNA-seq.) 29/39 were estrogen receptor (ER)/progesterone receptor (PR) positive and ERBB2 negative. 4/39 were ER + /PR-/ERBB2 negative. 2/39 were triple positive, 3/39 were ER+ /PR-/ERBB2 positive and 1/39 was ER/PR negative and ERBB2 positive. 32/39 had Ki-67 ≤ 20% (IQR 14 (9, 19.5). 35/39 IBC were T1 or T2. Three were T3 and one T4. 21/39 IBC were N0, 4/39 N1mic, 9/39 N1a, 3/39 N2a, 2/39 N3a. Comprehensive genomic profiling using a 648-gene panel (Tempus xT) specimens and whole transcriptome RNA-seq were performed on all paired DCIS and IBC. The mutational profiles for the paired specimens across the most frequent alterations can be seen in Fig. 1A. There was a much higher degree of inter-patient variation than intra, with the majority of cases 25/39 (64%) sharing at least one reported small variant between DCIS and IBC (Fig. 1A, B). 14/39 (36%) had no shared variants (5 cases had no detected variants). 20/39 (51%) shared at least one copy number variation (CNV). 19/39 (49%) had no shared copy number variations (3 had no copy number variations). 32/39 (82%) shared a variant or copy number variation. 7/39 (18%) shared no variants or copy number variations while considering all patients who had at least one small variant or copy number variation. From the figure we can also see a high degree of variability across the cohort, with only 1/39 (3%) of patients exhibiting an identical mutational profile between DCIS and IBC.
Global landscape of clinically significant variants and copy number variations across all 39 patients. A Discrete heatmap indicating presence or absence of variant and copy number variations (CNVs) for all 39 patients with paired DCIS and IBC DNA-seq results; see associated legend for the variant type color coding. White asterisks indicate the DCIS and IBC tumors for a given patient had mutations in the same gene but the mutation was different. B Barplot indicating the percentage of patients with shared genomic variants and copy number variations between DCIS and IBC tumors
Amongst the detected genomic variants, the most commonly mutated genes were quite similar between IBC and DCIS (Fig. 1A and Table 1). Despite this, no alterations were universal across the cohort, with even common driver alterations such as PIK3CA repeatedly exhibiting mutation in only one of a patient’s paired DCIS and IBC, perhaps suggestive of a non-clonal relationship between the lesions in these patients. For copy number variations (CNVs), we observed that the most frequently detected CNVs were in CKS1B, MCL1 and ERBB2 for both DCIS and IBC (Table 2). Moreover, CNVs were not always detected across both of a DCIS/IBC pair and ERBB2 was not seen in IBC without its presence in the corresponding DCIS (Table 3). A total of 130 CNV occurrences were observed across 53 unique CNVs amongst all DCIS tumors (0–12 patient). Similarly, we observed 121 CNV occurrences across 51 unique CNVs within invasive tumors (0–9/patient). There were 43 instances where the same CNV was reported in both the DCIS and paired IBC tumors (0–5/patient). Additionally, 87 instances of CNVs were detected in DCIS but not paired IBC (0–11/patient) and 75 CNVs were detected in IBC but not paired DCIS (0–9/patient). Furthermore, 73 unique small variants were observed across 94 variant occurrences within DCIS tumors (0–8/patient), while 76 unique variants were reported across a total of 87 variant occurrences within IBC tumors (0–6/patient). A total of 45 instances were observed where the same variant was detected in both the DCIS and paired IBC (0–4/patient) with an additional 44 instances of variants observed in DCIS but not paired IBC (0–5/patient), and 42 variants observed in IBC but not paired DCIS (0–5/patient). Tumor mutation burden was universally low across DCIS and IBC in this cohort with a median TMB of 2.1 (1.1, 3.2) for DCIS and 1.6 (1.1, 2.6) for IBC.
RNA-seq analysis
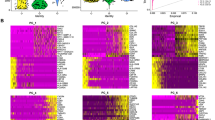
We next asked whether there were significant gene expression differences between DCIS and IBC. Looking at the groupwise level, we found 1209 genes were significantly differentially expressed (DE) between DCIS and IBC (FDR-corrected q-value < 0.001) (Fig. 2 A&B and Table 4), with the top DE genes upregulated in IBC largely related to extracellular matrix formation and comprising collagen-related genes (COL11A1, COL10A), matrix metallopeptidase subunits (MMP11, MMP13, MMP1, etc.) as well as genes associated with microfibril assembly (FN1, MFAP2, LRRC15, etc.). Meanwhile, top genes specifically associated with DCIS were related to muscle cell regulation (SMYD1, PDE1C, PAMR1), olfactory receptors (OR5P3, OR5P2) and immune related functions (IL33, LIFR, etc.). We evaluated gene expression differences on a patient-by-patient basis for each of the top DE genes (Fig. 2C) and noted that the difference between expression for DCIS and IBC largely trended in the same direction for each patient (with a few exceptions). However the overall magnitude of the difference is variable across each patient.
Differential expression of co-occurring DCIS versus IBC. A Heatmap of the 1209 genes that are significantly differentially expressed between DCIS and IBC (q < 0.001). B Volcano plot showing fold change versus -log10 p-value for the DCIS vs. IBC comparison. C Gene expression differences between DCIS vs IBC are shown for each patient for top DE genes plotted by log FPKM gene expression
We then asked whether certain a priori cellular pathways were significantly enriched as DE between DCIS and IBC. To do this we performed pathway enrichment analysis using multiple approaches including Gene Set Enrichment Analysis (GSEA) across a variety of gene set collections as well as DAVID functional annotation for DE genes. The GSEA results weighing heavily on KEGG, Gene Ontology and Reactome pathways, yielded significant differences between DCIS and IBC (Fig. 3), with IBC samples significantly overexpressing pathways associated with cycling cells and proliferation (e.g. “Replication Dependent Chromatin Organization” (GO), “DNA Replication” (KEGG) etc.) but also pathways associated with replication stress and activation of cell cycle checkpoints (“Activation of ATR in Response to Replication Stress” (Reactome), “Cell Cycle Checkpoints” (Reactome), consistent with active proliferation and dysregulated DNA replication. For DCIS, many of the enriched pathways were related to cytochrome (CYP) enzymes and xenobiotic metabolism (“Metabolism of xenobiotics by cytochrome P450” (KEGG) and immune responses (“Complement Cascade” (Reactome), “B-Cell Receptor Signaling” (GO)). We also assessed enrichment of specific transcription factor binding sites (TFBS) with the associated DE genes and observed a variety of enriched TFBS (Figure S1). The top site enriched in the IBC was a promoter regulatory element of unknown function (M120 motif KRCTCNNNNMANAGC, q < 0.001) [37] as well as sites regulated by heat shock transcription factor 4 (HSF4) (q < 0.02). Top TFBS enriched in DCIS included MEF2 (q < 0.005), HMEF2 (q < 0.006), AR (q < 0.002) and CEBPE (q < 0.003).
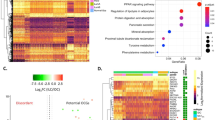
Ranked whole transcriptome level pathway enrichment for DCIS vs IBC. A Top 20 most significant pathways from the MSigDB C2 and C5 biological processes collections tested for pathway enrichment using DE genes from the DCIS vs IBC tumor contrast utilizing GSEA. B Cytoscape networks highlighting 3 major clusters from GSEA for DCIS vs IBC on MsigDB C2 all collection. The color of the node indicates the normalized enrichment score (NES) and directionality assigned by ranked GSEA
We also tested for pathway enrichment using DAVID, which uses only the significant DE genes as opposed to GSEA (which uses a transcriptome-wide enrichment approach). While there was significant overlap between the two approaches, including significant enrichment for pathways associated with DNA replication and cell cycle progression in IBC, the top pathways associated with IBC using DAVID were related to ECM organization and regulation (Fig. 4A). Interestingly, the top pathways associated with DCIS also included several pathways associated with ECM organization (Fig. 4B), leading us to hypothesize that ECM organization is mechanistically different between DCIS and IBC. To further characterize these differences, we plotted genes differentially expressed in DCIS vs. IBC on Reactome pathway diagrams for ECM formation (Fig. 5A, B). There we observed distinct components of ECM formation and organization enriched in DCIS and IBC lesions, with invadopodia formation components and proteoglycans specifically enriched in IBC, while DCIS shows specific enrichment for integrin/laminin signaling (Fig. 5).
Pathway enrichment for DCIS vs IBC. A Top significant DAVID pathways from the Reactome, KEGG and Gene Ontology (Biological Processes only) collections based on differentially expressed genes enriched in IBC resulting from differential expression analysis between DCIS and IBC tumors. B Top significant pathways from DAVID involving DE genes enriched in DCIS tumors
Differential expression of ECM regulatory networks by DCIS and IBC. A Genes overexpressed in IBC are highlighted in yellow overlayed on Reactome pathway R-HSA-1474244 ~ Extracellular matrix organization, with red boxes outlining key components of the ECM machinery enriched in IBC. B The same pathway diagram, with overexpressed genes in DCIS highlighted
Integration of DNA-seq and RNA-seq
To characterize the concordance/discordance between DNA-seq and RNA-seq profiles observed across the DCIS and IBC lesions we constructed a similarity matrix using Euclidean distance for gene expression across DCIS and IBC profiles. We then asked whether DCIS/IBC pairs with shared mutations were more similar in gene expression patterns than DCIS/IBC pairs without shared mutations. We see that for the most part gene expression patterns are more similar between an individual’s paired DCIS/IBC lesions than with DCIS or IBC from another individual (Fig. 6A). Comparing patients with shared variants in their DCIS/IBC pairs versus those without shared variants, we observed that gene expression was significantly more diverse when DCIS and IBC had no shared genomic variants (Euclidean distance of overall gene expression between paired DCIS and IBC, p < 0.005) (Fig. 6B), likely more evidence that these lesions developed independently versus via a common progenitor. However, even within lesions that had highly similar mutational genomes, we observed a subset of DCIS/IBC pairs that had highly variable gene expression. To characterize factors driving this, we subdivided mutantnome-specific tumors into two groups based on higher or lower gene expression similarity. We detected significant expression differences between these groups (Fig. 6C), with divergent DCIS/IBC pairs showing upregulation of pathways associated with B-cell receptor signaling (q < 0.001), heme scavenging (q < 0.001) and phagocytosis (q < 0.001). Pairs with highly similar transcriptomes showed enrichment for a variety of metabolic pathways (Fig. 6D).
Transcriptome similarity and the association with shared genomic variants. A Sample-to-sample matrix of transcriptome similarity via Euclidean distance. B Violin plot and Student’s T-test between transcriptome-wide gene expression Euclidean distance of pairwise samples for patients whose paired DCIS/IBC shared genomic variants versus those that did not. C For tumors with shared variants, transcriptional differences between those with high and low gene expression similarity. D Pathway enrichment analysis of DE genes between patients who had high versus low gene expression similarity via GSEA
We also asked whether significant mutational differences were associated with large-scale changes to the transcriptome. Given the frequency of MCL1 copy number variations in breast cancer, we asked whether there were transcriptional differences between MCL1-amplified versus wildtype lesions (Fig. 7A). From this we observed 357 genes that were significantly different between MCL1-CNV and MCL1-wt lesions (q < 0.05) with key differences in extracellular matrix organization and adaptive immune response pathways (Fig. 7B). Similarly, we asked whether there were significant differences between tumors with CKS1B copy number variations versus CKS1B wildtype. From differential expression analysis, we observed 627 differentially expressed genes (q < 0.05) between CKS1B-CNV and CKS1B-wt lesions (Fig. 7C), with the top targets related to endothelial function and angiogenesis (Fig. 7D). From these combined data, we conclude that co-occurring DCIS and IBC display a wide degree of heterogeneity across individuals, and within individuals there are varying degrees of heterogeneity that are correlated with specific somatic mutations and copy number variations.
Correlation between copy number variations and transcriptional differences. A Volcano plot showing differential expression between MCL1-CNV versus MCL1-wt tumors. B Pathway enrichment analysis of gene expression differences between MCL1-CNV versus MCL1-wt tumors using GSEA and highlighting MSigDB C2 and C5 biological processes collections. C Volcano plot showing differential expression between CKS1B-CNV versus CKS1B-wt tumors. D Pathway enrichment analysis of gene expression differences between CKS1B-CNV versus CKS1B-wt tumors
Finally, we compared ER+ /PR+ to ER+ /PR-patients. Comparisons were made between these groups for IBC vs IBC, DCIS vs DCIS, and differences between paired IBC/DCIS patients for ER+ /PR+ vs ER+ /PR-cohorts. The only marked difference was the finding of overexpression of ERBB2 in ER+ /PR-DCIS vs ER+ /PR+ DCIS (supplementary text). The number of ER+ /PR-patients is quite small in this series so the significance of these analyses requires confirmation in a larger cohort.
Discussion
Co-occurring DCIS and IBC lesions continue to represent a compelling model for understanding the evolution of DCIS to IBC. There are many advantages versus other systems as it allows us to directly interrogate these lesions in humans as opposed to model organisms as well as look in a common genetic background (i.e. within an individual). Indeed, co-occurring DCIS/IBC lesions are typically more similar to each other based on gene expression than to other lesions with the same histological subtype such as DCIS versus DCIS, likely showing that the original genetic background is the key driver for the majority of basal transcription (Fig. 6), even showing greater similarity than for different cases with the same oncogenic driver alterations (Fig. 1). The most frequent genomic variants in both DCIS and IBC were noted to be PIK3CA, TP53, KMT2C, MAP3K1, GATA3 and SF3B1, with KMT2C being more frequent in DCIS and TP53 and MAP3K1 more frequent in IBC, though the numbers are too small for definitive conclusions. The most frequent copy number variations were seen in MCL1, CKSB1 and ERBB2. ERBB2 changes were not seen in IBC unless present in the corresponding DCIS. As such, while there have been some advances in developing prognostic gene expression signatures for DCIS progression, differences in genetic background are likely a key factor currently limiting the power of such approaches [11, 38]. Studies involving significantly larger cohorts of DCIS patients with associated expression and outcome information will be key to fully account for these factors.
Overall, our data and that from others point to a model for co-occurring lesions where a subset of similar DCIS/IBC lesions seem to arise from a common somatic progenitor, while others with highly divergent mutational and expression profiles appear to arise mostly independently. Indeed, just under 20% of DCIS and IBC cases shared no variant or copy number changes. This finding is quite similar to that of Lips et al., who examined such changes in a comparison of DCIS and later recurrence of invasive disease [22]. In that study approximately 25% of subsequent but temporally distinct invasive recurrences appeared to be clonally distinct from the original DCIS. These data are supportive of a more general “field effect” hypothesis throughout the breast rendering it susceptible to subsequent carcinogenesis. Our work extends this current literature in providing a combined portrait of gene expression and mutational changes across these co-occurring lesions. Common intuition would suggest that co-occurring DCIS/IBC with similar somatic genomes would appear more similar in their gene expression programs, and indeed they do in our dataset (Fig. 6). However, note that even in DCIS/IBC pairs with similar somatic genomes, there is a large spread in transcriptional similarity/dissimilarity. Looking more deeply at those with transcriptional dissimilarity (Fig. 6C), reveals that a large proportion of the detected variation comprises immune-related pathways (Fig. 6D). Thus, it is likely that these differences are not necessarily related to a more divergent tumor transcriptome but related to more immunogenicity in this subset of lesions driving the infiltration of diverse immune cell types. As such, this metric may be useful for predicting which subset of lesions may be responsive to immunotherapeutic treatment modalities.
While these expression data showed several key phenotypes distinct to IBC in relation to DCIS, one key mechanism that stood out was the significant difference in expression for key subsets of extracellular matrix formation and signaling. DCIS ECM expression was hallmarked by laminin/integrin signaling (laminin 322, etc.) as markers for early stage disease. In contrast, IBC ECM gene expression featured fibronectin matrix formation (Fig. 5), which is corroborated by recent mechanistic work by Hayward et al. who show that DCIS to IBC progression is associated with deposition of fibronectin in the duct and duct expansion potentially as a mechanism supporting invasion [39]. IBC also showed expression of invadopodia formation machinery, likely representing a key step in transitioning to a metastatic state. In addition, a large number of additional detected genes did not fit into known breast cancer progression mechanisms (e.g. SYNDIG1, OVCH2, OR5P3, etc.) and further studies of these targets may yield new insights into additional mechanisms that support breast cancer progression.
We also took an integrative look at somatic mutations and transcriptional activity across paired DCIS/IBC lesions (Fig. 7). Notably, we observed a transcriptional program that was distinct to MCL1-amplified tumors (Fig. 7A). We noted upregulation of pathways related to immune response, cell migration and activation, chemotaxis and apoptosis in DCIS and IBC samples with increased CNVs for the MCL1 gene. Copy number variations for the MCL1 gene, which has been implicated in apoptotic functions, have been reported in many hematologic as well as solid malignancies, including breast cancer [40,41,42,43]. MCL1 copy number variations have been seen in triple negative, hormone receptor positive and ERBB2-positive breast cancer [40, 42,43,44] and have been correlated with both prognosis and resistance to various antitumor therapies [45, 46]. Campbell et al. examined MCL1 in the MMTV-PyMT genetic mouse model of breast carcinogenesis, which recapitulates features of breast cancer progressing through hyperplasia to metastasis [47, 48]. This study revealed that deletion of MCL1 in the mammary epithelium of the genetically engineered mice showed an “absolute requirement” for MCL1 in breast tumorigenesis [47]. We are unaware of prior data on MCL1 in preinvasive human breast cancer. While it is well established that MCL1 is important in anti-apoptotic activities [49], the data presented here suggest the need for additional evaluation of its role in modulating the tumor immune microenvironment.
Unlike MCL1-amplified tumors, CKS1B-altered tumors showed differences related to endothelial function and angiogenesis (Fig. 7C, D). CKS1B protein is known to bind to the catalytic subunit of the cyclin-dependent kinases and has a major role in cell cycle regulation and has been implicated in apoptosis inhibition as well [50,51,52,53]. CKS1B amplification and overexpression have been reported in breast cancer and multiple myeloma and has been associated with poor prognosis in both [51, 54, 55]. Slotky et al. has reported CKS1B in breast cancer was associated with patient’s age, estrogen, and progesterone receptor levels and increased with malignant degree while Shi et al. has implicated it in the development of drug resistance [55, 56]. Jia et al. noted an association of CKS1B activity and infiltration of breast tumors with cancer-associated fibroblasts and noted and association with cell cycle and kinase regulation [51].
These data suggest additional areas of opportunity for studies to be performed. The changes in transcriptomics and pathway analysis can be confirmed at the proteomic level and/or using metabolomics to assess the metabolic output of these pathways. In particular, the differences in extracellular matrix activities noted in this study and others [38, 39] and immune-related phenomena between DCIS and IBC [17,18,19] may yield clues to the seminal events in the change from non-invasive to invasive activity. Spatial genomic studies of the tumor microenvironment may allow for the assessment of morphological changes in the ECM that result from these pathway alterations. It is clear that the genomic changes that are found in IBC are already often present in DCIS, leading to the question of when in the evolution from normal breast tissue to invasive cancer these events occur. Studies analogous to those presented here in patients with atypical ductal hyperplasia as well as histologically normal breast tissue in these patients would be helpful in this regard and would address the “field effect” hypothesis.
In addition, with larger patient cohorts, tumor-specific alterations in genomics, transcriptomics and proteomics can be correlated with clinical prognostic parameters, such as hormone receptor and ERBB2 status, histologic and nuclear grade, Ki67 and clinically available genomic predictive assays such as OncotypeDx DCIS [57]. We hope this and other work may help usher in more clinically useful biomarkers and potential therapeutic targets to better predict and modify the evolution of these breast lesions.
Conclusion
Overall, the genomic and transcriptomic characterization of co-occurring DCIS/IBC lesions shows a diverse spectrum of patterns spanning close co-evolution to clonally independent lesions. Moreover, the specific genetic backgrounds underlying these tumors drives complex transcriptomic programs that may have key relevance for progression and/or response to treatment. Even within these subgroups there is considerable diversity, suggesting epigenetic or other levels of regulation are additional key contributors to tumorigenesis that have gone unmapped. Adding additional levels of characterization such as whole genome bisulfite sequencing, ATAC-seq, proteomics, detailed immune profiling and more may shed new light on tumor progression and lead to superior prognostic and/or predictive biomarker signatures as well as lead to novel treatment paradigms.
Data availability
The datasets generated during and /or analyzed during this study are available from the corresponding author on reasonable request.
References
Kim SY, Jung S-H, Kim MS et al (2015) Genomic differences between pure ductal carcinoma in situ and synchronous ductal carcinoma in situ with invasive breast cancer. Oncotarget. 6:7597–7607. https://doi.org/10.18632/oncotarget.3162
Wärnberg F, Yuen J, Holmberg L (2000) Risk of subsequent invasive breast cancer after breast carcinoma in situ. Lancet 355:724–725. https://doi.org/10.1016/S0140-6736(99)03703-4
Maxwell AJ, Clements K, Hilton B et al (2018) Risk factors for the development of invasive cancer in unresected ductal carcinoma in situ. Eur J Surg Oncol 44:429–435. https://doi.org/10.1016/j.ejso.2017.12.007
Ryser MD, Weaver DL, Zhao F et al (2019) Cancer outcomes in DCIS patients without locoregional treatment. J Natl Cancer Inst 111:952–960. https://doi.org/10.1093/jnci/djy220
Shaaban AM, Hilton B, Clements K et al (2021) Pathological features of 11,337 patients with primary ductal carcinoma in situ (DCIS) and subsequent events: results from the UK Sloane Project. Br J Cancer 124:1009–1017. https://doi.org/10.1038/s41416-020-01152-5
Rawal R, Lorenzo Bermejo J, Hemminki K (2005) Risk of subsequent invasive breast carcinoma after in situ breast carcinoma in a population covered by national mammographic screening. Br J Cancer 92:162–166. https://doi.org/10.1038/sj.bjc.6602250
Jansen S (2012) Biology of DCIS and progression to invasive disease. Med Radiol 63:35–48. https://doi.org/10.1007/174_2012_647
Gorringe KL, Hunter SM, Pang J-M et al (2015) Copy number analysis of ductal carcinoma in situ with and without recurrence. Mod Pathol 28:1174–1184. https://doi.org/10.1038/modpathol.2015.75
Rane SU, Mirza H, Grigoriadis A, Pinder SE (2015) Selection and evolution in the genomic landscape of copy number alterations in ductal carcinoma in situ (DCIS) and its progression to invasive carcinoma of ductal/no special type: a meta-analysis. Breast Cancer Res Treat 153:101–121. https://doi.org/10.1007/s10549-015-3509-x
Johnson CE, Gorringe KL, Thompson ER et al (2012) Identification of copy number alterations associated with the progression of DCIS to invasive ductal carcinoma. Breast Cancer Res Treat 133:889–898. https://doi.org/10.1007/s10549-011-1835-1
Strand SH, Rivero-Gutiérrez B, Houlahan KE et al (2022) DCIS genomic signatures define biology and clinical outcome: Human Tumor Atlas Network (HTAN) analysis of TBCRC 038 and RAHBT cohorts
Clark SE, Warwick J, Carpenter R et al (2011) Molecular subtyping of DCIS: heterogeneity of breast cancer reflected in pre-invasive disease. Br J Cancer 104:120–127. https://doi.org/10.1038/sj.bjc.6606021
Yu K-D, Wu L-M, Liu G-Y et al (2011) Different distribution of breast cancer subtypes in breast ductal carcinoma in situ (DCIS), DCIS with microinvasion, and DCIS with invasion component. Ann Surg Oncol 18:1342–1348. https://doi.org/10.1245/s10434-010-1407-3
Doebar SC, Sieuwerts AM, de Weerd V et al (2017) Gene expression differences between ductal carcinoma in situ with and without progression to invasive breast cancer. Am J Pathol 187:1648–1655. https://doi.org/10.1016/j.ajpath.2017.03.012
Adeyinka A, Emberley E, Niu Y et al (2002) Analysis of gene expression in ductal carcinoma in situ of the Breast1. Clin Cancer Res 8:3788–3795
Bergholtz H, Lien TG, Swanson DM et al (2020) Contrasting DCIS and invasive breast cancer by subtype suggests basal-like DCIS as distinct lesions. npj Breast Cancer 6:1–9. https://doi.org/10.1038/s41523-020-0167-x
Hendry S, Pang J-MB, Byrne DJ et al (2017) Relationship of the breast ductal carcinoma in situ immune microenvironment with clinicopathological and genetic features. Clin Cancer Res 23:5210–5217. https://doi.org/10.1158/1078-0432.CCR-17-0743
Gil Del Alcazar CR, Huh SJ, Ekram MB et al (2017) Immune escape in breast cancer during in situ to invasive carcinoma transition. Cancer Discov 7:1098–1115. https://doi.org/10.1158/2159-8290.CD-17-0222
Trinh A, Gil Del Alcazar CR, Shukla SA et al (2021) Genomic alterations during the in situ to invasive ductal breast carcinoma transition shaped by the immune system. Mol Cancer Res 19:623–635. https://doi.org/10.1158/1541-7786.MCR-20-0949
Hernandez L, Wilkerson PM, Lambros MB et al (2012) Genomic and mutational profiling of ductal carcinomas in situ and matched adjacent invasive breast cancers reveals intra-tumour genetic heterogeneity and clonal selection. J Pathol 227:42–52. https://doi.org/10.1002/path.3990
Casasent AK, Schalck A, Gao R et al (2018) Multiclonal invasion in breast tumors identified by topographic single cell sequencing. Cell 172:205-217.e12. https://doi.org/10.1016/j.cell.2017.12.007
Lips EH, Kumar T, Megalios A et al (2022) Genomic analysis defines clonal relationships of ductal carcinoma in situ and recurrent invasive breast cancer. Nat Genet 54:850–860. https://doi.org/10.1038/s41588-022-01082-3
Visser LL, Elshof LE, Van de Vijver K et al (2019) Discordant marker expression between invasive breast carcinoma and corresponding synchronous and preceding DCIS. Am J Surg Pathol 43:1574–1582. https://doi.org/10.1097/PAS.0000000000001306
Beaubier N, Bontrager M, Huether R et al (2019) Integrated genomic profling expands clinical options for patients with cancer. NAture Biotech 37(11):1351–1360. https://doi.org/10.1038/s41587-019-0259-z
Beaubier N, Tell R, Lau D et al (2019) Clinical validation of the Tempus xT next-generation targetedoncology sequencing assay. Oncotarget 10(24):2384–2396
Dobin A, Davis CA, Schlesinger F et al (2013) Star: ultrafast universal RNA-se1 aligner. Bioinformatics 29(1):15–21. https://doi.org/10.1093/bioinformatics/bts635
Signal B, Kahlke T (2022) How are we stranded here: quick determination of RNA-seq strandedness. BMC Bioinform 23:49. https://doi.org/10.1186/s12859-022-04572-7
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550. https://doi.org/10.1186/s13059-014-0550-8
Gu Z (2016) Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32(18):2847–9. https://doi.org/10.1093/bioinformatics/btw313
Kolde R (2019) pheatmap: Pretty Heatmaps_.R package version 1.0.12. <https://CRAN.R-project.org/package=pheatmap>
Blighe K, Rana S, Lewis M (2023) EnhancedVolcano: Publication-ready volcano plots with enhanced colouring and labeling. https://doi.org/10.18129/B9.bioc.EnhancedVolcano, R package version 1.18.0; <https//bioconductor.org/packages/EnhancedVolcano>
Subramaniam A, Tamayo P, Mootha VK et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 43:15545–15550. https://doi.org/10.1073/pnas.0506580102
Merico D, Isserlin R, Stueker O et al (2010) Enrichment map: a network-based method for gene-set enrichment visualization and interpretation. PLoS One 5(11):e13984. https://doi.org/10.1371/journal.pone.0013984
Sherman BT, Hao M, Qiu J et al (2022) DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res 50(w1):W216–W221. https://doi.org/10.1093/nar/gkac194
Sidiropoulos K, Viteri G, Sevilla C et al (2017) Reactome enhanced pathway visualization. Bioinformatics 33(21):3461–3467. https://doi.org/10.1093/bioinformatics/btw441
“Extracellular matrix organization.” Reactome release #81. https://reactome.org/content/detail/R-HAS-1474244 (2202-07-18)
Xie X, Lu J, Kulbokas EJ et al (2005) Systematic discovery of regulatory motifs in human promoters and 3’ UTRs by comparison of several mammals. Nature 434:338–345. https://doi.org/10.1038/nature03441
Rebbeck CA, Xian J, Bornelöv S et al (2022) Gene expression signatures of individual ductal carcinoma in situ lesions identify processes and biomarkers associated with progression towards invasive ductal carcinoma. Nat Commun 13:3399. https://doi.org/10.1038/s41467-022-30573-4
Hayward M-K, Allen MD, Gomm JJ et al (2022) Mechanostimulation of breast myoepithelial cells induces functional changes associated with DCIS progression to invasion. npj Breast Cancer 8:1–12. https://doi.org/10.1038/s41523-022-00464-4
Williams MM, Cook RS (2015) Bcl-2 family proteins in breast development and cancer: could Mcl-1 targeting overcome therapeutic resistance? Oncotarget 6:3519–3530
Beroukhim R, Mermel CH, Porter D et al (2010) The landscape of somatic copy-number alteration across human cancers. Nature 463:899–905. https://doi.org/10.1038/nature08822
Xiang W, Yang C-Y, Bai L (2018) MCL-1 inhibition in cancer treatment. Onco Targets Ther 11:7301–7314. https://doi.org/10.2147/OTT.S146228
Young AIJ, Law AMK, Castillo L et al (2016) MCL-1 inhibition provides a new way to suppress breast cancer metastasis and increase sensitivity to dasatinib. Breast Cancer Res 18:125. https://doi.org/10.1186/s13058-016-0781-6
Louault K, Bonneaud TL, Séveno C et al (2019) Interactions between cancer-associated fibroblasts and tumor cells promote MCL-1 dependency in estrogen receptor-positive breast cancers. Oncogene 38:3261–3273. https://doi.org/10.1038/s41388-018-0635-z
Lee K, Giltnane JM, Balko JM et al (2017) MYC and MCL1 cooperatively promote chemotherapy-resistant breast cancer stem cells via regulation of mitochondrial oxidative phosphorylation. Cell Metab 26:633-647.e7. https://doi.org/10.1016/j.cmet.2017.09.009
Ferrari P, Scatena C, Ghilli M et al (2022) Molecular mechanisms, biomarkers and emerging therapies for chemotherapy resistant TNBC. Int J Mol Sci 23:1665. https://doi.org/10.3390/ijms23031665
Campbell KJ, Mason SM, Winder ML et al (2021) Breast cancer dependence on MCL-1 is due to its canonical anti-apoptotic function. Cell Death Differ 28:2589–2600. https://doi.org/10.1038/s41418-021-00773-4
Lin EY, Jones JG, Li P et al (2003) Progression to malignancy in the polyoma middle T oncoprotein mouse breast cancer model provides a reliable model for human diseases. Am J Pathol 163:2113–2126. https://doi.org/10.1016/S0002-9440(10)63568-7
Widden H, Placzek WJ (2021) The multiple mechanisms of MCL1 in the regulation of cell fate. Commun Biol 4:1–12. https://doi.org/10.1038/s42003-021-02564-6
Shrestha S, Yang C-D, Hong H-C et al (2018) Integrated microRNA–mRNA analysis reveals miR-204 inhibits cell proliferation in gastric cancer by targeting CKS1B, CXCL1 and GPRC5A. Int J Mol Sci 19:87. https://doi.org/10.3390/ijms19010087
Jia Y, Tian Q, Yang K et al (2021) A pan-cancer analysis of clinical prognosis and immune infiltration of CKS1B in human tumors. Biomed Res Int 2021:5862941. https://doi.org/10.1155/2021/5862941
Zhan F, Colla S, Wu X et al (2007) CKS1B, overexpressed in aggressive disease, regulates multiple myeloma growth and survival through SKP2- and p27Kip1-dependent and -independent mechanisms. Blood 109:4995–5001. https://doi.org/10.1182/blood-2006-07-038703
Lin L, Fang Z, Lin H et al (2016) Depletion of Cks1 and Cks2 expression compromises cell proliferation and enhance chemotherapy-induced apoptosis in HepG2 cells. Oncol Rep 35:26–32. https://doi.org/10.3892/or.2015.4372
Santarius T, Shipley J, Brewer D et al (2010) A census of amplified and overexpressed human cancer genes. Nat Rev Cancer 10:59–64. https://doi.org/10.1038/nrc2771
Slotky M, Shapira M, Ben-Izhak O et al (2005) The expression of the ubiquitin ligase subunit Cks55in human breast cancer. Breast Cancer Res 7:R737. https://doi.org/10.1186/bcr1278
Shi W, Huang Q, Xie J et al (2020) CKS1B as drug resistance-inducing gene—a potential target to improve cancer therapy. Front Oncol 10:582451. https://doi.org/10.3389/fonc.2020.582451
Shee K, Muller KE, Marotti J et al (2019) Ductal carcinoma in situ biomarkers in a precision medicine era. Am J Pathol 2019:956–965. https://doi.org/10.1016/j.ajpath.2018.08.020
Funding
This work was supported by the Kaplan Cancer Research Fund and the Providence Foundation of Oregon.
Author information
Authors and Affiliations
Contributions
Kaplan, Berry, and Piening contributed to the study conception and design. Material preparation, data collection and analysis were performed by all authors. The first draft of the manuscript was written by Kaplan and Piening. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors report no conflicts of interest.
Ethical approval
This study was approved by the Swedish Institutional Review Board (STUDY2023000084).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kaplan, H.G., Dowdell, A.K., Berry, A.B. et al. Multi-omic profiling of simultaneous ductal carcinoma in situ and invasive breast cancer. Breast Cancer Res Treat 205, 451–464 (2024). https://doi.org/10.1007/s10549-024-07270-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-024-07270-5