Abstract
Purpose
Triple-negative breast cancer (TNBC) is a highly heterogeneous disease. Patients with early-stage TNBCs have distinct likelihood of distant recurrence. This study aimed to develop a prognostic signature of early-stage TNBC patients to improve risk stratification.
Methods
Using RNA-sequencing data, we analyzed 189 pathologically confirmed pT1-2N0M0 TNBC patients and identified 21 mRNAs that were highly expressed in tumor and related to relapse-free survival. All-subset regression program was used for constructing a 7-mRNA signature in the training set (n = 159); the accuracy and prognostic value were then validated using an independent validation set (n = 158).
Results
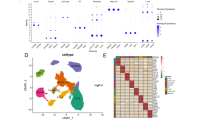
Here, we profiled the transcriptome data from 189 early-stage TNBC patients along with 50 paired normal tissues. Early-stage TNBCs mainly consisted of basal-like immune-suppressed subtype and had higher homologous recombination deficiency scores. We developed a prognostic signature including seven mRNAs (ACAN, KRT5, TMEM101, LCA5, RPP40, LAGE3, CDKL2). In both the training (n = 159) and validation set (n = 158), this signature could identify patients with relatively high recurrence risks and served as an independent prognostic factor. Time-dependent receiver operating curve showed that the signature had better prognostic value than traditional clinicopathological features in both sets. Functionally, we showed that TMEM101 promoted cell proliferation and migration in vitro, which represented a potential therapeutic target.
Conclusions
Our 7-mRNA signature could accurately predict recurrence risks of early-stage TNBCs. This model may facilitate personalized therapy decision-making for early-stage TNBCs individuals.




Similar content being viewed by others
Data availability
The datasets analyzed during the current study are available in the Gene Expression Omnibus (GSE76250, HTA2.0) and Sequence Read Archive (SRP157974, RNA-seq).
Abbreviations
- AIC:
-
Akaike information criterion
- AUC:
-
Area under curve
- BLIS:
-
Basal-like and immune-suppressed
- ER:
-
Estrogen receptor
- HER2:
-
Human epidermal growth factor receptor 2
- HR:
-
Hormone receptor
- HRD:
-
Homologous recombination deficiency
- IM:
-
Immunomodulatory
- LAR:
-
Luminal androgen receptor
- LASSO:
-
Least absolute shrinkage and selection operator
- MES:
-
Mesenchymal-like
- RFS:
-
Relapse-free survival
- ROC:
-
Receiver operating characteristic
- PR:
-
Progestogen receptor
- RT-qPCR:
-
Real-time quantitative-polymerase chain reaction
- siRNA:
-
Small interfering RNAs
- TNBC:
-
Triple-negative breast cancer
References
Foulkes WD, Smith IE, Reis-Filho JS (2010) Triple-negative breast cancer. N Engl J Med 363:1938–1948
Fan L, Strasser-Weippl K, Li J-J, St Louis J, Finkelstein DM, Yu K-D et al (2014) Breast cancer in China. Lancet Oncol 15:e279-289
Zeng H, Ran X, An L, Zheng R, Zhang S, Ji JS et al (2021) Disparities in stage at diagnosis for five common cancers in China: a multicentre, hospital-based, observational study. Lancet Public Health 6:e877–e887
Bao J, Donovan C, Amersi F, Zhang X, Giuliano AE, Chung A (2019) Outcomes in patients with small node-negative invasive breast cancer. Breast J 25:638–643
Parise CA, Caggiano V (2017) Risk of mortality of node-negative, ER/PR/HER2 breast cancer subtypes in T1, T2, and T3 tumors. Breast Cancer Res Treat 165:743–750
Amar S, McCullough AE, Tan W, Geiger XJ, Boughey JC, McNeil RB et al (2010) Prognosis and outcome of small (<=1 cm), node-negative breast cancer on the basis of hormonal and HER-2 status. Oncologist 15:1043–1049
de Nonneville A, Gonçalves A, Zemmour C, Cohen M, Classe JM, Reyal F et al (2017) Adjuvant chemotherapy in pT1ab node-negative triple-negative breast carcinomas: results of a national multi-institutional retrospective study. Eur J Cancer 84:34–43
Liu NQ, Stingl C, Look MP, Smid M, Braakman RBH, De Marchi T et al (2014) Comparative proteome analysis revealing an 11-protein signature for aggressive triple-negative breast cancer. J Natl Cancer Inst 106:djt376
Liu Y-R, Jiang Y-Z, Xu X-E, Yu K-D, Jin X, Hu X et al (2016) Comprehensive transcriptome analysis identifies novel molecular subtypes and subtype-specific RNAs of triple-negative breast cancer. Breast Cancer Res 18:33
Wang Y, Klijn JGM, Zhang Y, Sieuwerts AM, Look MP, Yang F et al (2005) Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet 365:9
Simon R, Altman DG (1994) Statistical aspects of prognostic factor studies in oncology. Br J Cancer 69:979–985
Xiao Y, Ma D, Zhao S, Suo C, Shi J, Xue M-Z et al (2019) Multi-omics profiling reveals distinct microenvironment characterization and suggests immune escape mechanisms of triple-negative breast cancer. Clin Cancer Res 25:5002–5014
Hammond MEH, Hayes DF, Wolff AC, Mangu PB, Temin S (2010) American Society of Clinical Oncology/College of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J Oncol Pract 6:195–197
Amin MB, Edge SB (2017) AJCC cancer staging manual. Springer, Cham
Tibshirani R (1997) The lasso method for variable selection in the Cox model. Stat Med 16:385–395
Calcagno V, de Mazancourt C (2010) glmulti: an R package for easy automated model selection with (generalized) linear models. J Stat Softw. https://doi.org/10.18637/jss.v034.i12
Youden WJ (1950) Index for rating diagnostic tests. Cancer 3:32–35
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102:15545–15550
Jin X, Xu X-E, Jiang Y-Z, Liu Y-R, Sun W, Guo Y-J et al (2019) The endogenous retrovirus-derived long noncoding RNA TROJAN promotes triple-negative breast cancer progression via ZMYND8 degradation. Sci Adv 5:eaat9820
Jiang Y-Z, Ma D, Suo C, Shi J, Xue M, Hu X et al (2019) Genomic and transcriptomic landscape of triple-negative breast cancers: subtypes and treatment strategies. Cancer Cell 35:428-440.e5
Timms KM, Abkevich V, Hughes E, Neff C, Reid J, Morris B et al (2014) Association of BRCA1/2 defects with genomic scores predictive of DNA damage repair deficiency among breast cancer subtypes. Breast Cancer Res. https://doi.org/10.1186/s13058-014-0475-x
Srikureja W, Kyulo NL, Runyon BA, Hu K-Q (2005) MELD score is a better prognostic model than Child-Turcotte-Pugh score or Discriminant Function score in patients with alcoholic hepatitis. J Hepatol 42:700–706
Panczykowski DM, Puccio AM, Scruggs BJ, Bauer JS, Hricik AJ, Beers SR et al (2012) Prospective independent validation of IMPACT modeling as a prognostic tool in severe traumatic brain injury. J Neurotrauma 29:47–52
Matsuda A, Suzuki Y, Honda G, Muramatsu S, Matsuzaki O, Nagano Y et al (2003) Large-scale identification and characterization of human genes that activate NF-kappaB and MAPK signaling pathways. Oncogene 22:3307–3318
Khan ZA, Labala RK, Yumnamcha T, Devi SD, Mondal G, Sanjita Devi H et al (2018) Artificial light at night (ALAN), an alarm to ovarian physiology: a study of possible chronodisruption on zebrafish (Danio rerio). Sci Total Environ 628–629:1407–1421
Fusella F, Seclì L, Busso E, Krepelova A, Moiso E, Rocca S et al (2017) The IKK/NF-κB signaling pathway requires Morgana to drive breast cancer metastasis. Nat Commun 8:1636
Rakha EA, El-Sayed ME, Green AR, Lee AHS, Robertson JF, Ellis IO (2007) Prognostic markers in triple-negative breast cancer. Cancer 109:25–32
Ren Y-X, Hao S, Jin X, Ye F-G, Gong Y, Jiang Y-Z et al (2019) Effects of adjuvant chemotherapy in T1N0M0 triple-negative breast cancer. Breast 43:97–104
Foekens JA, Atkins D, Zhang Y, Sweep FCGJ, Harbeck N, Paradiso A et al (2006) Multicenter validation of a gene expression-based prognostic signature in lymph node-negative primary breast cancer. J Clin Oncol 24:1665–1671
Perou CM, Sørlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA et al (2000) Molecular portraits of human breast tumours. Nature 406:747–752
Piccart M, van’t Veer LJ, Poncet C, Lopes Cardozo JMN, Delaloge S, Pierga J-Y et al (2021) 70-gene signature as an aid for treatment decisions in early breast cancer: updated results of the phase 3 randomised MINDACT trial with an exploratory analysis by age. Lancet Oncol 22:476–488
Liu Y-R, Jiang Y-Z, Xu X-E, Hu X, Yu K-D, Shao Z-M (2016) Comprehensive transcriptome profiling reveals multigene signatures in triple-negative breast cancer. Clin Cancer Res 22:1653–1662
Li L, Liu C, Amato RJ, Chang JT, Du G, Li W (2014) CDKL2 promotes epithelial-mesenchymal transition and breast cancer progression. Oncotarget 5:10840–10853
Pineda B, Diaz-Lagares A, Pérez-Fidalgo JA, Burgués O, González-Barrallo I, Crujeiras AB et al (2019) A two-gene epigenetic signature for the prediction of response to neoadjuvant chemotherapy in triple-negative breast cancer patients. Clin Epigenetics 11:33
Zhao SG, Chen WS, Das R, Chang SL, Tomlins SA, Chou J et al (2019) Clinical and genomic implications of luminal and basal subtypes across carcinomas. Clin Cancer Res 25:2450–2457
Dadhania V, Zhang M, Zhang L, Bondaruk J, Majewski T, Siefker-Radtke A et al (2016) Meta-analysis of the luminal and basal subtypes of bladder cancer and the identification of signature immunohistochemical markers for clinical use. EBioMedicine 12:105–117
Rodriguez-Pinilla SM (2006) Prognostic significance of basal-like phenotype and fascin expression in node-negative invasive breast carcinomas. Clin Cancer Res 12:1533–1539
van de Rijn M, Perou CM, Tibshirani R, Haas P, Kallioniemi O, Kononen J et al (2002) Expression of cytokeratins 17 and 5 identifies a group of breast carcinomas with poor clinical outcome. Am J Pathol 161:1991–1996
Rowlands D, Lensjø KK, Dinh T, Yang S, Andrews MR, Hafting T et al (2018) Aggrecan directs extracellular matrix-mediated neuronal plasticity. J Neurosci 38:10102–10113
Suna G, Wojakowski W, Lynch M, Barallobre-Barreiro J, Yin X, Mayr U et al (2018) Extracellular matrix proteomics reveals interplay of aggrecan and aggrecanases in vascular remodeling of stented coronary arteries. Circulation 137:166–183
Oskarsson T (2013) Extracellular matrix components in breast cancer progression and metastasis. Breast 22(Suppl 2):S66-72
Joensuu H, Kellokumpu-Lehtinen P-L, Huovinen R, Jukkola-Vuorinen A, Tanner M, Kokko R et al (2017) Adjuvant capecitabine in combination with docetaxel, epirubicin, and cyclophosphamide for early breast cancer: the randomized clinical FinXX trial. JAMA Oncol 3:793–800
Masuda N, Lee S-J, Ohtani S, Im Y-H, Lee E-S, Yokota I et al (2017) Adjuvant capecitabine for breast cancer after preoperative chemotherapy. N Engl J Med 376:2147–2159
Acknowledgements
This work was supported by the Program of Shanghai Academic/Technology Research Leader (20XD1421100), the Fok Ying-Tong Education Foundation for College Young Teachers (171034), the Shanghai Sailing Program (19YF1409000), the Innovation Team of Ministry of Education (IRT1223), and the Shanghai Key Laboratory of Breast Cancer (12DZ2260100). The funders had no role in the study design, data collection and analysis, or manuscript preparation.
Author information
Authors and Affiliations
Contributions
Conception and design: Y-SY, Y-XR, C-LL, XJ, X-EX, Y-ZJ, and Z-MS; Development of methodology: Y-SY, Y-XR, X-EX, and Y-ZJ; Acquisition of data: Y-SY, Y-XR, SH, and Y-ZJ; Analysis and interpretation of data: Y-SY, Y-XR, C-LL, XJ, and Y-ZJ; Writing, review, and/or revision of the manuscript: Y-SY, Y-XR, C-LL, SH, XJ, Y-ZJ, and Z-MS; Administrative, technical, or material support: Y-SY, Y-XR, SH, XJ, X-EX, and Y-ZJ; Study supervision: Y-ZJ, Z-MS.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
The independent ethics committee/institutional review board of FUSCC (Shanghai Cancer Center Ethics Committee) approved our study.
Consent to participate
Written informed consent from patients in our study was obtained before enrollment.
Consent for publication
All authors read and approved the manuscript as submitted.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yang, YS., Ren, YX., Liu, CL. et al. The early-stage triple-negative breast cancer landscape derives a novel prognostic signature and therapeutic target. Breast Cancer Res Treat 193, 319–330 (2022). https://doi.org/10.1007/s10549-022-06537-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-022-06537-z