Abstract
Trastuzumab is effective in the treatment of HER2/neu over-expressing breast cancer, but not all patients benefit from it. In vitro data suggest a role for HER3 in the initiation of signaling activity involving the AKT–mTOR pathway leading to trastuzumab insensitivity. We sought to investigate the potential of HER3 alone and in the context of p95HER2 (p95), a trastuzumab resistance marker, as biomarkers of trastuzumab escape. Using the VeraTag® assay platform, we developed a dual antibody proximity-based assay for the precise quantitation of HER3 total protein (H3T) from formalin-fixed paraffin-embedded (FFPE) breast tumors. We then measured H3T in 89 patients with metastatic breast cancer treated with trastuzumab-based therapy, and correlated the results with progression-free survival and overall survival using Kaplan–Meier and decision tree analyses that also included HER2 total (H2T) and p95 expression levels. Within the sub-population of patients that over-expressed HER2, high levels of HER3 and/or p95 protein expression were significantly associated with poor clinical outcomes on trastuzumab-based therapy. Based on quantitative H3T, p95, and H2T measurements, multiple subtypes of HER2-positive breast cancer were identified that differ in their outcome following trastuzumab therapy. These data suggest that HER3 and p95 are informative biomarkers of clinical outcomes on trastuzumab therapy, and that multiple subtypes of HER2-positive breast cancer may be defined by quantitative measurements of H3T, p95, and H2T.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Trastuzumab (Herceptin®, Genentech), a humanized monoclonal antibody that targets the extracellular domain of the ErbB2 (HER2) receptor, is an effective treatment of both early and metastatic HER2-positive breast cancer (MBC), particularly when used in combination with chemotherapy [1–6]. However, many patients treated with trastuzumab do not respond, or experience disease recurrence, despite disease classification as HER2-positive by immunohistochemistry (IHC) or fluorescence in situ hybridization (FISH).
Much attention has been focused on improving the prediction of clinical benefit, as well as overcoming the technical shortcomings of current diagnostic methods [7–10]. It is plausible that clinical outcomes can be improved simply through better selection of patient candidates for trastuzumab by increasing the accuracy of HER2 assessments, if the increased accuracy extends the treatment indication to cases which were otherwise excluded. However, assuming HER2 is the sole indicator of trastuzumab responsiveness ignores the complex biological context of the cell signaling system in which HER2 operates.
HER2 functions as a partner with other HER receptor family members in the formation of heterodimers, which initiate signaling through several key pathways that are known to drive proliferation and survival in epithelial cancers [11, 12]. Given the overlap and redundancy of cell signaling pathways, the measurement of HER2 alone is likely insufficient to accurately characterize trastuzumab sensitivity for any given tumor. While overexpression of HER2 protein may be necessary for trastuzumab activity in vivo, the concomitant expression of other HER-family receptors may provide opportunities for escape from trastuzumab antagonism via their role as dimerization partners of HER2. Unless methods are developed to measure additional biomarkers of trastuzumab response, it will remain difficult to distinguish HER2-positive tumors that are driven primarily by HER2 overexpression, and thus may be highly susceptible to trastuzumab, from HER2-positive tumors that are driven by HER2 heterodimers (e.g., HER2:HER3) and may not be easily antagonized by trastuzumab [13].
Although studies suggest that ligand-independent HER2:HER3 heterodimers represent a trastuzumab-sensitive oncogenic driver in HER2-positive breast cancer [14, 15], one can hypothesize that the concomitant high expression of HER3 or its ligands in HER2-positive tumors may confer escape from trastuzumab inhibition. Recent data suggests that upregulation of a set of receptor tyrosine kinases, including HER3, is pivotal in attenuating the anti-tumor effects of a number of inhibitors of the PI3K/AKT pathway [16], including those that act directly on HER2 [17, 18]. Furthermore, the down-regulation of HER3 mRNA has been postulated as a surrogate for activation of HER2 dimerization [19], and HER3 mRNA levels have been correlated with clinical benefit in ovarian tumors treated with the HER2:HER3 dimerization-inhibiting antibody, pertuzumab [20]. Lastly, in vitro data suggest that heregulin-induced HER2:HER3 heterodimers mediate trastuzumab insensitivity by activating signaling through the AKT–mTOR pathway, and that trastuzumab is ineffective in abrogating this signal [13]. Recent clinical data demonstrating the enhanced efficacy in both early and metastatic HER2-positive breast cancer with the addition of the heterodimer-blocking HER2 antibody pertuzumab to trastuzumab is consistent with ligand-activated and possibly ligand-independent HER2:HER3 signaling in HER2-positive breast cancer [21, 22].
In this study, we sought to explore the association between HER3 protein expression and clinical outcome following trastuzumab therapy in a cohort of MBC patients with HER2 positive tumors. We hypothesized that elevated HER3 protein expression in the setting of HER2 overexpression is a reasonable surrogate for the presence of HER2:HER3 heterodimers given a correlation between the two measurements [23, 24], and therefore may correlate with suboptimal response to trastuzumab. HER3 is expressed at relatively low levels and is difficult to quantify by traditional IHC procedures in FFPE sections; therefore we used the VeraTag platform [25, 26] to develop a proximity-based assay for the precise quantitation of HER3 protein expression in FFPE specimens. We then measured HER3 expression in a cohort of trastuzumab-treated MBC patients and performed statistical analyses to explore the relationship between HER3 expression and clinical outcomes following trastuzumab-based treatment.
Finally, having previously found a dependence of outcome on quantitative HER2 [27] and p95 (truncated HER2) levels [28] in the current cohort, we further examined the relationship of HER3 expression with clinical outcome in the context of HER2 protein expression and p95 expression using regression tree analysis.
Patients and methods
Materials
HER3 monoclonal antibody Ab6 was purchased from Lab Vision (Fremont, CA). HER3 monoclonal antibody B9A11 was generated in-house as described in the Supplementary Methods. Biotinylated and unconjugated mouse monoclonal anti-human IgG1 antibodies were purchased from BD Biosciences (Franklin Lakes, NJ). The fluorescent reporter Pro11 was synthesized and purified as described in US Patent 7,105,308. Antibody conjugates Ab6–Pro11, B9A11–biotin, and IgG1–Pro11 were conjugated and purified as described [25, 26]. Ab6–Pro11 and IgG1–Pro11 conjugations were matched to similar hapten derivatization ratios. All antibody conjugations were stored in 1× PBS with 1 mg/mL BSA and 0.001 % sodium azide (Sigma-Aldrich, St. Louis, MO). Streptavidin-conjugated methylene blue was synthesized and purified as described in US Patent 7,105,308.
Antibody characterization was performed using cell lysates and FFPE slides prepared from MDA-MB-453, MDA-MB-468, T47D, MDA-MB-231, SK-OV-3, and H1650 cell lines as described in the Supplementary Methods. The cross-validation of VeraTag assays using ELISA, flow cytometry, and IHC assays is described in the Supplementary Methods.
VeraTag assays
The H2T (HERmark® HER2 VeraTag assay) and p95 assays were performed as previously described [25, 26, 28]. The p95 VeraTag assay measures the highly active, membrane-bound Met611–p95 form as previously described [28]. Similar to the H2T assay (Fig. 1a, left panel), the H3T assay (center panel) utilizes a pair of monoclonal antibodies that bind epitopes in the C-terminal, intracellular region of the receptor. H3T assay details are described in the Supplementary Methods.
a HER3 total protein assay (H3T) design. The H3T assay (center panel) is a proximity binding assay that utilizes two monoclonal antibodies that recognize distinct epitopes in the intracellular domain of the HER3 receptor to generate a highly specific and quantifiable signal by capillary electrophoresis in FFPE tumor specimens. The H3T assay is similar in design to the H2T assay (left panel) described previously [26]. One antibody in a proximity pair is tethered to a fluorescent reporter group via a molecular moiety containing a cleavable thio-ether bond while the other contains a bound biotin–streptavidin–methylene blue complex that upon illumination (650 nm) results in the generation of singlet oxygen molecules which release the fluorescent reporter molecules from antibody–antigen complexes by cleaving the thio-ether linkage. Released reporter molecules are quantified using capillary electrophoresis. The p95 assay (right panel) uses a proprietary mouse monoclonal antibody specific for the N-terminal amino acid sequence of p95 that is recognized by a secondary anti-mouse antibody. This assay has also been described previously [28], and uses the reduction of a disulfide linkage by dithiothreitol (DTT) to liberate the fluorescent reporter molecule from antibody–antigen complexes. b H3T assay accuracy. HER3 protein measurements for a panel of cell lines that express variable amounts of HER3 were generated using the VeraTag H3T assay and compared to the HER3 measurements obtained using conventional technologies including IHC, ELISA, and flow cytometry. VeraTag H3T levels are expressed in units of relative fluorescence per mm2 of invasive tumor or control cell lines (RF/mm2). c H3T assay precision (intra-assay variation). Replicate H3T measurements for four cell lines that express variable amounts of HER3 protein were generated in the same assay run. Intra-assay variation (CV) was 29, 16, 12, and 10 % for the SK-OV-3, MDA-MB 231, MDA-MB 468, and MDA-MB 453 cell lines, respectively. d H3T assay reproducibility (inter-assay variation). Duplicate H3T measurements for 57 FFPE breast cancer specimens were generated in separate assay runs with a mean CV of 18 %. e H3T assay specificity and sensitivity of detection. The specificity of the release of the H3T fluorophore reporter [H3T (Pro-11)] and the lower limit of H3T detection were demonstrated in both tumor cell lines and FFPE specimens by substituting the photosensitizing (“scissor”) H3T antibody with an isotype matched control antibody. Assay criteria have been established to define specimens with H3T signals less than minimum (LTM) signal as negative for HER3 protein (see “Results” section). f H3T assay specificity in the presence of “interfering” substances. To evaluate the binding of HER3 antibodies to non-tumor tissue in FFPE tumor specimens the H3T assay was performed on two breast cancer specimens with and without exogenous fat and stroma as described in the Supplementary Methods. Non-tumor stroma and fat do not contribute to the H3T signal
Study population
This cohort has been thoroughly described in prior publications [27–29]; the characteristics of the specific patient set analyzed here are presented in Supplementary Table S1. Briefly, the cohort is comprised of patients initially identified as having HER2-positive MBC by standard pathologic criteria of HER2 IHC 3+ staining (HercepTest; DAKO Diagnostics) or HER2 FISH gene copy number to centromere 17 ratio of ≥2.0 and prospectively observed during the administration of trastuzumab-based therapy at the Medical University of Vienna between 1999 and 2006. ER and PR were considered positive with detectable nuclear staining in ≥10 % of cells. Five patients received endocrine therapy (exemestane) concurrently with trastuzumab. All treatment decisions were made by the treating physicians. Response to treatment was documented by review of all imaging studies according to SWOG criteria, and data on HER2 and hormone receptor status were retrieved from pathology reports. Tissue for analysis was derived from the primary tumor, five of which were core biopsies and the remainder excisional biopsies. All patients were re-tested by central laboratory FISH (PathVysion; Vysis/Abbott, Des Plaines, IL) and by HERmark for determination of HER2 gene copy number and protein expression levels (H2T), respectively. The concordance and discordance between the various HER2 assay results have been reported [29]. The expression of p95 in this cohort has also been characterized and reported [28]. Written informed consent was obtained prior to initiation of treatment and biomarker studies. The research protocol was approved by the Institutional Review Boards of Penn State University/Hershey Medical Center and the Medical University of Vienna.
Statistical methods
All HERmark H2T assays were performed retrospectively and in blinded fashion. Analyses of cutoffs were exploratory in nature. Kaplan–Meier analyses used the log rank test. All p values were two-sided. Progression-free survival (PFS) was defined as the time from the initiation of trastuzumab-based treatment to progression or censor, and overall survival (OS) was defined as the time from initiation of trastuzumab-based treatment to death or censor. Patients were classified as H2T-high or H2T-low (HER2-normal) expression measured by the HERmark test and based on cutoffs from previously published analyses [29]. The univariate cutoff discriminating high from normal H3T expression was defined using positional scanning and selection based on the lowest p value as previously described [29].
Recursive partitioning was used to correlate continuous H2T, H3T, and p95 measurements with PFS data. The probabilities of progression for the different subgroups derived from the optimal partition tree were then compared using Kaplan–Meier analysis. Regression analyses were carried out using the R package rpart version 4.1-0 [30].
Results
H3T assay design and performance characteristics
The H3T assay was developed to quantify the total expression of HER3 protein in FFPE specimens. The H3T assay is based on proximity binding of two derivatized antibodies and is similar in design to the VeraTag HER2 assay (HERmark), as previously described [25, 29] (Fig. 1a).
The performance characteristics of the VeraTag H3T assay are illustrated in Fig. 1b–f. H3T assay accuracy was cross-validated using a panel of cell lines expressing different levels of HER3 protein, prepared either as FFPE blocks, cell lysates or fixed cells, by comparing H3T measurements obtained using the VeraTag assay to measurements derived using several standard, well established technologies (Fig. 1b). ELISA and VeraTag measurements display a strong, linear correlation (R 2 = 0.97, p = 0.0002) across more than 2 orders of magnitude of HER3 expression. Based on flow cytometry data, HER3 receptor levels varied from several hundred to thirty-thousand receptors per cell in the selected cell lines. As expected, a strong correlation (R 2 = 0.89, p = 0.005) was found between VeraTag H3T measurements and the number of receptors per cell. HER3 IHC was also performed on FFPE sections prepared from cell line pellets. As expected, MB453 cells stained strongly, consistent with high HER3 expression. Staining for the other cell lines was much weaker, with the IHC limit of detection near the HER3 expression levels in MB468 cells. In contrast to IHC where only slight differences in staining were observed between the MB468, MB231, and SKOV3 cell lines, the VeraTag assay revealed a ~20-fold difference in H3T expression levels across these cell lines, consistent with FACS and ELISA results.
The precision (intra-assay variation) of the H3T assay was demonstrated by generating replicate measurements of H3T expression in cell pellets prepared by FFPE and tested in the same assay batch (Fig. 1c). Assay reproducibility (inter-assay variation) was demonstrated by performing duplicate measurements of H3T in 57 breast tumors prepared as FFPE specimens and tested in different assay batches (Fig. 1d). The excellent correlation between duplicate measurements (Spearman r = 0.94, p < 0.0001) showed a mean CV of 18 %.
The specificity of the VeraTag assay for HER3 detection was evaluated by substituting non-specific isotype matching control (ITC) antibodies for each of the HER3 specific antibodies (Ab6, B9A11) in the assay. This experiment was performed in both cell lines and breast tumors prepared as FFPE specimens (Fig. 1e). The overlap in assay signal between tumors at the low end of the H3T dynamic range and tumors at the high end of the ITC range is small and thus was used to establish a less than minimum (LTM) signal threshold. Samples that generate LTM H3T values were considered negative for HER3 within the sensitivity limits of the assay.
The potential contribution to the H3T assay signal by non-specific Ab6 and B9A11 binding to fat and non-tumor stroma was evaluated by comparing H3T measurements obtained using multiple paired tumor sections that were or were not macro-dissected to remove non-tumor tissue prior to testing. H3T measurements were not affected by the presence of fat or non-tumor stroma in the tumor section (Fig. 1f). The specificity of the two HER3 antibodies used in the assay was further confirmed by the absence of cross-reactivity in SDS-PAGE/western blot and VeraTag assays using cell lines expressing variable levels of HER3, HER2, and HER1 (data not shown).
Impact of HER3 expression on clinical outcomes in trastuzumab-treated MBC
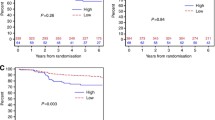
H3T results were generated for 89 specimens with available tissue in a cohort of patients with MBC that were treated with trastuzumab and had well-documented clinical outcomes (PFS and OS). The clinical characteristics of the patients are described in Supplementary Table S1, and correlation of clinical characteristics is reported in Table S2. HER2 tumor status and trastuzumab treatment were based largely on HER2 IHC results with the remainder by HER2 FISH. However, retrospective testing by HERmark, a quantitative HER2 protein assay, and central laboratory performed HER2 FISH revealed better outcomes in patients with HERmark HER2-positive tumors and, to a lesser extent, those with HER2 FISH-positive tumors [29]. In the current study, we sought to determine whether those patients with the most favorable outcomes as predicted by HERmark or HER2 FISH assays could be subdivided into groups of different outcomes by applying a quantitative HER3 cutoff. Sixty-one of the 89 patients that had tissue available for H3T analysis were assessed as HER2-high or H2T-high (H2T > 13.8) by the HERmark H2T assay, as previously reported [29]. We were unable to derive precise H3T values for four H2T-high cases. Based on the correlation between H3T expression levels and PFS for the remaining 57 H2T-high cases, we used positional scanning analyses of the hazard ratio and selection by the lowest p value to establish a H3T assay cutoff (H3T = 3.5) that best discriminated patient subgroups with significantly different outcomes. Using this cutoff, Kaplan–Meier analyses were performed to compare outcomes (PFS and OS) for the two H2T-high groups, i.e., HER3-low (H3T ≤ 3.5) and HER3-high (H3T > 3.5), against the H2T-low (HER2-normal) group (Fig. 2a, b; Table 1). Within the H2T-high group (H2T > 13.8), patients with HER3-low tumors experienced longer PFS than patients with HER3-high tumors (PFS: HR = 2.7, p = 0.0021), although no significant difference in OS was observed. Similar results were found when focusing on the HER2 FISH-positive subgroup (Fig. 2c, d; Table 1). The clinical outcomes of the H2T-high (H2T > 13.8)/HER3-high (H3T > 3.5) group were comparable to the H2T-low group (H2T ≤ 13.8; Table 1). Within the H2T-low group, the four patients with H3T > 3.5 had PFS (p = 0.6) and OS (p = 0.14) that were indistinguishable from the poor outcomes of the other H2T-low patients.
Clinical outcome based on HER2 and HER3 expression levels. Kaplan–Meier analyses of PFS (a, c) and OS (b, d) for MBC patients that received a trastuzumab containing regimen. Patient groups were classified based on low and high HER2 (H2T) (a, b) or HER2 FISH (c, d) and HER3 (H3T) expression levels. Discrimination of H3T high from H3T low expression was performed by positional scanning with selection of the H3T value with the most significant p value (see “Patients and methods” section for details)
Impact of HER3 expression on clinical outcomes independent of p95 and very high H2T levels
Next, a multivariate analysis was undertaken to determine the role of HER3 in the context of both H2T and p95. A regression tree analysis was used to ensure that previously established univariate cutoffs for H2T and p95 were similar when considering H3T, H2T, and p95 simultaneously. Using this same cohort of trastuzumab-treated MBC patients, we previously reported that within the confirmed HER2-positive group (H2T > 13.8 or FISH/CEP17 ≥ 2.0), patients with tumors that expressed elevated levels of p95 (p95-high) experienced less favorable outcomes compared to the subgroup of patients with tumors that expressed low p95 levels (p95-low) [28]. Also in the confirmed HER2-positive group, patients with tumors that expressed very high H2T levels of H2T > 68.5 in the current cohort [27] and H2T > 125 in an early breast cancer cohort [31] experienced less favorable outcomes compared to the rest of the HER2-positives. Having applied the VeraTag HER3 assay to measure H3T expression, we next considered HER3 expression in the context of p95 and quantitative HER2 expression to explore whether these additional measurements could further explain the varied clinical outcomes of patients within the confirmed HER2-positive group. No significant correlations were observed between either the levels of H3T and H2T (Spearman p = 0.7), or H3T and p95 (Spearman p = 0.6).
We used the regression tree method as a multivariate analysis to scan the continuum of H2T, H3T, and p95 measurements for significant association with PFS. Subgroups of patients defined by these biomarkers were identified that experienced different clinical outcomes in response to trastuzumab-based therapies. The optimal regression tree and the Kaplan–Meier curves derived from the subgroups defined by the tree are presented in Figs. 3, 4, and 5. PFS and OS for each group are given in Table 2. The first split of the tree is based on a H2T cutoff value of 16.1, which separates patients with low HER2 expression (H2T < 16.1) on the right from those with high HER2 expression (H2T ≥ 16.1) on the left. This cutoff value is similar to the H2T ≤ 13.8 clinical cutoff previously reported [29], and used in the univariate analysis of HER3 expression described above. These patients are segregated next by intermediate HER2 expression (16.1 ≤ H2T ≤ 68.3) versus very high HER2 expression (H2T > 68.3). It is important to note that this HER2-intermediate group identified by regression tree analysis is essentially the HER2-positives (i.e., not HER2-low), excluding only those expressing extremely high levels of HER2, greater than approximately fivefold above the cutoff for HER2-positivity. The patients within the HER2-intermediate subgroup were further separated based on H3T and p95 expression levels (Table 2; Fig. 3). Subgroup A, characterized as H2T-intermediate, H3T-low, and p95-low, experienced the best clinical outcomes (Fig. 5; Table 2). In this cohort of HER2-positive MBC patients treated with trastuzumab, multiple factors were found to be statistically significant independent correlates of shorter PFS and OS including elevated p95 expression (p95 > 3.75), elevated HER3 expression (H3T > 3.89), very high levels of HER2 expression (H2T > 68.3), and low (normal) levels of HER2 expression (H2T ≤ 16.1).
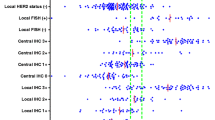
Distributions of H3T versus H2T (a) and p95 versus H2T (b). As mentioned in the “Methods” section, H3T measurements with less than the minimum quantifiable signal are designated LTM. Cutoffs from the univariate analysis are shown in red as shown in Ref. [29] for H2T, Fig. 3 for H3T, and in Ref. [28] for p95. Cutoffs derived from multivariate recursive partitioning are shown in blue
Clinical outcomes of HER2-positive subgroups established by recursive partitioning of HER2, HER3, and p95 expression levels. Kaplan–Meier plots of PFS (a) and OS (b) for multiple sub-populations of HER2 MBC patients based on recursive partitioning (Fig. 3) of HER2 (H2T), HER3 (H3T), and p95 expression levels. The groups are defined as shown in Fig. 3 and described in Table 2
Discussion
Multiple, large, prospectively randomized clinical trials have clearly demonstrated that trastuzumab is effective in the treatment of HER2-positive breast cancer in both the early and metastatic settings. However, many individuals with HER2-positive breast cancer fail to respond or derive sustainable benefit from this drug. To improve the clinical outcomes of patients with HER2-positive breast cancer, it is imperative that we understand why not all patients benefit equally from trastuzumab treatment. In vitro data have suggested that signaling through the AKT–mTOR pathway initiated by HER2:HER3 heterodimerization and mediated by PI3 kinase provides tumor cells that overexpress HER2 with an escape route from the antagonistic effects of trastuzumab. Indeed, pertuzumab was developed precisely to target this putative resistance mechanism. Recent data indicate that pertuzumab can abrogate signaling through AKT–mTOR in response to heregulin-mediated HER2:HER3 heterodimerization, while trastuzumab cannot [13, 14, 21, 22]. Thus, we reasoned that accurate measurements of HER3 protein expression might permit the identification of HER2-positive breast cancer patients whose suboptimal outcomes on trastuzumab therapy could be explained by HER3-mediated signaling.
The data presented here represent the first clinical description of trastuzumab escape mediated by the co-expression of HER2 and HER3 receptors in metastatic breast tumors. In the clinical cohort studied here, the PFS of HER2-positive MBC patients with higher HER3 expression levels was similar to patients with tumors expressing normal (low) levels of HER2 (median PFS 5.0 vs. 4.2 months, respectively) and classified as HER2 FISH-negative upon central laboratory re-testing [29]. By comparison, the median PFS of patients having tumors that over-expressed HER2 but with lower levels of HER3 expression was 12.1 months, supporting a correlation between high-HER2/high-HER3 co-expression and reduced sensitivity to trastuzumab.
While it may be true that some patients with tumors that overexpress HER2 fail trastuzumab because their tumors also highly express HER3, it is likely that not every patient that fails trastuzumab necessarily does so because of elevated HER3 expression. Previously, in this same cohort, we reported a correlation between elevated p95 expression and poor outcomes in response to trastuzumab treatment [28]. Furthermore, using this cohort, as well as an early breast cancer trastuzumab-treated cohort, we observed that patients with tumors that overexpress HER2 at very high levels fare considerably worse than patients with tumors that overexpress HER2 at intermediate levels [27, 31]. Building on our previous observations, we carried out a multivariate analysis to investigate whether the patient group with HER3-high tumors and poor trastuzumab treatment outcomes overlapped with patient groups with either p95-high or H2T-very high tumors. Kaplan–Meier analyses on groups identified by the regression tree analysis demonstrated that patients with tumors that expressed intermediate levels of HER2, essentially all HER2-positives except for the very high expressors, in the absence of high HER3 and/or high p95 expression were most responsive to trastuzumab treatment (median PFS 15.7 months, median OS 47.6 months) (Fig. 5; Table 2). While the tumors of these patients had been confirmed as HER2-positive (IHC 3+, central laboratory FISH-positive, HERmark H2T > 13.8) [29], they exhibited no other risk factors associated with trastuzumab treatment failure using currently accepted clinical criteria. It remains likely that even this optimally performing subgroup (HER2-intermediate, HER3-low, p95-low expression) is heterogeneous, and the further expansion of this analysis to include analytes such as HER1, PIK3CA mutations, and PTEN might identify additional subgroups with suboptimal outcomes that further diminish the overall response rates of HER2-positive MBC to trastuzumab containing therapies. Admittedly, such analyses will require significantly more patients in order to be statistically meaningful.
Conclusions
The observations reported here suggest the existence of multiple prognostic subtypes of HER2-positive breast cancer, characterized by the co-expression of HER2, HER3, and/or p95, and are consistent with current models of trastuzumab resistance. Furthermore, these data not only offer a potential explanation for the clinical failure of trastuzumab treatment despite confirmed HER2 overexpression, but also implicate particular subsets of HER2-positive breast cancer patients as candidates for combination drug therapy employing multiple targeted inhibitors of aberrant signaling mediated by HER2, HER3, and/or p95. Nevertheless, these findings represent exploratory retrospective analyses of a well-characterized, yet small and uncontrolled, clinical cohort of patients with MBC, and thus must be considered hypothesis-generating. To establish true clinical utility, these observations will require confirmation in larger, well-controlled studies using pre-defined clinical cutoffs for HER2, HER3, and p95 expression in blinded analyses.
References
Joensuu H, Kellokumpu-Lehtinen PL, Bono P et al (2006) Adjuvant docetaxel or vinorelbine with or without trastuzumab for breast cancer. N Engl J Med 354:809–820
Piccart-Gebhart MJ, Procter M, Leyland-Jones B et al (2005) Trastuzumab after adjuvant chemotherapy in HER2-positive breast cancer. N Engl J Med 353:1659–1672
Slamon DJ, Leyland-Jones B, Shak S et al (2001) Use of chemotherapy plus a monoclonal antibody against HER2 for metastatic breast cancer that overexpresses HER2. N Engl J Med 344:783–792
Vogel CL, Cobleigh MA, Tripathy D et al (2002) Efficacy and safety of trastuzumab as a single agent in first-line treatment of HER2-overexpressing metastatic breast cancer. J Clin Oncol 20:719–726
Cobleigh MA, Vogel CL, Tripathy D et al (1999) Multinational study of the efficacy and safety of humanized anti-HER2 monoclonal antibody in women who have HER2-overexpressing metastatic breast cancer that has progressed after chemotherapy for metastatic disease. J Clin Oncol 17:2639–2648
Romond EH, Perez EA, Bryant J et al (2005) Trastuzumab plus adjuvant chemotherapy for operable HER2-positive breast cancer. N Engl J Med 353:1673–1684
Wolff AC, Hammond ME, Schwartz JN et al (2007) American Society of Clinical Oncology/College of American Pathologists Guideline Recommendations for human epidermal growth factor receptor 2 testing in breast cancer. Arch Pathol Lab Med 131:18
Press MF, Sauter G, Bernstein L et al (2005) Diagnostic evaluation of HER-2 as a molecular target: an assessment of accuracy and reproducibility of laboratory testing in large, prospective, randomized clinical trials. Clin Cancer Res 11:6598–6607
Press MF, Slamon DJ, Flom KJ et al (2002) Evaluation of HER-2/neu gene amplification and overexpression: comparison of frequently used assay methods in a molecularly characterized cohort of breast cancer specimens. J Clin Oncol 20:3095–3105
Perez EA, Suman VJ, Davidson NE et al (2006) HER2 testing by local, central, and reference laboratories in specimens from the North Central Cancer Treatment Group N9831 intergroup adjuvant trial. J Clin Oncol 24:3032–3038
Yarden Y (2001) The EGFR family and its ligands in human cancer: signalling mechanisms and therapeutic opportunities. Eur J Cancer 37(Suppl 4):S3–S8
Yarden Y, Sliwkowski MX (2001) Untangling the ErbB signalling network. Nat Rev Mol Cell Biol 2:127–137
Ghosh R, Narasanna A, Wang SE et al (2011) Trastuzumab Has Preferential Activity against Breast Cancers Driven by HER2 Homodimers. Cancer Res 71:1871–1882
Junttila TT, Akita RW, Parsons K et al (2009) Ligand-independent HER2/HER3/PI3K complex is disrupted by trastuzumab and is effectively inhibited by the PI3K inhibitor GDC-0941. Cancer Cell 15:429–440
Lee-Hoeflich ST, Crocker L, Yao E et al (2008) A central role for HER3 in HER2-amplified breast cancer: implications for targeted therapy. Cancer Res 68:5878–5887
Serra V, Scaltriti M, Prudkin L et al (2011) PI3K inhibition results in enhanced HER signaling and acquired ERK dependency in HER2-overexpressing breast cancer. Oncogene 30:2547–2557
Garrett JT, Olivares MG, Rinehart C et al (2009) Transcriptional and posttranslational up-regulation of HER3 (ErbB3) compensates for inhibition of the HER2 tyrosine kinase. Proc Natl Acad Sci U S A 108:5021–5026
Sergina NV, Rausch M, Wang D et al (2007) Escape from HER-family tyrosine kinase inhibitor therapy by the kinase-inactive HER3. Nature 445:437–441
Nagumo Y, Faratian D, Mullen P et al (2009) Modulation of HER3 is a marker of dynamic cell signaling in ovarian cancer: implications for pertuzumab sensitivity. Mol Cancer Res 7:1563–1571
Makhija S, Amler LC, Glenn D et al (2010) Clinical activity of gemcitabine plus pertuzumab in platinum-resistant ovarian cancer, fallopian tube cancer, or primary peritoneal cancer. J Clin Oncol 28:1215–1223
Baselga J, Cortes J, Kim SB et al (2011) Pertuzumab plus trastuzumab plus docetaxel for metastatic breast cancer. N Engl J Med 366:109–119
Gianni L, Bianchini G, Kiermaier A et al (2011) Neoadjuvant pertuzumab (P) and trastuzumab (H): biomarker analyses of a 4-Arm randomized phase II study (NeoSphere) in patients (pts) with HER2-positive breast cancer (BC). Cancer Res 71:S5–S11
Mukherjee A, Badal Y, Nguyen X-T et al (2011) Profiling the HER3/PI3K pathway in breast tumors using proximity-directed assays identifies correlations between protein complexes and phosphoproteins. PLoS ONE 6(1):e16443
Wallweber J, Chenna A, Ravanera R, et al (2013) Profiling HER3/ErbB3 activation in formalin-fixed, paraffin-embedded (FFPE) breast tumor samples that express high and low HER2/ErbB2 levels using proximity-based immunoassays. AACR Annual Meeting, April 2013; 3029
Larson J, Goodman L, Tan Y, DeFazio-Eli L, Paquet AC, Cook JW, Rivera A, Frankson K, Bose J, Chen L, Cheung J, Shi Y, Irwin S, Kiss L, Huang W, Utter S, Sherwood T, Bates M, Weidler J, Parry G, Winslow J, Petropoulos CJ, Whitcomb J (2010) Analytical validation of a highly sensitive, accurate, and reproducible assay (HERmarkTM) for the measurement of HER2 total protein and HER2 homodimers in FFPE breast cancer tumor specimens. Pathol Res Int 2010: article ID 814176
Shi Y, Huang W, Tan Y et al (2009) A novel proximity assay for the detection of proteins and protein complexes: quantitation of HER1 and HER2 total protein expression and homodimerization in formalin-fixed, paraffin-embedded cell lines and breast cancer tissue. Diagn Mol Pathol 18:11–21
Bates M, Sperinde J, Kostler WJ et al (2011) Identification of a subpopulation of metastatic breast cancer patients with very high HER2 expression levels and possible resistance to trastuzumab. Ann Oncol 22:2014–2020
Sperinde J, Jin X, Banerjee J et al (2010) Quantitation of p95HER2 in paraffin sections by using a p95-specific antibody and correlation with outcome in a cohort of trastuzumab-treated breast cancer patients. Clin Cancer Res 16:4226–4235
Lipton A, Kostler WJ, Leitzel K et al (2010) Quantitative HER2 protein levels predict outcome in fluorescence in situ hybridization-positive patients with metastatic breast cancer treated with trastuzumab. Cancer 116:5168–5178
Therneau T, Atkinson B (2011) rpart: recursive partitioning. R package version 3.1-50. http://CRAN.R-project.org/package=rpart
Joensuu H, Sperinde J, Leinonen M et al (2011) Very high quantitative tumor HER2 content and outcome in early breast cancer. Ann Oncol 22:2007–2013
Acknowledgments
The authors would like to thank Sarah Irwin and the members of the Oncology Clinical Reference Laboratory at Monogram Biosciences, and Shannon Utter and the members of the Quality Department at Monogram Biosciences for their assistance in generating the VeraTag assay data. Most especially, we would like to thank the patients who selflessly agreed to participate in this study. This work was supported by Komen Grant BCTR 0707722 to AL, Mayor of the City of Vienna (MWFB 47-07) and Monogram Biosciences.
Conflict of interest
JC, JS, MH, WH, JMW, AN, AP, YT, AC, JL, YL, CP, and JW are employees of Monogram Biosciences, Inc., a subsidiary of Laboratory Corporation of America (LabCorp) and may from time to time hold stock in LabCorp. AL has received research support and has been an advisory board member of Monogram Biosciences. CFS has received research support and honoraria from Roche and Glaxo-SmithKline. WJK, EF, and CFS received research support from Monogram Biosciences. All remaining authors have declared no conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Diagram of samples with H2T, H3T and p95 data. Of the 98 patients with complete central FISH and H2T results used to generate the H2T clinical cutoff of 13.8 (ref #27), 93 had tissue available for the p95 assay (ref #25) and 89 had tissue available for the H3T assay. Three of the 89 cases were not in the set of 93 and therefore did not have p95 data available. Four of the 89 cases initially identified for H3T testing had insufficient tumor to generate an H3T result. Supplementary material 1 (TIFF 134 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Lipton, A., Goodman, L., Leitzel, K. et al. HER3, p95HER2, and HER2 protein expression levels define multiple subtypes of HER2-positive metastatic breast cancer. Breast Cancer Res Treat 141, 43–53 (2013). https://doi.org/10.1007/s10549-013-2665-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-013-2665-0