Abstract
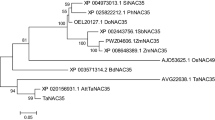
The gene, designated TaMADS2, was obtained from wheat leaves infected with the wheat stripe rust fungus by in silico cloning and RT-PCR. TaMADS2 encodes a predicted 159-amino-acid polypeptide that contains a highly conserved MADS domain. Phylogenetic analysis revealed that TaMADS2 is a type I MADS-box gene. The TaMADS2 transcript was detected in wheat leaves, stems, and roots. The expression of TaMADS2 was substantially down-regulated in the compatible interaction between wheat and Puccinia striiformis f. sp. tritici (Pst) at 36 and 48 h post-inoculation (hpi), whereas in the incompatible interaction the down-regulation was only observed at 48 hpi. Exogenous salicylic acid (SA) and abscisic acid (ABA) greatly induced the expression of TaMADS2 at 12 h post treatment (hpt), whereas methyl jasmonate (MeJA) down-regulated TaMADS2 at 6 hpt by approximately two-fold.
Similar content being viewed by others
Abbreviations
- hpi:
-
hour post inoculation
- hpt:
-
hour post treatment
- HR:
-
hypersensitive response
- ORF:
-
open reading frame
- qRT-PCR:
-
quantitative reverse transcription polymerase chain reaction
References
Alvarez-Buylla, E.R., Liljegren, S.J., Pelaz, S., Gold, S.E., Burgeff, C., Ditta, G.S., Vergara-Silva, F., Yanofsky, M.F.: MADS-box gene evolution beyond flowers: expression in pollen, endosperm, guard cells, roots and trichomes. — Plant J. 24: 457–466, 2000a.
Alvarez-Buylla, E.R., Pelaz, S., Liljegren, S.J., Gold, S.E., Burgeff, C., Ditta, G.S., Ribas de Pouplana, L., Martínez-Castilla, L., Yanofsky, M.F.: An ancestral MADS-box gene duplication occurred before the divergence of plants and animals. — Proc. nat. Acad. Sci. USA 97: 5328–5333, 2000b.
Ando, S., Sato, Y., Kamachi, S., Sakai, S.: Isolation of a MADS-box gene (ERAF17) and correlation of its expression with the induction of formation of female flowers by ethylene in cucumber plants (Cucumis sativus L.). — Planta 213: 943–952, 2001.
Bonhomme, F., Kurz, B., Melzer, S., Bernier, G., Jacqmard, A.: Cytokinin and gibberellin activate SaMADSA, a gene apparently involved in regulation of the floral transition in Sinapis alba. — Plant J. 24: 103–111, 2000.
Chen, X., Moore, M., Milus, E.A., Long, D.L., Line, R.F., Marshall, D., Jackson, L.: Wheat stripe rust epidemics and races of Puccinia striiformis f. sp. tritici in the United States in 2000. — Plant Dis. 86: 39–46, 2002.
Crampton, B.G., Hein I., Berger, D.K.: Salicylic acid confers resistance to a biotrophic rust pathogen, Puccina substriata, in pearl millet (Pennisetum glaucum). — Mol. Plant Pathol. 10: 291–304, 2009.
De Bodt, S., Raes, J., Florquin, K., Rombauts, S., Rouzé, P., Theissen, G., Van de Peer, Y.: Genome wide structural annotation and evolutionary analysis of the type I MADSbox genes in plants. — J. mol. Evol. 56: 573–586, 2003.
Duun, R.M., Hedden, P., Bailey, J.P.: A physiologicallyinduced resistance of Phaseolus vulgaris to a compatible race of Colletotrichum lindemuthianum is associated with increases in ABA content. — Physiol. mol. Plant Pathol. 36: 339–349, 1990.
Feys, B.J., Parker, J.E.: Interplay of signaling pathways in plant disease resistance. — Trends Genet. 16: 449–455, 2000.
Jack, T.: Plant development going MADS. — Plant mol. Biol. 46: 515–520, 2001.
Jones, J.D., Robert-Seilaniantz, A., Navarro, L., Bari, R.: Pathological hormone imbalances. — Curr. Opin. Plant Biol. 10: 372–379, 2007.
Kang, Z.S., Huang, L.L., Buchenauer H.: Ultrastructural changes and localization of lignin and callose in compatible and incompatible interactions between wheat and Puccinia striiformis. J. Plant Dis. Protect. 109: 25–37, 2002.
Kang, Z.S., Li, Z.Q., Chong, J., Rohringer, R.: [Ultrastructure and cytochemistry of haustorium of wheat stripe rust.] — Acta mycol. sin. 13: 52–57, 1994. [In Chin.]
Kunkel, B.N., Brooks, D.M.: Cross talk between signaling pathways in pathogen defense. — Curr. Opin. Plant Biol. 5: 325–331, 2002.
Lamb, C.J., Lawton, M.A., Dron, M., Dixon, R.A.: Signal and transduction mechanisms for activation of plant defenses against microbial attack. — Cell 56: 215–224, 1989.
Lee, S., Woo, Y.M., Ryu, S.I., Shin, Y.D., Kim, W.T., Park, K.Y., Lee, I.J., An, G.: Further characterization of a rice AGL12 group MADS-box gene, OsMADS26. — Plant Physiol. 147: 156–--, 2008.
Livak, K.J., Schmittgen, T.D.: Analysis of relative gene expression data using real-time quantitative PCR and the 2-[Delta][Delta] CT method. — Methods 25: 402–408, 2001.
Lorenzo, O., Piqueras, R., Sanchez-Serrano, J.J., Solano, R.: ETHYLENE RESPONSE FACTOR1 integrates signals from ethylene and jasmonate pathways in plant defense. — Plant Cell 15: 165–178, 2003.
Mao, L., Begum, D., Chuang, H., Budiman, M.A., Szymkowiak, E.J., Irish, E.E., Wing, R.A.: JOINTLESS is a MADS-box gene controlling tomato flower abscission zone development. — Nature 406: 910–912, 2000.
Mauch-Mani, B., Mauch, F.: The role of abscisic acid in plantpathogen interactions. — Curr. Opin. Plant Biol. 8: 409–414, 2005.
Münster, T., Pahnke, J., Di Rosa, A., Kim, J.T., Martin, W., Saedler, H., Theissen, G.: Floral homeotic genes were recruited from homologous MADS-box genes preexisting in the common ancestor of ferns and seed plants. — Proc. nat. Acad. Sci. USA 94: 2415–2420, 1997.
Nam, J., Kim, J., Lee, S., An, G., Ma, H., Nei, M.: Type I MADS-box genes have experienced faster birth-and-death evolution than type II MADS-box genes in angiosperms. — Proc. nat. Acad. Sci. USA 101: 1910–1915, 2004.
Ng, M., Yanofsky, M.F.: Function and evolution of the plant MADS-box gene family. — Nat. Rev. Genet. 2: 186–195, 2001.
Parenicova, L., De Folter, S., Kieffer, M., Horner, D.S., Favalli, C., Busscher, J., Cook, H.E., Ingram, R.M., Kater, M.M., Davies, B.: Molecular and phylogenetic analyses of the complete MADS-box transcription factor family in Arabidopsis: new openings to the MADS world. — Plant Cell 15: 1538–1541, 2003.
Portereiko, M.F., Lloyd, A., Steffen, J.G., Punwani, J.A., Otsuga, D., Drews, G.N.: AGL80 is required for central cell and endosperm development in Arabidopsis. — Plant Cell, 18: 1862–1872, 2006.
Robert-Seilaniantz, A., Navarro, L., Bari, R., Jones, J.D.G.: Pathological hormone imbalances. — Curr. Opin. Plant Biol. 10: 372–379, 2007.
Theissen, G.: Development of floral organ identity: stories from the MADS house. — Curr. Opin. Plant Biol. 4: 75–85, 2001.
Theissen, G., Saedler, H.: Plant biology: floral quartets. — Nature 409: 469–471, 2001.
Theissen, G., Becker, A., Di Rosa, A., Kanno, A., Kim, J.T., Münster, T., Winter, K.U., Saedler, H.: A short history of MADS-box genes in plants. — Plant mol. Biol. 42:115–149, 2000.
Ton, J., Flors, V., Mauch-Mani, B.: The multifaceted role of ABA in disease resistance. — Trends Plant Sci. 14: 310–317, 2009.
Wang, C.F., Huang, L.L., Buchenauer, H., Han, Q.M., Zhang, H.C., Kang, Z.S.: Histochemical studies on the accumulation of reactive oxygen species (O2 - and H2O2) in the incompatible and compatible interaction of wheat-Puccinia striiformis f. sp. tritici. — Physiol. mol. Plant Pathol. 71: 230–239, 2007.
Wang, Y.F., Qu, Z.P., Zhang, Y.H., Ma, J.B., Guo, J., Han, Q.M., Huang, L., Kang, Z.S.: [Construction of a cDNA library and analysis of expressed sequence tags in association with the incompatible interaction between wheat and Puccinia striiformis.] — Scientia agr. sin. 41: 3376–3381, 2008. [In Chin.]
Xu, H., Li, X., Li, Q., Bai, S., Lu, W., Zhang, X.: Characterization of HoMADS1 and its induction by plant hormones during in vitro ovule development in Hyacinthus orientalis L. — Plant mol. Biol. 55: 209–220, 2004.
Zeng, S.H., Xu, Y.Q., Wang, Y.: Isolation and characterization of two MADS-box genes from Lycium barbarum. — Biol. Plant. 55: 567–571, 2011.
Zhang, G., Li, Y.M., Sun, Y.F., Wang, J.M., Liu, B., Zhao, J., Guo, J., Huang, L.L., Chen, X.M., Kang, Z.S.: Molecular characterization of a gene induced during wheat hypersensitive reaction to stripe rust. — Biol. Plant. 55: 696–702, 2011.
Zhao, T., Ni, Z., Dai, Y., Yao, Y., Nie, X., Sun, Q.: Characterization and expression of 42 MADS-box genes in wheat (Triticum aestivum L.). — Mol. Genet. Genom. 276: 334–350, 2006.
Zhu, C., Perry, S.E.: Control of expression and autoregulation of AGL15, a member of the MADS-box family. — Plant J. 41: 583–594, 2005.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Acknowledgments: We thank Prof. L. Dunkle from the USDA-Agricultural Research Service at Purdue University for critically reading the manuscript. This work was supported by National Basic Research Program of China (No. 2013CB127700), National Natural Science Foundation of China (Nos. 30930064 and 31171795), the Fundamental Research Funds for the Central Universities of China (QN2009035) and the 111 Project from Ministry of Education of China (No. B07049).
Rights and permissions
About this article
Cite this article
Guo, J., Shi, X.X., Zhang, J.S. et al. A type I MADS-box gene is differentially expressed in wheat in response to infection by the stripe rust fungus. Biol Plant 57, 540–546 (2013). https://doi.org/10.1007/s10535-012-0297-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10535-012-0297-6