Abstract
The transformation and turnover time of medium- to long-chain dicarboxylic acids (DCA) in soil is regulated by microbial uptake and mineralization. However, the chain length of n-alkyl lipids may have a remarkable influence on its microbial utilization and mineralization and therefore on the formation of stable soil organic carbon from e.g. leave- needle- and root-derived organic matter during decomposition. To investigate their size dependent mineralization and microbial incorporation, four DCA of different chain lengths (12–30 carbon atoms), that were 13C labeled at each of their terminal carboxylic groups, were applied to the Ah horizon of a Fluvic Gleysol. Incorporation of 13C into CO2 and in distinct microbial groups classified by phospholipid fatty acid (PLFA) analysis was investigated. Mineralization of DCA and incorporation into PLFA decreased with increasing chain length, and the mineralization rate was highest during the first days of incubation. Half-life time of DCA carbon in soil increased from 7.6 days for C12 DCA to 86.6 days for C18 DCA and decreased again to 46.2 days for C22 DCA, whereas C30 DCA had the longest half-life time. Rapid and efficient uptake of C12 DCA as an intact molecule was observable. Gram-negative bacteria incorporated higher amounts of DCA-derived 13C compared to other microbial groups, especially compared to actinomycetes and fungi during the first phase of incubation. However, the incorporation of C12 DCA derived 13C into the PLFA of actinomycetes, and fungi increased steadily during the entire incubation time, suggesting that those groups take up the 13C label from necromass of bacteria that used the C12 DCA for formation of their lipids before.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
An increasing number of studies emphasizes the significance of root-derived organic matter inputs as precursor material for soil organic carbon (Altmann et al. 2021a). One distinct class of compounds present in root-litter are long-chain dicarboxylic acids (DCA). DCA are known to be a major moiety (up to 50%) of suberin (van Bergen et al. 1998; Watanabe et al. 2013), a polyester which accumulates at the hypodermal/exodermal cell layers of the roots and that can make up to 62 µg cm− 1 of a mature root, whereas lignin contributes in some cases only less than 50 µg cm− 1, depending on the plant species (Watanabe et al. 2013). Despite their quantitative importance in suberin, the fate of medium- to long-chain DCA in soil has received little attention, whereas the degradation of lignin and lignin monomers, as well as their contribution to SOM has been studied extensively (e.g. Zimmermann 1990; Bahri et al. 2006; Datta et al. 2017). Moreover, suberin-derived DCA have been used for decades as root-specific biomarkers to identify and quantify root inputs to sediments and soils (e.g. Nierop 1998; Rasse et al. 2005; Mendez-Millan et al. 2011; Altmann et al. 2021a). However, their use as root-biomarkers in soil requires appropriate knowledge about the fate of individual indicator molecules during degradation, which is still not well understood (Feng et al. 2010; Altmann et al. 2021a, b) determined the change of concentrations of root- and leaf-derived ester-bound lipids within one year of leave, needle and root degradation and found no linear trend, however, they focused solely on the depolymerization of the polymers cutin and suberin but did not study the degradation of individual monomers. The further cycling and turnover of DCA in soil, once the ester-bounds are cleaved is still not fully understood, which would be essential to their utilization as biomarkers for root-derived organic matter input and turnover.
The cleavage of ester-bonds and therefore release of cutin and suberin derived DCA by microbial cutinases and suberinases during litter decomposition is one possible source of DCA in the detritus sphere (Kolattukudy 1980). The composition of lipid monomers (e.g. ω-hydroxy fatty acids, dihydroxy fatty acids and DCA) released by cutinase CcCut1 showed a similar pattern to the monomers released by alkaline hydrolysis (Järvinen et al. 2009; Kontkanen et al. 2009). However, secondary formation via enzymatic oxidation of precursor compounds could also be a major, yet underestimated, source of DCA in soil (Ambles et al. 1994; Coon 2005; Rojo 2009). Microbial residues consist of long-chain n-alkanes, n-alkanols, and n-alkanoic acids. In the case of n-alkanes aerobic degradation usually starts with the oxidation of a terminal methyl group to render primary alcohols, which are further oxidized to the corresponding aldehydes, and finally converted into fatty acids (Rojo 2009). Fatty acids are conjugated to coenzyme A (CoA) and further processed by β-oxidation to generate acetyl-CoA (Watkinson and Morgan 1990; Ashraf et al. 1994; van Hamme et al. 2003; Wentzel et al. 2007). Both ends of the alkane molecule may be oxidized through ω-hydroxylation of fatty acids at the terminal methyl group, rendering an ω-hydroxy fatty acid. The latter is further converted into DCA which can subsequently be processed by β-oxidation (Watkinson and Morgan 1990; Coon 2005). This synthesis of medium- to long-chain DCA and cutin- or suberin-derived monomers by microorganisms has been described for pure cultures of some yeasts (e.g. Candida tropicalis or Yarrowia lipolytica), filamentous fungi and some bacteria present in soils and litter layers (Mauersberger et al. 1992; Eschenfeldt et al. 2003; van den Brink et al. 1998; Scott 2008; Van Bogaert et al. 2011). In the course of three enzymatic reactions, fatty acids are oxidized to DCA (Mauersberger et al. 1992; Eschenfeldt et al. 2003).
Additionally, microorganisms also possess the biochemical degradation of DCA in soil via β-oxidation, a process similarly demonstrated in pure cultures for preventing e.g. the accumulation of DCA in naturally occurring yeasts (Mauersberger et al. 1992; Eschenfeldt et al. 2003). The degradation of DCA occurred slower for long-chain DCA (C18, C20, C22), which showed a higher resistance to microbial degradation than those with short and moderate chain lengths (C2 − 12) (Olivella et al. 2015; Saxena and Hildemann 1996). Degradation time may be expressed as half-life, defining the time required to degrade half the amount of the specific compound, which was applied and released as CO2 (Jones and Shannon 1999; Boddy et al. 2007; Pearson and Eglinton 2000) estimated mean terrigenous residence times for higher plant wax (C24 − 33) derived long-chain n-alkanes and DCA from sediments to be less than 100 years. Average terrestrial residence times of vascular plant biomarkers deduced from n-C28 − 30 alkanes and from stations directly in front of the river mouths range from 900 to 4400 years (Kusch et al. 2010).
Bacteria (According to PLFA analysis, especially gram-negative bacteria) were highly competitive for the uptake of some organic acids over other microbial groups (Gunina et al. 2014; Garcia-Pausas and Paterson 2011; Waldrop and Firestone 2004). The contribution of fungi to carboxylic acid-C utilization was less than that of bacteria, due to their lower competitiveness for water soluble substances (Gunina et al. 2014; Waldrop and Firestone 2004). To unravel the fate and transformation/incorporation of DCA-C by specific microbial groups, we established an incubation experiment using position-specifically labeled D13CA. Position-specific labeling is a tool that has recently reached an increasing consideration in soil science (Fischer and Kuzyakov 2010; Dijkstra et al. 2011; Apostel et al. 2013; Dippold and Kuzyakov 2013). It overcomes the limitations of uniform labeling because it allows differentiation between the incorporation of molecule fragments vs. the incorporation of entire intact molecules (Apostel et al. 2015). PLFA analysis allows the reconstruction of the microbial community composition (Zelles et al. 1995; Zelles 1999), but in combination with 13C labeling- i.e. 13C-PLFA analysis – it enables the tracing of substrate incorporation into microbial cell membranes and therefore the reconstruction of individual microbial groups (Glaser 2005). We selected four carboxylic group enriched fatty diacids as monomers, which we expected to be quite abundant in soils since they have either been derived from microbial synthesis or suberin decomposition (Banfield et al. 2018). The incorporation of the terminal enrichment into hydrolysable lipids like PLFA will reveal that at one of the two carboxylic groups is still present in this lipid, most likely involved in the ester bond (Deamer 2017).
We hypothesized that (1) DCA half-life time increases with chain-length of DCA due to decreased microbial accessibility, (2) bacterial groups are most competitive in the uptake of the DCA with lower chain length, but this uptake competitive advantage decreases with increasing DCA chain length, and (3) with increasing turnover time of DCA, the incorporation of DCA-C into actinomycetes would increase as they are known as a highly competitive group for microbial necromass utilization.
Materials and methods
Sampling site
Soil samples were collected from the Ah horizon of a Fluvic Gleysol located in the Middle-Elbe flood plain used as grassland at the end of October 2016 from 0–10 cm depth. The geographical location of the site was 12°75’ northern latitude; 51°83’ eastern longitude and 107 m above sea level, respectively (Gallin, Germany). The average annual temperature has been recorded as 8 °C (Weather station Zarrentin am Schaalsee). The collected soil samples were sieved to < 2 mm, and stored for one day at + 5 °C, except for subsamples used to determine the soil water content (dried at 105 °C following standard procedures). Physico-chemical properties of the soil sample are given in Table 1.
Experimental setup
Four position specifically 13 C labeled isotopomers of DCA with different chain lengths were used as tracer compounds, and two non-labeled DCA served as natural background substances. A total of 72 microcosms (6 DCA treatments × 4 replicates × 3 sampling dates) were prepared. The DCA applied to the microcosms were dodecane-dioic acid (13C12H22O4), octadecane-dioic acid (13C2C16H22O4), docosane-dioic acid (13C2C20H22O4) and triacontane-dioic acid (13C2C28H22O4). The rationale behind our approach to use DCAs with similar chemical structure (homologous series) but increasing chain length is that the mean nominal oxidation state of the C atoms (average NOSC) in the respective molecule decreases with increasing chain length (Chakrawal et al. 2020). Hence, the average NOSC of the applied DCAs decreases in the order C12 DCA, C18 DCA, C22 DCA to C30 DCA. Molecules containing carbon atoms that are more reduced on average (i.e. have a lower average NOSC) tend to have higher oxidation activation energies (i.e. higher Gibbs energies for the oxidation half reaction). As enzymes catalyze reactions by decreasing the energy of activation, we assume that microorganisms incur higher metabolic costs when decomposing long-chain molecules from a homologous series compared to their shorter-chain representatives (Gunina et al. 2017; Chakrawal et al. 2020).
To produce the applied DCA, a novel synthetic protocol based on chain elongation of various precursors followed by a terminal 13C introduction using K13CN and subsequent hydrolysis to produce the respective DCA was designed (Schink et al. 2020).
Labeled DCA were weighed according to their molar mass to achieve an application rate of 100 µmol fatty acids to 1 kg soil and were mixed with 100 mg silt-sized quartz (Quarzwerke GmbH, Frechen, Germany).
Soil incubation was done in 50 ml microcosms. For this purpose, 20 g of dry soil with a mean bulk density of 1.5 gcm− 3 was weighed and transferred to microcosms in four different layers (6 g, 6 g, 5 g, and 3 g). These four soil layers were separated by three layers of DCA compounds. The final acid layer was transferred with two times 0.5 ml ultrapure water, to achieve a weight of 20.1 g±0.2 g. Soil water content was kept at 45% water holding capacity (WHC). The microcosms were incubated for 14 days at 8 °C, which is the mean annual temperature of the sampling site.
Additionally, microcosms in 100 ml crimped bottles were installed for 13CO2 incubation. They received similarly the four layers of soil (totally 20 g), the interlayer of sand and DCA, and were adjusted to a soil water content of 45% (WHC).
13CO2 sampling took place after 1, 2, 3, 4, 6, 8, 10, 15, 20, 25, 30, and 35 days. A gas chromatography syringe was used to transfer trapped CO2 to 15 ml pre-evacuated exetainers at the same time for all four replicates. Thereafter, syn air flushing was done for 45 s on crimped bottles, and then they were transferred to the fridge to be kept at 10 °C until the next sampling time. The destructive soil sampling was done after 3, 10, and 40 days and soils were frozen at −20 °C until PLFA analysis and freeze-dried just before extraction.
Quantification of soil mineralization
The CO2 of the gas samples was quantified using GC-FID-ECD (Thermo Fisher Scientific, Bremen, Germany). In addition to the samples, four test gases with a known CO2 concentration were measured as calibration gases. These external standards contained 400, 1500, 3000, and 4000 ppm CO2. The starting temperature of the measurements was 50 °C. This was held for five minutes. The temperature was then increased to 130 °C at a rate of 100 °C per minute. The final temperature was held for two minutes for column reconditioning. Following the measurements, the CO2 peaks of each chromatogram were integrated, and the areas determined using Agilent Chem-Station 10 (Agilent, Waldbronn, Germany). To quantify the CO2 of the gas samples, a quadratic calibration curve was created using the four external standards of increasing CO2 concentration. Finally, the amount of CO2 contained in the microcosms was calculated using the quantified CO2 production and the general gas law (Schink 2019).
Determination of 13C enrichment in respired CO2
The determination of the respired δ13C values was carried out at the center for stable isotope analysis (KOSI) at Georg-August University Göttingen. Briefly, 5.5 ml of the sampled gas was injected into a GC-C-IRMS equipped with an autosampler unit (AS 200) (Thermo Fisher Scientific, Bremen, Germany). We identified the peak areas and integrated these via Isodat 2.0, and referenced the measurements by a reference gas, which was injected before each sample application.
Calculation of 13C incorporation into CO2
13C in CO2 derived from the applied DCA was calculated according to a linear mixing model (Eqs. 1 and 2) (Gearing et al. 1991) where the CO2 amount of the background sample in Eq. 1 was determined by Eq. 2.
where \({\left[{C}_{{CO}_{2}}\right]}_{LAB; BG; appDCA}\) is the amount of C released in CO2 of labelled samples, background samples, or derived from applied DCA, respectively (all in µg C g− 1 soil). \({r}_{{CO}_{2}-LAB; {CO}_{2} BG; appDCA}\) is the at% 13C enrichment of the CO2-eflux from labelled microcosms, background microcosms or derived from the applied DCA, respectively.
Double exponential kinetics was applied to calculate the corresponding estimation or 13CO2 efflux rates from individual pools at time t (Eq. 3) (Kuzyakov 2011).
where 13CO2 (t) is the percent of 13C incorporation into CO2 at any time t; 13Cmax_a/b is the maximal utilization of the fast and slowly mineralizable DCA pool, respectively; and ka and kb are the fast and slow decomposition rate, with
where \({\lambda }_{a/b}\) is the mineralization half-time of the fast and the slowly mineralizable DCA pool (Gunina et al. 2017). The cumulative 13CO2 efflux with time results as integral of Eq. 3.
PLFA extraction and preparation
PLFA extraction and purification was done according to Frostegard et al. (1991), modified by Gunina et al. (2014). Before extraction, 1 µg per µl solution of the first internal standard (phosphatidylcholine-dinonadecanoic acid-IS1) was added to 6 g of each soil sample and extracted with a Bligh-and-Dyer extraction solution buffered by a citric acid buffer. After liquid-liquid and solid-phase purification steps, the fatty acids were saponified to free fatty acids and derivatized into fatty acid methyl esters (FAME) following the description of Knapp (1979). 15 µL of the second internal standard (tridecanoate methyl ester - IS2, 1 µg per µl) and 185 µL of toluene were added to the extracted FAME and transferred to 200 µl inserts in 1.5 ml vials for gas chromatography (GC) analysis. In parallel to the samples, 5 external standards with 28 fatty acids were hydrolyzed, derivatized, and measured on both GC together with the samples.
PLFA identification
FAME-contents were identified and quantified via gas chromatograph-mass spectrometry (GC–MS) (GC 5890 with MS 5971 A, Agilent, Waldbronn, Germany) with a 30 m DB1-MS column, in the selected ion mode. FAME-contents were calculated by relating their areas to the area of IS2 followed by calibration via linear regression of the external standards. Extracted PLFA-contents were corrected based on the recovery of IS1. Agilent’s MassHunter Software was used for integration.
Determination of 13 C values of PLFA
Measuring the FAME δ13C analyzed by GC-C-IRMS consisting of an autosampler unit (AS 200) and combustion unit (Combustion Interface III) interfaced to a Delta plus isotope ratio mass spectrometer (all units from Thermo Fisher Scientific, Bremen, Germany). One µl sample were injected with 1 min splitless time into a liner heated to 280 °C. Peak separation was accomplished with two capillary columns (30 m DB-5MS followed by a 15 m DB-1MS, Agilent Waldbronn Germany) with helium (99.9996% pure) as carrier gas at a flow rate of 1.7 mL/min. Isodat 2.0 software was used for the calculation of δ13C values (‰) after the integration of FAME peaks. CO2 reference gas with 99.995% purity was injected four times per chromatogram for a first calibration and drift correction.
Calculation of tracer 13C incorporation
The isotopic mixing model suggested by Gearing et al. (1991) was used to calculate the amount of 13C incorporation into PLFA (in % of applied tracer) (Eqs. 5 and 6).
where \({\left[{C}_{PLFA}\right]}_{LAB; BG; appDCA }\)is the amount of C in PLFA of the labelled sample, or in the background sample or derived from the applied DCA (µg C per g− 1 soil), respectively. rPLFA_LAB; PLFA_BG; appDCA is the at% enrichment of the PLFA of the labelled microcosm or the background microcosm or derived from the applied DCA, respectively.
These equations quantify the incorporation of 13C DCA into the respective fatty acids. Then the gram of 13C incorporation has been divided by the amount of applied 13C and multiplied with 100% to get the relative incorporation to the applied 13C, also called tracer 13C recovery in the respective fatty acid. This calculation is critical for the comparison between the individual DCA as it corrects the minimal differences in 13C application of the individual tracer amendments. The incorporation is a quantitative measure of the total 13C allocated into the pool of this membrane lipid classes.
We determined the relative incorporation into PLFA (F-value), reflecting a rather ecological parameter that describes the relative incorporation of the fatty acids of a distinct microbial group irrespective of its abundance, describing the replacement of 12C by 13C (12C/13Crepl) (Gunina et al. 2014) (Eq. 7).
where \({({}_{X}{}^{12}C/{}_{X}{}^{13}C)}_{repl}\) is the % of replacement of the unlabeled PLFA-pool by DCA-derived 13C-PLFA (µg 13C per soil); and \({\left[{C}_{PLFA}\right]}_{total}\) is the total amount of PLFA in the respective PLFA pool (µg C per soil).
Statistical analysis
A Nalimov outlier test at a 95% significance level was performed for mineralization and incorporation values (Lozán and Kausch 1998). The number of replicates has been considered before performing the outlier test. To disentangle the fate of the position-specific labeled D13CA the respired 13C–CO2, and incorporation into PLFA was investigated at different time points. The significance of differences was tested by repeated-measures analysis of variance (ANOVA) (Supplementary Table S1, S2). Tukey’s post-hoc test was done after considering the assumptions of the ANOVA test (homogeneity and normal distribution) at 95% confidence level. The microbial grouping was conducted using factor analysis of PLFA contents. Microbial grouping was achieved if PLFA loaded on the same factor higher than 0.5 and if literature data on pure cultures revealed a common origin of the PLFA loading on the same factor (Apostel et al. 2013; Zelles 1999; Zelles et al. 1995). Statistica 12.0 software was used for all this statistical analysis.
Results
13 CO 2 release from DCA mineralization
The mineralization of the terminal carboxylic groups of the DCA to CO2 decreased with increasing chain-length of DCA for both, the cumulative (Fig. 1, top) and the rate of 13CO2 (Fig. 1, bottom). The total 13CO2 released during the incubation was considerably lower in C18 DCA compared to the C12 DCA (30-fold) and then further decreased for longer-chained DCA.
Mineralization of DCA to 13CO2 was observed immediately after the application of compounds, and the rate of 13CO2 released peaked within the first 24 h. The initial (maximal) respiration rate yielded 8, 0.12, 0.009 and 0.006% of applied tracer per day for C12, C18, C22, and C30 DCA, respectively. The DCA mineralization rate to 13CO2 decreased in general rapidly after this initial peak, more intensively for C12 DCA (320 times) compared to the other long-chain DCA (in average 9 times) (Fig. 1, bottom). In consequence, most of the 13CO2 was respired during a short initial period.
Half-life time of DCA varied strongly between the two mineralization phases. During the fast mineralization, half-life time (\({\lambda }_{a}\)) of DCA ranged between 1.8 and 2.3 d for C12 and C22 DCA, respectively, indicating no clear chain length effect for the first phase of DCA mineralization. The half-life time during the slow (second) mineralization phase (\({\lambda }_{b}\)) was noticeably longer compared to that of the fast (first) one (in average 239-fold). The half-life time of the second mineralization time varied widely between 7.62 and 1732.87 d for C12 and C30 DCA, respectively, and at least showed from shortest to longest DCA a pronounced effect of the chain-length (Table 2).
Incorporation of 13C labeled DCA into PLFA
The incorporation of 13C into the sum of all PLFA was highest for C12 DCA (p < 0.05) and had its maximum at the first harvesting time point (0.14% of applied tracer), thereafter decreasing significantly with time (Fig. 2). Overall, the highest 13C incorporation into PLFA was found in shorter-chain DCA compounds and always within the first few days of incubation. However, with increasing chain length the effect of time on the 13C incorporation disappeared. C18 DCA was incorporated 12-fold, C22 45-fold and C30 32-fold lower than C12 DCA into the sum of all PLFA, respectively (Fig. 2).
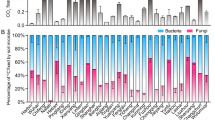
By combining the PCA based grouping of the fatty acids results with fatty acid fingerprints of pure-cultures we classified six microbial groups consisting of four groups of bacteria, including gram-positive 1 (G + 1), gram-positive 2 (G + 2), gram-negative 1 (G-1), and gram-negative 2 (G-2); one group of actinobacteria (Act), and one group of arbuscular mycorrhizal fungi (AMF) (Supplementary Table S3).
13C incorporation into PLFA for gram-negative bacteria was on average 6.9, 8.46 and 2.75 times higher than that of the gram-positive 1, gram positive 2 and gram negative 2 groups, respectively (Fig. 3). Overall, the incorporation of 13C12 into PLFA of all bacterial groups was 21 times higher than 13C18. Moreover, we determined a significant decrease of ~ 170% of 13C12 DCA-derived C incorporation into PLFA of all bacterial groups from day 3 to day 40 (Fig. 3). The decrease in incorporation with increasing incubation time was also visible for most higher chain length but in many cases not significant.
13C incorporation into the filamentous microorganisms changed only for 13C12 DCA significantly over time, but, in contrast to the incorporation into single cell prokaryotes, increased over the duration of the experiment. Uptake of 13C12 DCA into PLFA doubled for actinobacteria and arbuscular mycorrhizal fungi between day 10 and 40 (Fig. 4). Actinobacteria are the only group, where C12 and C18-derived 13C was incorporated in similar amounts however with contrasting temporal pattern: whereas C12-derived 13C incorporation increased with time, that of C22 DCA decreased with time resembling more the temporal pattern observed for free living prokaryotes. Filamentous microbial groups seemed to avoid the use of C22 DCA, which was lower than that of C30 DCA. Relative incorporation of 13C from DCA into PLFA (the isotopic F-value = fraction of tracer-derived 13C from total C of a pool) showed similar patterns for all microbial groups (supplementary Figure S1). Solely the filamentous actinobacteria did not show a decrease of relative incorporation of 13C from DCA into PLFA from the short-chain C12 DCA to the longer chain ones.
13C incorporation into PLFA (in % of applied tracer) (mean ± SEM) of actinobacteria (Act) and fungi/arbuscular mycorrhizal fungi microbial groups; data from position-specific labeled DCA at different time points. Lower-case letters above error bars indicate significant differences between the treatments and time of individual microbial groups at a level of p < 0.05, n = 4
Discussion
Mineralization and total utilization
Most of the DCA mineralization was observed during the first days of the experiment leading to remarkably higher decomposition constants for the rapid phase of DCA mineralization. This suggests that the mineralization controlling factors are very different between the first and the second phase of decomposition. One often described phenomenon when adding substances to soils is the fact that mechanisms reducing microbial access to a potential C source are temporally slower than microbial uptake and utilization (Fischer and Kuzyakov 2010; Dippold and Kuzyakov 2013). This implies that during the initial phase mainly microbial uptake and metabolic potential to mineralize the DCA might have been the controlling factor for their decomposition, whereas their reduced access by e.g. hydrophobic interaction with hydrophobic parts of the soil organic matter might have added up on that increasingly with increasing incubation time leading to the reduced decomposition kinetics as observed in the second phase of the experiment (Kuzyakov 2011).
Mineralization of DCA displays a clear chain length effect, with cumulative mineralization over 40 days and the long-term decomposition constant Kb decreasing remarkably with increasing chain length. Compounds with a high hydrophobicity and thus a low water solubility are less susceptible to decomposition in soils (Miao et al. 2017). It was discovered that water-insoluble material reduces microbial activity as well as the rate of decomposition, particularly in organic soils (Spaccini et al. 2002; Miao et al. 2017). This suggests that hydrophobicity of added substances not only affects their own mineralization but also the mineralization of the adjacent SOC. So, decreasing mineralization of the DCA compounds with increasing number of carbon atoms can be explained by the increasing hydrophobicity with increasing chain length. The average NOSC is likely the other important factor in mineralization of DCA. Increasing the chain length in DCA compounds leads to the longer chains of –(CH2)– bonded carbon atoms that decreases the average NOSC of the overall molecule and consequently decreases its mineralization (Gunina et al. 2017). Nevertheless, both factors would suggest a continued decrease/increase with chain length but instead the chain-length effect was highly pronounced comparing the very long chain C30 DCA with the short-chain C12 DCA but was not visible any more or not consistent when comparing the two mid-chain DCA C18 and C22. This suggests that besides mere biophysical (hydrophobicity) and chemical (oxidation state) parameters, also clear adaptive mechanisms of microbial decomposition metabolism to defined chain length seem to exist, which partially overlay the availability mechanisms.
Several mechanisms are involved in cellular uptake of hydrophobic substrates in different microorganisms, including substrate solubilization by extracellular emulsifying agents (Lee et al. 2018; Atshaves et al. 2010), cell surface alteration to promote adhesion (Beopoulos et al. 2009), passive diffusion through the cell membrane (Huf et al. 2011), and transporter-mediated uptake (Thevenieau et al. 2007). However, it appears that the degradation pathways and transporters for middle-chain and long-chain fatty acids are distinct (Schaffer 2002; Werner and Zibek 2017).
Long-chain fatty acids are thought to enter cells through a mechanism that includes adsorption to the membrane, protonation, membrane flip-flopping, and desorption into the cytosolic space (Hamilton and Kamp 1999). Long-chain fatty acid uptake through cellular membranes is linked to esterification by acyl-CoA synthetases, according to studies in both prokaryotic and eukaryotic organisms (Schaffer 2002). Long-chain fatty acids are activated to acyl-CoA and must be transported to the mitochondrial interior via the carnitine shuttle in eukaryotic cells. In contrast, the common feature of short-chain and middle-chain fatty acids is their carnitine-independent uptake and, in case of eukaryotes, intramitochondrial activation to acyl-CoA thioesters (Schönfeld and Wojtczak 2016). Furthermore, unlike long-chain fatty acids, short-chain and middle-chain fatty acids’ cellular metabolism is less dependent on fatty acid binding proteins (Schönfeld and Wojtczak 2016). Some evidences emerged that proteins are important mediators and/or regulators of trafficking of long-chain fatty acids into and within cells (Schaffer 2002; Atshaves et al. 2010). As a result, uptake of long-chain fatty acids needs more energy than middle-chain fatty acids (Schönfeld and Wojtczak 2016) – a potentially highly relevant factor explaining the reduced microbial utilization of the very long chain C30 DCA. We suppose that the lack of a chain-length effect in the range of our mid-chain DCA i.e. the observation that the increased hydrophobicity of C22 DCA compared to C18 DCA does not go along with a decreased decomposition, is linked to an alteration of chain-length specific uptake, intracellular activation or decomposition mechanisms not yet described in detail.
Utilization of DCA by the soil microbial community
Microorganisms, by producing microbial biomass and necromass have significant influence on SOM formation and cycling (Powlson et al. 2001; Koyama et al. 2018). Organic acids like DCA can be taken up by soil microorganisms whose residues in turn can be sorbed to soil particles or form biofilms on aggregate and particle surfaces, one key mechanism of SOM stabilization (Jones et al. 2003). The hydrophobicity of the cell surface is very important in the attachment to, or detachment from the surfaces (Wu et al. 2012). High cell surface hydrophobicity enables microorganisms to attach to hydrocarbon droplets on the surface or cells (Krasowska and Sigler 2014). Gram-negative bacteria increase cell surface hydrophobicity and their ability to form biofilms by releasing outer membrane vesicles (Baumgarten et al. 2012) which is most likely one of the key mechanisms explaining their high ability to use the DCA carbon for their metabolism. In contrast, Gram Positive bacteria’s outer membrane is comprised of a peptidoglycan layer (Schaffer 2002), which is rather hydrophilic and has no general adaptation improving access to hydrophobic substances.
Fungi/arbuscular mycorrhizal fungi, as the eukaryotic group, was unable to take up similar amounts of the applied DCA compared to the prokaryotic groups in our experiment. Fungi are considered to be more hydrophobic than bacteria due to their hydrophobic rodlet layer (Kwan et al. 2006) and effective production of hydrophobins (Linder et al. 2005; Bayry et al. 2012). Therefore, microorganisms with hydrophobic surface preferentially interact with hydrophobic compounds, and likewise for hydrophilic microorganisms and compounds (Katsikogianni and Missirlis 2004). Regardless of molecular properties of an added substrate, the predominant bacterial biomass in the microbial community of our soil leads generally to a higher overall contribution of bacterial substrate use (Qiu et al. 2017). Additionally, eukaryotes also have a slower metabolism than that of prokaryotes because of their larger cell-size and higher cellular complexity (Bååth 1998; Rousk and Bååth 2007). The larger size of the eukaryotes´ cell leads to a smaller surface to volume ratio (Apostel et al. 2013), and consequently, the uptake and incorporation of monomeric substrates into eukaryotic cell components takes longer (Moore et al. 2005; Reischke et al. 2014) found that loadings of small amounts of complex substrates did not support fungal growth, which may explain the low activity of fungal biomass in DCA utilization in this study. Bacteria rapidly take up labile organic compounds, whereas fungi are often associated with recalcitrant forms (Brant et al. 2006; Paterson et al. 2008; Gunina et al. 2017), a very general statement which we cannot confirm here for the case of hydrophobic long-chain DCA, which were still taken up to a higher degree by prokaryotes than eukaryotes in our study.
The lower uptake of applied DCA into PLFA by actinobacteria compared to other Gram positives indicated the preference of osmotrophic groups living at least partially as single cells in the soil solution for highly water-soluble, simple substrates (Gunina et al. 2014). Furthermore, actinobacteria and proteobacteria with the alkB gene were shown to be able to grow on n-alkanes with lengths ranging from C10 to C19 as their sole source of carbon and energy (Ratajczak et al. 1998; Gibu et al. 2019). This can explain the higher uptake of C12 and C18 DCA in actinobacteria than in fungi/AMF. Furthermore, a second specific pattern was found for this group: Incorporation of 13C of C12 DCA into actinobacteria’s PLFA increased significantly over the course of the experiment. This can be explained by the uptake of microbial necromass as a nutritional source of organics (Apostel et al. 2018). The high and rapid consumption of C12 DCA by the osmotrophic living gram-positive and gram-negative accompanied with a rapid turnover of their biomass results in an accumulation of considerable amounts of 13C-enriched necromass during the 40 days of the experiment. Actinobacteria, by producing lytic enzymes, have a wider metabolic ability than other bacteria in the uptake and use of necromass C (Ryckeboer et al. 2003). The increasing incorporation of DCA-C into actinobacteria PLFA in the second phase of the experiment demonstrates the beginning of this necromass recycling process.
Overall, we could demonstrate distinct mechanisms of DCA turnover in soils, which are controlled by chain-length and thus hydrophobicity at the one hand, but also by substrate-specific metabolic processes. These processes are likely driven by individual microbial groups which could induce divergent DCA pathways in soils. Moreover, our results indicate that the persistence of DCA in soil and therefore their applicability as biomarkers for root derived OC input, cycling and stabilization, might be a function of chain lengths. Altmann et al. (2021b) found a strong decrease of concentrations of cutin-derived dihydroxy fatty acids from Pinus needles after one year, that was less profound with increasing chain length. They explained this observation either by preferential degradation of the ester-bound compounds located on the surface of the cutin-polymers (Beaulieu et al. 2016; Altmann 2021b) or with the atomic oxygen-to-carbon ratio, which influences compound degradation rates (Kleber 2010; Altmann 2021b). Our results show that a chain-length dependent degradation occurs even if the cross-linking ester-bounds are already hydrolyzed and the fatty acids are available as monomers. This challenges the theory of a preferential degradation caused by location within the cutin- or suberin-polymer. However, general assumptions about the applicability of DCA as biomarkers and their stability in soil, solely based on incubation studies, have to be considered carefully and should be complemented e.g. with stable isotope approaches under field conditions. A schematic overview over these processes is illustrated in Fig. 5.
Conclusion
Position-specific labeling combined with 13C-PLFA and 13C–CO2-analysis have revealed new information about the microbial transformation of DCA in soil. The biochemical degradation of DCA differing in chain length demonstrated that C mineralization and cellular uptake of C12 DCA are higher than that of other long-chained DCA. The water-solubility and lower molecular weight but higher oxidation state of C12 DCA can be recognized as the most important factors accelerated its entry into metabolic cycles. The transformation pathway of long-chain DCA can differ from that of middle-chain DCA.
In most cases, mineralization to CO2 and incorporation into PLFA decreased with increasing incubation time and a clear two-phase kinetic with altering factors controlling the DCA metabolism could be identified. Necromass utilization and lower general activity of actinomycetes and fungi/arbuscular mycorrhizal fungi changed this temporal pattern for the filamentous groups. Generally, the cellular structure and adaption mechanisms of the microbial groups played a crucial role in their ability to use the applied DCA, as reflected in the higher uptake of DCA into outer-membrane and thus hydrophobic gram-negative bacteria than into other microbial groups.
In conclusion, we found clear microbial preferences in the use of long chain DCA-derived C. Thus, microbial chain-length specific utilization preferences may significantly shape the DCA pattern observed within the free lipid fraction extracted from soils and the DCA composition after hydrolysis of cutin and suberin polymers. Hence, the specific degradation pattern of DCA in soil has to be considered when using them as biomarkers for root organic matter contribution to different SOM pools and should be complemented with stable isotope approaches.
Our results furthermore show that the longer-term stability of DCA derived carbon in soil is neither depending on the esterification of DCA in suberin-like structures nor on their cycling via the microbial carbon pump into microbial necromass. Instead it is rather a function of their chain-length dependent hydrophobicity and as such a recalcitrance based on their own aliphatic structure.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on request but will be uploaded during the revision process to be available for readers when the publication gets online.
Change history
31 March 2023
A Correction to this paper has been published: https://doi.org/10.1007/s10533-023-01034-x
References
Altmann JG, Jansen B, Jungkunst HF, Kalbitz K (2021) Dynamics of leaf- and root-specific biomarkers during 1-year of litter decomposition. Forests 12:1732. https://doi.org/10.3390/f12121732
Altmann JG, Jansen B, Palviainen M, Kalbitz K (2021b) Stability of needle- and root-derived biomarkers during litter decomposition. J Plant Nutr Soil Sci 184:65–75. https://doi.org/10.1002/jpln.201900472
Ambles A, Jambu P, Parlantin E, Joffre J, Riffe C (1994) Incorporation of natural monoacids from plant residues into an hydromorphic forest podzol. Eur J Soil Sci 45:175–182. https://doi.org/10.1111/j.1365-2389.1994.tb00499.x
Apostel C, Dippold MA, Glaser B, Yakov K (2013) Biochemical pathways of amino acids in soil: assessment by position-specific labeling and 13 C-PLFA analysis. Soil Biol Biochem 67:31–40. https://doi.org/10.1016/j.soilbio.2013.08.005
Apostel C, Dippold MA, Yakov K (2015) Biochemistry of hexose and pentose transformations in soil analyzed by position-specific labeling and 13 C-PLFA. Soil Biol Biochem 80:199–208. https://doi.org/10.1016/j.soilbio.2014.09.005
Apostel C, Herschbach J, Bore EK, Spielvogel S, Kuzyakov Y, Dippold MA (2018) Food for microorganisms: position-specific 13 C labeling and 13 C-PLFA analysis reveals preferences for sorbed or necromass C. Geoderma 312:86–94. https://doi.org/10.1016/j.geoderma.2017.09.042
Ashraf W, Mihdhir A, Murrell JC (1994) Bacterial oxidation of propane. FEMS Microbiol Lett 122:1–6. https://doi.org/10.1111/j.1574-6968.1994.tb07134.x
Atshaves BP, Martin GG, Hostetler HA, McIntosh AL, Kier AB, Schroeder F (2010) Liver fatty acid-binding proteins and obesity. J Nutr Biochem 21:1015–1032. https://doi.org/10.1016/j.jnutbio.2010.01.005
Bååth E (1998) Growth rates of bacterial communities in soils at varying pH: a comparison of the thymidine and leucine incorporation techniques. Microb Ecol 36:316–327. https://doi.org/10.1007/s002489900118
Bahri H, Dignac MF, Rumpel C, Rasse DP, Chenu C, Mariotti A (2006) Lignin turnover kinetics in an agricultural soil is monomer specific. Soil Biol Biochem 38:1977–1988. https://doi.org/10.1016/j.soilbio.2006.01.003
Banfield CC, Pausch J, Kuzyakov Y, Dippold MA (2018) Microbial processing of plant residues in the subsoil-the role of biopores. Soil Biol Biochem 125:309–318. https://doi.org/10.1016/j.soilbio.2018.08.004
Baumgarten T, Vazquez J, Bastisch C, Veron W, Feuilloley MG, Nietzsche S, Wick LY, Heipieper HJ (2012) Alkanols and chlorophenols cause different physiological adaptive responses on the level of cell surface properties and membrane vesicle formation in Pseudomonas putida DOT-T1E. Appl Microbiol Biotechnol 93:837–845. https://doi.org/10.1007/s00253-011-3442-9
Bayry J, Aimanianda V, Guijarro JI, Sunde M, Latge JP (2012) Hydrophobins-unique fungal proteins. PLoS Pathog 8:e1002700. https://doi.org/10.1371/journal.ppat.1002700
Beaulieu C, Sidibé A, Jabloune R, Simao-Beaunoir AM, Lerat S, Monga E, Bernards MA (2016) Physical, chemical and proteomic evidence of potato suberin degradation by the plant pathogenic bacterium Streptomyces scabiei. Microbes Environ 31:427–434. https://doi.org/10.1264/jsme2.ME16110
Beopoulos A, Chardot T, Nicaud JM (2009) Yarrowia lipolytica: a model and a tool to understand the mechanisms implicated in lipid accumulation. Biochimie 9:692–696. https://doi.org/10.1016/j.biochi.2009.02.004
Boddy E, Hill PW, Farrar J, Jones DL (2007) Fast turnover of low molecular weight components of the dissolved organic carbon pool of temperate grassland field soils. Soil Biol Biochem 39:827–835. https://doi.org/10.1016/j.soilbio.2006.09.030
Brant JB, Sulzman EW, Myrold DD (2006) Microbial community utilization of added carbon substrates in response to long-term carbon input manipulation. Soil Biol Biochem 38:2219–2232. https://doi.org/10.1016/j.soilbio.2006.01.022
Chakrawal A, Herrmann AM, Šantrůčková H, Manzoni S (2020) Quantifying microbial metabolism in soils using calorespirometry – a bioenergetics perspective. Soil Biol Biochem 148:107945. https://doi.org/10.1016/j.soilbio.2020.107945
Coon MJ (2005) Omega oxygenases: nonheme-iron enzymes and P450 cytochromes. Biochem Biophys Res Commun 338:378–385. https://doi.org/10.1016/j.bbrc.2005.08.169
Datta R, Kelkar A, Baraniya D, Molaei A, Moulick A, Meena RS, Formanek P (2017) Enzymatic degradation of lignin in soil: a review. Sustainability 9:1163. https://doi.org/10.3390/su9071163
Deamer D (2017) The role of lipid membranes in life’s origin. Life 7:5. https://doi.org/10.3390/life7010005
Dijkstra P, Blankinship JC, Selmants PC, Hart SC, Koch GW, Schwartz E, Hungate BA (2011) Probing carbon flux patterns through soil microbial metabolic networks using parallel position-specific tracer labeling. Soil Biol Biochem 43:126–132. https://doi.org/10.1016/j.soilbio.2010.09.022
Dippold MA, Kuzyakov Y (2013) Biogeochemical transformations of amino acids in soil assessed by position-specific labeling. Plant Soil 373:385–401. https://doi.org/10.1007/s11104-013-1764-3
Eschenfeldt WH, Zhang Y, Samaha H, Stols L, Eirich LD, Wilson CR, Donnelly MI (2003) Transformation of fatty acids catalyzed by cytochrome P450 monooxygenase enzymes of Candida tropicalis. Appl Environ Microbiol 69:5992–5999. https://doi.org/10.1128/AEM.69.10.5992-5999.2003
Feng X, Xu Y, Jaffé R, Schlesinger WH, Simpson MJ (2010) Turnover rates of hydrolysable aliphatic lipids in duke Forest soils determined by compound specific. Org Geochem 41:573–579. https://doi.org/10.1016/j.orggeochem.2010.02.013
Fischer H, Kuzyakov Y (2010) Sorption, microbial uptake and decomposition of acetate in soil. transformations revealed by position-specific 14 C labeling. Soil Biol Biochem 42:186–192. https://doi.org/10.1016/j.soilbio.2009.10.015
Frostegard A, Tunlid A, Baath E (1991) Microbial biomass measured as total lipid phosphate in soils of different organic content. J Microbiol Methods 14:151–163. https://doi.org/10.1016/0167-7012(91)90018-L
Garcia-Pausas J, Paterson E (2011) Microbial community abundance and structure are determinants of soil organic matter mineralisation in the presence of labile carbon. Soil Biol Biochem 43:1705–1713. https://doi.org/10.1016/j.soilbio.2011.04.016
Gearing PJ, Gearing JN, Maughan JT, Oviatt CA (1991) Isotopic distribution of carbon from sewage-sludge and eutrophication in the sediments and food web of estuarine ecosystems. Environ Sci Technol 25:295–301. https://doi.org/10.1021/es00014a012
Gibu N, Kasai D, Ikawa T, Akiyama E, Fukuda M (2019) Characterization and transcriptional regulation of n-alkane hydroxylase gene cluster of Rhodococcus jostii RHA1. Microorganisms 7:479. https://doi.org/10.3390/microorganisms7110479
Glaser B (2005) Compound-specific stable-isotope (δ13C) analysis in soil science. J Plant Nutr Soil Sci 168:633–648. https://doi.org/10.1002/jpln.200521794
Gunina A, Dippold MA, Glaser B, Kuzyakov Y (2014) Fate of low molecular weight organic substances in an arable soil: from microbial uptake to utilization and stabilization. Soil Biol Biochem 77:304–313. https://doi.org/10.1016/j.soilbio.2014.06.029
Gunina A, Smith AR, Kuzyakov Y, Jones DL (2017) Microbial uptake and utilization of low molecular weight organic substrates in soil depend on carbon oxidation state. Biogeochemistry 133:89–100. https://doi.org/10.1007/s10533-017-0313-1
Hamilton JA, Kamp F (1999) How are free fatty acids transported in membranes? is it by proteins or by free diffusion through the lipids? Diabetes 48:2255–2269. https://doi.org/10.2337/diabetes.48.12.2255
Huf S, Krügener S, Hirth T, Rupp S, Zibek S (2011) Biotechnological synthesis of long-chain dicarboxylic acids as building blocks for polymers. Eur J Lipid Sci Technol 113:548–561. https://doi.org/10.1002/ejlt.201000112
Järvinen R, Silvestre AJD, Holopainen U, Kaimainen M, Nyyssölä A, Gil A, Neto CP, Lehtinen P, Buchert J, Kallio H (2009) Suberin of potato (Solanum tuberosum var Nikola): comparison of the effect of cutinase CcCut1 hydrolysis with chemical depolymerization. J Agric Food Chem 57:9016–9027. https://doi.org/10.1021/jf9008907
Jones DL, Shannon D (1999) Mineralization of amino acids applied to soils: impact of soil sieving, storage, and inorganic nitrogen additions. Soil Sci Soc Am J 63:1199–1206. https://doi.org/10.2136/sssaj1999.6351199x
Jones DL, Dennis PG, Owen AG, van Hees PAW (2003) Organic acids behavior in soils-misconceptions and knowledge gaps. Plant Soil 248:31–41. https://doi.org/10.1023/A:1022304332313
Katsikogianni M, Missirlis YF (2004) Concise review of mechanisms of bacterial adhesion to biomaterials and of techniques used in estimating bacteria-material interactions. Eur Cells Mater 8:37–57. https://doi.org/10.22203/ecm.v008a05
Kleber M (2010) What is recalcitrant soil organic matter? Environ Chem 7:320–332. https://doi.org/10.1071/EN10006
Knapp DR (1979) Handbook of analytical derivatization reagents. Wiley, New York
Kolattukudy PE (1980) Cutin, suberin and waxes. In: Stumpf PK, Conn (eds) EE Comprehensive biochemistry of plants, vol 4. Academic Press, New York, pp 571–645
Kontkanen H, Westerholm-Parvinen A, Saloheimo M, Bailey M, Rättö M, Mattila I, Mohsina M, Nakari-Setälä T, Buchert J (2009) Novel Coprinopsis cinerea Polyesterase that hydrolyzes cutin and suberin. Appl Environ Microbiol 75:2148–2157. https://doi.org/10.1128/AEM.02103-08
Koyama A, Harlow B, Kuske CR, Belnap J, Evans RD (2018) Plant and microbial biomarkers suggest mechanisms of soil organic carbon accumulation in a Mojave Desert ecosystem under elevated CO2. Soil Biol Biochem 120:48–57. https://doi.org/10.1016/j.soilbio.2018.01.033
Krasowska A, Sigler K (2014) How microorganisms use hydrophobicity and what does this mean for human needs? Front Cell Infect Microbiol 4:112. https://doi.org/10.3389/fcimb.2014.00112
Kusch S, Rethemeyer J, Schefuß E, Mollenhauer G (2010) Controls on the age of vascular plant biomarkers in Black Sea sediments. Geochim Cosmochim Ac 74:7031–7047. https://doi.org/10.1016/j.gca.2010.09.005
Kuzyakov Y (2011) How to link soil C pools with CO2 fluxes? Biogeosciences 8:1523–1537. https://doi.org/10.5194/bg-8-1523-2011
Kwan AHY, Winefield RD, Sunde M, Matthews JM, Haverkamp RG, Templeton MD, Mackay JP (2006) Structural basis for rodlet assembly in fungal hydrophobins. Proc Natl Acad Sci USA 103:3621–3626. https://doi.org/10.1073/pnas.0505704103
Lee H, Han C, Lee HW, Park G, Jeon W, Ahn J, Lee H (2018) Development of a promising microbial platform for the production of dicarboxylic acids from biorenewable resources. Biotechnol Biofuels 11:310. https://doi.org/10.1186/s13068-018-1310-x
Linder MB, Szilvay GR, Nakari-Setälä T, Penttilä ME (2005) Hydrophobins: the protein-amphiphiles of filamentous fungi. FEMS Microbiol Rev 29:877–896. https://doi.org/10.1016/j.femsre.2005.01.004
Lozán JL, Kausch H (1998) Angewandte statistik für naturwissenschaftler. Parey Buchverlag, Singhofen
Mauersberger S, Drechsler H, Oehme G, Müller HG (1992) Substrate specificity and stereoselectivity of the fatty alcohol oxidase from the yeast Candida maltosa. Appl Microbiol Biotechnol 37:66–73. https://doi.org/10.1007/BF00174205
Mendez-Millan M, Dignac MF, Rumpel C, Derenne S (2011) Can cutin and suberin biomarkers be used to trace shoot and root-derived organic matter? a molecular and isotopic approach. Biogeochemistry 106:23–38. https://doi.org/10.1007/s10533-010-9407-8
Miao SJ, Ye RZ, Qiao YF, Zhu-Barker X, Doane TA, Horwath WR (2017) The solubility of carbon inputs affects the priming of soil organic matter. Plant Soil 410:129–138. https://doi.org/10.1007/s11104-016-2991-1
Moore JC, McCann K, de Ruiter PC (2005) Modeling trophic pathways, nutrient cycling, and dynamic stability in soils. Pedobiologia 49:499–510. https://doi.org/10.1016/j.pedobi.2005.05.008
Nakamura MT, Yudell BE, Loor JJ (2014) Regulation of the energy metabolism by long-chain fatty acid. Prog Lipid Res 53:124–144. https://doi.org/10.1016/j.plipres.2013.12.001
Nierop KGJ (1998) Origin of aliphatic compounds in a forest soil. Org Geochem 29:1009–1016. https://doi.org/10.1016/S0146-6380(98)00165-X
Olivella MA, Bazzicalupi C, Bianchi A, del Río JC, Fiol N, Villaescusa I (2015) Binding interactions between suberin monomer components and pesticides. Sci Total Environ 527–528:159–164. https://doi.org/10.1016/j.scitotenv.2015.04.118
Paterson E, Osler G, Dawson LA, Gebbing T, Sim A, Ord B (2008) Labile and recalcitrant plant fractions are utilised by distinct microbial communities in soil: independent of the presence of roots and mycorrhizal fungi. Soil Biol Biochem 40:1103–1113. https://doi.org/10.1016/j.soilbio.2007.12.003
Pearson A, Eglinton TI (2000) The origin of n-alkanes in santa monica basin surface sediment: a model based on compound-specific ∆14 C and δ13C data. Org Geochem 31:1103–1116. https://doi.org/10.1016/S0146-6380(00)00121-2
Powlson DS, Hirsch PR, Brookes PC (2001) The role of soil microorganisms in soil organic matter conservation in the tropics. Nutr Cycl Agroecosyst 61:41–51. https://doi.org/10.1023/A:1013338028454
Qiu H, Zheng X, Ge T, Dorodnikov M, Chen X, Hu Y, Kuzyakov Y, Wu J, Su Y, Zhang Z (2017) Weaker priming and mineralisation of low molecular weight organic substances in paddy than in upland soil. Eur J Soil Biol 83:9–17. https://doi.org/10.1016/j.ejsobi.2017.09.008
Rasse DP, Rumpel C, Dignac MF (2005) Is soil carbon mostly root carbon? mechanisms for a specific stabilisation. Plant Soil 269:341–356. https://doi.org/10.1007/s11104-004-0907-y
Ratajczak A, Geissdörfer W, Hillen W (1998) Expression of alkane hydroxylase from acinetobacter sp. Strain ADP1 is induced by a broad range of n-alkanes and requires the transcriptional activator AlkR. J Bacteriol 180:5822–5827. https://doi.org/10.1128/JB.180.22.5822-5827.1998
Reischke S, Rousk J, Bååth E (2014) The effects of glucose loading rates on bacterial and fungal growth in soil. Soil Biol Biochem 70:88–95. https://doi.org/10.1016/j.soilbio.2013.12.011
Rojo F (2009) Degradation of alkanes by bacteria. Environ Microbiol 11:2477–2490. https://doi.org/10.1111/j.1462-2920.2009.01948.x
Rousk J, Bååth E (2007) Fungal biomass production and turnover in soil estimated using the acetate-in-ergosterol technique. Soil Biol Biochem 39:2173–2177. https://doi.org/10.1016/j.soilbio.2007.03.023
Ryckeboer J, Mergaert J, Vaes K, Klammer S, De Clercq DA, Coosemans J, Insam H, Swings J (2003) A survey of bacteria and fungi occurring during composting and self-heating processes. Ann Microbiol 53:349–410
Saxena P, Hildemann LM (1996) Water-soluble organics in atmosphere particles: a critical review of the literature and application of thermodynamics to identify candidate compounds. J Atmos Chem 24:57–109. https://doi.org/10.1007/BF00053823
Schaffer JE (2002) Fatty acid transport: the roads taken. Am J Physiol Endocrinol Metab 282:E239–E246. https://doi.org/10.1152/ajpendo.00462.2001
Schink C, Spielvogel S, Imhof W (2020) Synthesis of 13 C-labelled cutin and suberin monomeric dicarboxylic acids of the general formula HO213C–(CH2)n-13CO2H (n = 10, 12, 14, 16, 18, 20, 22, 24, 26, 28). J Label Compd Radiopharm 64:14–29. https://doi.org/10.1002/jlcr.3885
Schönfeld P, Wojtczak L (2016) Short- and medium-chain fatty acids in energy metabolism: the cellular perspective. J Lipid Res 57:943–954. https://doi.org/10.1194/jlr.R067629
Scott JG (2008) Insect cytochrome P450s: thinking beyond detoxification. In: Li N (ed) Recent Advances in Insect Physiology. Toxicology and Molecular Biology, Research Signpost, Kerala, India, pp 117–124
Spaccini R, Piccolo A, Conte P, Haberhauer G, Gerzabek MH (2002) Increased soil organic carbon sequestration through hydrophobic protection by humic substances. Soil Biol Biochem 34:1839–1851. https://doi.org/10.1016/S0038-0717(02)00197-9
Thevenieau F, Le Dall MT, Nthangeni B, Mauersberger S, Marchal R, Nicaud JM (2007) Characterization of Yarrowia lipolytica mutants affected in hydrophobic substrate utilization. Fungal Genet Biol 44:531–542. https://doi.org/10.1016/j.fgb.2006.09.001
van Bergen PF, Nott CJ, Bull ID, Poulton PR, Evershed RP (1998) Organic geochemical studies of soils from the rothamsted classical experiments – IV. preliminary results from a study of the effect of soil pH on organic matter decay. Org Geochem 29:1779–1795. https://doi.org/10.1016/S0146-6380(98)00188-0
van Bogaert INA, Groeneboer S, Saerens K, Soetaert W (2011) The role of cytochrome P450 monooxygenases in microbial fatty acid metabolism. FEBS J 278:206–221. https://doi.org/10.1111/j.1742-4658.2010.07949.x
van den Brink HM, van Gorcom RFM, van den Hondel CAMJJ, Punt PJ (1998) Cytochrome P450 enzyme systems in fungi. Fungal Genet Biol 23:1–17. https://doi.org/10.1006/fgbi.1997.1021
van Hamme JD, Singh A, Ward OP (2003) Recent advances in petroleum microbiology. Microbiol Mol Biol Rev 67:503–549. https://doi.org/10.1128/MMBR.67.4.503-549.2003
Waldrop MP, Firestone MK (2004) Microbial community utilization of recalcitrant and simple carbon compounds: impact of oak-woodland plant communities. Oecologia 138:275–284. https://doi.org/10.1007/s00442-003-1419-9
Watanabe K, Nishiuchi S, Kulichikhin K, Nakazono M (2013) Does suberin accumulation in plant roots contribute to waterlogging tolerance? Front Plant Sci 4:178. https://doi.org/10.3389/fpls.2013.00178
Watkinson RJ, Morgan P (1990) Physiology of aliphatic hydrocarbon-degrading microorganisms. Biodegradation 1:79–92. https://doi.org/10.1007/BF00058828
Wentzel A, Ellingsen TE, Kotlar HK, Zotchev SB, Throne-Holst M (2007) Bacterial metabolism of long‐chain n‐alkanes. Appl Microbiol Biotechnol 76:1209–1221. https://doi.org/10.1007/s00253-007-1119-1
Werner N, Zibek S (2017) Biotechnological production of bio-based long-chain dicarboxylic acids with oleogenious yeasts. World J Microbiol Biotechnol 33:194. https://doi.org/10.1007/s11274-017-2360-0
Wu C, Peng Y, Wang R, Zhou Y (2012) Understanding the granulation process of activated sludge in a biological phosphorus removal sequencing batch reactor. Chemosphere 86:767–773. https://doi.org/10.1016/j.chemosphere.2011.11.002
Zelles L (1999) Fatty acid patterns of phospholipids and lipopolysaccharides in the characterisation of microbial communities in soil: a review. Biol Fertil Soils 29:111–129. https://doi.org/10.1007/s003740050533
Zelles L, Bai QY, Rackwitz R, Chadwick D, Beese F (1995) Determination of phospholipid-derived and lipopolysaccharide-derived fatty-acids as an estimate of microbial biomass and community structures in soils. Biol Fertil Soils 19:115–123. https://doi.org/10.1007/BF00336146
Zimmermann W (1990) Degradation of lignin by bacteria. J Biotechnol 13:119–130. https://doi.org/10.1016/0168-1656(90)90098-V
Funding
Open Access funding enabled and organized by Projekt DEAL. This project was carried out in the framework of the priority program 2322 “SoilSystems, Systems ecology of soils – energy discharge modulated by microbiome and boundary conditions” funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) – Project numbers: SP 943/8 − 1 and DI 2136/17 − 1. Part of the work was furthermore funded within the DFG projects DI 2136/1–1 and DI 2136/4 − 1.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation was performed by Jennifer Herschbach, Carina Schink and Wolfgang Imhof. Data collection and analysis were performed by Hamed Kashi, Michaela A. Dippold and Sandra Spielvogel. The first draft of the manuscript was written by Hamed Kashi and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Responsible editor: Edward Brzostek
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised as the phrase “Furthermore, actinobacteria with the alkB gene were shown to be able to grow on n-alkanes with lengths ranging from C10 to C19 as their sole source of carbon and energy (Ratajczak et al. 1998; Gibu et al. 2019)” should be corrected to “Furthermore, actinobacteria and proteobacteria with the alkB gene were shown to be able to grow on n-alkanes with lengths ranging from C10 to C19 as their sole source of carbon and energy (Ratajczak et al. 1998; Gibu et al. 2019)”.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kashi, H., Loeppmann, S., Herschbach, J. et al. Size matters: biochemical mineralization and microbial incorporation of dicarboxylic acids in soil. Biogeochemistry 162, 79–95 (2023). https://doi.org/10.1007/s10533-022-00990-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10533-022-00990-0