Abstract
Terrestrial carbon (C)-climate feedbacks depend strongly on how soil organic matter (SOM) decomposition responds to temperature. This dependency is often represented in land models by the parameter Q10, which quantifies the relative increase of microbial soil respiration per 10 °C temperature increase. Many studies have conducted paired laboratory soil incubations and inferred “active” and “slow” pool Q10 values by fitting linear two-pool models to measured respiration time series. Using a recently published incubation study (Qin et al. in Sci Adv 5(7):eaau1218, 2019) as an example, here we first show that the very high parametric equifinality of the linear two-pool models may render such incubation-based Q10 estimates unreliable. In particular, we show that, accompanied by the uncertain initial active pool size, the slow pool Q10 can span a very wide range, including values as high as 100, although all parameter combinations are producing almost equally good model fit with respect to the observations. This result is robust whether or not interactions between the active and slow pools are considered (typically these interactions are not considered when interpreting incubation data, but are part of the predictive soil carbon models). This very large parametric equifinality in the context of interpreting incubation data is consistent with the poor temporal extrapolation capability of linear multi-pool models identified in recent studies. Next, using a microbe-explicit SOM model (RESOM), we show that the inferred two pools and their associated parameters (e.g., Q10) could be artificial constructs and are therefore unreliable concepts for integration into predictive models. We finally discuss uncertainties in applying linear two-pool (or more generally multiple-pool) models to estimate SOM decomposition parameters such as temperature sensitivities from laboratory incubations. We also propose new observations and model structures that could enable better process understanding and more robust predictive capabilities of soil carbon dynamics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Predicting the fate of the large amount of soil carbon stored globally [more than twice as in the current atmosphere (Ciais et al. 2013)] is critical for quantifying feedbacks between terrestrial biogeochemistry and the Earth’s climate. In particular, land models need accurate quantifications of soil organic matter (SOM) decomposition sensitivity to soil temperature under a warming climate. To address this need, thousands of empirical studies have inferred SOM decomposition temperature sensitivities, often characterized by the parameter Q10 that measures the relative increase in microbial SOM respiration per 10 °C temperature increase (e.g., Fang and Moncrieff 2001; Haddix et al. 2011; Hamdi et al. 2013; Lloyd and Taylor 1994; Schadel et al. 2013). However, debate continues regarding the laboratory-based Q10 values for incorporation into SOM models, or whether alternative model structures are more appropriate (Conant et al. 2011; Davidson and Janssens 2006; Davidson et al. 2006; Hemingway et al. 2019; Tang and Riley 2015).
The use of Q10 (or alternatively activation energy in the Arrhenius representation of SOM turnover) is inherent in the substrate quality (SQ) conceptual model of SOM decomposition (e.g., Hartley and Ineson 2008; Parton et al. 1988; Wetterstedt et al. 2010). In these SQ-based models, SOM is assumed to consist of several abstract (i.e., not measurable) pools, each of which has its own decomposition turnover time that varies as a function of temperature, moisture, and other edaphic conditions. Despite their popularity, SQ-based models cannot explain the observation that old carbon (which by assumption is recalcitrant and therefore of long turnover time) can be decomposed rapidly when made accessible to active microbes (e.g., Kleber et al. 2011; Nowinski et al. 2010; Schuur et al. 2009).
Another problem with SQ-based models is their usually very high parametric equifinality, i.e., for a given set of observations, many parameter combinations are able to produce model outputs that fit the observations equally well (e.g., Beven and Freer 2001; Tang and Zhuang 2008), while their predictions often diverge wildly. For instance, using the APSIM model, which has six SQ-based SOM pools, Luo et al. (2015) showed that “convergent modeling of past soil organic carbon stocks” is followed by “divergent projections” of twenty-first century SOM dynamics. In a subsequent study, Luo et al. (2017) deduced the same conclusion for a SQ-based linear two-transfer-pool model (that considers interactions between active and slow pools; see supplemental material for a description and solutions for these two types of linear two-pool models). These findings therefore motivate an analysis of whether the high parametric equifinality of SQ-based models may lead to large uncertainties in temperature sensitivities inferred from incubation experiments.
Notwithstanding these conceptual defects, the simplicity of SQ-based models has led them to be widely applied to interpret laboratory soil incubations. For instance, Liang et al. (2015) compared results from several linear multiple-pool models to infer temperature sensitivity parameters from respiration time series measured during SOM incubations. They found that the two-discrete-pool model, which does not consider interactions between active and slow pools, was able to fit the soil respiration data very well under many conditions, and Schadel et al. (2013) drew the same conclusion from a similar analysis. In a recent study, which we analyze here, Qin et al. (2019; hereafter Q2019) used a linear two-discrete-pool model to fit their 330-day incubation of topsoil (0–10 cm) and subsoil (30–50 cm) samples collected from three sites on the northeastern Tibetan Plateau. They inferred that topsoil is more sensitive than subsoil to temperature warming and concluded that SOM models can be improved by incorporating such a contrast in temperature sensitivity.
Meanwhile, to address the conceptual difficulties with SQ-based models (e.g., “old carbon” can often behave just like “active” pool carbon), new model structures have been developed to attempt explicit representations of the myriad interactions between microbes, enzymes, substrates, and various abiotic soil processes (Abramoff et al. 2019; Dwivedi et al. 2017, 2019; Riley et al. 2014; Sulman et al. 2018; Tang and Riley 2015; Wang et al. 2013; Wieder et al. 2013). These new models then predict that SOM decomposition emerges from these interactions. In general, including more processes requires more model parameters and will potentially increase a model’s parametric equifinality. Nevertheless, these new models enable more direct correspondence between their simulations and empirical measurements, and provide mechanistic insights that are otherwise not available from SQ-based models. Moreover, with improved mathematical formulations (Tang and Riley 2013, 2017, 2019b), these new models may enable first-principle based parameterization that is not feasible with SQ-based models (e.g., Tang and Riley 2019a, b).
Here, by reanalyzing the Q2019 incubation data using the linear two-discrete-pool model (e.g., Liang et al. 2015; Qin et al. 2019; Schadel et al. 2013) and the linear two-transfer-pool model (Luo et al. 2017), we first show that the high parametric equifinality makes it almost impossible to obtain a robust inference of Q10 values and other related parameters for SQ-based SOM pools. Next, using a microbe-explicit model, i.e., the REaction‐network‐based model of Soil Organic Matter and microbes (RESOM) (Tang and Riley 2015), we show that apparent active and slow pools can emerge from the decomposition of a single substrate with a single temperature sensitivity (i.e., activation energy). We then discuss why mechanistic SOM decomposition models like RESOM should be more extensively explored and how laboratory incubations can be better used to inform the development of these new models.
Large parametric equifinality in linear two-pool models leads to uncertain Q10 and other parameters
Q2019 conducted a long-term (330-day) laboratory incubation for topsoil (0–10 cm) and subsoil (30–50 cm) samples collected from three sites on the northeastern Tibetan Plateau. They then applied a linear two-discrete-pool model to derive the temperature sensitivities of active and slow SOM pools, and asserted that topsoil has higher temperature sensitivity than subsoil for bulk soil, active, and slow pools. We digitized their respiration time series data using the MATLAB script grabit (https://www.mathworks.com/matlabcentral/fileexchange/7173-grabit), and conducted Markov chain Monte Carlo (MCMC) based inversions (Vrugt 2016) for a linear two-discrete-pool model using these respiration data. We note that Q2019 set their Q10 parameter prior ranges to [1, 2] and [2, 4] for the active and slow pool, respectively (reported in Table S4 of their Supplemental Material), thereby strongly limiting the range of their possible posterior values. Analogously Schadel et al. (2013) imposed upper and lower limits to the decay rates of active and slow pools. However, some observationally-based studies have reported bulk soil Q10 values as high as a few hundred (e.g., Hamdi et al. 2013). Therefore, if we interpret their approach correctly, the Q2019 parameter inversion results could have been biased methodologically, because these unjustified prior constraints force their inferred Q10 values to fall within an unrealistically small range. We further note that emergent Q10 values have been observed to vary across a wide range and be hysteretic with respect to temperature over time (e.g., Davidson et al. 2006; Pingintha et al. 2010), and microbe-explicit models are better at resolving such hysteresis by accounting for hysteresis in microbial biomass and kinetics (e.g., Tang and Riley 2015).
We performed seven MCMC inversions to quantify how sensitive the inferred Q10 values are to the imposed parameter ranges for both active and slow pools. For both pools, the seven inversions used maximum Q10 values of 4, 10, 20, 40, 50, 70, and 100; the experiments are named accordingly as IQ4, IQ10, IQ20, IQ40, IQ50, IQ70, and IQ100, where IQx refers to an inversion with Q10 bounded at a maximum value of x. Additionally, since Q10 has been reported to be as low as 0.5 (Hamdi et al. 2013), we allowed Q10 prior values to extend to 0, but none of the MCMC inversions resulted in Q10 values smaller than 1.
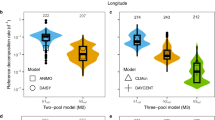
When the maximum Q10 is bounded at 4 (for both pools), our MCMC inversions (Fig. 1) corroborate Q2019s inference that Q10 values of both the active and slow pools in topsoil (\(2.06\pm 0.012\) and \(3.89\pm 0.092\), respectively) are higher than those in subsoil (\(1.51\pm 0.031\) and \(2.86\pm 0.23\), respectively), although these Q10 values differ from those in Q2019, who reported mean values of about 1.8 and 2.4 for topsoil active and slow pools, respectively; and 1.6 and 2.2 for subsoil active and slow pools, respectively. These differences may exist because (1) we did not exactly recover their data and the inversion is sensitive to data uncertainty characterization, and (2) we used different, but still restrictive, parameter ranges. Further, when we imposed the same upper prior ranges as Q2019 did, the inversion inferred topsoil Q10 values for the active and slow pool of \(2.00\pm 0.003\) and \(3.95\pm 0.045\), respectively, and for the subsoil, of \(1.53\pm 0.030\) and \(3.69\pm 0.26\), respectively. However, for inversions with larger maximum Q10 values, we find very wide ranges of slow pool Q10 values (i.e., a manifestation of high parametric equifinality; Fig. 1c, d), while all predicted respiration time series match the observations equally well (Fig. 1a, b). Moreover, this large parametric equifinality is higher for topsoil than for subsoil. For example, the most uncertain inference occurs for topsoil when Q10 is bounded at 100, where the posterior maximum and minimum slow pool Q10 values are 97.5 and 4.6, respectively; whereas for subsoil the most uncertain inference occurs when Q10 is bounded at 70, where the posterior maximum and minimum slow pool Q10 values are 40.9 and 2.80, respectively. In summary, our results are consistent with those of Luo et al. (2017), even though they instead used time series of soil carbon stocks as constraints in their parameter inversions for the linear two-transfer-pool model, which has one more parameter than does the linear two-discrete-pool model (see supplemental material).
For the incubation time series from Qin et al. (2019), many MCMC based inversions of two-discrete-pool model parameters are able to fit the data equally well (panels a and b), but their slow pool Q10 values are quite different (panels c and d). The inversions are labelled as IQx, i.e., an inversion with a maximum Q10 value of x for both active and slow pools. Panels a and c are for topsoil, b and d are for subsoil
When the posterior distributions of slow pool decay rates at different temperatures and initial active pool fractions are compared for inversions with Q10 values bounded at 4 and 100 (i.e., IQ4 and IQ100 in Fig. 2), we find that the wider range of topsoil slow pool Q10 for IQ100 (than for IQ4; Fig. 2a, c, e) occurs because (1) the inferred slow pool decay rates at 10 °C and 20 °C are left edge-hitting and more center-peaked, respectively, and (2) the initial active pool is larger in IQ100 than in IQ4 (Fig. 2g, h). For subsoil, the posterior differences between IQ100 and IQ4 are smaller, but high parametric equifinality remains (Fig. 2b, d, f).
For topsoil, when the MCMC inversion applies a maximum Q10 value of 100 (i.e., IQ100), the slow pool Q10 distribution (panel a) has a longer tail (than IQ4) because the distribution of IQ100 slow pool decay rate (\({k}_{s}\)) at 10 °C (panel c) does not align well with the distribution at 20 °C (panel e) and the initial active pool is larger (panel g). Differences between IQ100 and IQ4 for subsoil also exist but are comparatively smaller (panels b, d, f, h)
Our analyses above show that more than 12,500 combinations (out of the 500,000 MCMC samples for each inversion) of posterior parameters fit the incubation soil respiration observations equally well. This high parametric equifinality and the resultant wide range of Q10 values indicate that (1) it is difficult to use a linear two-discrete-pool model to robustly derive the temperature sensitivity of SOM decomposition from laboratory incubation soil respiration data alone, and (2) the initial active and slow pool SOM stocks are difficult to robustly quantify. For assertion (2), although the differences in the fraction of initial SOM as active pool between inversions (Fig. 2g, h) are relatively small (about 0.09 vs. 0.11 on average), they force significant uncertainty into the slow pool decay rates, making those small differences consequential (which is reflected in the wide range of slow pool Q10).
Parameters inferred from linear two-discrete-pool models are often applied to inform more complex transfer-pool models (e.g., CENTURY; Parton et al. (1988)). However, the difference in these two model structures makes it difficult to use the discrete-pool model to demonstrate the impacts of the high parametric equifinality on model projections. To facilitate such a demonstration, we conducted an inversion using a linear two-transfer-pool model (see supplemental material for model formulation) for the topsoil with maximum Q10 values of 4 and 100, and performed forward simulations using the inferred posterior parameters. We found that the full set of inversions using the linear two-transfer-pool model produced very similar results (not shown) to those shown in Figs. 1 and 2.
We find the MCMC inversions with the linear two-transfer-pool model are equally uncertain (Fig. 3) as those for the linear two-discrete-pool model (Figs. 1, 2). Moreover, the associated 25-year projection with no warming demonstrates a very strong divergence through amplifying effects from the carbon input flux on the posterior parameters after year 2 (Fig. 4). If warming is considered, we expect that the divergence shown in Fig. 4 will become even larger because it will include additional uncertainty from Q10 (Feller 1968). Given these results and previous studies (Luo et al. 2015; Tang and Zhuang 2008), we conclude that linear two-pool models (and probably multiple-pool models) are of such high parametric equifinality that they cannot be well-constrained by incubation time series observations of either respiration or soil carbon stocks.
MCMC inversion with the linear two-transfer-pool model for topsoil with the maximum Q10 set to 4 and 100, respectively. Panel a is model comparison to respiration time series; b is posterior active and slow pool Q10 values; c, d, e, f and g are probability distribution of the fraction of decomposed active pool into slow pool, slow pool Q10, slow pool \({k}_{s}\) at 10 °C, slow pool \({k}_{s}\) at 20 °C and initial active pool fraction of the soil sample, respectively
A demonstration of the parametric equifinality problem of the linear two-transfer-pool model for soil carbon projections using posterior parameters from Fig. 3. The carbon input to the model begins from year 2, consistent with the beginning of the wide uncertainty range that increases over time
We also investigated whether additional data could help better constrain the linear two-pool models. We tested this idea by dramatically increasing the sampling frequency and reducing the observational error of respired CO2. This synthetic inversion experiment with the linear two-transfer-pool model indicates that, although sampling respiration daily at 2% relative error (which is rarely achievable in reality) does constrain Q10 values quite well, it is still insufficient to robustly infer other model parameters (i.e., initial active pool fraction and fraction of decomposed active pool into slow pool) that also must be used for predictions (Fig. 5). Increasing the relative error of daily observations to a more realistic yet still challenging-to-achieve 5% would have resulted in an even more biased posterior of the initial fraction of SOM as active pool (0.027 for IQ4, and 0.21 for IQ100), and transfer coefficient (0.02 for IQ4, and 0.83 for IQ100) between active and slow pools during decomposition (Figure S1). Analogously, we suspect that adding multi-site bulk SOM stocks or radiocarbon respiration data as observational constraints is unlikely to improve the inversion, because of the weak signal of the slow pool as can be triggered in an incubation experiment (which will lead to biased estimation of the fraction of decomposed active pool that enters slow pool, as shown in Fig. 5g).
MCMC parameter inversion using synthetic observations of respiration sampled daily with a relative error 2%. True model parameter values of the linear two-transfer-pool model are annotated in all panels of the corresponding posterior histogram. Panel a is model comparison to respiration time series; b, c, d, e, f and g are probability distributions of active pool Q10, slow pool Q10, initial active pool fraction, active pool \({k}_{f}\) at 10 °C, slow pool \({k}_{s}\) at 10 °C, and the fraction of decomposed active pool into slow pool
Multiple apparent SOM pools can emerge from a single substrate
We next conducted incubation experiments with the microbe-explicit RESOM model (Abramoff et al. 2019; Tang and Riley 2015) to show that apparent active and slow pools may be inferred from incubation data, even though a single substrate with a single activation energy of \(55\) kJ (mol C)−1 was actually undergoing microbial decomposition. For this demonstration of commonly-observed SOM decomposition patterns, we configured RESOM with one microbial population (with a substrate assimilation efficiency of 0.27 for topsoil, and 0.50 for subsoil, respectively), one extracellular enzyme, one mineral soil surface (with surface area of 575 g C m−2 and 1800 g C m−2 for topsoil and subsoil, respectively), and one substrate (which can exist in both aqueous and solid phases). The substrate assimilation efficiency and mineral soil surface were obtained through trial-and-error by comparing the simulated respiration data to that of Q2019, and other parameters are from previous publications (Abramoff et al. 2019; Tang and Riley 2015). Following Tang and Riley (2015), we first generated initial conditions from a spinup to equilibrium and then ran RESOM for 330 days at the two temperatures to mimic the incubation. After normalizing the respiration to initial SOM, these RESOM simulations reproduced the Q2019 respiration time series for both topsoil and subsoil within the observational uncertainty (Fig. 6a, b), suggesting that the predictive capability of RESOM is scientifically reasonable to demonstrate the potential to mis-identify multiple substrate pools from incubation data. We also note that when the temperature forcing has higher temporal variability, like that observed in field conditions, the emergent temperature sensitivity is dynamic and the corresponding instantaneous Q10 can vary across a wide range. This dynamics occurs because temperature sensitivity of SOM decomposition depends on all the interactions between microbes, substrates, enzymes, and mineral surfaces represented in the model, as discussed extensively in Tang and Riley (2015).
The one-substrate RESOM model accurately fits the laboratry incubation data (panels a and b). The model predictions and observations show why more than one apparent SOM pool could be mistakenly inferred (panels c and d). In particular, calibrating (from MCMC inversion) the two-discrete-pool model fits the RESOM simulations for subsoil almost perfectly (panel d) and reasonably well for topsoil (panel c). Panels a and c are for topsoil, b and d are for subsoil
When the natural logarithm of the RESOM-predicted respiration is plotted against time (Fig. 6c, d), nonlinear curves are found for both topsoil and subsoil at 10 °C and 20 °C. This commonly-observed pattern, which can also be produced with other model parameters or by other microbial models (e.g., Grant et al. 1993; Smith 1979; Wieder et al. 2014), is a primary reason that researchers assume SOM has multiple pools with different turnover times. In this one-substrate-pool RESOM version, these nonlinear-curves occur because, as the incubation progresses, (1) substrate is gradually depleted, and (2) microbial biomass is reduced accordingly. Indeed, by conducting linear two-discrete-pool model parameter MCMC inversions with the RESOM predicted respiration time series data, the inferred model is found to fit almost perfectly to the RESOM subsoil simulation (Fig. 6d), and relatively accurately for the topsoil (more than two pools are required to fit the RESOM simulation for topsoil) (Fig. 6c). This contrast of model fitting to RESOM simulations for topsoil and subsoil suggests that definitions of SQ pools are not unique even for a single substrate. In real soils, substrate diversity is likely very high, and the forcing temperature is much more variable. Thus, depending on the relative strength of interactions (and likely the imposed temperature forcing in incubation experiments), SQ-based linear multiple-pool models may succeed in one situation (still at the expense of uncertain parameters) while fail in another, even for similar overall system properties.
Improving new SOM model structures from better use of observations and cross-disciplinary knowledge
Our analyses demonstrate that, because of their high parametric equifinality, SQ-based linear two-pool models provide inaccurate inferences of SOM decomposition temperature sensitivity (and other related parameters). We also showed that the oft-inferred active and slow pools could be artificial constructs that emerge even when only a single substrate is being decomposed. Therefore, improving SOM decomposition models and temperature sensitivity inferences from observations requires more explicit representation of the interactions between the many biotic and abiotic processes that contribute to SOM dynamics. Intuitively, one could argue that models with more mechanistic representations may require more parameters and may not resolve the curse of high parametric equifinality. However, more mechanistic SOM models have at least several advantages, including that they:
-
(1)
can incorporate first-principle based representations of kinetic parameters, e.g., substrate affinity parameter, maximum substrate uptake rate, or even substrate assimilation efficiency, whose variability is unlikely to be resolved through multiplier functions as often used in SQ-based models (Tang and Riley 2019a, b);
-
(2)
can provide mechanistic insights into important systems dynamics, including why some old carbon (i.e., with relatively negative Δ14C values) can relatively easily be decomposed when exposed to microbes and how mineral protection affects SOM persistence (Hemingway et al. 2019; Tang and Riley 2015);
-
(3)
can produce more realistic dynamic predictions of how temperature and moisture affect SOM dynamics, which have been statically represented using multiplier functions in SQ-based models (Tang and Riley 2019b), allowing for observed bulk soil Q10 values to vary over a wide range or be hysteretic (Pingintha et al. 2010);
-
(4)
enable consistent representation of the myriad biogeochemical processes that control the cycling of various important chemicals (e.g., CO2, CH4, N2O, etc.), a need evidenced by the SQ-based models’ ever-increasing introduction of new response functions for new processes (Sierra et al. 2015; Zhuang et al. 2004);
-
(5)
enable a direct link between models and the vast amount of knowledge accumulated in related fields like microbiology, microbial ecology, aqueous chemistry, physical chemistry, etc., so that more diverse data (e.g., microbial cell size) can be used for model constraints [e.g., Jin and Kirk (2018a, b), la Cecilia et al. (2019), LaRowe and Van Cappellen (2011), and Tang and Riley (2019a, b)];
-
(6)
may allow the use of parametric equifinality to reduce predictive divergence. This counter-intuitive idea is analogous to the approach used to link kinetic gas theory and Navier–Stokes equations in fluid mechanics (Batchelor 1967), except that in microbial models parametric equifinality could be represented with bio-diversity. When bio-diversity is further integrated with known ecological principles (e.g., competition and symbiosis), the many parameters can compensate each other to reduce predictive uncertainty and meanwhile enhance the models’ resilience to perturbations (see Sect. 6 of Tang and Riley (2017) for a discussion of this concept).
Finally, with the advantages described above, we emphasize that mechanistic models will benefit from laboratory respiration measurements when biogeochemical and biogeophysical processes affecting respiration are also measured. Although we demonstrated that the inferred temperature sensitivities in Q2019 are likely biased because of their assumed SOM model structure and inversion approach, their other measurements characterizing soil aggregates, mineral soil protection, and relative microbial abundances and composition are valuable resources for future analyses. As experiments begin collecting such critical information, a suite of process-rich predictive models can be developed which in turn enable better interpretation of laboratory incubation studies and improve predictions of terrestrial biogeochemical interactions at larger scales.
Conclusions
SQ-based SOM models have been argued to be self-consistent (Blankinship et al. 2018) and have provided useful understanding of SOM dynamics, yet we show here that their high parametric equifinality prevents them from reliably being used to infer temperature sensitivity and other related parameters of assumed SOM pools from laboratory incubation data. This problem is serious, since many global land models used in climate assessments have SOM decomposition models parameterized from these types of observations. New and more mechanistic models of SOM decomposition should be explored to represent the myriad biotic and abiotic entities and interactions that affect temperature [and moisture; e.g., Tang and Riley (2019b)] sensitivity of SOM decomposition. By combining observations and model structures that more explicitly represent the underlying processes affecting SOM decomposition, we can expect improved understanding and predictions of how SOM cycling may respond to climate change.
Change history
05 September 2022
The original online version of this article was revised: formatting codes were removed from Figure citations in the text.
References
Abramoff RZ, Torn MS, Georgiou K, Tang JY, Riley WJ (2019) Soil organic matter temperature sensitivity cannot be directly inferred from spatial gradients. Glob Biogeochem Cycles 33(6):761–776
Batchelor GK (1967) An introduction to fluid dynamics. Cambridge University Press, Cambridge
Beven K, Freer J (2001) Equifinality, data assimilation, and uncertainty estimation in mechanistic modelling of complex environmental systems using the GLUE methodology. J Hydrol 249(1–4):11–29
Blankinship JC, Berhe AA, Crow SE, Druhan JL, Heckman KA, Keiluweit M, Lawrence CR, Marin-Spiotta E, Plante AF, Rasmussen C, Schadel C, Schimel JP, Sierra CA, Thompson A, Wagai R, Wieder WR (2018) Improving understanding of soil organic matter dynamics by triangulating theories, measurements, and models. Biogeochemistry 140(1):1–13
Ciais P, Sabine C, Bala G, Bopp L, Brovkin V, Canadell J, Chhabra A, DeFries R, Galloway J, Heimann M, Jones C, Le Quere C, Myneni RB, Piao SL, Thornton P (2013) Carbon and other biogeochemical cycles. In: Stocker TF, Qin D, Plattner G-K, Tignor M, Allen SK, Boschung J, Nauels A, Xia Y, Bex V & Midgley PM (eds) Climate Change 2013: the physical science basis. Contribution of working group I to the Fifth Assessment Report of the Intergovernmental Panel on Climate Change. Cambridge University Press, Cambridge, United Kingdom and New York, NY, USA
Conant RT, Ryan MG, Agren GI, Birge HE, Davidson EA, Eliasson PE, Evans SE, Frey SD, Giardina CP, Hopkins FM, Hyvonen R, Kirschbaum MUF, Lavallee JM, Leifeld J, Parton WJ, Steinweg JM, Wallenstein MD, Wetterstedt JAM, Bradford MA (2011) Temperature and soil organic matter decomposition rates - synthesis of current knowledge and a way forward. Glob Change Biol 17(11):3392–3404
Davidson EA, Janssens IA (2006) Temperature sensitivity of soil carbon decomposition and feedbacks to climate change. Nature 440(7081):165–173
Davidson EA, Janssens IA, Luo YQ (2006) On the variability of respiration in terrestrial ecosystems: moving beyond Q(10). Glob Change Biol 12(2):154–164
Dwivedi D, Riley WJ, Torn MS, Spycher N, Maggi F, Tang JY (2017) Mineral properties, microbes, transport, and plant-input profiles control vertical distribution and age of soil carbon stocks. Soil Biol Biochem 107:244–259
Dwivedi D, Tang JY, Bouskill NJ, Georgiou K, Chacon SS, Riley WJ (2019) Abiotic and biotic controls on soil organo-mineral interactions: developing model structures to analyze why soil organic matter persists. Rev Miner Geochem 85(1):329–348
Fang C, Moncrieff JB (2001) The dependence of soil CO2 efflux on temperature. Soil Biol Biochem 33(2):155–165
Feller W (1968) An introduction to probability theory and its applications. Wiley, Hoboken
Grant RF, Juma NG, Mcgill WB (1993) Simulation of carbon and nitrogen transformations in soil - mineralization. Soil Biol Biochem 25(10):1317–1329
Haddix ML, Plante AF, Conant RT, Six J, Steinweg JM, Magrini-Bair K, Drijber RA, Morris SJ, Paul EA (2011) The role of soil characteristics on temperature sensitivity of soil organic matter. Soil Sci Soc Am J 75(1):56–68
Hamdi S, Moyano F, Sall S, Bernoux M, Chevallier T (2013) Synthesis analysis of the temperature sensitivity of soil respiration from laboratory studies in relation to incubation methods and soil conditions. Soil Biol Biochem 58:115–126
Hartley IP, Ineson P (2008) Substrate quality and the temperature sensitivity of soil organic matter decomposition. Soil Biol Biochem 40(7):1567–1574
Hemingway JD, Rothman DH, Grant KE, Rosengard SZ, Eglinton TI, Derry LA, Galy VV (2019) Mineral protection regulates long-term global preservation of natural organic carbon. Nature 570(7760):228–231
Jin QS, Kirk MF (2018a) pH as a primary control in environmental microbiology: 1. Thermodynamic perspective. Front Environ Sci-Switz 6:21
Jin QS, Kirk MF (2018b) pH as a primary control in environmental microbiology: 2. Kinetic perspective. Front Environ Sci-Switz 6:101
Kleber M, Nico PS, Plante AF, Filley T, Kramer M, Swanston C, Sollins P (2011) Old and stable soil organic matter is not necessarily chemically recalcitrant: implications for modeling concepts and temperature sensitivity. Glob Change Biol 17(2):1097–1107
la Cecilia D, Riley WJ, Maggi F (2019) Biochemical modeling of microbial memory effects and catabolite repression on soil organic carbon compounds. Soil Biol Biochem 128:1–12
LaRowe DE, Van Cappellen P (2011) Degradation of natural organic matter: a thermodynamic analysis. Geochim Cosmochim Acta 75(8):2030–2042
Liang JY, Li DJ, Shi Z, Tiedje JM, Zhou JZ, Schuur EAG, Konstantinidis KT, Luo YQ (2015) Methods for estimating temperature sensitivity of soil organic matter based on incubation data: a comparative evaluation. Soil Biol Biochem 80:127–135
Lloyd J, Taylor JA (1994) On the temperature-dependence of soil respiration. Funct Ecol 8(3):315–323
Luo Z, Wang E, Zheng H, Baldock JA, Sun OJ, Shao Q (2015) Convergent modelling of past soil organic carbon stocks but divergent projections. Biogeosciences 12(14):4373–4383
Luo ZK, Wang EL, Sun OJ (2017) Uncertain future soil carbon dynamics under global change predicted by models constrained by total carbon measurements. Ecol Appl 27(3):1001–1009
Nowinski NS, Taneva L, Trumbore SE, Welker JM (2010) Decomposition of old organic matter as a result of deeper active layers in a snow depth manipulation experiment. Oecologia 163(3):785–792
Parton WJ, Stewart JWB, Cole CV (1988) Dynamics of C, N, P and S in grassland soils - a model. Biogeochemistry 5(1):109–131
Pingintha N, Leclerc MY, Beasley JP, Zhang GS, Senthong C (2010) Assessment of the soil CO2 gradient method for soil CO2 efflux measurements: comparison of six models in the calculation of the relative gas diffusion coefficient. Tellus B 62(1):47–58
Qin SQ, Chen LY, Fang K, Zhang QW, Wang J, Liu FT, Yu JC, Yang YH (2019) Temperature sensitivity of SOM decomposition governed by aggregate protection and microbial communities. Sci Adv 5(7):eaau1218
Riley WJ, Maggi F, Kleber M, Torn MS, Tang JY, Dwivedi D, Guerry N (2014) Long residence times of rapidly decomposable soil organic matter: application of a multi-phase, multi-component, and vertically resolved model (BAMS1) to soil carbon dynamics. Geosci Model Dev 7(4):1335–1355
Schadel C, Luo YQ, Evans RD, Fei SF, Schaeffer SM (2013) Separating soil CO2 efflux into C-pool-specific decay rates via inverse analysis of soil incubation data. Oecologia 171(3):721–732
Schuur EAG, Vogel JG, Crummer KG, Lee H, Sickman JO, Osterkamp TE (2009) The effect of permafrost thaw on old carbon release and net carbon exchange from tundra. Nature 459(7246):556–559
Sierra CA, Trumbore SE, Davidson EA, Vicca S, Janssens I (2015) Sensitivity of decomposition rates of soil organic matter with respect to simultaneous changes in temperature and moisture. J Adv Model Earth Syst 7(1):335–356
Smith OL (1979) Application of a model of the decomposition of soil organic-matter. Soil Biol Biochem 11(6):607–618
Sulman BN, Moore JAM, Abramoff R, Averill C, Kivlin S, Georgiou K, Sridhar B, Hartman MD, Wang GS, Wieder WR, Bradford MA, Luo YQ, Mayes MA, Morrison E, Riley WJ, Salazar A, Schimel JP, Tang JY, Classen AT (2018) Multiple models and experiments underscore large uncertainty in soil carbon dynamics. Biogeochemistry 141(2):109–123
Tang JY, Riley WJ (2013) A total quasi-steady-state formulation of substrate uptake kinetics in complex networks and an example application to microbial litter decomposition. Biogeosciences 10(12):8329–8351
Tang JY, Riley WJ (2015) Weaker soil carbon-climate feedbacks resulting from microbial and abiotic interactions. Nat Clim Change 5(1):56–60
Tang JY, Riley WJ (2017) SUPECA kinetics for scaling redox reactions in networks of mixed substrates and consumers and an example application to aerobic soil respiration. Geosci Model Dev 10(9):3277–3295
Tang JY, Riley WJ (2019a) Competitor and substrate sizes and diffusion together define enzymatic depolymerization and microbial substrate uptake rates. Soil Biol Biochem 139:107624
Tang JY, Riley WJ (2019b) A theory of effective microbial substrate affinity parameters in variably saturated soils and an example application to aerobic soil heterotrophic respiration. J Geophys Res-Biogeo 124(4):918–940
Tang JY, Zhuang QL (2008) Equifinality in parameterization of process-based biogeochemistry models: a significant uncertainty source to the estimation of regional carbon dynamics. J Geophys Res-Biogeo. https://doi.org/10.1029/2008JG000757
Vrugt JA (2016) Markov chain Monte Carlo simulation using the DREAM software package: theory, concepts, and MATLAB implementation. Environ Modell Softw 75:273–316
Wang GS, Post WM, Mayes MA (2013) Development of microbial-enzyme-mediated decomposition model parameters through steady-state and dynamic analyses. Ecol Appl 23(1):255–272
Wetterstedt JAM, Persson T, Agren GI (2010) Temperature sensitivity and substrate quality in soil organic matter decomposition: results of an incubation study with three substrates. Glob Change Biol 16(6):1806–1819
Wieder WR, Bonan GB, Allison SD (2013) Global soil carbon projections are improved by modelling microbial processes. Nat Clim Change 3(10):909–912
Wieder WR, Grandy AS, Kallenbach CM, Bonan GB (2014) Integrating microbial physiology and physio-chemical principles in soils with the MIcrobial-MIneral Carbon Stabilization (MIMICS) model. Biogeosciences 11(14):3899–3917
Zhuang Q, Melillo JM, Kicklighter DW, Prinn RG, McGuire AD, Steudler PA, Felzer BS, Hu S (2004) Methane fluxes between terrestrial ecosystems and the atmosphere at northern high latitudes during the past century: a retrospective analysis with a process-based biogeochemistry model. Glob Biogeochem Cycles. https://doi.org/10.1029/2004GB002239
Acknowledgements
This research was supported by the RUBISCO Scientific Focus Area in the Regional Global Climate Modeling Program by the Director, Office of Science, Office of Biological and Environmental Research, of the U.S. Department of Energy under contract DE‐AC02‐05CH11231. JYT and WJR conceived the study, JYT conducted the numerical analysis, JYT and WJR together wrote the paper. The authors declare no competing interest. All data needed to evaluate the conclusions in this paper are present in the paper and in the cited references. Additional scripts and data related to this paper are available upon request from the authors. We also thank two reviewers and the editor Prof. Stuart Grandy to help us improve the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Responsible Editor: Stuart Grandy.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tang, J., Riley, W.J. Linear two-pool models are insufficient to infer soil organic matter decomposition temperature sensitivity from incubations. Biogeochemistry 149, 251–261 (2020). https://doi.org/10.1007/s10533-020-00678-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10533-020-00678-3