Abstract
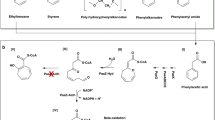
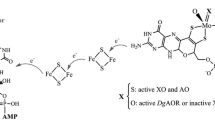
The secondary structure of the trimeric protein 4-chlorobenzoyl coenzyme A dehalogenase from Arthrobacter sp. strain TM-1, the second of three enzymes involved in the dechlorination of 4-chlorobenzoate to form 4-hydroxybenzoate, has been examined. EmM for the enzyme was 12.59. Analysis by circular dichroism spectrometry in the far uv indicated that 4-chlorobenzoyl coenzyme A dehalogenase was composed mostly of α-helix (56%) with lesser amounts of random coil (21%), β-turn (13%) and β-sheet (9%). These data are in close agreement with a computational prediction of secondary structure from the primary amino acid sequence, which indicated 55.8% α-helix, 33.7% random coil and 10.5% β-sheet; the enzyme is, therefore, similar to the 4-chlorobenzoyl coenzyme A dehalogenase from Pseudomonas sp. CBS-3. The three-dimensional structure, including that of the presumed active site, predicted by computational analysis, is also closely similar to that of the Pseudomonas dehalogenase. Study of the stability and physicochemical properties revealed that at room temperature, the enzyme was stable for 24 h but was completely inactivated by heating to 60°C for 5 min; thereafter by cooling at 1°C min−1 to 45°C, 20.6% of the activity could be recovered. Mildly acidic (pH 5.2) or alkaline (pH 10.1) conditions caused complete inactivation, but activity was fully recovered on returning the enzyme to pH 7.4. Circular dichroism studies also indicated that secondary structure was little altered by heating to 60°C, or by changing the pH from 7.4 to 6.0 or 9.2. Complete, irreversible destruction of, and maximal decrease in the fluorescence yield of the protein at 330–350 nm were brought about by 4.5 M urea or 1.1 M guanidinium chloride. Evidence was obtained to support the hypothetical three-dimensional model, that residues W140 and W167 are buried in a non-polar environment, whereas W182 appears at or close to the surface of the protein. At least one of the enzymes of the dehalogenase system (the combined 4-chlorobenzoate:CoA ligase, the dehalogenase and 4-hydroxybenzoyl coenzyme A thioesterase) appears to be capable of association with the cell membrane.







Similar content being viewed by others
Notes
Erratum: The value reported for pI by Zhou et al. (2004) was incorrectly given as pH 6.42. The correct value is 5.70, compared with a predicted value of 5.54 given in UniProtKB/TrEMBL entry 085078 for the dehalogenase of Arthrobacter strain TM-1.
Abbreviations
- CD:
-
Circular dichroism
- 4-CB:
-
4-Chlorobenzoate
- 4-CBCoA:
-
4-Chloro-benzoyl CoA
- GdmCl:
-
Guanidinium chloride
- 4-HB:
-
4-Hydroxybenzoate
- 4-HBCoA:
-
4-Hydroxybenzoyl CoA
References
Benning MM, Taylor KL, Liu R et al (1996) Structure of 4-chlorobenzoyl coenzyme A dehalogenase determined to 1.8 Å resolution: an enzyme catalyst generated via adaptive mutation. Biochemistry 35:8103–8109. Amino-acid sequence from Protein Data Bank, ID code 1NZY
Boehm G, Muhr R, Jaenicke R (1992) Quantitative analysis of protein far UV circular dichroism spectra by neural networks. Protein Eng 5:191–195
Chang K-H, Liang P-H, Beck W et al (1992) Isolation and characterisation of the three polypeptide components of 4-chlorobenzoate dehalogenase from Pseudomonas sp. strain CBS-3. Biochemistry 31:5605–5610
Creighton TE (1989) Disulphide bonds between cysteine residues. In: Creighton TE (ed) Protein structure: a practical approach. IRL Press, Oxford, pp 155–167
Crooks GP, Copley SD (1994) Purification and characterisation of 4-chlorobenzoyl CoA dehalogenase from Arthrobacter sp. strain 4-CB1. Biochemistry 33:11645–11649
Dong J, Xiang H, Luo L et al (1999) Modulating electron density in the bound product, 4-hydroxybenzoyl CoA, by mutation in 4-chlorobenzoyl-CoA dehalogenase near the 4-hydroxy group. Biochemistry 38:4198–4206
Fan Y-X, Zhou J-M, Kihara H, Tsou C-L (1998) Unfolding and refolding of dimeric creatine kinase equilibrium and kinetic studies. Prot Sci 7:2631–2641
Gartemann K-H, Fiedler J, Schmitz A et al (1998) GenBank Accession Number: AF042490
Gartemann K-H, Fiedler J, Schmitz A et al (1999) Hydrolytic dechlorination of 4-chlorobenzoate. Cloning and characterisation of the 4-chlorobenzoate dehalogenase operon from Arthrobacter spp. shows a duplication of the operon, association with a trans-membrane protein, and the occurrence of insertion elements upstream of the operon. Unpublished. NCBI accession no. AAF78820; resubmitted 2001
Guex N, Peitsch MC (1997) SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modelling. Electrophoresis 18:2714–2723
Jones DT (1999) Protein secondary structure prediction based on position-specific scoring matrices. J Mol Biol 292:195–202 (http://bioinf.cs.ucl.ac.uk/psipred)
Löffler F, Lingens F, Müller R (1995) Dehalogenation of 4-chlorobenzoate, characterisation of 4-chlorobenzoyl-coenzyme A dehalogenase from Pseudomonas sp. CBS3. Biodegradation 6:203–212
Marks TS (1986) The microbial degradation of chlorobenzoic acids. Ph.D. thesis, CNAA; Thames Polytechnic, London
Marks TS, Smith ARW, Quirk AV (1984) Degradation of 4-chlorobenzoic acid by Arthrobacter sp. Appl Environ Microbiol 48:1020–1025
McGuffin LJ, Bryson K, Jones DT (2000) The PSIPRED protein structure prediction server. Bioinformatics 16:404–405
Peitsch MC (1995) Protein modeling by E-mail. Bio/Technology 13:658–660
Schmid FX (1989) Spectral methods of characterising protein conformation and conformational changes. In: Creighton TE (ed) Protein structure: a practical approach. IRL Press, Oxford, pp 251–285
Schwede T, Kopp J, Guex N, Peitsch MC (2003) SWISS-MODEL: an automated protein homology-modeling server. Nucleic Acids Res 31:3381–3385
SWISS-MODEL (Automated Protein Modelling Server) Model Ref. No. AAAaOI-Nk for 4-CBCoA dehalogenase from Arthrobacter strain TM-1 (O85078) based on coordinates of PDB 1NZY, 4-CBCoA dehalogenase from Pseudomonas strain CBS-3
Tanford C (1968) Protein denaturation C, Theoretical Models for the Mechanism of Denaturation. Adv Protein Chem 23:121–282
Tanford C (1970) Protein denaturation. Adv Protein Chem 24:1–95
Taylor KL, Liu RQ, Liang PH et al (1995) Evidence for electrophilic catalysis in the 4-chlorobenzoyl CoA dehalogenase reaction: UV, Raman, and 13C-NMR spectral studies of dehalogenase complexes of benzoyl CoA adducts. Biochemistry 34:13881–13888
Taylor KL, Xiang H, Liu R et al (1997) Investigation of substrate activation by 4-chloro-benzoyl coenzyme A dehalogenase. Biochemistry 36:1349–1361
Wang ZX, Wu JW, Tsou CL (1998) The inactivation kinetics of papain by guanidine hydrochloride: a re-analysis. Biochim Biophys Acta – Protein Struct Mol Enzymol 1388:84–92
Xiang H, Dong J, Carey PR et al (1999) Product catalyses the deamidation of D145N dehalogenase to produce the wild-type enzyme. Biochemistry 38:4207–4213
Xiao J, Liang SJ, Tsou CL (1993) Inactivation before significant conformational change during denaturation of papain by guanidine-hydrochloride. Biochim Biophys Acta 1164:54–60
Yu XC, Wang CC, Tsou CL (1994) Association and dissociation of protein disulphide isomerase. Biochim Biophys Acta – Protein Struct Mol Enzymol 1207:109–113
Zhou L, Marks T, Poh PCR et al (2004) The purification and characterisation of 4-chloro-benzoate:CoA ligase and 4-chlorobenzoyl CoA dehalogenase from Arthrobacter sp. strain TM-1. Biodegradation 15:97–109
Acknowledgements
L. Z. thanks the research community of the University of Greenwich for the award of a bursary. Particular thanks are due to Dr. Ronan O’Brien for his valuable assistance in securing CD spectra.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhou, L., Poh, R.P.C., Marks, T.S. et al. Structure and denaturation of 4-chlorobenzoyl coenzyme A dehalogenase from Arthrobacter sp. strain TM-1. Biodegradation 19, 65–75 (2008). https://doi.org/10.1007/s10532-007-9115-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-007-9115-9