Abstract
The distribution of marine sponges in the tropical Southwest Pacific Ocean is largely unexplored despite the vital ecological role of sponges in coral reefs and their value as sources of metabolites for drug design. Several collection campaigns to the French Polynesian archipelagos (Society, Marquesas, Tuamotu, Gambier, and Austral) were conducted to assess the bio- and chemodiversity of the island groups. In the course of these scientific expeditions, more than 200 identified sponge specimens were acquired, for which we were able to assign 102 Molecular Operational Taxonomic Units (MOTUs). Based on these MOTUs, we assessed, in the largest analysis of its kind for this area to date, the sponge composition and faunistic overlaps of the marine province Southeast Polynesia with Marquesas and Central Polynesia. We also compared the sponge fauna of these Eastern Indo-Pacific provinces with marine provinces of the adjacent Central Indo-Pacific realm. Our findings corroborate that sponge faunal similarity within marine realms is higher than among realms, and follows the marine barriers to gene flow observed for other taxa. We detected high levels of provincial endemism for marine sponges, consistent with findings from other Indo-Pacific regions. At the level of province, geographical distance and ocean surface currents influence faunal similarity, and constitute the primary factors for the connectivity of sponge faunas between the disjunct and remote island groups in the tropical Southwest Pacific Ocean.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The islands and archipelagos in the Eastern Indo-Pacific host a multitude of diverse terrestrial and marine environments, often still largely undisturbed by humans. Their faunal and floral assemblages are highly diverse, originating both from Australasia and the Americas, with ocean currents, geological history and recent human influence shaping the unique faunal composition of the different islands (Gillespie et al. 2008; Neall and Trewick 2008; Aswani and Allen 2009; Hall et al. 2013).
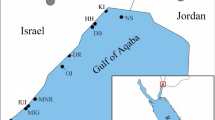
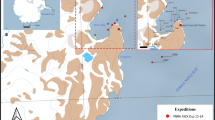
The 118 islands and atolls of French Polynesia are among the most isolated regions on Earth, and with only about half of them being inhabited, large parts of their natural environments remain pristine (Lecchini et al. 2021). The 2000 km region spanning French Overseas Collectivity is divided into the five main archipelagos of Society (Windward & Leeward Islands), Tuamotu, Marquesas, Austral, and Gambier. The region hosts a wide variety of endemic terrestrial and marine lifeforms (e.g., Delrieu-Trottin et al. 2015; Petek et al. 2017; Ramage 2017) (Fig. 1). As defined by the Marine Ecoregions of the World (MEOW, Spalding et al. 2007), a hierarchical classification of marine regions into realms, provinces and ecoregions, these island systems form the two Marine Provinces (“MP”s) Southeast Polynesia and the MP Marquesas within the Eastern Indo-Pacific realm.
Map of French Polynesia (Eastern Indo-Pacific realm), covering MPs Southeast Polynesia and Marquesas (highlighted in red), displaying major island groups and other notable islands. Border colors reflect positions of map and inset on miniature globe on the top right. Inset: Detail of Wallis and Futuna islands with the adjacent Samoa archipelago in the MP Central Polynesia (highlighted in blue).
The islands of Wallis and Futuna constitute a further French Overseas Collectivity in the Eastern Indo-Pacific realm. They are located west of Samoa (Fig. 1), as part of the MP Central Polynesia. Wallis and Futuna adjoin the border between the Central Indo-Pacific realm and the Eastern Indo-Pacific realm, making them a junction for faunal exchange in the region and emphasizing their significance for biodiversity studies.
While the biodiversity of many reef ecosystems in the Eastern Indo-Pacific realm has been surveyed in several independent studies, most of the research has focused on molluscs (Legendre and Salvat 2015), corals (Pratchett et al. 2013), and fishes (Galzin et al. 1994). Despite other formerly underrepresented organismal groups with important roles in the maintenance of reef ecology, like sponges (Phylum Porifera), having been explored more thoroughly in the past decade, they still remain understudied in the Eastern Indo-Pacific realm in comparison (e.g., Van Soest et al. 2012; Hall et al. 2013; Petek et al. 2017). The ability of sponges to cycle abundant dissolved organic matter into particulate organic matter, which then becomes available to higher trophic levels, is a key component of the nutrient cycle in oligotrophic coral reef ecosystems (de Goeij et al. 2013; Pawlik and McMurray 2020). Likewise, their capacity to substantially alter different types of marine substrates contributes to the shaping of reefs, while also acting as important providers of micro- and macrohabitats (Bell 2008; Maldonado et al. 2015; Folkers and Rombouts 2020). Sophisticated biochemical defense mechanisms present in many sponges constitute an important source of marine bioactive compounds with a large potential for novel pharmaceutical applications, making bioprospecting for these compounds another major source of data on global sponge distribution and biodiversity (e.g., Amade et al. 1982; El-Demerdash et al. 2019; Galitz et al. 2021). One of the most challenging aspects of chemo- and biodiversity studies on sponges is frequently uncertain or unreliable identification of the sponges due to phenotypic plasticity and convergent characters (see Erpenbeck et al. 2006; Andreakis et al. 2012; Galitz et al. 2021). In most cases molecular data are indispensable for the correct identification and delineation of ambiguous sponge taxa, although major taxonomic challenges still apply in certain orders, e.g., Haplosclerida (López-Legentil et al. 2010; DeBiasse and Hellberg 2015; Vicente et al. 2019).
Hall et al. (2013) analyzed the biodiversity of sponges of the Society and Marquesas Islands based on 75 morphospecies, but data on the other island groups of French Polynesia remain comparatively sparse, with the Tuamotu archipelago, Austral Islands and Gambier Islands in particular being understudied (see also: Moorea, Society Islands: Freeman and Easson 2016; Calcarea only: Klautau et al. 2020; Society Islands, Marquesas (submarine caves): Schuster et al. 2021). To date, only 16 valid sponge species have been confirmed for MP Marquesas, with an additional 39 species for the remainder of the numerous French Polynesian islands in MP Southeast Polynesia, and another 14 species known from MP Central Polynesia (see de Voogd et al. 2023). These numbers appear to be exceptionally low, when compared to the recorded sponge species biodiversity in some of the marine provinces of the Central Indo-Pacific realm (Tropical Southwestern Pacific: 412 species; Sahul Shelf: 321 species; Northeast Australian Shelf: 321 species). This vast and remote geographic area of French Polynesia with its multitude of different reef ecosystems might be prone to biases, such as incomplete sampling or the omission of cryptic (i.e., reef matrix) habitats. Consequently, additional data are needed to obtain a more comprehensive understanding of the sponge fauna in the Eastern Indo-Pacific realm and of the extent of sponge biodiversity in this region (Van Soest et al. 2012).
In this study, we report on the sponge fauna of Tuamotu and other major French Polynesian archipelagos, as collected during different campaigns in the area between 2009 and 2013, and for the first time, with an emphasis on molecular taxonomy. We aim to investigate the molecular biodiversity of demosponges (Class Demospongiae) in the Eastern Indo-Pacific realm, understand the connectivity of its marine provinces to those of the adjacent Central Indo-Pacific realm, and discuss the results in relation to previous findings from these regions. This study aims to fill gaps in the knowledge of biodiversity and species distribution of Indo-Pacific sponges, and to provide information necessary for the adoption of appropriate measures to increase effectivity of species conservation and bioprospecting (Glover et al. 2018).
Materials and methods
The sponge material processed in this study was collected in the course of multiple scientific expeditions to the French Polynesian archipelagos, and had been morphologically described and classified into Queensland Museum morpho-OTUs (QM####) (here also referred to as “morpho-IDs”) in Hall et al. (2013) and Petek et al. (2017). The present study aimed to generate independent corresponding molecular taxonomic units (referred to as MOTUs in the following text) from the available sponge material, for the use in biodiversity and phylogenetic analyses, which were not necessarily restricted to the previously assigned morpho-IDs. Specimens with conflicting results between morpho-ID and subsequent MOTU underwent additional morphological re-identification, where possible.
Sponge material
In total, 371 samples (Supplementary Table S1) were collected by SCUBA in the course of different collecting campaigns in the Eastern Indo-Pacific. The campaigns aimed to assess the bio- and chemodiversity of marine invertebrates around the Polynesian archipelagos and islands. The applied sampling strategy focussed on the collection of macroscopic sponges on the reef surface, consequently cryptic spaces, such as overhangs, caves or internal reef matrix, were not explicitly sampled in the course of these expeditions.
Surveys of the Marquesas Islands were conducted in 2009 (BSM-PF1 expedition Leg 2, Debitus et al. 2009) and 2011 (Pakaihi I Te Moana cruise Leg 2), while during the Tuam 2011 expedition, Tuamotu and a part of the Society archipelago were sampled (Debitus et al. 2011). Note that for samples and data from Marquesas Islands primarily sponge material with relevancy for biochemical studies was to our disposal, consequently deviating from the sampling scheme of other expeditions, and resulting in a specimen bias of a numerical overrepresentation of a small number of species (see Supplementary Table S1). Further samples from the Society Archipelago were obtained during the Tahiti Iti expedition (Debitus et al. 2013b), during surveys for Hall et al. (2013), and Leg 1 of the BSM-PF1 expedition (Debitus et al. 2009). Additional specimens from the Austral and Gambier islands were collected during the Coralspot and Tuhaa Pae (Debitus et al. 2013a) surveys respectively. Most of the sponge specimens collected in the course of these expeditions and used in this study are also depicted in Petek et al. (2017), with relevant metadata, classification into (morphological) OTUs, and, for some, morphological descriptions. Comparative sequences of sponge specimens from MP Central Polynesia (Wallis and Futuna), as collected in 2016 during the Tara Pacific expedition (Futuna, see Planes et al. 2019) and the WALLIS 2018 expedition (Petek et al. 2018), were added to this study. The aim of these scientific expeditions was to investigate the chemo- and biodiversity of those islands, which will be covered in greater detail in a following study.
DNA extraction, amplification, and sequencing
For DNA extraction, the protocol for PALL plate extraction was used, which allows for fast processing of large amounts of samples (Vargas et al. 2012). Widely used barcoding primers for fragments of the C-region of the nuclear ribosomal large subunit (28S) (Chombard et al. 1998) and the cytochrome oxidase subunit 1 (CO1) standard barcoding fragment (Meyer and Paulay 2005) were used in the PCR amplification process of the samples (Table 1). The combination of these two markers (ribosomal and mitochondrial) has been successfully employed in previous biodiversity studies on the sponge communities of other Indo-Pacific sponge faunas (Erpenbeck et al. 2016, 2020). In this study we primarily focused on the use of 28S sequences for MOTU generation, biodiversity analyses, and supplemental phylogenetic tree reconstruction, due to their higher phylogenetic resolution in most sponge taxa. The comparatively more conservative mitochondrial CO1 marker was employed to confirm the generated 28S sequences and identify potential ambiguities.
PCR reactions were performed in volumes of 25 µL, comprising 5 µL 5× green GoTaq ® PCR Buffer (Promega Corp, Madison, WI), 4 µL 25 mM MgCl2 (Promega Corp, Madison, WI), 2 µL 10 mM dNTPs, 1 µL of primer (5 µM), 2 µL BSA (100 µg/ml), 7.8 µL water, 0.2 µL GoTaq® DNA polymerase (5u/μL, Promega Corp, Madison, WI), and 2 µL DNA template per reaction.
Amplification followed established temperature profiles of initial denaturation (3 min/95 °C), followed by 35 cycles of 95 °C/30 s (denaturation), followed by 51 °C (28S) or 40 °C (CO1) for 30 s (annealing) and 72 °C/1 min (extension), concluded by a final extension step at 72 °C for 5 min (Erpenbeck et al. 2016). The examination of PCR products was conducted on a 1% TAE agarose gel stained with peqGREEN (peqlab) fluorescence dye. The processed products were subsequently sequenced with a BigDye® Terminator v3.1 (Applied Biosystems®) following the manufacturer’s protocol at the Sequencing Service of the Department Biology, LMU—Genomics Service Unit (Martinsried, Munich, Germany) on an ABI 3730 capillary sequencing machine. Raw sequences were basecalled and trimmed in CodonCode Aligner v9.0.4 (www.codoncode.com), and assembled in Geneious Prime 2019 (v2019.2.3) (www.geneious.com). Individual sequences were checked for intragenomic polymorphisms, which were subsequently corrected to corresponding IUPAC ambiguity codes and disregarded in the subsequent biodiversity analyses. To check for contamination and to pre-classify unidentified samples, sequences were compared against the databases of the European Nucleotide Archive (ENA) using BLASTn (Altschul et al. 1990), and against NCBI Genbank. Verified Porifera sequences were further processed and corrected in Geneious. Final sequences are deposited in the European Nucleotide Archive (ENA) database under the accession number range OX451516-OX451734 and the Sponge Barcoding Project (www.spongebarcoding.org) (Wörheide and Erpenbeck 2007) (SBD#2345-2565).
Phylogenetic analyses and taxonomic evaluation
Sequences were aligned using the MAFFT v7.450 (Katoh and Standley 2013) plugin for Geneious Prime® (v2019.2.5) and applying the FFT-NS-i x2 alignment algorithm. Comparative sponge sequences from other marine provinces were added from the Sponge Barcoding Project. These included MP Central Polynesia (Galitz et al. unpublished), and three further MPs of the adjacent Central-Indo Pacific realm (MPs Tropical Southwestern Pacific, Sahul Shelf, and Northeast Australian Shelf). Maximum-likelihood reconstructions were generated with RAxML 8 (Stamatakis 2014) as implemented in Geneious Prime® 2019.2.5 under the GTR GAMMA I model as suggested by Modeltest 2.1.10 (Darriba et al. 2012) and 1000 rapid bootstrap replicates.
Molecular Biodiversity analyses
For the biodiversity analysis, 28S sequences, which comprised the largest set of Central Indo-Pacific sequences for comparison, were divided into their respective MPs sensu Spalding et al. (2007) using QGIS 3.10. (QGIS Development Team 2019, https://qgis.org/). While CO1 sequences were generated in addition to 28S, the smaller size of the available mitochondrial datasets from the studied MPs, and lower resolution compared to the nuclear marker, made the CO1 sequences less suitable for biodiversity analyses. Consequently, in the following “MOTU” will always refer to 28S sequences as their origin. Sequences longer than 350 bp were aligned with ClustalW (Thompson et al. 1994) as implemented in the msa package for R (Bodenhofer et al. 2015). DECIPHER 2.0 (Wright 2016) was used to cluster sequences into respective MOTUs. MOTUs will be used as the molecular equivalent to the previously assigned morpho-IDs, and in this context as baseline for molecular biodiversity assessments. Note that MOTUs are inherently independent from their original morpho-IDs, with conflicting results, however, being further investigated. To delineate MOTUs, we used the unweighted pair group method with arithmetic mean (UPGMA) algorithm (see Cowman et al. 2017; Hadiyanto et al. 2021), which has been shown to have the best performance in hierarchical cluster analyses (Kreft and Jetz 2010). The threshold for 28S MOTU differentiation was set to 0.3% base pairs difference over a minimal sequence length of 350 bp in this fragment, equal to a genetic difference of ≤ 1 bp, which follows Erpenbeck et al. (2016, 2020), but accounts for additional, undetected intragenomic polymorphisms. Since false MOTUs may occur due to undetected sequencing errors, despite careful basecalling and sequence quality control, we also repeated the analysis with single-specimen-MOTUs (= singletons) omitted for additional quality control.
Biodiversity analyses were performed in R v4.1.1 using the packages VEGAN (v4.2.4), picante (v1.8.2), and iNEXT (Dixon 2003; Kembel et al. 2010; Chao et al. 2016; R Core Team 2021). Visualization of MOTU overlap or uniqueness between regions was conducted using the package UpSetR (Conway et al. 2017). For biodiversity analyses within and between regions commonly used indices were computed, with additional Chao1 species richness and Pielou’s evenness estimates (Pielou 1966; Chao 1984): Local (alpha) biodiversity is represented by Shannon–Wiener and Fisher’s alpha indices, although differences in sampling strategy, sample size and sampling biases might make Shannon values less suitable for comparing one area’s regional biodiversity to another. Fisher’s alpha should be less sensitive in this regard (Beck and Schwanghart 2010). Beta diversity between the selected marine provinces was calculated by the Jaccard dissimilarity index (Jaccard 1912). Beta diversity estimates can be sensitive to differences in sample size. Among the plethora of beta-diversity indices, the Jaccard index is discussed as comparatively invulnerable to errors of taxonomy and enumeration, and with low error rates in geographic undersampling (see e.g., Williams 1949; Pos et al. 2014; Schroeder and Jenkins 2018; Hadiyanto et al. 2021). The significance of the calculated beta diversity indices was subsequently verified by applying a pairwise PERMANOVA (permutational multivariate analysis of variance) using a custom variation of the adonis function of VEGAN for pairwise comparison of regions, with 999 permutations and adjustments for false discovery rate (FDR) taken into account (Anderson 2001, 2017). In addition to the computation of biodiversity indices, rarefaction analyses were conducted to assess and extrapolate both the species richness and the sampling completeness of the respective provinces. These were computed and visualized with iNEXT (Chao et al. 2016).
Morphological species identification
For supplemental morphological re-identification of conflicting results between prior morphological analyses and molecular genetic data generated in the course of this study, identical analytical procedures to Hall et al. (2013) and Petek et al. (2017) were conducted where necessary:
For morphological analyses and subsequent categorization into morphospecies and morpho-IDs of the collected Queensland Museum specimens, the fresh sponge material was immediately preserved in 70% ethanol. For light microscopy analyses, thin sections through the ectosome and choanosome of the tissue were cut with a scalpel. The sections were cleared in phenol-xylene overnight and embedded in Fluka Durcopan™ (Sigma-Aldrich Co., St. Louis, MO, USA). Additional preparations of the spicules were made for light microscopy by digesting small portions of sponge in nitric acid; the sponge-acid mix was heated over a flame and the remaining spicules mounted in Canada balsam.
Separate preparations of fibers were conducted by the addition of 12.5% sodium hypochlorite to remove soft tissue. The dissolution of tissue was monitored and facilitated by the removal of excess collagen with forceps, with subsequent neutralization of the reaction with distilled water, and finally two-fold rinsing in 70% ethanol and then 100% ethanol. Fibers were examined using an Olympus SZ60 dissection microscope with a Tucsen 3.0 camera. The fixed section and spicule microscope slides were examined using an Olympus BH2 with an optical stage micrometer and photographed with a Nikon CoolPix 5400 mounted camera. Preserved specimens were photographed with a Canon G5X. Surface characteristics were examined using an Olympus SZ60 dissection microscope with a Tucsen 3.0 camera.
Spicule preparations for Scanning Electron Microscope (SEM) were made by dissolving the tissue in 12.5% sodium hypochlorite for the removal of soft tissue, followed by neutralization in distilled water, rinsing twice in 70% ethanol, and finally rinsing twice in 98% ethanol and air-drying the preparations. SEM preparations were sputter-coated in gold to improve resolution. The scanning electron micrograph photos and measurements were made using a Hitachi TM-1000 SEM.
Results
Sequencing and species identification
From a total of 371 available sponge specimens from the Eastern Indo-Pacific MP Marquesas and MP Southeast Polynesia, 242 28S (65.2%) and 196 CO1 (52.8%) sequences were successfully obtained, with 137 specimens being covered by both fragments (36.9%). Maximum length of complete sequences was 536 bp for 28S (shortest fragmented: 264 bp) and 682 bp for CO1 (shortest fragmented: 288 bp). Of the initial 371 sponge samples, 327 specimens (88.1%) were morphologically identified to species, or at least to genus level, corresponding to 130 different morpho-IDs and 104 MOTUs respectively.
Rarefaction analyses of the MP Southeast Polynesia samples indicated high species diversity and high potential for yet undiscovered species (Fig. 2). The MP Marquesas was not included in the final rarefaction analysis, as part of the MP Marquesas specimen data were primarily forwarded for biochemical studies, which deviates from the sampling scheme of the other regions included in this study, which is also reflected in the low evenness value for MP Marquesas (Table 2). Due to this specimen bias of a numerical overrepresentation of a small number of species (i.e., Ptilocaulis sp. (QM1640), Monanchora sp. (QM4696), Suberea sp. (QM2093), Suberea ianthelliformis; see Supplementary Table S1) in the available Marquesas data subsets, they were unsuitable for further biodiversity evaluation (see “Discussion”).
Biodiversity analyses
After the addition of comparative material published in the Sponge Barcoding Project (430 28S sequences) from the adjacent MPs Northeast Australian Shelf, Tropical Southwestern Pacific, Sahul Shelf, and Central Polynesia, the final dataset comprised a total of 966 sequences, of which 894 sequences exceeded the predefined threshold of 350 bp and were included in the following analysis steps. The total for unique MOTUs across the entire dataset amounted to 388, of which singletons accounted for 61.3% (238) (Table 2, see Table S1 in supplementary material for a complete list of MOTUs). The number of MOTUs per province strongly differed between 13 MOTUs from MP Marquesas to up to 104 MOTUs from MP Central Polynesia. The ratio of MOTUs unshared with other provinces (= endemic in the dataset) ranged between 63.9% (MP Tropical Southwestern Pacific) and 86.5% (MP Sahul Shelf) (Table 2). 179 MOTUs from the dataset are exclusive to the Eastern Indo-Pacific realm (MP Central Polynesia = 84, MP Southeast Polynesia = 76, MP Marquesas = 9). They comprised 46.1% of the total 388 unique MOTUs in this study.
Within the Eastern Indo-Pacific realm, the largest MOTU overlap between marine provinces was between MPs Southeast Polynesia and Central Polynesia (7 MOTUs), followed by MPs Southeast Polynesia and Marquesas (2 MOTUs), and no shared MOTUs between MPs Central Polynesia and Marquesas. Only one MOTU was shared between all three provinces (Figs. 3, 4). The remaining 169 MOTUs were not shared with any other province of the Eastern Indo-Pacific realm. The MOTUs restricted to the Central Indo-Pacific realm amounted to 193 (49.7%) (Fig. 4). In total, only 16 (4.1%) MOTUs were shared between the Central Indo-Pacific realm (= MPs Northeast Australian Shelf + Tropical Southwestern Pacific + Sahul Shelf) and the Eastern Indo-Pacific realm (= MPs Central Polynesia + Southeast Polynesia + Marquesas).
MOTU count (incl. singletons) per marine province and overlap between provinces. Histogram on the left: Number of MOTUs per MP. To its right: Black dots without connecting lines represent the number of unique MOTUs in a MP, while dots connected by lines indicate the number of MOTUs present in two or more MPs. Respective numbers of MOTUs for each scenario depicted with histogram on the top. Abbreviated marine province names (MPs) adopted from Spalding et al. (2007): Marquesas (MAR), Southeast Polynesia (SEP), Central Polynesia (CEP), Tropical Southwestern Pacific (TSP), Sahul Shelf (SAH), Northeast Australian Shelf (NEA)
a Number of shared and unshared MOTUs as unique to the MPs Central Polynesia, Marquesas and Southeast Polynesia within the Eastern Indo-Pacific realm. Graphical overlap between Marquesas and Central Polynesia removed due to the absence of shared MOTUs; b Left: MOTU overlap between the Central Indo-Pacific realm and Eastern Indo-Pacific MPs Central Polynesia, Southeast Polynesia + Marquesas shown separately. Right: MOTU overlap between Central and Eastern Indo-Pacific realms only. Circle sizes relative to absolute MOTU numbers. Created with BioVenn (Hulsen et al. 2008)
The Eastern Indo-Pacific realm MP with the highest Central Indo-Pacific realm MOTU overlap was MP Central Polynesia (Agelasida: 3; Dictyoceratida: 2; Scopalinida: 1; Tetractinellida: 1; Verongiida: 1; Biemnida: 1), compared to MP Southeast Polynesia (Agelasida: 1; Biemnida: 1; Haplosclerida: 1) and MP Marquesas (Verongiida: 1) (Figs. 3, 4). These respective MOTUs were not shared with another MP of the Eastern Indo-Pacific realm, and consequently constitute independent faunal links to the Central Indo-Pacific realm, with the genera Agelas (4 MOTUs) and Suberea (2 MOTUs) being the only ones representing their order with more than one species.
The alpha biodiversity analysis results of the studied MPs displayed comparatively high values for the Shannon–Wiener index, largely coinciding with the respective estimated Chao1 species richness, with the lowest in the MP with the smallest total MOTU numbers (MP Tropical Southwestern Pacific, Table 2). Fisher’s alpha analysis results indicate generally high biodiversity, but show strong variations between provinces, with no clear relationship to location, richness estimates, or numbers of specimens or MOTUs. Despite the varying results of the alpha diversity analyses, evenness across the studied MPs appears high, displaying values > 0.9 for all datasets. Indices for MP Marquesas are excluded in this assessment due to the above-mentioned specimen bias (See “Materials and methods”).
The beta diversity analyses recovered Jaccard dissimilarity indices between the studied provinces mostly ranging between 0.95 and 1.0, coinciding with the low number of MOTU overlaps between the MPs of the study area and high endemism. All p values (FDR adjusted) of Jaccard indices were recovered as being highly significant (Fig. 5). In a few instances, dissimilarity values were below 0.95: between MPs Central Polynesia and MP Southeast Polynesia in the Eastern Indo-Pacific realm (11 MOTUs), and also between MPs Northeast Australian Shelf and Tropical Southwestern Pacific in the Central Indo-Pacific realm (10 MOTUs) (Fig. 3), reflecting a relatively higher overlap compared to the other regions (see also Figs. 5, 6). The majority of overlapping OTUs belonged to sponges of the orders Verongiida, Dictyoceratida and Haplosclerida in both cases, however no genera stood out as prominently represented for their respective orders. Higher values of similarity between MPs are in line with the biogeographical classification on the realm level, with highest congruence between the faunas in the MPs of the Eastern Indo-Pacific realm and of the Central Indo-Pacific realm respectively (Fig. 5). Similarity values on the province level may not (always) reflect the affiliation of an MP with their respective realm, as geographical distance (generally) correlated well with lower Jaccard indices.
Values on bottom right of the pairwise comparison diagram indicate 28S gene sequence Jaccard dissimilarity (in percent) of the studied marine provinces; higher values correspond to lower faunistic similarity between MPs, with 100 representing no overlap of MOTUs; graduated in 1.5% increments. Values on top right of the pairwise comparison diagram indicate false discovery rate (FDR) adjusted p values of respective 28S Jaccard indices generated through pairwise PERMANOVA. Values for MP Marquesas under reserve due to potential distortion by specimen bias. The dendrogram on the left depicts faunal similarity between two given regions based on their respective Jaccard indices; numbers on branches are bootstrap replicates. Background color corresponds to Central Indo-Pacific marine realms (yellow) and Eastern Indo-Pacific marine realms (green). Abbreviated MP names: Marquesas (MAR), Southeast Polynesia (SEP), Central Polynesia (CEP), Tropical Southwestern Pacific (TSP), Sahul Shelf (SAH), Northeast Australian Shelf (NEA)
Geographical plot of Jaccard dissimilarities according to values in Fig. 5. Coloured lines display the degree of dissimilarity between the MPs as denoted in the legend; higher Jaccard-indices correspond to lower faunistic similarity between MPs, with 100 representing no overlap of MOTUs; graduated in 1.5% increments. Marine realms of the Central Indo-Pacific (yellow) and Eastern Indo-Pacific (green) are highlighted. Abbreviated MP names: Marquesas (MAR), Southeast Polynesia (SEP), Central Polynesia (CEP), Tropical Southwestern Pacific (TSP), Sahul Shelf (SAH), Northeast Australian Shelf (NEA). Dashed lines to MP Marquesas (MAR) indicating a data/specimen bias, and consequently a potential ambiguity of the respective results (see “Discussion” for details)
Discussion
Our results reveal high biodiversity and high levels of endemism among the previously largely unknown sponge faunas of the investigated Eastern Indo-Pacific MPs, but also highlight the substantial differences between marine provinces regarding species richness and composition. Most of these differences appear to be independent of geographical distance, but rather are influenced by factors like local geography, sea surface currents, and animal biology which shape the regional biodiversity (for similar results of other marine taxa see also Briggs and Bowen 2012, 2013; Kulbicki et al. 2013; Veron et al. 2015; Crandall et al. 2019).
Molecular taxonomic diversity within MPs Southeast Polynesia and Marquesas
MP Southeast Polynesia: Our MP Southeast Polynesia data (92 MOTUs) exceeds the study of Hall et al. (2013, 41 (morpho-)species), by inclusion of Austral, Gambier, and Tuamotu samples. While Hall et al. (2013) exclusively discussed samples from the Society archipelago, our MP Southeast Polynesia data comprises a wider geographic range, in which now 53% originated from French Polynesian archipelagos other than Society Islands (77 of 146).
Our data display similarities in taxonomic composition of the MP Southeast Polynesia sponge fauna to those of Hall et al. for Society Islands. Dictyoceratida (36 MOTUs) constitute the dominant sponge order, followed by Haplosclerida (18) and Verongiida (10). Such dominance of Dictyoceratida corroborates taxonomic studies across the Indo-Pacific, including the Western Indian Ocean (e.g., Erpenbeck et al. 2016, 2020), and can therefore be regarded as a general pattern for shallow water tropical reefs. It is interpreted as the consequence of a complex interaction of biotic and abiotic factors favoring the occasionally photosymbiotic Dictyoceratida, such as types of substrate, exposure to light and ocean currents, as well as individual reproductive and dispersal potential over short and long distances (see for details Wilkinson 1988; Duckworth et al. 2008; Wulff 2012).
MP Marquesas: In contrast to the number of morphospecies (38; incl. 15 verified to species level) from MP Marquesas reported in Hall et al. (2013), the data of our molecular study recovers a lower number of MOTUs (13 MOTUs). The taxonomic composition of our MP Marquesas MOTUs differs from Hall et al. (2013) in that Poecilosclerida, Verongiida and Haplosclerida (4 each) account for the majority of MOTUs. This large disparity is mainly caused by an apparent specimen bias of the available data subsets from the respective Marquesas expeditions, with a numerical overrepresentation of a small number of species (e.g., BSM-PF1) (see “Materials and methods”). The bias is reflected in the skewed ratio of molecular MOTUs among the three Eastern Indo-Pacific realm MPs Marquesas [~ 6.7% of the total 195 MOTUs (including MOTUs shared with all MPs)], Southeast Polynesia (~ 47.2%), and Central Polynesia (~ 53.3%), despite comprising 15.6% of the total specimen numbers from this realm.
Our molecular phylogenetic analyses initially also recovered several morphospecies (sensu Petek et al. 2017) as potential cryptic species based on non-monophyletic occurrences in the tree. After careful re-examination of the original material, it was possible to assign new morpho-IDs to the majority of these assumed cryptic species based on minute differences of morphological characters. The ambiguous species complexes in question comprise a divergent clade of Monanchora sp. (QM4696) in Poecilosclerida, now regarded as Poecilosclerida indet. (SNSB-BSPG.GW9929, GW9930, GW9933), due to morphological ambiguities. Likewise, a seemingly cryptic complex of Spongia sp. (QM1983) in Dictyoceratida could be resolved into Spongia sp. (QM4490) and Hyattella sinuosa. Lastly, a single verongiid Suberea sp. (QM2093) exhibited molecular identity to Suberea sp. (QM2121), grouping apart from the majority of other Suberea sp. (QM2093) specimens. Initial morphological re-identification yet did not recover morphological differences but will be continued (See Supplementary Table S1, Fig. S2). In this respect, molecular data repeatedly had revealed cryptic speciation in sponges of the Indo-Pacific (e.g., Wörheide et al. 2002, 2008; Andreakis et al. 2012) and particularly morpho-OTU based taxonomic assessments (i.e., prior to the full description of a species, e.g., Hall et al. 2013) can be prone to overlooked cryptic species (Pöppe et al. 2010).
An important, but frequently ignored component of biodiversity of benthic organisms in tropical reefs is the cryptofauna living within the reef matrix, which sponges are major occupants of (Richter et al. 2001). Novel metabarcoding approaches on its cryptobenthic community in Hawaiʻi complemented traditional collection and barcoding efforts and significantly increased the taxonomic knowledge on Eastern-Indo Pacific cryptofauna and its differences to the benthic epifauna (Vicente et al. 2021; Timmers et al. 2022). These studies on cryptofaunal diversity highlighted once more the vast taxonomic potential that is hidden within the reef, which traditional sampling efforts, as applied in our study, cannot assess.
Evaluation of differences in local (alpha) biodiversity
The studied MPs of the Central and Eastern Indo-Pacific realms display high local biodiversity, as implied by their alpha diversity indices and richness estimates, with values comparable to other studies investigating the diversity of Indo-Pacific sponge faunas (e.g., Powell et al. 2014; Fromont et al. 2016; Rovellini et al. 2019). Shannon–Wiener indices for most MPs exceed values of 4, with MP Tropical Southwestern Pacific displaying the lowest value of ~ 3.5, and MP Marquesas exhibiting a specimen bias (Table 2). The notable drop for MP Tropical Southwestern Pacific coincides with the dataset for this province being the smallest, which can pose a bias for the Shannon–Wiener index calculation, despite the relative evenness calculated for all unbiased MPs. Alpha biodiversity values of the Fisher’s index display larger variations between MPs, with values ranging from ~ 57.8 (MP Central Polynesia) up to 113.4 (MP Sahul Shelf), and MP Marquesas again being a distinct outlier. The Fisher’s alpha values locally deviate from the predicted Shannon–Wiener and Chao1 indices, suggesting a comparatively lower biodiversity for MPs Central Polynesia and Northeast Australian Shelf instead (Table 2). The large provincial variations, however, also imply the possibility of index bias due to differences in the respective dataset structures, as the different collections were not initially conducted with detailed local or large scale biodiversity estimation in mind.
Indices measuring the alpha biodiversity can be beneficial to determine an estimation of the local species richness of a given region, but they can also have some major drawbacks and variance depending on the index used and the type and quality of data provided for the respective locality. Shannon–Wiener indices in particular have to be interpreted carefully, as coupling of species richness and evenness is discussed as an increasingly inadequate measure of actual biodiversity (Strong 2016; MacDonald et al. 2017). Notable outliers in our results of Shannon–Wiener and Fisher’s alpha indices as assessed for MP Marquesas (Table 2) are due to the high number of specimens with concurrently low taxonomic diversity, with a single genus (Suberea) making up almost half of the total specimens. Considering the respective biodiversity indices of MP Southeast Polynesia and other MPs (Table 2), these values are unlikely to represent accurate biodiversity, but can be attributed to the large proportion of mono-specific specimens present in the available data subset of the BSM-PF1 expedition, while sponges of the AMP-Marquises expedition only represent 10% of the total specimen count from Marquesas.
Conclusively, the large variations in specimen and MOTU counts between the regions make an objective comparison of alpha biodiversity difficult. Despite high evenness for the majority of datasets, the variances of both Shannon–Wiener and Fisher’s alpha biodiversity indices may signify a lack of sampling completeness, which is supported by rarefaction projections and richness estimations of the respective MPs (Table 2).
Sponge beta diversity in Central- and French Polynesia
The Jaccard dissimilarity values indicate low MOTU overlap between the marine provinces studied, with most provinces displaying dissimilarity values > 95 and consequently emphasizing high provincial endemism (Fig. 5). Similar levels of endemism in sponge faunas have been found during molecular biodiversity assessments of the Red Sea and Persian Gulf sponge fauna (Erpenbeck et al. 2016, 2020); these studies corroborated restricted species distribution in sponges as earlier detected in taxon-level studies (Wörheide et al. 2008; Pöppe et al. 2010; Xavier et al. 2010; Reveillaud et al. 2011; Setiawan et al. 2015; Erpenbeck et al. 2017).
Despite the Jaccard index being less prone to certain types of common biases (see “Materials and methods”), it is not invulnerable to biased sampling or datasets, as seen in the results for MP Marquesas. With MP Marquesas disregarded, we find Jaccard dissimilarity values among MPs of the same marine realm mostly lower than inter-realm (e.g., MPs Central Polynesia to Southeast Polynesia; MP Northeast Australian Shelf to Tropical Southwestern Pacific or Sahul Shelf, Figs. 5, 6), which supports the biogeographic classification of Spalding et al. (2007). Our findings also support the morphospecies multivariate clustering of Hall et al. (2013), that found the sponge faunas of the Eastern Indo-Pacific MPs Southeast Polynesia and Marquesas closer to each other compared to Central Indo-Pacific provinces.
We recover a comparatively strong molecular overlap for Eastern Indo-Pacific's MP Central Polynesia with Central Indo-Pacific realm provinces with shared MOTUs unshared with any other MP of the Eastern Indo-Pacific realm (See Fig. 4). Orders Agelasida (Agelas spp.) and Dictyoceratida constitute the majority of shared MOTUs. The apparent abundance of Agelasida MOTUs in both MP Central Polynesia and the Central Indo-Pacific realm is notable, given that the genus Agelas is yet unreported for MP Southeast Polynesia (Petek et al. 2017).
Consequently, our results indicate that the sponge fauna of MP Central Polynesia, situated as a geographic midpoint between the realms of Central and Eastern Indo-Pacific, is comparatively strongly influenced by the sponge faunas of adjacent MPs of both realms, like Southeast Polynesia and Tropical Southwestern Pacific, turning it into a small-scale “melting pot” of regional sponge species biodiversity.
Influence of ocean surface currents and geography on sponge dispersal and distribution
Geographical proximity between marine provinces generally correlates well with faunal similarity according to our data. However, our results also show that the similarity of sponge faunal assemblages between different biogeographical regions in the Central and Eastern Indo-Pacific realms is dependent on the scale of the spatial biogeographical units examined. Computed relatedness of MPs generally corresponds well to their respective marine realm (Fig. 5), however in some cases similarities between singular MPs may be higher across realms than within them, e.g. for MPs Tropical Southwestern Pacific, Northeast Australian Shelf and Central Polynesia, when compared to their connectivity to Sahul Shelf and, under reserve, lower faunal similarity between Marquesas and Central Polynesia, when compared to Tropical Southwestern Pacific (Figs. 5, 6).
Understanding biodiversity and connectivity of biota between isolated regions like the French Polynesian islands, MP Central Polynesia, and the ecoregions of the Central Indo-Pacific realm, thus requires the consideration of multiple factors besides geographical distance (Fig. 6): These factors include (a) ocean surface currents as a primary factor influencing the dispersal range, directions, speed and seasonal fluctuations of animal larvae, and (b) larval movement speed, motility, and pelagic larval duration (Maldonado 2006).
Ocean surface currents
The principal current system regulating the regional ocean movements in the Southern Pacific is the South Pacific Gyre, with the comparatively slow, westward flowing South Equatorial Current to its North (10° S) and the extensions of the South Pacific Current in the South (~ 25° S). This gyre also includes the East Australian Current in the West, which is one of the main drivers of dispersal of tropical marine fauna in the Central Indo-Pacific realm (Fig. 7) (e.g., James and Scandol 1992; Booth et al. 2007; Condie et al. 2011).
Marine provinces of the Central Indo-Pacific marine realm (yellow background) and Eastern Indo-Pacific marine realm (green background) after Spalding et al. (2007) in the southwestern Pacific. The shape of the marine realms has been simplified. The major sea surface currents are indicated by arrows, redrawn after satellite data which was produced by the OSCAR system (NASA/NOAA) (Dohan and Maximenko 2010). Abbreviated points of interest: Fi Fiji, NC New Caledonia, Sa Samoa, So Society Islands, Tu Tuamotu Islands, Wa Wallis and Futuna
The relative isolation of the marine sponge fauna of the Marquesas Islands, as discussed in Hall et al. (2013) and corroborated in this study, can be attributed to the comparatively strong influence of the South Equatorial Current in this region, with a constant WSW flow directed towards the North of the French Polynesian Islands (Fig. 7). This current system largely limits faunal exchange in one direction and only peripherally links to the northern islands of MP Southeast Polynesia, after a travel distance of 500–1500 km. Exchange between MP Marquesas and Hawaii or the Eastern Pacific (Americas) is further extremely limited due the strong Equatorial Countercurrent, both North and South Equatorial Currents, and the Eastern Pacific Barrier, an uninterrupted > 6000 km stretch of open ocean (Romero-Torres et al. 2018; Crandall et al. 2019).
Our data reveal higher faunal similarities between the distant MPs Southeast Polynesia, Central Polynesia (3000 km), and the provinces of the Central Indo-Pacific realm, as examined here, are, compared to the two adjacent French Polynesian provinces MP Southeast Polynesia and MP Marquesas (min. 500 km) (Fig. 6). Still, the total number of shared species is low, largely coinciding with the marine barrier hypotheses as proposed by Vermeij (1987). Geographic distance, and to a minor degree geologic history, appear to have the largest impact on gene flow within and between the continental and uplifted islands of the Central Indo‐Pacific realm and the volcanic islands of the Eastern Indo-Pacific realm (Crandall et al. 2019).
The main promoters of specimen exchange within and between the marine realms of Central and Eastern Indo-Pacific are the South Equatorial Current and the stronger South Pacific Current. The South Equatorial Current in particular is an important stimulus for faunal connectivity and dispersal of sponges between MP Central Polynesia and MP Southeast Polynesia (distance: ~ 3000 km). Facilitated by the comparatively stronger South Pacific Current, a number of sponge species from the Central Indo-Pacific realm may have also bypassed the MP Central Polynesia, and partially MP Tropical Southwestern Pacific, towards MP Southeast Polynesia, in rare cases of long-distance travel and settlement. The South Pacific Current and its numerous small gyres and turbulences to the South of Fiji and Tonga extend up to the islands of Wallis and Futuna and Samoa (MP Central Polynesia). These current systems are likewise in part responsible for the higher faunal overlap of MP Central Polynesia (Eastern Indo-Pacific realm) with the MPs in the Central Indo-Pacific realm, compared to the number of MOTUs shared with the French Polynesian provinces (Eastern Indo-Pacific realm) (Fig. 4). Another possible factor further impacting the biodiversity in MP Central Polynesia is the seasonal variation of the Pacific currents, causing strong eastward directed flows from the Solomon Islands and Papua New Guinea towards Wallis and Futuna during the warm El Niño events of the El Niño-Southern Oscillation (ENSO), which are not present during neutral or cold (La Niña) phases (Steele et al. 2010).
Pelagic larval duration
While ocean currents are the main forces driving larval dispersal, the survival time of animal larvae in the water column, Pelagic Larval Duration (PLD), plays a further pivotal role. A high PLD does not always guarantee a high larval distribution range, but it does dictate the effective range of animal larvae in combination with marine current systems (Alzate et al. 2019). This is particularly important for the lecithotrophic sponge larvae, as they have one of the shortest PLD among other benthic and pelagic reef organisms, ranging between few minutes and up to 20 days (van der Molen et al. 2018). Unlike planktotrophic larvae, sponge larvae are limited in their energy supply by the amount of yolk available, which consequently also limits their potential PLD (Maldonado 2006). The disadvantage of short PLDs is partially offset by the unusually high swimming speeds of sponge larvae, which are significantly faster than those of other marine organisms (Montgomery et al. 2019, 2020; Lanna and Riesgo 2020). Despite the combination of short PLD with high mobility, the high levels of endemism in the Southern Pacific realms, as recovered in our study and in Hall et al. (2013), highlight the poor connectivity among most of the ecoregions in the Central and Eastern Indo-Pacific realms, due to the comparatively long travel times and subsequent high levels of species filtration (Mora et al. 2012).
Anthropogenic influences on sponge dispersal
A further current-independent factor with potential impact on the dispersal capabilities of sponges is anthropogenic influence (e.g., ballast water), which can facilitate the crossing of larger distances or naturally impossible ranges, and thus contribute to the introduction of novel invasive species into foreign ecosystems (Hutchings et al. 2002; Carballo et al. 2013; Seebens et al. 2013). The influence of anthropogenic organism dispersal on sponge species in the Central and Eastern Indo-Pacific is yet insufficiently investigated. However, previous studies from other maritime regions and different taxa found this mode of transportation to also be a plausible factor for artificial dispersal: Ballast water and hull fouling of maritime vessels can aid sessile organisms to cross otherwise insurmountable marine barriers (Godwin 2003; Carballo et al. 2013), with long-range trade of aquacultures and related commercial goods also being a potential vector for the dispersal of invasive species (Henkel and Janussen 2011). Anthropogenic influence on species dispersal can also occur as a consequence of natural events, such as storms or tsunamis, creating floating debris of man-made structures as a hard substrate for the settlement of sessile organisms, and consequently expanding their potential dispersal range (Carlton et al. 2017).
Dispersal capabilities of other Indo-Pacific reef organisms
The dispersal capacities and biodiversity of other shallow water marine taxa in the Indo-Pacific can show a greater degree of variation compared to sponges, expressed in a variety of differing distribution patterns, which not always seem to directly translate from their effective potential dispersal ranges:
The Crown-of-Thorns starfish (Acanthaster planci Linnaeus), despite being widespread with a high dispersal range, tends to adhere to localized populations, with long-range expansions being comparatively uncommon (Timmers et al. 2012; Vogler et al. 2013; Pratchett et al. 2014). Populations in the South Pacific also show patterns of genetic isolation by distance between the Central and Eastern Indo-Pacific realms (Yasuda et al. 2009). Dispersal across the East Pacific Barrier appears to be possible according to molecular analyses, however sufficient data for this mode of migration is not yet available (Vogler et al. 2008; Haszprunar et al. 2017).
Scleractinian corals show similar distribution patterns to sponges in regard to the affinity to specific ecoregions, but their levels of endemism are magnitudes lower in comparison, while geographical ranges of coral species at the same time are vastly higher (Veron et al. 2015). While endemism of coral species in the Central and Eastern Indo-Pacific realms is low, Veron et al. (2015) also emphasize the clear ecological differentiation of both realms, which is further highlighted in other studies (e.g., Oury et al. 2021), but also note the lack of available data for the Eastern Indo-Pacific realm in particular. Coral communities of the Central and Eastern Indo-Pacific realms are generally isolated from the Tropical Eastern Pacific realm by the East Pacific Barrier, but with the possibility of rare long-range dispersal and settlement events (Romero-Torres et al. 2018).
Indo-Pacific reef fish show distributions very similar to those of corals, in that they display equally high or even higher levels of both species diversity and taxon overlap between ecoregions across the Central and Eastern Indo-Pacific realms. Maximum endemism levels of ~ 6% occur in those realms, supporting a hypothesis of high dispersal and colonization capability (Briggs and Bowen 2012, 2013; Kulbicki et al. 2013). Connectivity of shore fish species across the East Pacific Barrier is elevated in both directions in comparison to the low dispersal capabilities of benthic organisms across this barrier (Robertson et al. 2004).
In contrast to the observed distributions for fishes and corals, Meyer et al. (2005) showed, by using the example of marine gastropods, that fine-scale endemism of morphological species complexes on archipelago or even island scale may not be an uncommon occurrence in Indo-Pacific reef taxa, especially among those with limited larval dispersal capabilities. This appears to be the case for sponges in particular, as confirmed by the high rates of endemism and low numbers of shared species between regions found in this study. This finding is in agreement with earlier investigations of the sponge biogeography in the Indo-Pacific (Hooper and Lévi 1994), and is also evidenced in regional barcoding studies (Pöppe et al. 2010).
Conclusion
The sponge fauna of the Eastern Indo-Pacific realm is unique in displaying both high species diversity and high endemism. Species diversity appears to be increasing from French Polynesia westwards, with Marquesas being the most isolated and least diverse ecoregion. Wallis and Futuna, on the border to the Central Indo-Pacific ecoregions, displays among the highest levels of species richness and taxa shared with the adjacent realm, especially when taking in account the size of the regions investigated. A detailed look at the marine provinces also reveals differences in their biodiversity with some sponge orders being dominant in one region, but nearly absent in the others. These differences can be traced back to the local reef and island structure, the geographic distances between the studied regions, as well as the complex and seasonally impacted current systems. These factors have a large influence on the dispersal and settlement abilities of sponge taxa and are therefore responsible for the high rates of endemism and low numbers of shared taxa between marine provinces in this region, compared to other marine organisms. The Central Polynesian border region (e.g., Wallis and Futuna) plays a critical role in this regard, as it has a larger variety of those factors impacting biodiversity compared to the islands of French Polynesia, making it a “melting pot” of sponge species diversity between the two major marine realms of the Central and Eastern Indo-Pacific.
Data availability
The datasets generated during and/or analyzed during the current study are available in the EMBL Nucleotide Sequence Database (ENA) repository under the accession number range OX451516-OX451734, with additional specimen information available in the Sponge Barcoding Database (SBD) under the accession number range SBD#2345-2565.
References
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Alzate A, van der Plas F, Zapata FA et al (2019) Incomplete datasets obscure associations between traits affecting dispersal ability and geographic range size of reef fishes in the Tropical Eastern Pacific. Ecol Evol 9:1567–1577
Amade P, Pesando D, Chevolot L (1982) Antimicrobial activities of marine sponges from French Polynesia and Brittany. Mar Biol 70:223–228
Anderson MJ (2001) A new method for non-parametric multivariate analysis of variance. Austral Ecol 26:32–46
Anderson MJ (2017) Permutational multivariate analysis of variance (PERMANOVA). Wiley StatsRef: Statistics Reference Online 1–15
Andreakis N, Luter HM, Webster NS (2012) Cryptic speciation and phylogeographic relationships in the elephant ear sponge Ianthella basta (Porifera, Ianthellidae) from northern Australia. Zool J Linn Soc 166:225–235
Aswani S, Allen MS (2009) A Marquesan coral reef (French Polynesia) in historical context: an integrated socio-ecological approach. Aquat Conserv 19:614–625
Beck J, Schwanghart W (2010) Comparing measures of species diversity from incomplete inventories: an update. Methods Ecol Evol 1:38–44
Bell JJ (2008) The functional roles of marine sponges. Estuar Coast Shelf Sci 79:341–353
Bodenhofer U, Bonatesta E, Horejš-Kainrath C, Hochreiter S (2015) msa: an R package for multiple sequence alignment. Bioinformatics 31:3997–3999
Booth DJ, Figueira WF, Gregson MA et al (2007) Occurrence of tropical fishes in temperate southeastern Australia: role of the East Australian Current. Estuar Coast Shelf Sci 72:102–114
Briggs JC, Bowen BW (2012) A realignment of marine biogeographic provinces with particular reference to fish distributions. J Biogeogr 39:12–30
Briggs JC, Bowen BW (2013) Marine shelf habitat: biogeography and evolution. J Biogeogr 40:1023–1035
Carballo JL, Aguilar-Camacho JM, Knapp IS, Bell JJ (2013) Wide distributional range of marine sponges along the Pacific Ocean. Mar Biol Res 9:768–775
Carlton JT, Chapman JW, Geller JB et al (2017) Tsunami-driven rafting: transoceanic species dispersal and implications for marine biogeography. Science 357:1402–1406
Chao A (1984) Nonparametric estimation of the number of classes in a population. Scand Stat Theory Appl 11:265–270
Chao A, Ma KH, Hsieh TC (2016) User’s guide for iNEXT online: software for interpolation and extrapolation of species diversity. CoDesign 30043:1–14
Chombard C, Boury-Esnault N, Tillier S (1998) Reassessment of homology of morphological characters in tetractinellid sponges based on molecular data. Syst Biol 47:351–366
Condie SA, Mansbridge JV, Cahill ML (2011) Contrasting local retention and cross-shore transports of the East Australian Current and the Leeuwin current and their relative influences on the life histories of small pelagic fishes. Deep Sea Res Part 2(58):606–615
Conway JR, Lex A, Gehlenborg N (2017) UpSetR: an R package for the visualization of intersecting sets and their properties. Bioinformatics 33:2938–2940
Cowman PF, Parravicini V, Kulbicki M, Floeter SR (2017) The biogeography of tropical reef fishes: endemism and provinciality through time. Biol Rev Camb Philos Soc 92:2112–2130
Crandall ED, Riginos C, Bird CE et al (2019) The molecular biogeography of the Indo-Pacific: testing hypotheses with multispecies genetic patterns. Glob Ecol Biogeogr 28:943–960
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9:772
DeBiasse MB, Hellberg ME (2015) Discordance between morphological and molecular species boundaries among Caribbean species of the reef sponge Callyspongia. Ecol Evol 5:663–675
Debitus C, Folcher E, Petek S et al (2009) BSMPF-1 cruise, R/V Alis, (Society and Marquesas archipelago is.). https://doi.org/10.17600/9100030
Debitus C, Folcher E, Petek S et al (2011) Tuam’2011 cruise, R/V Alis, (Tuamotu archipelago is.). https://doi.org/10.17600/11100010
Debitus C, Folcher E, Petek S et al (2013a) Tuhaa Pae 2013a cruise, R/V Alis, (Austral archipelago is.). https://doi.org/10.17600/13100030
Debitus C, Petek S, Ekins M et al (2013b) Tahiti iti cruise, R/V Alis, (Tahiti is.). https://doi.org/10.17600/13100040
de Goeij JM, van Oevelen D, Vermeij MJA et al (2013) Surviving in a marine desert: the sponge loop retains resources within coral reefs. Science 342:108–110
Delrieu-Trottin E, Williams JT, Bacchet P et al (2015) Shore fishes of the Marquesas Islands, an updated checklist with new records and new percentage of endemic species. CheckList 11:1–13
de Voogd NJ, Alvarez B, Boury-Esnault N et al (2023) World Porifera Database. http://www.marinespecies.org/porifera. Accessed 23 Jan 2023
Dixon P (2003) VEGAN, a package of R functions for community ecology. J Veg Sci 14:927–930
Dohan K, Maximenko N (2010) Monitoring ocean currents with satellite sensors. Oceanography 23:94–103
Duckworth AR, Wolff C, Evans-Illidge E et al (2008) Spatial variability in community structure of Dictyoceratida sponges across Torres Strait, Australia. Cont Shelf Res 28:2168–2173
El-Demerdash A, Atanasov AG, Horbanczuk OK et al (2019) Chemical diversity and biological activities of marine sponges of the genus Suberea: a systematic review. Mar Drugs 17:115
Erpenbeck D, Breeuwer JAJ, Parra-Velandia FJ, van Soest RWM (2006) Speculation with spiculation? Three independent gene fragments and biochemical characters versus morphology in demosponge higher classification. Mol Phylogenet Evol 38:293–305
Erpenbeck D, Voigt O, Al-Aidaroos AM et al (2016) Molecular biodiversity of Red Sea demosponges. Mar Pollut Bull 105:507–514
Erpenbeck D, Aryasari R, Benning S et al (2017) Diversity of two widespread Indo-Pacific demosponge species revisited. Mar Biodivers 47:1035–1043
Erpenbeck D, Gholami A, Hesni MA et al (2020) Molecular biodiversity of Iranian shallow water sponges. Syst Biodivers 18:192–202
Folkers M, Rombouts T (2020) Sponges revealed: a synthesis of their overlooked ecological functions within aquatic ecosystems. In: Jungblut S, Liebich V, Bode-Dalby M (eds) YOUMARES 9—the oceans: our research, our future. Springer, Cham, pp 181–193
Freeman CJ, Easson CG (2016) Sponge distribution and the presence of photosymbionts in Moorea, French Polynesia. Peerj 4:e1816
Fromont J, Abdul Wahab MA, Gomez O et al (2016) Patterns of sponge biodiversity in the Pilbara, Northwestern Australia. Diversity 8:21
Galitz A, Nakao Y, Schupp PJ et al (2021) A soft spot for chemistry-current taxonomic and evolutionary implications of sponge secondary metabolite distribution. Mar Drugs 19:448
Galzin R, Planes S, Dufour V, Salvat B (1994) Variation in diversity of coral reef fish between French Polynesian atolls. Coral Reefs 13:175–180
Gillespie RG, Claridge EM, Goodacre SL (2008) Biogeography of the fauna of French Polynesia: diversification within and between a series of hot spot archipelagos. Philos Trans R Soc Lond B 363:3335–3346
Glover AG, Wiklund H, Chen C, Dahlgren TG (2018) Managing a sustainable deep-sea “blue economy” requires knowledge of what actually lives there. eLife 7:e41319
Godwin LS (2003) Hull fouling of maritime vessels as a pathway for marine species invasions to the Hawaiian Islands. Biofouling 19(Suppl):123–131
Hadiyanto H, Hovey RK, Glasby CJ et al (2021) Marine ecoregions and subecoregions within Indo-West Australian waters: a statistical approach based on species distributions. J Biogeogr 48:2246–2257
Hall KA, Sutcliffe PR, Hooper JNA et al (2013) Affinities of Sponges (Porifera) of the Marquesas and Society Islands, French Polynesia. Pasc 67:493–511
Haszprunar G, Vogler C, Wörheide G (2017) Persistent gaps of knowledge for naming and distinguishing multiple species of crown-of-thorns-seastar in the Acanthaster planci species complex. Diversity 9:22
Henkel D, Janussen D (2011) Redescription and new records of Celtodoryx ciocalyptoides (Demospongiae: Poecilosclerida)—a sponge invader in the north east Atlantic Ocean of Asian origin? J Mar Biol Assoc U K 91:347–355
Hooper JNA, Lévi C (1994) Biogeography of indo-west pacific sponges: Microcionidae, Raspailiidae, Axinellidae. Sponges Time Space 191:212
Hulsen T, de Vlieg J, Alkema W (2008) BioVenn—a web application for the comparison and visualization of biological lists using area-proportional Venn diagrams. BMC Genomics 9:488
Hutchings PA, Hilliard RW, Coles SL (2002) Species introductions and potential for marine pest invasions into tropical marine communities, with special reference to the Indo-Pacific. Pac Sci 56:223–233
Jaccard P (1912) The distribution of the flora in the alpine zone. New Phytol 11:37–50
James MK, Scandol JP (1992) Larval dispersal simulations: correlation with the Crown-of-Thorns starfish outbreaks database. Mar Freshw Res 43:569–581
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Kembel SW, Cowan PD, Helmus MR et al (2010) Picante: R tools for integrating phylogenies and ecology. Bioinformatics 26:1463–1464
Klautau M, Lopes MV, Debitus C (2020) Calcareous sponges from the French Polynesia (Porifera: Calcarea). Zootaxa 4748:zootaxa.4748.2.3
Kreft H, Jetz W (2010) A framework for delineating biogeographical regions based on species distributions. J Biogeogr 37:2029–2053
Kulbicki M, Parravicini V, Bellwood DR et al (2013) Global biogeography of reef fishes: a hierarchical quantitative delineation of regions. PLoS ONE 8:e81847
Lanna E, Riesgo A (2020) Sponge larvae do not swim that fast: a reply to Montgomery et al. (2019). J Mar Biol Assoc U K 100:181–183
Lecchini D, Bertucci F, Fogg L et al (2021) Marine biodiversity of a pristine coral reef in French Polynesia. Isl Stud J 16:292–307
Legendre P, Salvat B (2015) Thirty-year recovery of mollusc communities after nuclear experimentations on Fangataufa atoll (Tuamotu, French Polynesia). Proc Biol Sci 282:20150750
López-Legentil S, Erwin PM, Henkel TP et al (2010) Phenotypic plasticity in the Caribbean sponge Callyspongia vaginalis (Porifera: Haplosclerida). Sci Mar 74:445–453
MacDonald ZG, Nielsen SE, Acorn JH (2017) Negative relationships between species richness and evenness render common diversity indices inadequate for assessing long-term trends in butterfly diversity. Biodivers Conserv 26:617–629
Maldonado M (2006) The ecology of the sponge larva. Can J Zool 84:175–194
Maldonado M, Aguilar R, Bannister RJ, et al (2015) Sponge grounds as key marine habitats: a synthetic review of types, structure, functional roles, and conservation concerns. In: Rossi S, Bramanti L, Gori A, Orejas Saco del Valle C (eds) Marine animal forests: the ecology of benthic biodiversity hotspots. Springer, Cham, pp 1–39
Meyer CP, Paulay G (2005) DNA barcoding: error rates based on comprehensive sampling. PLoS Biol 3:e422
Meyer CP, Geller JB, Paulay G (2005) Fine scale endemism on coral reefs: archipelagic differentiation in turbinid gastropods. Evolution 59:113–125
Montgomery EM, Hamel J-F, Mercier A (2019) Larval nutritional mode and swimming behaviour in ciliated marine larvae. J Mar Biol Assoc U K 99:1027–1032
Montgomery EM, Hamel J-F, Mercier A (2020) Sponge larvae are not that fast, but still the fastest: Reply to: “Sponge larvae do not swim that fast.” J Mar Biol Assoc U K 100:185–187
Mora C, Treml EA, Roberts J et al (2012) High connectivity among habitats precludes the relationship between dispersal and range size in tropical reef fishes. Ecography 35:89–96
Neall VE, Trewick SA (2008) The age and origin of the Pacific islands: a geological overview. Philos Trans R Soc Lond B 363:3293–3308
Oury N, Gélin P, Magalon H (2021) High connectivity within restricted distribution range in Pocillopora corals. J Biogeogr 48:1679–1692
Pawlik JR, McMurray SE (2020) The emerging ecological and biogeochemical importance of sponges on coral reefs. Ann Rev Mar Sci 12:315–337
Petek S, Debitus C, Alencar A et al (2017) Sponges of Polynesia. Papeete (PYF): IRD. https://horizon.documentation.ird.fr/exl-doc/pleins_textes/divers17-07/010070137.pdf
Petek S, Folcher E, Ekins M et al (2018) WALLIS 2018 cruise, R/V Alis, (Wallis is.). https://doi.org/10.17600/18000524
Pielou EC (1966) The measurement of diversity in different types of biological collections. J Theor Biol 13:131–144
Planes S, Allemand D, Agostini S et al (2019) The Tara Pacific expedition—a pan-ecosystemic approach of the “-omics” complexity of coral reef holobionts across the Pacific Ocean. PLoS Biol 17:e3000483
Pöppe J, Sutcliffe P, Hooper JNA et al (2010) COI Barcoding reveals new clades and radiation patterns of Indo-Pacific sponges of the Family Irciniidae (Demospongiae: Dictyoceratida). PLoS ONE 5:e9950
Pos E, Guevara Andino JE, Sabatier D et al (2014) Are all species necessary to reveal ecologically important patterns? Ecol Evol 4:4626–4636
Powell A, Smith DJ, Hepburn LJ et al (2014) Reduced diversity and high sponge abundance on a sedimented Indo-Pacific reef system: implications for future changes in environmental quality. PLoS ONE 9:e85253
Pratchett MS, McCowan D, Maynard JA, Heron SF (2013) Changes in bleaching susceptibility among corals subject to ocean warming and recurrent bleaching in Moorea, French Polynesia. PLos ONE 8:e70443
Pratchett MS, Caballes CF, Rivera-Posada JA, Sweatman HPA (2014) Limits to understanding and managing outbreaks of Crown-of-Thorns Starfish (Acanthaster spp.). In: Hughes RN, Hughes DJ, Smith PI (eds) Oceanography and marine biology. CRC Press, pp 133–200
Ramage T (2017) Checklist of the terrestrial and freshwater arthropods of French Polynesia (Chelicerata; Myriapoda; Crustacea; Hexapoda). Zoos 39:213–225
R Core Team (2021) R: a language and environment for statistical computing. Version 4.1.1. R Foundation for Statistical Computing, Vienna. https://www.R-project.org/
Reveillaud J, van Soest RWM, Derycke S et al (2011) Phylogenetic relationships among NE Atlantic Plocamionida Topsent (1927) (Porifera, Poecilosclerida): Under-estimated diversity in reef ecosystems. PLoS ONE 6:e16533
Richter C, Wunsch M, Rasheed M et al (2001) Endoscopic exploration of Red Sea coral reefs reveals dense populations of cavity-dwelling sponges. Nature 413:726–730
Robertson DR, Grove JS, McCosker JE (2004) Tropical transpacific shore fishes. Pac Sci 58:507–565
Romero-Torres M, Treml EA, Acosta A, Paz-García DA (2018) The Eastern Tropical Pacific coral population connectivity and the role of the Eastern Pacific Barrier. Sci Rep 8:9354
Rovellini A, Dunn MR, Fulton EA et al (2019) Decadal variability in sponge abundance and biodiversity on an Indo-Pacific coral reef. Mar Ecol Prog Ser 620:63–76
Schroeder PJ, Jenkins DG (2018) How robust are popular beta diversity indices to sampling error? Ecosphere 9:e02100
Schuster A, Pisera A, Ekins M, Debitus C (2021) New genus and species of lithistid demosponges from submarine caves in Nuku Hiva (Marquesas Islands) and Tahiti Iti (Society Islands), French Polynesia. Eur Zool J 88:749–770
Seebens H, Gastner MT, Blasius B (2013) The risk of marine bioinvasion caused by global shipping. Ecol Lett 16:782–790
Setiawan E, De Voogd NJ, Swierts T, et al (2015) MtDNA diversity of the Indonesian giant barrel sponge Xestospongia testudinaria (Porifera: Haplosclerida)—implications from partial cytochrome oxidase 1 sequences. J Mar Biol Assoc U K
Spalding MD, Fox HE, Allen GR et al (2007) Marine ecoregions of the world: a bioregionalization of coastal and shelf areas. Bioscience 57:573–583
Steele JH, Thorpe SA, Turekian KK (2010) Ocean currents. Academic Press, New York
Strong WL (2016) Biased richness and evenness relationships within Shannon-Wiener index values. Ecol Indic 67:703–713
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Timmers MA, Bird CE, Skillings DJ et al (2012) There’s no place like home: crown-of-thorns outbreaks in the central pacific are regionally derived and independent events. PLoS ONE 7:e31159
Timmers MA, Vicente J, Webb M et al (2022) Sponging up diversity: evaluating metabarcoding performance for a taxonomically challenging phylum within a complex cryptobenthic community. Environ DNA 4:239–253
van der Molen J, García-García LM, Whomersley P et al (2018) Connectivity of larval stages of sedentary marine communities between hard substrates and offshore structures in the North Sea. Sci Rep 8:14772
Van Soest RWM, Boury-Esnault N, Vacelet J et al (2012) Global diversity of sponges (Porifera). PLoS ONE 7:e35105
Vargas S, Schuster A, Sacher K et al (2012) Barcoding sponges: an overview based on comprehensive sampling. PLoS ONE 7:e39345
Vermeij GJ (1987) The dispersal barrier in the tropical Pacific: implications for molluscan speciation and extinction. Evolution 41:1046–1058
Veron J, Stafford-Smith M, DeVantier L, Turak E (2015) Overview of distribution patterns of zooxanthellate Scleractinia. Front Mar Sci 1:81
Vicente J, Ríos JA, Zea S, Toonen RJ (2019) Molecular and morphological congruence of three new cryptic Neopetrosia spp in the Caribbean. PeerJ 7:e6371
Vicente J, Webb MK, Paulay G et al (2021) Unveiling hidden sponge biodiversity within the Hawaiian reef cryptofauna. Coral Reefs 1–16
Vogler C, Benzie J, Lessios H et al (2008) A threat to coral reefs multiplied? Four species of crown-of-thorns starfish. Biol Lett 4:696–699
Vogler C, Benzie JAH, Tenggardjaja K et al (2013) Phylogeography of the crown-of-thorns starfish: genetic structure within the Pacific species. Coral Reefs 32:515–525
Wilkinson CR (1988) Foliose Dictyoceratida of the Australian great barrier reef. Mar Ecol 9:321–327
Williams CB (1949) Jaccard’s generic coefficient and coefficient of floral community, in relation to the logarithmic series and the index of diversity. Ann Bot 13:53–58
Wörheide G, Erpenbeck D (2007) DNA taxonomy of sponges—progress and perspectives. J Mar Biol Assoc U K 87:1629–1633
Wörheide G, Degnan BM, Hooper JNA, Reitner J (2002) Phylogeography and taxonomy of the Indo-Pacific reef cave dwelling coralline demosponge Astrosclera “willeyana”: new data from nuclear internal transcribed spacer sequences. In: Proceedings of the Ninth International Coral Reef Symposium. Ministry for Environment, Indonesian Institute of Sciences, International Society for Reef Studies Jakarta, pp 339–346
Wörheide G, Epp LS, Macis L (2008) Deep genetic divergences among Indo-Pacific populations of the coral reef sponge Leucetta chagosensis (Leucettidae): founder effects, vicariance, or both? BMC Evol Biol 8:24
Wright ES (2016) Using DECIPHER v2. 0 to analyze big biological sequence data in R. R J 8
Wulff J (2012) Ecological interactions and the distribution, abundance, and diversity of sponges. Adv Mar Biol 61:273–344
Xavier JR, Rachello- Dolmen PG, Parra-Velandia FJ et al (2010) Molecular evidence of cryptic speciation in the “cosmopolitan” excavating sponge Cliona celata (Porifera, Clionaidae). Mol Phylogenet Evol 56:13–20
Yasuda N, Nagai S, Hamaguchi M et al (2009) Gene flow of Acanthaster planci (L.) in relation to ocean currents revealed by microsatellite analysis. Mol Ecol 18:1574–1590
Acknowledgements
We thank the French Polynesian and the Wallis & Futuna authorities, as well as the communities, for allowing us to collect in their country. We acknowledge the crews of the R/V ALIS, Braveheart, Claymore II and IRD’s diving team (SEOH IRD Noumea, New Caledonia) for their essential contribution to all the field trips. We furthermore would like to thank Dominique Fleurisson, Armelle Renaud, and Bertrand Bourgeois for their contributions in underwater photography. Support and permission to undertake the study on Futuna samples were provided by Atoloto Malau (Service de l’Environnement, Wallis and Futuna) to the Tara Pacific Foundation and NUI Galway. We would like to express our gratitude to the administrative and customary authorities of the Territoire des iles Wallis et Futuna for their support in the sample collection. Special thanks to the Tara Ocean Foundation, the R/V Tara crew and the Tara Pacific Expedition Participants (https://doi.org/10.5281/zenodo.3777760). We are keen to thank the commitment of the following institutions for their financial and scientific support that made the unique Tara Pacific Expedition possible: CNRS, PSL, CSM, EPHE, Genoscope, CEA, Inserm, Université Côte d’Azur, ANR, agnès b., UNESCO-IOC, the Veolia Foundation, the Prince Albert II de Monaco Foundation, Région Bretagne, Billerudkorsnas, AmerisourceBergen Company, Lorient Agglomération, Oceans by Disney, L’Oréal, Biotherm, France Collectivités, Fonds Français pour l’Environnement Mondial (FFEM), Etienne Bourgois, and the Tara Ocean Foundation teams. Tara Pacific would not exist without the continuous support of the participating institutes. The authors also particularly thank Serge Planes, Denis Allemand, and the Tara Pacific consortium.
Funding
Open Access funding enabled and organized by Projekt DEAL. We acknowledge French Polynesian Authorities and Délégation à la Recherche-France in Tahiti (projects Marquesas and Biopolyval), Netbiome project (ANR-11-EBIM-0006 “POMARE”), the French Oceanographic Fleet and IRD for funding the R/V ALIS trips in French Polynesia. The Wallis 2018 oceanographic cruise was funded by the French Oceanographic Fleet, IRD, MNHN, Labex Mer and the Wallis and Futuna Environment Service. AG acknowledges financial support from LMU via the LMUmentoring program.
Author information
Authors and Affiliations
Contributions
AG and DE conceived and designed the study. Material preparation, data collection and morphological identification were performed by CD, ME, EF, KH, JNAH, SP, and OPT. GB, DE, AG, MMR, SS, and GW performed molecular data generation and analysis, DE, AG, and GW processed and evaluated the generated data. The first draft of the manuscript was written by AG and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
No approval of research ethics committees was required to accomplish the goals of this study because experimental work was conducted with an unregulated invertebrate species.
Sampling and field studies
All necessary permits for sampling and observational field studies have been obtained by the authors from the competent authorities and are mentioned in the acknowledgements, if applicable. The study is compliant with CBD and Nagoya protocols.
Additional information
Communicated by Khor Waiho.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Galitz, A., Ekins, M., Folcher, E. et al. Poriferans rift apart: molecular demosponge biodiversity in Central and French Polynesia and comparison with adjacent marine provinces of the Central Indo-Pacific. Biodivers Conserv 32, 2469–2494 (2023). https://doi.org/10.1007/s10531-023-02613-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10531-023-02613-y