Abstract
Understanding the historical context of biological invasions can improve weed management outcomes. In this study, we aim to identify the introduction pathway of bitou bush (Chrysanthemoides monilifera subsp. rotundata) into Australia and its biogeographical origin in southern Africa by combining multiple lines of evidence from genomic tools and historical documentation. Geographic structure of genomic diversity based on SNPs supported the previous analysis of the invasion pathway of bitou bush between the two countries and within Australia, namely that all Australian material originated from the southern part of the South African distribution. Our synthesis of historical records points to the introduction of this plant into eastern Australia in Newcastle, New South Wales, from its native range in South Africa, via dry shipping ballast in about 1900. Variation in the chloroplast genome was also informative as to the biogeographical origin of Australian material and the context of the introduction. Ten unique haplotypes were discovered in South Africa with only one occurring throughout Australia, indicating an introduction from a single source population to eastern Australia. The matching haplotype was from East London, a port in South Africa with documented shipping connections to Newcastle in eastern Australia, where the weed was first recorded. Historical records suggest that the most plausible explanation for the origins of the isolated bitou bush population in Western Australia is via the shipping of steel billets or landscape plantings associated with shipping companies. The most likely introduction pathway linked the eastern Australian steel processing ports of Newcastle or Port Kembla to the Western Australian port of Kwinana in 1995. Discovering the origin and pathway of bitou bush invasions in Australia opens new opportunities for sourcing biological control agents with a higher chance of impact as well as identifying additional quarantine measures to improve outcomes and reduce long-term costs to management.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The ever-increasing cost of managing non-native invasive species as they progress along the invasion curve, (sensu Blackburn et al. 2011) represents both a challenge and an opportunity. That is, the best return on investment is to prevent the arrival of the organism in the first place, with swift and intense action to eradicate the species as soon after arrival as possible representing the next best approach. The longer a non-native species is present in the landscape, the more challenging it is to achieve eradication, the harder it is to mitigate its impacts, and the greater the investment required to achieve these goals, if they are indeed possible. This reality makes it important to understand the factors that facilitate the arrival of non-native invasive species in new areas.
Globalisation has expanded trade and travel between countries and with this increase in contact comes an increased risk of plant, animal and pathogen invasions (Hulme 2009). Humans exert pressure on the environment by facilitating the movement, introduction and establishment of species beyond their native range either by deliberate actions or unwitting dispersal (Kueffer 2017). Anthropogenic pathways, therefore, require more scrutiny if we are to detect introduced populations in the early stages of the invasion curve. Discovering predictable ways by which these species reach and establish in novel environments offers insight that can be applied to improve biosecurity prevention and control measures (Blackburn et al. 2011; Novoa et al. 2020; Simberloff et al. 2013). Thus, elucidating pathways of introduction can improve biosecurity programs and provide solutions for controlling invaders (e.g. Scott et al. 2017).
Introduction context and invasion history can also provide critical information for identifying classical biological control agents. There are numerous examples where this information has helped to find cryptic species (Forno and Harley 1979), clonal genotypes (Cullen 2012), or hybrids (Gaskin and Schaal 2002), which need to be considered in any risk assessment for biological control. Establishing introduction context can even lead to the identification of species that were incorrectly identified as introduced but are actually native, and where biological control would normally not be an option (e.g. Scott 2012). Furthermore, when the origins of a species are unknown (native or non-native), this can become a major constraint on establishing an effective biological control program (Gildenhuys et al. 2013).
Whether or not introductions have been the result of a single accidental introduction, or a more determined deliberate collection of material from one or more locations in the native or non-native range (e.g., buffel grass, Cenchrus ciliaris; Friedel 2020; Marshall et al. 2012), can significantly alter the likelihood of establishment and invasion after introduction. Sufficient information on introduction pathways has now been amassed to enable the categorising of pathways. In Hulme et al. (2008), six primary invasion pathways were proposed (release, escape, contaminants, stowaways, corridors and unaided). Faulkner et al. (2020) took this further by ascribing further dispersal context that can influence the outcomes of introductions, including leading-edge dispersal, corridors, jump dispersal, extreme long-distance dispersal, mass dispersal and cultivation.
Early biogeographical literature used historical data to elucidate pathways of anthropogenic plant dispersal and introduction, including those that result in invasive populations (Ridley 1930). Ships records and customs manifests have been valuable sources of information for which to pinpoint the arrival time and location, as such vessels are often the likely source of the introduced material (Wilson et al. 2009). With the rapid digitisation and online open access availability of libraries, herbaria and historical literature, accessing this trove of information has become far more feasible, providing novel insight for past introductions that have remained unexplained up to now (Austin 2014). However, historical data alone can be prone to bias, misinterpretation and errors. For example, species names change over time and without herbarium material to assign an updated determination, the identity of complex introductions can remain obscured and misunderstood (Alpern 2008; Webber et al. 2011). Moreover, specimens lodged in herbaria come with their own inherent taxonomic and spatial sampling biases (Daru et al. 2018).
In more recent times, genetic and genomic techniques have played an important role in providing a more detailed analysis of the introduction and invasion process and to help inform how management can best adapt control to suit these factors (Cristescu 2015; Goolsby et al. 2006). Population genetics can help to pinpoint a more precise location for the origin of the introduced material in the native range, which can be particularly helpful for targeting biocontrol agent searches in species with a broad native range (Cullen 2012). Comparing population genetics within and between populations across the native and introduced range can help identify variation and spatial structuring of genetic diversity, such as the loss of alleles during the invasion process (Estoup and Guillemaud 2010). Greater genetic variation generally increases fitness for introduced populations yet is frequently reduced following introductions through the founder effect (Dlugosch and Parker 2008; Estoup et al. 2016; Sax and Brown 2000). However, the genetic bottleneck associated with introductions is often transient (Colautti et al. 2017). Genetic techniques can determine the mating system of invasive species, which is important because sexual reproduction assists in the generation of genetic variation, and invasion mediated by self-fertilisation and asexual reproduction can lead to very different genetic outcomes (Roman and Darling 2007).
When combined, historical data sources and genomic analysis can provide the most powerful evidence in reconstructing the introduction and invasion context of cryptic invasions. This evidence can be particularly invaluable in regions where introductions are ancient, complex and frequent, yet any historical context is poorly documented. In Australia, many non-native species have incomplete documentation of introduction histories, global trade continues with countries that have a long history of biotic exchange with Australia (i.e., a biosecurity risk remains), and Australia remains a global leader in investing in classical biological control as a solution for managing weeds (Julien et al. 2012). This trifecta of opportunity applies to some of Australia’s most threatening non-native plants, known as Weeds of National Significance (WoNS; Thorp and Lynch 2000).
Bitou bush (Chrysanthemoides monilifera (L.) T.Norl. subsp. rotundata (DC.) T.Norl. (Asteraceae) is listed as a WoNS and is present in four mainland states—Queensland, New South Wales, Victoria and Western Australia (Scott and Batchelor 2014; Weiss et al. 2008). The first Australian record of bitou bush dates to a herbarium specimen collected in 1908 at Stockton in Newcastle, New South Wales (Fig. 1). It has been long hypothesised that this introduction resulted from shipping activity and the deposition of founding material in discarded ship’s dry ballast. From the 1950s, bitou bush was actively planted along the New South Wales coast to stabilise dunes after mining activity (Barr and Atkinson 1970; Love 1986; Mort and Hewitt 1953). It is not known where the planting material for this activity came from—was it sourced locally or from the native range in South Africa, and was the sampling from few or many plants and from systematic or random sampling? In 2012, when bitou bush was discovered for the first time in Western Australia, established in the land base of a port (Kwinana), the hypothesis of a shipping-mediated introduction was again suggested (Scott and Batchelor 2014). However, again no evidence was assembled to test this claim, nor the properties of the material first introduced or if the introduction source was from eastern Australia or the native range in South Africa.
Introduction location of bitou bush (Chrysanthemoides monilifera subsp. rotundata) into Australia. a Ballast ground in Stockton, Newcastle, New South Wales showing land reclaimed by 1902 (indicated by red shading; Map: Newcastle University cultural collection). b Barques (sailing ships) waiting to be loaded with coal, having discarded the ballast. The ballast ground is located between the road and the water (Photo: Ralph Snowball, c 1900). Figure content sourced from https://livinghistories.newcastle.edu.au/
Along the east coast of Australia, spanning a range between Gippsland in Victoria to Fraser Island in Queensland (some 1600 km), bitou bush has disrupted many native ecosystems since its introduction (French and Watts 2015). Bitou bush replaces native plants as a food source and habitat, negatively impacting the native community composition (Behrendorff et al. 2019). Where the weed has not been controlled in a timely fashion, the damage has been complex, possibly irreversible, and a return to native vegetation is unlikely without significant restoration effort (Lindenmayer et al. 2013, 2015). The isolated Western Australian population has remained essentially contained within four industrial sites, due to fencing and site procedures limiting access, thereby providing fewer inadvertent human dispersal vectors to enhance the invasion process (Scott and Batchelor 2014). This population is presently being targeted for eradication (Scott et al. 2019b). Given the variation in invasion histories across the non-native range and the highly variable success of management strategies currently deployed, a better understanding of the invasion history for bitou bush in Australia would significantly improve management outcomes.
In a recent study, Byrne et al. (2022) used a single nucleotide polymorphism (SNP) approach to produce a partial reconstruction of the invasion history and a preliminary analysis into genetic diversity and structure within and between populations in eastern Australia, Western Australia and South Africa. However, that study could not determine the biogeographical origin of bitou bush from genomic DNA. Moreover, the high mutation rate of the SNPs and recombination of nuclear DNA documented by Byrne et al. (2022) made it difficult to detect evolutionary relationships between populations. No historical evidence was used to test the validity of these molecular results or to reinforce the robustness of the overall study findings.
To address these priority knowledge gaps, here we aim to clarify the introduction history for bitou bush in Australia. We seek to achieve these aims with an interdisciplinary approach, combining historical data and genetic analysis to establish an evidence-based case for the timing, pathway and context for the introduction history of bitou bush in Australia. This work will help inform reintroduction risks and refine existing and novel control efforts based on the influence of genetic novelty and diversity. More specifically, this study will:
-
1.
Clarify the source of bitou bush populations in Australia, including testing hypotheses for single or multiple introductions and redistribution;
-
2.
Identify the introduction pathway(s) for the Australian populations of bitou bush; and
-
3.
Assess the implications of source population and introduction pathway for the management of bitou bush.
Methods
Study system
Biology and taxonomy
The genus Chrysanthemoides is native to southern Africa and comprises two species and an uncertain number of subspecies, of which two are introduced to Australia (Griffioen 1995; Norlindh 1943). In this study we focus on bitou bush (Chrysanthemoides monilifera subsp. rotundata) and have included the related subspecies boneseed (Chrysanthemoides monilifera subsp. monilifera), which has also been introduced to Australia, and Chrysanthemoides monilifera subsp. pisifera (hereafter referred to as ‘pisifera’) as out-groups in phylogenetic analyses.
Bitou bush is a perennial shrub found in coastal dunes. Plants can be up to 3 m high and 6 m wide at maturity in the non-native range (Fig. 1 in Byrne et al. 2022), but plants are generally significantly smaller in the native range (Scott 1996; Scott et al. 2019a). Bitou bush can reproduce within twelve months of germination under suitable conditions. The plant has multiple yellow florets packed into each inflorescence, which can occur all year. Pollination is facilitated by insects but is not essential (Weiss et al. 2008). The taxon is an obligate outcrosser, which means that genomic DNA recombines with each seed formation (Gross et al. 2017; Scott et al. 2019b). Bitou bush produces fleshy fruits which are eaten by animals, meaning they can be dispersed over a long distance. The shrub forms dense stands that are known to modify dune environments in the non-native range, stabilising certain areas while altering dune mobility to make other areas more susceptible to erosion (French et al. 2008; Weiss and Noble 1984).
Distribution and introduction history
Globally, bitou bush is only known from its native range in South Africa and as a non-native introduction to Australia (Weiss et al. 2008). In its native range it is found in coastal environments of the Eastern Cape and KwaZulu-Natal provinces. The climate in the native range is “temperate with no dry season and hot summers” (Köppen-Geiger climate classification: Cfa; Kriticos et al. 2012). Its distribution along the east coast of Australia is primarily the entire coastline of the state of New South Wales with an extension northwards into the coasts of Queensland and southwards into Victoria (Weiss et al. 2008). An isolated but established population of bitou bush was discovered in 2012 at the Western Australian port of Kwinana, over 3000 km from the main infestations in eastern Australia (Scott and Batchelor 2014). From historical aerial photography it was determined that bitou bush was present as early as 1995 (Scott et al. 2019a). The areas where it is found in eastern Australia are similar in climate and environment to the native range along the east coast of South Africa (Weiss et al. 2008). In contrast, the area occupied by bitou bush in Western Australia has a Mediterranean-type climate (i.e., temperate, dry hot or warm summers; Köppen-Geiger climate classification Csa, Csb), which is similar to the climates of the Western Cape province in South Africa, where the closely related boneseed (Chrysanthemoides monilifera (L.) T.Norl. subsp. monilifera) is found (Kriticos et al. 2012).
Historical data synthesis
The primary research sources for information on the “in ballast” shipping vectors between South Africa and Australia were the scanned newspapers and shipping news found on the Trove digital database (https://trove.nla.gov.au/) and the tables for 1904 in Great Britain Board of Trade (1906). Trove is one of the most important primary sources of historical information in Australia and is particularly comprehensive for the late 1800s and early 1900s. The Great Britain Board of Trade summaries other than 1904 do not appear to be available in hard copy or online in Australia. Fortunately, the available trade summaries of 1904 were suitable for analysis, given the arrival of bitou bush sometime before 1908.
To identify useful secondary source material, we consulted the online library catalogue of the University of Newcastle (https://livinghistories.newcastle.edu.au/), the State Library of Western Australia (https://slwa.wa.gov.au/; particularly for information on Kwinana), all available volumes of the BHP Review periodical post-1950, and online libraries in South Africa. The Museum of East London (http://www.elmuseum.za.org/East-London-Museum/) provided access to documents and maps of the port area of East London, while Museum staff added additional local interpretation to the online material.
Genomics
Population coverage
Leaf samples from bitou bush plants across the native and non-native ranges were drawn from a larger collection of bitou bush samples assembled as part of the broader research program into bitou bush control in Western Australia, including those sampled by Byrne et al. (2022), with additional sampling conducted in localities without existing samples. Herbarium specimens were subsampled from localities where fresh material could not be obtained. Extracted DNA from native-range specimens analysed by Barker et al. (2015) were obtained from Nigel Barker (University of Pretoria). Collection dates of herbarium specimens ranged from 2008 to 2010. Collection dates of leaf samples collected for our research ranged from 2012 to 2020. Collection details are provided in Supplementary Table S1 and S2 and the location of sampling effort shown in Fig. 2.
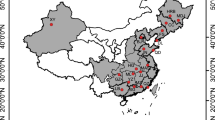
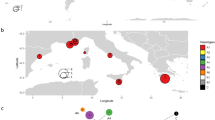
Locations of bitou bush (Chrysanthemoides monilifera subsp. rotundata) and boneseed (C. m. subsp. monilifera) samples from South Africa and Australia used for molecular analyses. a The Western Australian Kwinana population used for both chloroplast DNA (cpDNA) and sequence and genomic SNPs (gDNA), b, d Samples used for chloroplast DNA analysis. c, e Samples used for genomic SNP analysis, including the samples from the previous study by Byrne et al. (2022). The global bitou bush distribution is based on records sourced GBIF.org (15 September 2022) GBIF Occurrence Download https://doi.org/10.15468/dl.nabx5p (basis of record: preserved specimen) with post-download cleaning and quality control applied
We filled gaps in the previous whole-genome single nucleotide polymorphism (SNP) analysis by Byrne et al. (2022) to address the specific questions of this study. Additional samples (48) were processed using double-digest restriction site-associated DNA sequencing (ddRAD) sequencing and analysis of SNPs. These samples were obtained from recent surveys and collections in South Africa during 2019 and New South Wales, Australia, during 2020. This included 34 samples from the east coast of Australia (predominantly New South Wales populations) where there were spatial gaps in the previous study. Of these, 29 were identified as bitou bush based on morphology and 5 specimens were identified as either boneseed (Chrysanthemoides monilifera subsp. monilifera) or a suspected hybrid based on morphology. These latter samples were included to provide an outgroup for the work, and for confirmation of the subspecies identification. The 14 South African samples were comprised of 4 from Gqeberha (formerly known as Port Elizabeth), 4 from St Francis Bay, and 5 more from populations found further north between Gqeberha and East Beach, the southern-most sampling point of Byrne et al. 2022; Fig. 2). Samples sequenced by ddRAD were collected no later than 2017, except for one DNA extraction from Kenton on Sea (South Africa), provided by Nigel Barker.
For chloroplast sequencing, samples from across the native and non-native ranges were drawn from our existing collections, with additional sampling from across the non-native range in Australia and from representative populations across the native range in South Africa (Fig. 2). Herbarium specimens were subsampled from localities where fresh material could not be obtained. Additional details on the samples are provided in Supplementary Table S2.
DNA extraction for all samples
Following a preliminary assessment against a CTAB method and the DNeasy Plant Kit (Qiagen, Hilden, Germany), we used Synergy 2.0 Plant DNA Extraction Kit (OPS Diagnostics, New Jersey, USA) to obtain genomic DNA from 20 mg of dried plant tissue, according to the manufacturer’s instructions. DNA quality (fragment size) was assessed on a QIAxcel Advanced (Qiagen, Hilden, Germany) and quantity on an Invitrogen Qubit 4 fluorometer using the High Sensitivity assay (ThermoFisher Scientific, Waltham, Massachusetts, USA).
Reduced-representation genome sequencing and SNP analysis
The ddRAD libraries were prepared following the same procedure as described in Byrne et al. (2022) based on the method described by Severn-Ellis et al. (2020). For each sample, 200 ng of genomic DNA was digested with 5 units (0.5 µL) of each of the restriction enzymes HpyCh4IV and Hinfl (New England Biolabs, Ipswich, USA) and ligated to the barcode sequences using T4 DNA ligase (Invitrogen, Carlsbad, USA). Size selected fragments (250–800 bp) were amplified and indexed using Phusion Hot-Start High-Fidelity Polymerase Master Mix (Thermo Fisher Scientific, Waltham, USA) and 0.5 µM each of the indexed PCR2 and PCR1 primers of Peterson et al. (2012). The amplified libraries were purified, and the DNA concentrations were determined by HS Qubit assay. Equimolar amounts of the libraries were pooled and purified to remove any remaining DNA fragments < 200 bp using 0.8 X Ampure XP beads (Beckman Coulter, Brea, CA, USA). The final library was assessed for quality, size distribution, and concentration with the LabChip GX Touch (Perkin Elmer, Waltham, Massachusetts, USA) and Qubit HS assay. The final ddRADseq libraries were submitted to the Australian Genome Research Foundation (AGRF; Melbourne, Australia), for 150 bp paired-end sequencing using Illumina NovaSeq (Illumina, San Diego, California, USA).
The raw ddRAD sequencing data generated in this study was combined with the raw ddRAD data from Byrne et al. (2022) after which data analysis was carried out following the method of Severn-Ellis et al. (2020), available at https://github.com/ascheben/RAD_analysis_workflow, with minor modifications as described in Byrne et al. (2022). First, the new raw data set was filtered and demultiplexed using the STACKS2 process_radtags function (Rochette et al. 2019). The length limit was set to 95 bp to match existing data from the Byrne et al. (2022) study. Trimmed and filtered sequences were deposited at the National Center for Biotechnology Information (NCBI) under the BioProject PRJNA813745. SNP calling of the combined data set was carried out using STACKS2 (Rochette et al. 2019) with parameters -m = 2, -M = 2, -m = 2 for the bitou bush only and bitou-pisifera data sets and -m = 3, -M = 3, -n = 3 for the combined bitou-pisifera-boneseed data set as described by Byrne et al. (2022) whereafter all SNPs were exported to VCF using STACKS-populations.
Identified SNPs were filtered using VCFTOOLS (Danecek et al. 2011). Individuals with > 90% missingness data were first removed, followed by genotype calls with a read depth < 5. Biallelic SNPs with < 20% missing genotypes and minor allele frequency (MAF) > 0.05 were retained. The impact of linked SNPs from the same RAD locus on population genetic analyses was reduced through the random selection of a single SNP from each locus and the discarding of the other SNPs. Finally, a post-filtering check for missingness was carried out to remove individuals with > 50% missing genotypes. Population summary statistics, including pairwise FST and AMOVA-based statistics (ΦST and FST) were calculated using STACKS populations for all populations except for KOS for which there was only one individual.
Relationships and structure within the bitou bush and outgroup populations were determined as described in the Byrne et al. (2022) study. Identified SNPs were converted to Phylip format using a Python script (Ortiz 2019), whereafter the filterHets.py Python script from https://github.com/ascheben/RAD_analysis_workflow was used to remove all SNPs without homozygous alternate allele genotypes, to generate an input file suitable for RAXML. Maximum likelihood phylogeny was inferred with RAXML 8.2.11 (Stamatakis 2014) using a model with ascertainment bias correction (ASC_GTRGAMMA) and rapid bootstrapping with 1000 bootstraps. The resulting tree was visualised using Figtree v1.4.4 (https://github.com/rambaut/figtree/releases).
fastSTRUCTURE v 1.0 (Pritchard et al. 2000; Raj et al. 2014) was used to establish the population genetic structure and runs for 1–10 populations (K) using the default simple prior and convergence criterion, while the optimal value of K was selected by using fastSTRUCTURE chooseK. Data was visualised using the POPHELPER package (Francis 2017). Finally, SNPrelate 1.24.0 (Zheng et al. 2012), was used to carry out principal component analysis (PCA) in R.
Chloroplast DNA sequencing and assembly
DNA libraries were prepared for genome skimming using NEBNext Ultra II FS DNA Library Prep Kit for Illumina (New England Biolabs, Ipswich, Massachusetts, USA). The protocol for > 100 μg of DNA was used for all samples except four with a lower yield, for which we followed the protocol for < 100 μg of DNA. The digestion step was run for 20 min to obtain fragments of 200–450 bp. Purification steps were done using AMPure XP purification beads (Beckman Coulter, Brea, USA). The DNA libraries were assessed for fragment size and concentration on the QIAxcel and Qubit and pooled to equal concentrations. Libraries were sequenced by AGRF, Melbourne. The first pool of 24 samples was sequenced using Illumina NextSeq (Illumina, San Diego, California, USA). The second pool of 24 samples was sequenced on the newer Illumina NovaSeq (Illumina, San Diego, California, USA) and included three samples from herbarium collections to fill gaps in the distribution of samples. We included sample S621, from Kwinana, Western Australia, in both batches so that a comparison could be made and allow for adjustments should any discrepancy occur.
The DNA sequences were analysed using the high-performance computing (HPC) facilities at the University of Western Australia. Searches of the NCBI GenBank and other databases at the time of analysis (May 2020) did not reveal a published chloroplast genome for any relative of bitou bush within the tribe Calendulae so the chloroplast assembly was conducted de novo. The chloroplast genomes were assembled following the approach described in Nevill et al. (2020) using the package NOVOPlasty (Dierckxsens et al. 2017) designed specifically for plastomes around 150 kbp or less (https://github.com/ndierckx/NOVOPlasty). Several de novo assemblies of sample S154 from the Kwinana population were carried out to generate a circularised reference. The chloroplast was annotated using the software package Chloe (https://chloe.plantenergy.edu.au/annotate.html). The cpDNA sequences were imported as FASTA files into Geneious (Kearse et al. 2012), where all the fully circularised sequences were aligned. A haplotype parsimony network was created using TCS software (Clement et al. 2000), allowing connections with branch lengths of up to 20 nucleotide substitutions.
Results
Historical data synthesis
Introduction into Australia
The first confirmed Australian record of bitou bush remains the herbarium specimen collected in 1908 at Stockton in Newcastle, New South Wales, probably from the ballast field (= ballast ground; Fig. 1). This specimen was collected by the Mayor of Stockton and identified as Osteospermum monilifera (NSW133399; Gray 1976). A second specimen was collected on 1 October 1915, also from Stockton, Newcastle (AD98428122, https://www.gbif.org/occurrence/2828118979; GBIF.org; 15 September 2022; Download https://doi.org/10.15468/dl.nabx5p).
At the time, dry ballast was well known as a source of plant introductions (Ridley 1930). Dry ballast (sand, rocks, building rubble) was used in cargo-free sailing ships to provide stability and was unloaded at the ballast ground, Stockton, Newcastle, to prepare the sailing ships to take on coal, which was then transported mostly to the Americas. Newcastle was and continues to be a major port for the export of coal from Australia. In 1904, half (i.e., 51% of 793,629 tons) of the ballast arriving in Australia was landed from 228 vessels (sail 54%, steam 46%) in Newcastle (Great Britain Board of Trade 1906). In 1907, 2 million tons of ballast were deposited at Newcastle, comprising most of the ballast arriving in Australia (Scanlon 2021). Such a vast amount of ballast was dumped that the land reclaimed by the deposits now forms a large area of the suburb of Stockton. In contrast, ships arriving at other Australian ports brought cargo and departed with cargo and therefore did not need ballast.
Over the previous decade (1898–1908), Stockton town councillors had complained that the ballast field contained potentially invasive weeds, so it is possible that bitou bush was present 10 years before, but not identified. Indeed, bitou bush was declared a noxious weed by the Stockton council in 1909 (Hamilton et al. 2012). In summary, the evidence above suggests that the initial introduction of bitou bush from its native region of South Africa to Australia happened in Newcastle, New South Wales via accidental arrival by ship’s ballast.
Ports of departure
Bitou bush material could have come from anywhere in its native range, but not as a secondary introduction from a non-native population given that there were no known introductions anywhere else in the world. However, the association with ballast meant that the plant must have in some way entered the ballast transport system at a port and was dispersed via this mechanism to Australia. Ports for shipping activity on the east coast of South Africa in 1904 that overlap the distribution of bitou bush were located in (from south to north along the eastern coast) Port Elizabeth (now known as Gqeberha), East London, Port St John, Port Natal (Durban), and Lourenço Marques (Maputo, Mozambique; Table 1). Active investigation did not produce any information on the latter port; however, the location is at the northern most extreme of the bitou bush distribution. Port St John had a very small volume of trade and is therefore less likely to be the source of bitou bush (Table 1). This scenario leaves Port Elizabeth, East London and Durban as the most likely potential sources. Ships arriving in Port Elizabeth and East London carried mostly cargo, whereas many ships arriving in Durban were loaded with ballast (39%, presumably looking for cargo). Overall, more ships (57% of 1494 departures) were leaving the eastern coast of South Africa empty, aside from ballast.
Ballast provisioning
Evidence for dry ballast provisioning in South African ports was hard to find, despite it being a common practice and typically sourced from a quarry located close to the port. At Port Elizabeth, the source of ballast was from a local quarry, with rocks initially transported by trolley over land, then subsequently carried by surfboat out to the ships at large of the port. Later a depot was established for ballast at the end of the breakwater (http://thecasualobserver.co.za/port-elizabeth-of-yore-quarries-for-development/), however we found no record of exactly how ballast provisioning worked in the period of interest (c. 1900). It is not known if bitou bush was growing nearby the ballast quarry. All traces of the quarry have now vanished under expanding urban development. In Durban, ballast consisted of sand extracted directly from Sandy Bay (today called South Beach and North Beach or Durban Bight) or from a quarry on The Bluff, at the entrance to the harbour. These locations could have been an ideal habitat for bitou bush, although the area has become extensively urbanised today. Likewise, no detailed description of ballast provisioning processes was found for Durban in the time period of interest. By 1902 the Anglo-Boer War had ended and most outgoing transport was by military ships from Durban to Australian capital cities and not Newcastle (and therefore did not involve dry ballast; Plowman 2003).
The clearest example of a shipping association with bitou bush was at the port of East London, which occupies the mouth of the Buffalo River in Eastern Cape Province (Fig. 3a). Only East London has historical evidence that could be found relating to facilities for loading ballast in the first decade of the 1900s. A map of the Port of East London from 1920 shows a ballast quarry, a tramway for cocopans (hoppers) connecting the quarry to the ballast staith for loading ballast onto ships (Fig. 3b). Photos held by the East London Museum show this infrastructure to be present in 1905. Some of these ships (mostly sailing ships) would have unloaded coal into the coal sheds, ready to fuel steam-driven vessels. Ships sailed “in ballast” from East London to Newcastle in the decades before and after 1900. The historical shipping records of 1890–1910 (sourced from Trove.nla.gov.au) documented the travel of at least 18 ships sailing “in ballast” or without cargo (i.e., inferring the use of ballast) from East London, South Africa to Newcastle, Australia. These sailing ships, mostly barques (Fig. 1), rounded Tasmania and did not stop at intervening ports. However, the Trove repository only holds Australian newspapers and documents and not the records of East London. Therefore, the number of ships moving between the two ports could be a substantial underestimation.
Significant geographic locations for the invasion history of bitou bush (Chrysanthemoides monilifera subsp. rotundata). a Mouth of Buffalo River and Port of East London, South Africa, circa 1920. b Likely source location of bitou bush in and around the East London port quarry connected to the ballast staith (= wharf; Map source: East London Museum, South Africa). c Introduction point (red dot) for bitou bush in Kwinana, Western Australia (Photo taken 18 Feb 1986. Sourced from the collections of the State Library of Western Australia and reproduced with the permission of the Library Board of Western Australia). d The likely original bitou bush plant for Kwinana, Western Australia (Photo: John K. Scott, CSIRO)
Introduction context
Assuming the distribution of bitou bush in East London has not substantially changed, the transfer of seed and sections of stem of bitou bush into ballast would have been feasible, given that the plant grows next to and inside the now abandoned ballast quarry. Sections of bitou bush stem could also have been transported, providing an alternative introduction mechanism. Bitou bush layers (i.e., establishes from soil-covered stem sections) and the voyage to Newcastle from South Africa only took 36 days. Our observations at Kwinana in Western Australia are that stem sections will last and regrow over a much longer period of burial. The volume of ballast transported could allow for a significant quantity of seeds or stem material to be transported in the ballast.
Spread in eastern Australia
Bitou bush was originally documented as confined to a limited area of the coastal environment adjacent to the Hunter River and Newcastle in New South Wales (Love 1986) in the mid 1940s, confirming the likely area of initial establishment. From the 1950s onwards, bitou bush was actively planted along the New South Wales coast to stabilise dunes after mining activity (Barr and Atkinson 1970; Love 1986; Mort and Hewitt 1953; Weiss et al. 2008). Seedlings were collected from the field and grown on in a nursery prior to field planting in preference to planting seed (Barr 1965). There is no record of additional material being imported from South Africa to be used in these plantings. The active planting of bitou bush ceased in about 1971 when its invasive potential was recognised (Cooney et al. 1982). By 2000, 80% of the New South Wales coast had been planted or invaded by bitou bush and it had spread into areas of native vegetation. The New South Wales Department of Environment and Conservation subsequently issued a threat abatement plan in 2006 (Department of Environment and Conservation (NSW) 2006).
Introduction into Western Australia
Dry ballast in ships had ceased to be used by the time of the introduction of bitou bush into Kwinana, Western Australia. There are at least three alternative scenarios for an introduction pathway: plants introduced from eastern Australia for landscaping or dune stabilisation, material introduced as a shipping-related contaminant via vessels originating in eastern Australia, or by direct introduction from the native range (South Africa). Analysing high-resolution historical aerial photography enabled us to extrapolate as to the presence of bitou bush in the Kwinana industrial area to as early as 1995 (Scott et al. 2019a). However, seed can persist without germination for up to seven years (Scott et al. 2019b), so the actual year of introduction could have been earlier than 1995. From our annual surveys and an extensive aerial photographic record, we identified two plants as the likely initial invaders (Fig. 3c, d). Dead plants leave a recognisable skeleton and only one large and three smaller skeletons were found in the initial population delimitation. Consequently, we think that we have found every plant that ever survived at Kwinana past seedling stage. While it is possible that plants were transported off site in earth moving equipment this seems unlikely as very extensive searches (Batchelor et al. 2021; Scott et al. 2019b) have not found bitou bush outside of the controlled access site at Kwinana. These original individuals have since been removed as part of the eradication program.
The first scenario is that the bitou bush was planted in the sand dunes at Kwinana. All the surrounding land and jetty at this time was controlled by the steel manufacturing company, Broken Hill Proprietary (BHP). Construction of the Kwinana plant began following agreement with the state of Western Australia in 1952 and by 1954 the factory was rolling merchant sections (fence posts) from feedstock shipped from Newcastle, New South Wales (BHP Steel Rod and Barr Productions Division Kwinana Market Mill April 1956–April 1995; Sawer 1985). A blast furnace was added to the activities in the late 1960s and was supplied with coke from Newcastle and Port Kembla (ports with bitou bush). Evidence of sand dune accretion on the shores of Cockburn Sound, the region surrounding the Kwinana site, was present from 1984. BHP carried out extensive revegetation at other sites over long periods, maintained inhouse plant nurseries and revegetated extensive areas (e.g., Whyalla and Broken Hill; Anon. 1935; Stirling 1962).
Photographic evidence shows sand dune accretion (1986 shoreline from photo; Fig. 3c), in relation to the shoreline in Google Earth (2018) where the dunes have increased by some 10–15 m on the eastern shores of Cockburn Sound. A photo of the Kwinana site from 1986 (Fig. 3c) also shows lines of vegetation (small trees or shrubs, oleanders, eucalypts) some of which have survived until recently. Unfortunately, we were unable to track down records of any plant nursery and landscaping activities for the Kwinana area.
A second plausible scenario is that two plants colonised the sand dune near a jetty used to unload steel billets (crude material for feedstock for a rolling mill). BHP had a fleet of ships servicing ports including Newcastle, Port Kembla and Kwinana (e.g., Iron Monarch and Iron Duke; Sawer 1985). It is therefore possible that bitou bush was introduced via ships as a contaminant associated with steel billet shipments.
The least-likely scenario is direct transport to Kwinana from South Africa in the second half of the twentieth century. As far as can be ascertained, there was no trade between these two ports post World War 2. A 1992 map of BHP’s shipping routes reveals no destination ports in South Africa (BHP Transport Limited 1992). Furthermore, BHP ceased operations at Kwinana in 1995 and terminated its agreements with the state of Western Australia in 2005 before vacating the site (Chance 2006). Thereafter the site has been classified as a mine site, with a high level of security and controlled access (Scott and Batchelor 2014). Meanwhile the increase of the bitou bush population to some 1,700 plants went unnoticed until 2012.
Reduced-representation genomics and SNP analyses
Sequencing, SNP calling, and population genetic statistics
A total of 441.7 million reads were generated for the bitou bush samples sequenced in the current study. After low-quality reads and reads without RAD-tags were removed, 323.9 million filtered reads were retained for processing. Roughly, the de novo SNP calling identified 1,972,195 variant sites from the 191 samples in the extended bitou-pisifera-boneseed data set. After filtering, 177 samples remained, and 13,054 variable sites were retained. For the bitou-pisifera-only data set, de novo SNP calling identified 2,186,722 variant sites from the 166 individuals, of which 157 individuals and 10,486 variable sites were retained after filtering.
The expected heterozygosity (He) of filtered SNPs in the bitou-pisifera dataset (excluding the Kenton-on-Sea population from South Africa with only one sample) ranged from 0.096 ± 0.002 for the population of pisifera and 0.139 ± 0.002 in St Lucia (South Africa) to 0.288 ± 0.002 for eastern Australia (Supplementary Table S3). However, the values for South African populations may be inaccurate due to small sample sizes. For measures of pairwise genetic differentiation, the ΦST between bitou bush populations (excluding Kenton-on-Sea) was generally higher between populations in the native range, and lower between populations in the non-native range (Supplementary Table S4). The ΦST between Western Australia and eastern Australia was 0.117. The highest ΦST between South African sites was 0.368 between the geographically widespread St Lucia and St Francis Bay, and the lowest was 0.0401 between Hluleka and Port St John. The ΦST between bitou bush and pisifera populations was high (0.312–0.464). The ΦST between native and non-native populations ranged from 0.195 between Gqeberha and eastern Australia, to 0.430 between Tugela Mouth and Western Australia.
Phylogenetic and population structure analyses
Phylogenetic analysis of the extended data set (Fig. 4) revealed no major phylogenetic rearrangements in comparison to the phylogeny of Byrne et al. (2022). As noted previously, the Australian populations formed a clade, and this clade was most closely related to a clade of samples from southern and central sites in South Africa (including Gqeberha, East London, Qolora Mouth and Hluleka; Fig. 4). Samples from the more northerly locations around Durban and Saint Lucia were more distantly related. The phylogeny showed that the Western Australian samples formed a clade within a paraphyletic group of eastern Australian samples. Additional native range samples from the southern range of the subspecies distribution (Kenton-on-Sea, St Francis Bay, and Sardinia Bay) formed a paraphyletic group at the base of the phylogeny, showing a close relationship to the outgroup subspecies pisifera.
Phylogeny based on SNPs of bitou bush (Chrysanthemoides monilifera subsp. rotundata) in South Africa and Australia using Chrysanthemoides monilifera subsp. pisifera as an outgroup. South African samples are labelled according to population of origin (dark blue dots). Samples from the putative introduction point to Australia, Stockton, are also shown (light blue dots)
The PCA of bitou bush and pisifera shows the samples separated into east and west in Australia and a cline from south to north in the South African samples (Fig. 5). With the addition of boneseed, the PCA separates the subspecies along PC1 with bitou bush grouped together on the left and boneseed on the right, with pisifera in between but closer to bitou bush (Supplementary Fig. S1).
Molecular analyses of bitou bush (Chrysanthemoides monilifera subsp. rotundata) populations in South Africa and Australia. a PCA analysis. b fastSTRUCTURE analysis for the optimal K-value of 6. Populations: DUR; Durban, DWE; Dwesa, EAB; East Beach, EAU; Eastern Australia (states not separated), ELD; East London, HLU; Hluleka, HOL; Hole in the Wall, KOS; Kenton-on-Sea, MZN; Mtunzini, PEL; Gqeberha (formerly known as Port Elizabeth), PSJ; Port St John, QMO; Qolora Mouth, SFB; St Francis Bay, STL; Saint Lucia, TMO; Tugela Mouth, WAU; Western Australia
fastSTRUCTURE analysis of bitou bush only samples found the optimal K-value, according to the method of (Puechmaille 2016), to be K = 6 (Supplementary Fig. S2). At K = 6, genetic clusters corresponded to one population in Western Australia, one in eastern Australia and four in South Africa. The four clusters in South Africa (north to south) were: 1) St Lucia to Durban, 2) Hluleka and Dwesa, 3) Qolora Mouth and East London, and 4) East Beach to St Francis Bay. The clustering of this study is consistent with the clustering of bitou bush documented in Byrne et al. (2022), with the southern-most cluster representing the additional samples included in our analysis.
Chloroplast DNA molecular analyses
Fully circularised chloroplast genomes were assembled for 33 samples, of which 32 were bitou bush and one was boneseed (Supplementary Table S2). Of these assemblies, 24 were 151,881 bp, nine of them were slightly different lengths, while the boneseed sample was 151,911 bp in size. The remaining nine samples were partial assemblies of varying lengths. However, these partial assemblies could be aligned without evident SNPs to the complete chloroplast assemblies. Only the complete chloroplast assemblies were used in subsequent analyses.
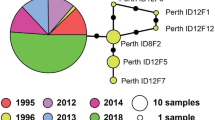
The haplotype parsimony network, created using TCS software, shows strong geographic structure throughout the native range (Fig. 6a, b). Four samples, representing three unique haplotypes, from East Beach to St Francis Bay, the southern limits of the subspecies distribution, were strongly divergent and unable to be connected to the remainder of the network within the threshold of 20 substitutions. The remaining native range samples show a unique haplotype in each location, with the network structured geographically along a north–south axis. The Australian samples all share the same haplotype as the sample from East London.
Chrysanthemoides monilifera subsp. rotundata haplotype network. a The haplotype network between the native range of South Africa and the non-native range of Australia for chloroplast DNA sequences. Cross bars are nodes of difference, each a definite SNP, the dotted line is more than 20 SNPs. Groups of similar haplotypes are coloured the same. The size of the dots relates to the number of sequences obtained. b Geographic distribution of haplotype groups (orange dotted lines) for sampled populations (pink dots) in South Africa in relation to large rivers (potential geographic barriers)
Discussion
Using multiple lines of inquiry, including historical accounts of dispersal pathways between regions, plant biology, and DNA-based molecular ecology, we have been able to reveal the origins and invasion history of bitou bush in Australia. We have greatly narrowed down the origin of the introduced material from the native range and have identified the introduction pathway from this origin in South Africa to (and within) the non-native range in Australia. That these diverse sources of information coalesce around a single coherent story serves to improve confidence in our interpretations and to improve the utility of these findings for guiding management.
Biogeographical origin
From the molecular analysis, past work on ddRAD analysis of populations from the native range points to southern populations as the putative origin of plant material that was introduced from South Africa into Australia (Byrne et al. 2022). Our study builds on the data set used by Byrne et al. (2022), expanding it by adding a further 48 samples. However, our genomic analysis did not reveal the biogeographical origin; the chloroplast DNA was more informative for this purpose, as surmised from the conserved nature of this plastid. The phylogenetic reconstruction from genomic DNA SNPs of different samples supported evidence that the East London population is the closest relative to the Australian material. The genetic differentiation between populations (ϕST) was low between East London and the eastern Australian populations at 0.08, but slightly lower between eastern Australia and the populations of Gqeberha (0.078) and Kenton-on-Sea (0.07). The PCA of the genome of bitou bush alone shows differentiation between the samples in eastern and Western Australia, distinct from the native range (Fig. 5a). However, genetic differentiation between populations was lowest between the eastern and Western Australian populations. Considering that the Kwinana population establishment in Western Australia is relatively recent, and most likely introduced from eastern Australia, this agrees with both the SNP-based phylogeny and the identical chloroplast genome.
Contextualising this genetic variation in both the native and non-native range provides further understanding into whether this single location of origin for material in Australia is a robust conclusion. Our results were consistent with those of Byrne et al. (2022), in showing considerable genetic structure using the genomic DNA in bitou bush’s native range in South Africa, as well as differentiation between native and non-native populations. The highest ϕST was between two populations in South Africa—Kenton-on-Sea and St Lucia—which are geographically far apart (800 km). The greatest differences occurred between populations that are spatially distant or those that are separated by a geographic barrier, such as a large river (Fig. 6b). The distribution of the South African locations, using only the x-axis of the PCA, is suggestive of a cline where samples are spatially distributed from south to north (Fig. 5a). We have not made any biological interpretations based on the y-axis of the PCA, as this appears to be compromised by the Guttman effect (also known as the horseshoe or arch effect), where the y-axis information could be misinterpreted, or overinterpreted, where the pattern is created based on how the data were input (Diaconis et al. 2008; Goodall 1954; Podani and Miklós 2002). This effect has also been shown to occur with SNP data, as a result of how the SNPs are coded (i.e., which alleles are coded as 0 and 1) and the PCA method used for data transformation and graphical representation (Gauch et al. 2019).
Our genetic analyses of chloroplast DNA showed that the population structure was high in the native range of South Africa and low in eastern Australia. Ten unique haplotypes were determined by chloroplast analysis, with only one of the haplotypes present in both countries. Each site in South Africa had a different chloroplast sequence (except Kenton-on-Sea and Sardinia Bay sharing a haplotype), with branch lengths on the network from 1 to 6 substitutions and greater than 20 substitutions between the main network and the samples from Kenton-on-Sea, Sardinia Bay, St Francis Bay, and East Beach (Fig. 6a). This finding indicates a high degree of diversity in the native range, despite the plant being widespread and with seed dispersed by birds. While a single chloroplast haplotype across the introduced range could be the result of a selective sweep eliminating other haplotypes that may have been introduced (e.g., Muir and Filatov 2007), we did not observe any significant conflict in phylogenetic relationships between the data from the chloroplast genome and the genomic SNPs to support this scenario. Further investigation of the ecology of the weed is required to identify the limits to dispersal, which seem to be present in bitou bush in its native range.
From a historical records perspective, many of the elements that enabled the transport of bitou bush to Australia are still present at the port of East London or captured in historical photos and maps of the port. Plants, including bitou bush, now overgrow the long disused ballast quarry. Indeed, the sample used for the molecular studies was collected just 1400 m from the ballast quarry (Fig. 3b). There were no confirmed records of introduction into Australia (similar to the detailed ships logs of plants that were intentionally distributed), which strongly supports the conclusion that the introduction was accidental. This cryptic dispersal pathway makes historical records more challenging for understanding accidental introductions, such as ballast contamination. Because of this challenge, historical information on pathways of dispersal can sometimes provide greater understanding into the putative origin for cryptic introductions.
Introduction pathways to Australia
Our genetic analyses revealed that only one chloroplast haplotype is present in all the Australian samples, yet ten are present across the native range in South Africa. The single maternal lineage present in Australia appears in all the samples and locations across four states. This would be consistent with the introduction from a single population in the native range, without additional introductions from other populations. This haplotype was identical to that of the sample from East London in South Africa, so we would expect the introduction into Australia to be from this population.
Strong genetic bottlenecks were detected in all Australian bitou bush populations (Byrne et al. 2022). Despite these bottlenecks, bitou bush successfully invaded Australia at least once in eastern Australia and in Kwinana in Western Australia, an area previously modelled to be climatically unsuitable for this shrub (Adair et al. 2012). The breeding system of the plant and the number of maternal lineages can influence the likelihood of establishment after introduction, for example if there was high genetic variation amongst limited founders or if there was little variation but a higher number of individuals (Barker et al. 2015; Cristescu 2015; Estoup and Guillemaud 2010).
Consistent with Byrne et al. (2022), SNP data showed that Kwinana was a distinct population without evidence of ongoing gene flow with other populations. There was a subsequent reduction in private alleles with each founding event. However, there were no indications of inbreeding, even in the Western Australian population that has endured two founding events. This is likely due to the outcrossing mating system, and few generations post-introduction in Western Australia (Byrne et al. 2022). There was less structure in the eastern states of Australia, compared to the native range. There is usually a reduction in genetic variation within populations and differentiation between populations in the non-native range, compared to that found in the native range (Dlugosch and Parker 2008). The results suggest that this reduction occurred with the founding event of bitou bush in Australia.
Considering the evidence together, we conclude that the most plausible explanation is that one introduction (e.g., multiple seeds or stem fragments) of bitou bush occurred from South Africa to Australia. Much later, a different introduction extended the population from eastern Australia to Western Australia as seeds or stem fragments, seedlings or as landscape plantings. The establishment of a naturalised introduction from a single introduction event is an unusual occurrence amongst non-native invasive plants. Wilson et al. (2009) compared the genetic diversity in native and introduced plant populations and found that only Olea europaea cuspidata, one of 14 species, was derived from a single introduction, so in this aspect bitou bush is unusual.
Available historical information from the literature and other sources from the 1900s, suggests that bitou bush material as either seed or stem sections may have become incorporated in the ballast extracted from the quarry on the west bank of the Buffalo River in East London, South Africa (there was only one ballast quarry, Fig. 3a, b). The extracted ballast was transported from the quarry into a sailing ship waiting at the ballast wharf (Fig. 3a, b). Bitou bush seeds accumulate under plants producing seed (despite being bird dispersed; Scott 1996; Scott et al. 2019b). The ballast would have consisted of rocks and soil containing seeds. The ship then likely sailed directly to Stockton, New South Wales, Australia, without stopping. This introduction is likely to have been nearer to 1900, to allow for germination and establishment, and for the plant to be noticed, instead of 1908 when the first herbarium specimen was made in Australia. More than one genetically unique individual was involved, making it more likely that seeds made the voyage (rather than stem fragments or plants).
The value of bitou bush for dune stabilisation in Australia was soon realised and consequently, the human-assisted spread escalated. Eventually the entire New South Wales coastline was infested. Further spread in the eastern states (human-assisted and natural spread) occurred to Melbourne, Victoria and southern Queensland and to Kinchega National Park (Menindee Lakes) in inland New South Wales (Weiss et al. 2008). While historical evidence was limited for highlighting where this additional material came from, the genomic analysis was able to provide clarity. Both the genomic DNA and cpDNA results point to there being one introduction into Australia. We are, therefore, able to conclude with confidence that no additional importations were made, despite the extensive propagation and spread of the plant in eastern Australia by human means. Given the proactive redistribution of bitou bush for dune stabilisation across the Australian east coast, it is surprising that only one chloroplast haplotype was introduced from the native range. Our findings suggest that past deliberate dispersal of bitou bush by humans within the introduced eastern Australian range was probably more important than currently realised.
Regarding the Western Australian population at Kwinana, no documented evidence was found to reveal a deliberate introduction, nor was there any evidence that could have suggested an introduction pathway direct from South Africa. The pathway from eastern Australia to Western Australia is not clear, as the possibility of human-assisted (but undocumented) planting or contamination of steel billet shipments are both plausible. If we were to assume that the latter pathway was more likely, given the lack of formal garden landscaping in the heavily industrialised Kwinana area, then the pathway of “stowaway” (in ballast) followed by “escape” as is escaping from plantations and finally “contaminant” (Hulme et al. 2008) would apply.
Outcrossing and hybrids
Beyond the two primary questions that set out the scope for this work, our findings have provided additional information on the population dynamics for bitou bush that can inform management in both the native and non-native regions. First, while there was a reduction in diversity in the introduced range, there was no reduction in heterozygosity. This provides further support to bitou bush being an obligate outcrossing species based on glasshouse studies showing a lack of pollination (Gross et al. 2017), and an absence of seed production in isolated plants (Scott et al. 2019b). The latter is an important element in any eradication attempt due to the additional time available for controlling spatially isolated plants (i.e., no seed production). Second, and based on the phylogenetic patterns, the southern St Francis Bay and Port Elizabeth genetic samples from South Africa are clustering between bitou bush and pisifera. This finding suggests the material could be a hybrid between these two subspecies. For cpDNA they would be expected to have either the bitou or pisifera haplotype because there is no recombination (Fig. 6a). The sampling did not include pisifera cpDNA, so further analysis would be required to resolve this matter.
Implications for management
Given that bitou bush can establish in areas that were previously thought to be climatically unsuitable, such as Kwinana in Western Australia, an expanded understanding of biosecurity risk is warranted. For example, improved awareness at entry points along invasion pathways, such as trade routes, is essential for preventing new incursions within and beyond the current Australian range. This global risk is equally applicable to other countries that have both trade links with South Africa and comparable climates. We recognise that the risk of dry ballast as a mediator of accidental introductions has reduced considerably in the past 100 years. From the end of the 1890s up to World War 1 shipping transferred from wooden sailing ships to iron-clad steamers. This led to a major change in ballast, from “dry” ballast to wet ballast (i.e., sea and fresh water), which is used today. The change to wet ballast has created its own set of problems due to its role as a dispersal vector for aquatic invasive species (Robinson et al. 2020). Either way, there remains a diverse set of pathways for accidental plant introductions, and proactive caution remains an essential priority for any biosecurity program looking to mitigate introduction risks (Hulme et al. 2008).
The current likelihood of eradication of bitou bush in Western Australia shows that successful management of established populations is a realistic outcome if control strategies are appropriately assessed and actioned (Behrendorff et al. 2019; Scott et al. 2019b). Eradication has the advantage of a finite cost, whereas containment and control methods, which are the only feasible options available to the eastern states of Australia, remain ongoing indefinitely into the future (Bradshaw et al. 2021; Diagne et al. 2020, 2021).
To improve control in regions where eradication is not possible, further work on novel biological control agents from bitou bush’s biogeographical origin is worth pursuing. If successful, such a control approach would likely lead to a significant reduction in management costs and less damage to native ecosystems from harmful treatments such as fire and herbicide (Behrendorff et al. 2019; van Wilgen et al. 2020). Up to now, biological control efforts with bitou bush have mostly failed due to prospective agents for bitou bush not establishing or lacking impact (Adair et al. 2012). Few of these agents were sourced from the southern parts of the range and came from northern regions around Durban, largely due to easier logistics. Our findings provide a rationale to target the southern end of the bitou bush distribution around East London (i.e., this is the likely biogeographical origin of the bitou bush population in Australia) where new agents may be revealed. For example, one possible agent overlooked until now is the flea beetle, Serraphula elongata Jacoby (Coleoptera: Chrysomelidae: Alticinae). This insect has a distribution including East London and is known to feed on Chrysanthemoides (Biondi and D'Alessandro 2010). This type of insect has been used successfully in the biological control of a range of weed species (Syrett et al. 1996). Investigating S. elongata populations from near East London should be a high priority in any new agent surveys.
Caveats and limitations
Applying an integrated approach to elucidating invasion context by drawing on both genomic and historical evidence can provide unprecedented insight, however we are aware that both sources of information have inherent limitations and biases. Regarding genomics, while the evidence is strong that East London is the South African source population for the Australian introduction, it is not possible to entirely exclude Durban and Gqeberha (Port Elizabeth) from the historical evidence, as modifications to the port areas due to urbanisation may have erased evidence of ballast production and resident populations of bitou bush. Given the strong geographic variation of bitou bush in the native range, it would be unlikely for these populations to have a similar genotype to those from East London and Australia. However, this scenario could still be plausible if there was movement of plant material between these ports due to shipping activities, or deliberate planting of genotypes from elsewhere. Sequencing additional herbarium specimens from South Africa could resolve this issue.
Historical evidence gathering is often an iterative process. While we were systematic and thorough in our approach to information gathering, we recognise there may remain additional information that we overlooked. The synthesis provided here could well act as a prompt for others to reveal further data sources. Moreover, we are aware that some sources of historical information do not necessarily give an accurate or unbiased rendering of the past. An example using the Trove digital database (also the major historical source used in this study) is the accuracy of newspaper records of house sparrow (Passer domesticus) introductions into Australia, where the number of sparrows released and their origin are contested (Andrew and Griffith 2016). In contrast, the records of ship movements in the shipping news component of Trove are more robust and can generally be independently verified using multiple sources. The advantage of our integrated approach is that both data sources can fill information gaps of the other, thereby working together in a complementary way to provide a more complete invasion context (e.g., Fischer et al. 2015).
Future research
While our research provides novel findings, and improved resolution for the introduction history of bitou bush from South Africa into Australia (and subsequent movements within Australia), our work has raised three points of note where further research would be worth pursuing.
First, getting a clearer understanding of the within-Australia movements between the east coast and Western Australia could be clarified by a pathway analysis test. Bitou bush arrived and established in Western Australia near a steel rolling mill in the industrial port of Kwinana. An opportunity to begin a post hoc test of the generality of this introduction was recently pursued by examining for bitou bush presence at another isolated steel rolling mill serviced by the same ships coming from the same putative source populations in New South Wales (i.e., Newcastle, Port Kembla) via the same companies and in the same era as the Kwinana introduction. This ‘twin’ is Whyalla in South Australia. In a preliminary pathway analysis, we (a) consulted weeds officers from Whyalla in 2022 to enquire about the presence of bitou bush, and (b) undertook visual analyses of high-resolution aerial photography (Google Earth, Nearmap) for vegetation in the town that could be bitou bush. Neither investigative avenue revealed the presence of bitou bush in Whyalla. This prima facie evidence maybe the first example of a pathway analysis test in the field of plant biosecurity and invasion ecology and warrants further investigation given the potential climatic suitability of the area.
Second, while our understanding of genetic diversity for bitou bush in its native range has been greatly improved, analysis of further specimens from the native and introduced range, including historical collections, could provide additional insights. Our phylogenomic analyses have detected the possibilities of potential biogeographic barriers in the native range. Further sampling throughout the range, particularly on both sides of the major geographic divisions that exist throughout the range, could improve understanding of the genetic structure in the native range and test hypotheses of biogeographic barriers (Fig. 6b; e.g., the east–west flow of rivers as potential barriers, a north–south cline). Such understanding could confirm the findings of the cpDNA analysis as more SNPs between samples coincided with larger rivers or obstructions between sites. Greater sampling resolution would also investigate if the maternal lineage from East London was widespread in the native range or localised. Including historical herbarium specimens from the initial introduction to Australia, along with contemporary samples from South Africa could be informative on introduction history and post-introduction evolution (Lopez et al. 2020). For example, this could determine whether additional genotypes were introduced but subsequently lost through selective sweeps. There is established methodology for genome skimming for chloroplast genome assembly of historical specimens (e.g., Bakker et al. 2016) and recently methods have been developed to obtain data comparable to ddRAD-seq from historical specimens (Lang et al. 2020). In addition, if a chromosomal-level reference genome assembly could be generated for bitou bush or a close relative, this would be invaluable in enabling the mapping of SNPs to chromosome positions for the detection of selective sweeps in the nuclear genome and detection of variation associated with invasive traits (e.g., Turner et al. 2021).
Last, our work points to further information on the significant historical introduction pathway that was dry ballast into Australia, particularly in the Newcastle region of New South Wales. Heyligers (2008) surveyed the flora of the North Stockton area of Newcastle (about 2 km north of the ballast grounds that are now playing fields) and found that 18 of the 102 plant taxa found in the local flora could be associated with an initial introduction via dry ballast. He also encountered abundant bitou bush. Not listed was fireweed (Senecio madagascariensis), a significant species that on circumstantial evidence we propose took the same route as bitou bush (Dormontt et al. 2014; Sindel et al. 1998). A reanalysis of the chloroplast variation for this species from the Eastern Cape Province, South Africa, could help confirm if this introduction pathway is shared.
Conclusion
Our study adds to an emerging number of studies that take an integrated approach to recreating the invasion context of non-native introductions to novel regions (Alvesa et al. 2022), including for introductions mediated by dry ship’s ballast (Brawley et al. 2009). By combining historical, and two types of genetic data, it was possible to resolve the question of the origins of bitou bush in Australia. This should lead to more effective management for the species now and into the future. The two contrasting genomic approaches (genomic SNPs and chloroplast DNA) provided complementary results for both the location of origin as well as the invasion pathway, where either approach on its own would have been insufficient. When combined with our third line of evidence, derived from historical shipping and industrial development accounts, we were able to create a robust interpretation of the invasion context for bitou bush in Australia, and better understand the biogeography of the native range in South Africa. Turning this information into improved management, via refocusing where to search for novel biocontrol agents, is now the next logical step to manage this Weed of National Significance in Australia. More broadly, this work also highlights the ongoing importance of identifying the high-risk invasion pathways, including weak points in biosecurity perimeters, which can improve management outcomes.
Data availability
The high-throughput sequencing data from ddRAD and genome skimming are available from NCBI short-read archive under BioProjects PRJNA814435 and PRJNA813745, respectively. The whole chloroplast genomes are available from NCBI GenBank under accession numbers OP604609-OP604641. Other datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Adair RJ, Morley T, Morin L (2012) Chrysanthemoides monilifera (L.) T.Norl. - bitou bush and boneseed. In: Julien M, McFadyen R, Cullen J (eds) Biological control of weeds in Australia. CSIRO Publishing, Collingwood, pp 170–183
Alpern SB (2008) Exotic plants of Western Africa: Where they came from and when. Hist Af 35:63–102. https://doi.org/10.1353/hia.0.0018
Alvesa J, Carneiro M, Day J et al (2022) A single introduction of wild rabbits triggered the biological invasion of Australia. PNAS 119:e2122734119. https://doi.org/10.1073/pnas.2122734119
Andrew SC, Griffith SC (2016) Inaccuracies in the history of a well-known introduction: a case study of the Australian House Sparrow (Passer domesticus). Avian Res 7:1–15. https://doi.org/10.1186/s40657-016-0044-3
Anon. (1935) Successful tree cultivation. BHP Review Jubilee Number: 39–40
Austin DF (2014) Sendera-clandi (Xenostegia tridentata (L.) D.F. Austin & Staples, Convolvulaceae): a medicinal creeper. Ethno Res App 12:433–454
Bakker FT, Lei D, Yu J et al (2016) Herbarium genomics: plastome sequence assembly from a range of herbarium specimens using an Iterative Organelle Genome Assembly pipeline. Biol J Lin Soc 117:33–43. https://doi.org/10.1111/bij.12642
Barker NP, Paterson I, Howis S (2015) “Barcoding” and ISSR data illuminate a problematic infraspecific taxonomy for Chrysanthemoides monilifera (Calenduleae; Asteraceae): lessons for biocontrol of a noxious weed. Biochem Syst Ecol 61:541–551. https://doi.org/10.1016/j.bse.2015.07.021
Barr DA (1965) Restoration of coastal dunes after beach mining. J Soil Conserv Ser NSW 21:199–209
Barr DA, Atkinson WJ (1970) Stabilization of coastal sands after mining. J Soil Conserv Ser NSW 26:89–107
Batchelor KL, Scott JK, Webber BL (2021) Boneseed management in Western Australia A review and synthesis of past management and its outcomes for Chrysanthemoides monilifera subsp. monilifera, with recommendations for future control. A report prepared for the Western Australian Department of Primary Industries and Regional Development, pp 114
Behrendorff L, Harris SM, Muirhead IF (2019) Towards eradication: the history and management of bitou bush on K’gari-Fraser Island, Australia. Ecol Manag Rest 20:92–100. https://doi.org/10.1111/emr.12349
BHP Steel Rod and Barr Productions Division Kwinana Market Mill (April 1956–April 1995) A place in history. BHP
BHP Transport Limited (1992) The Iron ships: a maritime history of BHP 1885–1992. BHP Transport Limited, Melbourne
Biondi M, D’Alessandro P (2010) Revision of the Afrotropical flea beetle genus Serraphula Jacoby and description of Bechynella, a new genus from Western and Central Africa (Coleoptera: Chrysomelidae: Alticinae). Zootaxa 2444:1–44. https://doi.org/10.5281/zenodo.195007
Blackburn TM, Pysek P, Bacher S et al (2011) A proposed unified framework for biological invasions. Trends Ecol Evol 26:333–339. https://doi.org/10.1016/j.tree.2011.03.023
Bradshaw C, Hoskins A, Haubrock P et al (2021) Detailed assessment of the reported economic costs of invasive species in Australia. NeoBiota 67:511–550. https://doi.org/10.3897/neobiota.67.58834
Brawley S, Coyer J, Blakeslee A et al (2009) Historical invasions of the intertidal zone of Atlantic North America associated with distinctive patterns of trade and emigration. PNAS 106:8239–8244. https://doi.org/10.1073/pnas.0812300106
Byrne D, Scheben A, Scott JK et al (2022) Genomics reveals the history of a complex plant invasion and improves the management of a biological invasion from the South African-Australian biotic exchange. Ecol Evol 12:e9179. https://doi.org/10.1002/ece3.9179
Chance K (2006) BHP Billiton (termination of agreements) agreement bill 2005. Government of Western Australia Hansard 12 April 2006: 1559b–1560a
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1659. https://doi.org/10.1046/j.1365-294x.2000.01020.x
Colautti RI, Alexander JM, Dlugosch KM et al (2017) Invasions and extinctions through the looking glass of evolutionary ecology. Philos Trans R Soc B 372:20160031. https://doi.org/10.1098/rstb.2016.0031
Cooney PA, Gibbs DG, Golinski KD (1982) Evaluation of the herbicide “Roundup” for control of bitou bush (Chrysanthemoides monilifera). J Soil Conserv Ser NSW 38:6–12
Cristescu ME (2015) Genetic reconstructions of invasion history. Mol Ecol 24:2212–2225. https://doi.org/10.1111/mec.13117
Cullen J (2012) Chondrilla juncea L. skeleton weed. In: Julien M, McFadyen R, Cullen J (eds) Biological control of weeds in Australia. CSIRO Publishing, Collingwood, pp 150–161
Danecek P, Auton A, Abecasis G et al (2011) The variant call format and VCFtools. Bioinformatics 27:2156–2158. https://doi.org/10.1093/bioinformatics/btr330
Daru P, Park DS, Primack RB, Willis CG et al (2018) Widespread sampling biases in herbaria revealed from large-scale digitization. New Phytol 217:939–955. https://doi.org/10.1111/nph.14855
Department of Environment and Conservation (NSW) (2006) NSW Threat Abatement Plan—invasion of native plant communities by Chrysanthemoides monilifera (bitou bush and boneseed). Department of Environment and Conservation (NSW), Hurstville, pp 85
Diaconis P, Goel S, Holmes S (2008) Horseshoes in multidimensional scaling and local kernel methods. Ann App Stat 2:777–807. https://doi.org/10.1214/08-AOAS165
Diagne C, Leroy B, Gozlan RE et al (2020) InvaCost, a public database of the economic costs of biological invasions worldwide. Sci Data 7:277. https://doi.org/10.1038/s41597-020-00586-z
Diagne C, Leroy B, Vaissiere AC et al (2021) High and rising economic costs of biological invasions worldwide. Nature 592:571–576. https://doi.org/10.1038/s41586-021-03405-6
Dierckxsens N, Mardulyn P, Smits G (2017) NOVOPlasty: de novo assembly of organelle genomes from whole genome data. Nucleic Acids Res 45:e18. https://doi.org/10.1093/nar/gkw955
Dlugosch K, Parker I (2008) Founding events in species invasions: genetic variation, adaptive evolution, and the role of multiple introductions. Mol Ecol 17:431–449. https://doi.org/10.1111/j.1365-294X.2007.03538.x
Dormontt EE, Gardner MG, Breed MF et al (2014) Genetic bottlenecks in time and space: Reconstructing invasions from contemporary and historical collections. PLoS ONE 9:e106874. https://doi.org/10.1371/journal.pone.0106874
Estoup A, Guillemaud T (2010) Reconstructing routes of invasion using genetic data: why, how and so what? Mol Ecol 19:4113–4130. https://doi.org/10.1111/j.1365-294X.2010.04773.x
Estoup A, Ravigné V, Hufbauer R et al (2016) Is there a genetic paradox of biological invasion? Ann Rev Ecol Evol Syst 47:51–72. https://doi.org/10.1146/annurev-ecolsys-121415
Faulkner KT, Hulme PE, Pagad S et al (2020) Classifying the introduction pathways of alien species: are we moving in the right direction? NeoBiota 62:143–159. https://doi.org/10.3897/neobiota.62.53543
Fischer M, Hochkirch A, Heddergott M et al (2015) Historical invasion records can be misleading: Genetic evidence for multiple introductions of invasive iaccoons (Procyon lotor) in Germany. PLoS ONE 10:e1025441. https://doi.org/10.1371/journal.pone.0125441
Forno IW, Harley KLS (1979) The occurrence of Salvinia modesta in Brazil. Aquat Bot 6:185–187. https://doi.org/10.1016/0304-3770(79)90061-5
Francis RM (2017) POPHELPER: an R package and web app to analyse and visualize population structure. Mol Ecol Res 17:27–32. https://doi.org/10.1111/1755-0998.12509
French K, Watts E (2015) Patterns of loss of biodiversity associated with invasion by Chrysanthemoides monilifera subsp monilifera (boneseed) across a large geographic region. Weed Res 55:537–545. https://doi.org/10.1111/wre.12168
French K, Ens E, Gosper CR et al (2008) Management implications of recent research into the effect of bitou bush invasion. Plant Prot Q 23:24–28
Friedel MH (2020) Unwelcome guests: a selective history of weed introductions to arid and semi-arid Australia. Aust J Bot 68:75–99. https://doi.org/10.1071/BT20030
Gauch HG, Qian S, Piepho HP et al (2019) Consequences of PCA graphs, SNP codings, and PCA variants for elucidating population structure. PLoS ONE 14:e0218306. https://doi.org/10.1371/journal.pone.0218306
Gaskin JF, Schaal BA (2002) Hybrid Tamarix widespread in U.S. invasion and undetected in native Asian range. PNAS 99:11256–11259. https://doi.org/10.1073/pnas.132403299
Gildenhuys E, Ellis A, Carroll S et al (2013) The ecology, biogeography, history and future of two globally important weeds: Cardiospermum halicacabum Linn. and C. grandiflorum Sw. NeoBiota 19:45–65. https://doi.org/10.3897/neobiota.19.5279
Goodall D (1954) Objective methods for the classification of vegetation. III. An essay in the use of factor analysis. Aust J Bot 2:304–324. https://doi.org/10.1071/BT9540304
Goolsby JA, De Barro PJ, Makinson JR et al (2006) Matching the origin of an invasive weed for selection of a herbivore haplotype for a biological control programme. Mol Ecol 15:287–297. https://doi.org/10.1111/j.1365-294X.2005.02788.x
Gray M (1976) Miscellaneous notes on Australian plants. 2. Chrysanthemoides (Compositae). Contributions from Herbarium. Australiense 16:1–5
Great Britain Board of Trade (1906) Statistical tables relating to British colonies, possessions, and protectorates, Part XXIX 1904, London, Printed for H.M. Stationary Office, by Darling & Son Ltd
Griffioen RC (1995) A taxonomic study of Chrysanthemoides Tourn. ex Medik. (Compositae). Botany Department. University of Cape Town, pp 177
Gross CL, Whitehead JD, de Souza CS et al (2017) Unsuccessful introduced biocontrol agents can act as pollinators of invasive weeds: Bitou bush (Chrysanthemoides monilifera ssp. rotundata) as an example. Ecol Evol 7:8643–8656.https://doi.org/10.1002/ece3.3441
Hamilton MA, Winkler MA, Cherry H et al (2012) Changes in the distribution and density of bitou bush (Chrysanthemoides monilifera subsp. rotundata (DC.) T.Norl.) in eastern Australia. Plant Prot Quart 27:23–30
Heyligers PC (2008) Flora of the Stockton and Port Hunter sandy foreshores with comments on fifteen notable introduced species. Cunninghamia 10:493–511
Hulme PE (2009) Trade, transport and trouble: managing invasive species pathways in an era of globalization. J Appl Ecol 46:10–18. https://doi.org/10.1111/j.1365-2664.2008.01600.x
Hulme PE, Bacher S, Kenis M et al (2008) Grasping at the routes of biological invasions: a framework for integrating pathways into policy. J Appl Ecol 45:403–414. https://doi.org/10.1111/j.1365-2664.2007.01442.x
Julien M, McFadyen R, Cullen J (2012) Biological control of weeds in Australia. CSIRO Publishing, Collingwood
Kearse M, Moir R, Wilson A et al (2012) Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. https://doi.org/10.1093/bioinformatics/bts199
Kriticos DJ, Webber BL, Leriche A et al (2012) CliMond: global high-resolution historical and future scenario climate surfaces for bioclimatic modelling. Method Ecol Evol 3:53–64
Kueffer C (2017) Plant invasions in the Anthropocene. Science 358:724–725. https://doi.org/10.1126/science.aao6371
Lang PLMM, Weiß CL, Kersten S et al (2020) Hybridization ddRAD-sequencing for population genomics of nonmodel plants using highly degraded historical specimen DNA. Mol Ecol Resour 20:1228–1247. https://doi.org/10.1111/1755-0998.13168
Lindenmayer DB, MacGregor C, Dexter N et al (2013) Booderee National Park management: connecting science and management. Ecol Manag Rest 14:2–10. https://doi.org/10.1111/emr.12027
Lindenmayer DB, Wood J, MacGregor C et al (2015) A long-term experimental case study of the ecological effectiveness and cost effectiveness of invasive plant management in achieving conservation goals: Bitou bush control in Booderee National Park in eastern Australia. PLoS ONE 10:e0128482. https://doi.org/10.1371/journal.pone.0128482
Lopez L, Turner KG, Bellis ES et al (2020) Genomics of natural history collections for understanding evolution in the wild. Mol Ecol Resour 20:1153–1160. https://doi.org/10.1111/1755-0998.13245
Love A (1986) Bitou bush and boneseed current control techniques and programmes. In: Love A, Dyason R (eds) Bitou bush and boneseed: proceedings of a conference on Chrysanthemoides monilifera. NSW National Parks and Wildlife Service and NSW Department of Agriculture, Sydney, 8–9 August 1984, Port Macquarie, NSW, pp 95–104
Marshall VM, Lewis MM, Ostendorf B (2012) Buffel grass (Cenchrus ciliaris) as an invader and threat to biodiversity in arid environments: a review. J Arid Environ 78:1–12. https://doi.org/10.1016/j.jaridenv.2011.11.005
Mort GW, Hewitt BR (1953) Vegetation survey of the marine sand drifts of New South Wales some remarks on useful stabilising species. J Soil Conserv Ser NSW 9:59–69
Muir G, Filatov D (2007) A selective sweep in the chloroplast DNA of dioecious silene (section Elisanthe). Genetics 177:1239–1247. https://doi.org/10.1534/genetics.107.071969
Nevill PG, Zhong X, Tonti-Filippini J et al (2020) Large scale genome skimming from herbarium material for accurate plant identification and phylogenomics. Plant Methods 16:8. https://doi.org/10.1186/s13007-019-0534-5
Norlindh T (1943) Studies in the Calenduleae. 1. Monograph of the genera Dimorphotheca, Castalis, Osteospermum, Gibbaria and Chrysanthemoides. CWK Gleerup, Lund
Novoa A, Richardson DM, Pysek P et al (2020) Invasion syndromes: a systematic approach for predicting biological invasions and facilitating effective management. Biol Invas 22:1801–1820. https://doi.org/10.1007/s10530-020-02220-w
Ortiz EM (2019) vcf2phylip v2.0: convert a VCF matrix into several matrix formats for phylogenetic analysis. https://zenodo.org/record/2540861#.XSLdYU17nrc. Accessed 8 July 2019
Peterson BK, Weber JN, Kay EH et al (2012) Double digest RADseq: an inexpensive method for de novo SNP discovery and genotyping in model and non-model species. PLoS ONE 7:e37135. https://doi.org/10.1371/journal.pone.0037135
Plowman P (2003) Across the sea to war. Rosenberg Publishing Pty Ltd
Podani J, Miklós I (2002) Resemblance coefficients and the Horseshoe Effect in Principal Coordinates Analysis. Ecology 83:3331–3343. https://doi.org/10.2307/3072083
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1093/genetics/155.2.945
Puechmaille S (2016) The program structure does not reliably recover the correct population structure when sampling is uneven: subsampling and new estimators alleviate the problem. Mol Ecol Res 16:608–627. https://doi.org/10.1111/1755-0998.12512
Raj A, Stephens M, Pritchard JK (2014) fastSTRUCTURE: variational inference of population structure in large SNP data sets. Genetics 197:573–589. https://doi.org/10.1534/genetics.114.164350
Ridley HN (1930) The dispersal of plants throughout the world. L. Reeve and Co. Ltd, Ashford
Robinson TB, Peters K, Brooker B (2020) Coastal invasions: the South African context. In: van Wilgen BW, Measey J, Richardson DM, Wilson JR, Zengeya TA (eds) Biological invasions in South Africa. Invading nature—Springer Series in Invasion Ecology, pp 229–247
Rochette NC, Rivera-Colón AG, Catchen JM (2019) Stacks 2: analytical methods for paired-end sequencing improve RADseq-based population genomics. Mol Ecol 28:4737–4754. https://doi.org/10.1111/mec.15253
Roman J, Darling JA (2007) Paradox lost: genetic diversity and the success of aquatic invasions. Trends Ecol Evol 22:454–464. https://doi.org/10.1016/j.tree.2007.07.002
Sawer D (1985) Australians in Company. The Broken Hill Propriety Company Limited in its 100th year. The Pot Still Press Pty Limited, Artarmon
Sax DF, Brown JH (2000) The paradox of invasion. Glob Ecol Biogeogr 9:363–371. https://doi.org/10.1046/j.1365-2699.2000.00217.x
Scanlon M (2021) What secrets are burried in Stockton's ballast ground? In: Newcastle Herald. https://www.newcastleherald.com.au/story/7189136/standing-on-foreign-soil-in-stockton/. 2 Apr 2021
Scott JK (1996) Population ecology of Chrysanthemoides monilifera in South Africa: implications for its control in Australia. J Appl Ecol 33:1496–1508. https://doi.org/10.2307/2404788
Scott JK (2012) Tribulus terrestris L. caltrop. In: Julien M, McFadyen R, Cullen J (eds) Biological control of weeds in Australia. CSIRO Publishing, Collingwoo, pp 576–580
Scott JK, Batchelor KL (2014) Management of Chrysanthemoides monilifera subsp. rotundata in Western Australia. Invasive Plant Sci Manag 7:190–196. https://doi.org/10.1614/ipsm-d-13-00052.1
Scott JK, McKirdy SJ, van der Merwe J et al (2017) Comprehensive biosecurity protects high-conservation-value island nature reserve from intensive development. Sci Rep 7:1–9. https://doi.org/10.1038/s41598-017-00450-y
Scott JK, Batchelor KL, Jucker T et al (2019a) Aerial photography and dendrochronology as tools for recreating invasion histories: do they work for bitou bush (Chrysanthemoides monilifera subsp. rotundata)? Biol Invas 21:2983–2996. https://doi.org/10.1007/s10530-019-02026-5
Scott JK, Batchelor KL, Webber BL (2019b) Long term monitoring of recruitment dynamics determines eradication feasibility for an introduced coastal weed. NeoBiota 50:31–53. https://doi.org/10.3897/neobiota.50.35070
Severn-Ellis AA, Scheben A, Neik TX et al (2020) Genotyping for species identification and diversity assessment using double-digest restriction site-associated DNA sequencing (ddRAD-seq). In: Jain M, Garg R (eds) Legume genomics: methods and protocols, methods in molecular biology. Springer Nature, pp 159–187
Simberloff D, Martin J-L, Genovesi P et al (2013) Impacts of biological invasions: what’s what and the way forward. Trends Ecol Evol 28:58–66. https://doi.org/10.1016/j.tree.2012.07.013
Sindel BM, Radford IJ, Holtkamp RH et al (1998) The biology of Australian weeds. 33. Senecio madagascariensis Poir. Plant Prot Q 13:2–15
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Stirling FS (1962) Whyalla's western myall trees. BHP Review April 1962:Inside cover, 10–12
Syrett P, Fowler SV, Emberson RM (1996) Are chrysomelid beetles effective agents for biological control of weeds? In: Moran VC, Hoffmann JH (eds) Proceedings of the IX international symposium on biological control of weeds. Stellenbosch, South Africa, 19–26 January 1996, pp. 399–407
Thorp JR, Lynch R (2000) The determination of weeds of national significance. National Weeds Strategy Executive Committee, Launceston
Turner KG, Ostevik KL, Grassa CJ et al (2021) Genomic analyses of phenotypic differences between native and invasive populations of diffuse knapweed (Centaurea diffusa). Front Ecol Evol 8:577635. https://doi.org/10.3389/fevo.2020.577635
van Wilgen BW, Raghu S, Sheppard AW et al (2020) Quantifying the social and economic benefits of the biological control of invasive alien plants in natural ecosystems. Curr Opin Insect Sci 38:1–5. https://doi.org/10.1016/j.cois.2019.12.004
Webber BL, Born C, Conn BJ et al (2011) What is in a name? That which we call Cecropia peltata by any other name would be as invasive? Plant Ecol Diver 4:289–293. https://doi.org/10.1080/17550874.2011.610372
Weiss PW, Noble IR (1984) Status of coastal dune communities invaded by Chrysanthemoides monilifera. Aust J Ecol 9:93–98. https://doi.org/10.1111/j.1442-9993.1984.tb01347.x
Weiss PW, Adair RJ, Edwards PB et al (2008) Chrysanthemoides monilifera subsp. monilifera (L.) T.Norl. and subsp. rotundata (DC.) T.Norl. Plant Prot Q 23:3–14
Wilson JRU, Dormontt EE, Prentis PJ et al (2009) Something in the way you move: dispersal pathways affect invasion success. Trends Ecol Evol 24:136–144. https://doi.org/10.1016/j.tree.2008.10.007
Zheng X, Levine D, Shen J et al (2012) A high-performance computing toolset for relatedness and principal component analysis of SNP data. Bioinformatics 28:3326–3328. https://doi.org/10.1093/bioinformatics/bts606
Acknowledgements
We thank Reece Luxton, Peter Michael, Ruth Armstrong, Kim Waterhouse, Hellen Haapakoski, John Hodgon, Hillary Cherry, Brendon Yetman, Stacy Harris, Lynette Willsher, Mark Thomas, Andrew Johnson and Roger Cousens who assisted with the sampling of bitou bush across eastern Australia. We thank Barbara Murphy, Team Leader Landscape Operations East, Peninsula Landscape Board, Whyalla for checking bitou bush presence at Whyalla, South Australia. We thank Vincent Ralph Clark and Nigel Barker for sampling bitou bush from South Africa. We thank Jacqui Batley and David Edwards at the University of Western Australia for provision of laboratory facilities and advice on experimental design. We thank Ian Small and Ian Castleden for arranging bioinformatics advice and access to HPC and bioinformatics software. Also, thanks to David Edwards and Phillipp Bayer for tutorials on Linux and bioinformatics at UWA. We especially thank William Martinson and the staff at the East London Museum for sharing information on ballast at the port of East London. We thank Mariana Campos and Gavin Hunter for their comments on a draft of the manuscript.
Funding
Open access funding provided by CSIRO Library Services. This work was funded by CSIRO Health and Biosecurity, the University of Western Australia, and Fremantle Ports Authority.
Author information
Authors and Affiliations
Contributions
AME, JKS, BLW and KLB designed the study and organised sample material. AME, AAS-E and KLB carried out the molecular genomics studies. AME, KLB, JKS and BLW drafted the initial manuscript. JKS drafted the history analysis which was reviewed by BLW. All authors contributed critically to drafts and gave final approval for publication.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Consent for publication
All authors consent to publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Emmett, A.M., Scott, J.K., Webber, B.L. et al. Combining multiple lines of evidence to elucidate the origin and introduction pathway of bitou bush (Chrysanthemoides monilifera subsp. rotundata) in Australia. Biol Invasions 25, 1881–1905 (2023). https://doi.org/10.1007/s10530-023-03017-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-023-03017-3