Abstract
Our ability to predict the outcome of invasion declines rapidly as non-native species progress through intertwined ecological barriers to establish and spread in recipient ecosystems. This is largely due to the lack of systemic knowledge on key processes at play as species establish self-sustaining populations within the invaded range. To address this knowledge gap, we present a mathematical model that captures the eco-evolutionary dynamics of native and non-native species interacting within an ecological network. The model is derived from continuous-trait evolutionary game theory (i.e., Adaptive Dynamics) and its associated concept of invasion fitness which depicts dynamic demographic performance that is both trait mediated and density dependent. Our approach allows us to explore how multiple resident and non-native species coevolve to reshape invasion performance, or more precisely invasiveness, over trait space. The model clarifies the role of specific traits in enabling non-native species to occupy realised opportunistic niches. It also elucidates the direction and speed of both ecological and evolutionary dynamics of residing species (natives or non-natives) in the recipient network under different levels of propagule pressure. The versatility of the model is demonstrated using four examples that correspond to the invasion of (i) a horizontal competitive community; (ii) a bipartite mutualistic network; (iii) a bipartite antagonistic network; and (iv) a multi-trophic food web. We identified a cohesive trait strategy that enables the success and establishment of non-native species to possess high invasiveness. Specifically, we find that a non-native species can achieve high levels of invasiveness by possessing traits that overlap with those of its facilitators (and mutualists), which enhances the benefits accrued from positive interactions, and by possessing traits outside the range of those of antagonists, which mitigates the costs accrued from negative interactions. This ‘central-to-reap, edge-to-elude’ trait strategy therefore describes the strategic trait positions of non-native species to invade an ecological network. This model provides a theoretical platform for exploring invasion strategies in complex adaptive ecological networks.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biotic interactions are ubiquitous and essential for the survival of any species (von Humboldt 1807; Darwin 1859; Thompson 2013). This applies equally to non-native species in novel environments where invasiveness is mediated by habitat suitability, propagule pressure, and the payoffs resulting from interactions with other species within the recipient ecological network (Hui and Richardson 2017). In this regard, the reasons for the success or failure of a non-native species cannot be assessed in isolation but depend on the co-occurrence and interactions with resident species (Enders et al. 2020; Hui et al. 2020; Traveset and Richardson 2020). The invasiveness of a non-native species and the invasibility of the recipient ecological network are, therefore, interlinked. Considering biological invasions with reference to ecological networks has the potential to transform our theoretical models, resulting in improved predictability of both invasiveness and invasibility (Hui and Richardson 2019). In particular, a network approach shifts the emphasis away from the current species-centric view that overly emphasises the roles of ecological and environmental barriers imposed sequentially during the invasion progression of an introduced species (i.e., the unified invasion framework; Blackburn et al. 2011).
Here, we consider an adaptive ecological network under eco-evolutionary feedbacks (Minoarivelo and Hui 2016a; Guimarães Jr et al. 2017) in order to expand the concept of invasion fitness and to capture the complex response of the network to biological invasions. We do so by introducing a simple model, with specific examples, to capture the eco-evolutionary dynamics in an adaptive network subjected to biological invasions. Our model is rooted in the progressive understanding of the fitness concept in evolutionary ecology. Although the concept of fitness pre-dates the theory of natural selection (Darwin 1859), the concept of fitness landscape was only proposed about 90 years ago (Wright 1932). It remains a valuable visual and conceptual tool for depicting the relationship between genotype and fitness. For a single-locus diallelic gene in a large and stable population, the fitness landscape can be described as the relationship between allele frequency and mean fitness. The slope of this relationship determines the rate of change in allele frequencies due to natural selection, often leading to allele fixation. The hill-climbing process of evolution via natural selection would drive the genetic makeup of a population to a local peak (Wright 1988; Gavrilets 2004). Mechanisms forming such slopes and driving changes in allele frequency in a population constitute the crux of ‘population genetics’. Following the framework of ‘quantitative genetics’ (Lynch and Walsh 1998), one can consider continuously varying phenotypic traits of a population, as the direct attribute and mapping of its genetic makeups (e.g. Norman et al. 2019). Therefore the fitness landscape can depict the relationship between phenotypic traits and their mean fitness (Gavrilets 2004). In its classic setting, only random genetic drift or temporal changes in the fitness landscape itself can allow the population to escape the local maxima, thereby potentially reaching a higher fitness peak. Evolutionary ecology has focused on uncovering the mechanisms that unlock populations from their local suboptimal fitness peaks. There has been substantial progress in elucidating the dimensions of invasion fitness over the last few decades (Lehmann et al. 2016).
Both population and quantitative genetics, however, have only given scant attention to the ecological and environmental contexts in which evolution takes place (Metz et al. 1992). In our view, capturing the eco-evolutionary dynamics of an ecological network subject to biological invasions requires us to follow a new paradigm. In particular, population size is often assumed to be large and fixed in the old paradigm, which implies that the fitness landscape is a static surface that can only explain local adaptation and optimization of fitness. However, fitness is not static and is intimately tied to an ecological context. To survive and complete its life cycle, an organism must constantly ‘play games’ with co-occurring individuals from the same and other species for resources and opportunities. According to evolutionary game theory, the payoff relies not only on the player’s own strategy (and its associated traits) but also the strategies and abundance of its opponents (Maynard Smith and Price 1973). Moreover, the received payoff only makes sense for a specific environmental context over a relevant spatial and temporal scale (von Neumann and Morgenstern 1944). For this reason, density-dependent fitness was introduced and advocated in the early 1990s (Hofbauer and Sigmund 1990; Metz et al. 1992, 1996). In the new paradigm of Adaptive Dynamics, invasion fitness is considered as a demographic measure of population performance, where trait changes are still mostly considered via incremental evolution with limited mutation rates and steps. As a result, the commonness and rarity of a trait relative to those of others in an ecological network can further affect the strengths of eco-evolutionary feedbacks. With the dynamic demographic performance as invasion fitness (Meszena et al. 2001; Dieckmann and Ferrière 2004; Nowak and Sigmund 2004; Waxman and Gavrilets 2005), this new paradigm has successfully explained a plethora of evolutionary outcomes, especially regarding the evolutionary branching and sympatric speciation via disruptive selection (Geritz et al. 1997; Dercole et al. 2016). Adaptive evolution under this new paradigm does not necessarily drive a species towards the highest fitness in the trait space, but can follow dynamic and interactive, open-ended trajectories (e.g. Red Queen dynamics, evolutionary suicide and traps; Gyllenberg and Parvinen 2001; Kisdi et al. 2002; Zhang et al. 2013). This new paradigm of considering invasion fitness as demographic performance paves the way for us to explore the eco-evolutionary dynamics of an ecological network subject to biological invasions.
This paper expands several elements of the Adaptive Dynamics paradigm to accommodate the unique circumstances in an ecological network subject to invasion by non-native species. Biological invasions represent a special type of disruption to an ecological network. First, suites of traits of non-native species do not necessarily resemble those of resident species. In classic Adaptive Dynamics, novel traits invading a system only appear through small-step mutations via incremental trait evolution. In contrast, studies have shown that invasive non-native species with traits distinct from those of resident species may have a greater chance of establishing and causing substantial impacts (Minoarivelo and Hui 2016b; Divíšek et al. 2018), related to, albeit different from, the classic limiting similarity theory (MacArthur and Levins 1967; Leimar et al. 2013). Second, propagule pressure provides the umbrella effect for explaining invasiveness (Catford et al. 2009; Simberloff 2009), which deviates from Adaptive Dynamics that assumes a negligible rate of novel trait incursion. Third, whereas the classic definition of evolutionary fitness relates to long-term performance, demographic performance can be dynamic and reflects the realised ecological niches for prospective invaders in the novel environment. We could therefore consider the instantaneous demographic performance as a fitness measure. Finally, although eco-evolution time separation is often assumed to differentiate ecological events operating at much faster temporal scales from evolutionary events that play out more slowly, such an assumption often breaks down in the face of non-equilibrium invasion dynamics and potential rapid evolution (Guimarães Jr et al. 2017; Hui and Richardson 2017).
We embrace these unique circumstances in an ecological network subject to invasion by developing a simple model based on the demographic concept of invasion fitness (section "Eco-evolutionary dynamics of ecological networks under invasion"). We show that these features of biotic interactions, being both trait-mediated and density-dependent, are important components to consider when formulating invasiveness and ecosystem invasibility, both of which involve the combined and intertwined eco-evolutionary dynamics of recipient ecosystems and invading species. We also provide four examples that demonstrate the immediate implications of our model for several ecological networks, including a horizontal competitive network, a bipartite mutualistic network, an antagonistic network, and a multitrophic food web (section "Trait-mediated interactions and invasiveness"). Simple analyses of these model networks allow us to infer invasiveness based on the position of non-native species in the feasible trait space in relation to the trait dispersion of other resident species. We also discuss how the model can be used to address questions related to the stability of ecological networks facing biological invasions (section "Discussion"). For instance, what processes could explain the formation of ecological and evolutionary barriers that dictate the survival of related resident species and the performance of non-native species? These complex challenges we face in invasion science (Ricciardi et al. 2017) highlight the need for within-discipline synthesis and cross-discipline integration (Courchamp et al. 2017; Vaz et al. 2017). To this end, our simple mathematical model does not provide the ultimate solution, but, rather, a “hitchhiker’s guide” to formulating the challenge of predicting biological invasions within a tractable framework.
Eco-evolutionary dynamics of ecological networks under invasion
The influx of non-native propagules into an ecological network defined by trait-mediated biotic interactions between species can impose two non-exclusive effects on a resident species: by ecologically modifying its population size (e.g. by increasing predation pressure or competition for common resources), and by changing the trait composition of its individuals through natural selection. We present a simple mathematical model to capture the eco-evolutionary dynamics of all resident species and the incoming non-native species. To reduce the mathematical complexity in what follows, we present the model in three logic steps, representing sequentially (1) the population dynamics of resident species, (2) the trait evolutionary dynamics of resident species, and (3) the invasiveness of an introduced species in the ecological network and the invasibility of the latter. For transparency, we provide necessary details for each step on the reasoning and interpretation behind model formulation. All variables and parameters, as well as their meanings, are listed in Table 1. In this section we discuss the generic model formulation. We then exemplify the model for specific networks.
Trait-mediated population dynamics
Let us assume that there are \(S\) distinct resident species (natives or non-natives) in a network, with species \(i\) described by four time-varying state variables: its population size (\({n}_{i}\)), mean trait value (\({x}_{i}\)), influx rate of propagules (\({\gamma }_{i}\)) from outside the network with their mean trait value (\({z}_{i}\)). Note, for a short span of time, \(\tau\), a number of \({\gamma }_{i}\tau\) propagules are introduced into this network. Without considering the propagule influx, the per-capita population change rate of species \(i\), \({f}_{i}\), is a function of the abundances of all resident species and can be Taylor expanded at zero abundances to its simplest linear form:
where \({r}_{i}\) is the intrinsic rate of increase (i.e. per-capita growth rate in an empty network), and \({\alpha }_{ij}\) (\(=\partial {f}_{i}/\partial {n}_{j}\) at zero abundances) the per-capita interaction impact of species \(j\) on the intrinsic rate of species \(i\), often called the per-capita interaction strength. This forms a typical Lotka-Volterra model. The intrinsic rate of increase of species \(i\) is often assumed to be solely dependent on its trait, \({r}_{i}=r({x}_{i})\). In most cases of biological invasions, it suffices to assume that per-capita interaction strength is solely mediated by traits of involved species (e.g. Doebeli and Dieckmann 2000; Toju 2011), \({\alpha }_{ij}=\alpha ({x}_{i},{x}_{j})\).
When including propagule pressure from external sources, as a constant inflow of individuals at rate \({\gamma }_{i}\) (Simberloff 2009), we could estimate the amount of population change after a short span of time as \({n}_{i}\left(t+\tau \right)-{n}_{i}\left(t\right)={\gamma }_{i}\left(t\right)\tau +{n}_{i}\left(t\right){f}_{i}\left(t\right)\tau\). Dividing both sides by \(\tau\) and letting \(\tau \to 0\), we have the following population dynamics of species \(i\):
This means that, in a closed network (\({\gamma }_{i}=0\)) with trait-mediated interactions, the identity of a species is masked by its trait value in relation to the distribution of traits of other species (\({f}_{i}={f}_{j}\) if \({x}_{i}={x}_{j}\)). In an open network (\({\gamma }_{i}\ne 0\)), the propagule pressure affects the population dynamics of a species and can also affect its trait evolutionary dynamics, as shown below in section “Trait evolutionary dynamics”.
Trait evolutionary dynamics
The web of trait-mediated biotic interactions in an ecological network imposes eco-evolutionary feedbacks and selection forces on the traits of involved species. The traits of a species, relative to others, could therefore affect not only how species perform over ecological time scales, but also how such traits evolve over much longer (evolutionary) time scales. Adaptive Dynamics has largely been developed to depict how traits evolve under such a complex setting (Dieckmann and Law 1996; Geritz et al. 1998). To express the evolutionary dynamics of the trait, we need to formulate the per-capita growth rate of mutants, \({f}_{i}^{^{\prime}}\). For incremental evolution, the mutant should inherit the ecological function of its source population; that is, mutant individuals of \({x}_{i}^{^{\prime}}\) should interact with others just like species \(i\) with trait \({x}_{i}\) but with minor quantitative differences. In particular, \({f}_{i}^{^{\prime}}\) should resemble, but not be identical to, \({f}_{i}\). Moreover, we here only consider mutants of \({x}_{i}^{^{\prime}}\) with \({n}_{i}^{^{\prime}}\to 0\) arising within the resident population \({n}_{i}\), while individuals with different traits from propagule influx will be considered below as the introduction of individuals with different traits. Following Meszena et al. (2005), we do not consider complete time separation between ecological and evolutionary processes, and residents are thus not necessarily at the ecological equilibrium. This allows us to account for the potential rapid evolution that often accompanies biological invasions (Prentis et al. 2008). We therefore have the following per-capita growth rate of mutants (also called the invasion fitness),
Accordingly, we define the selection gradient here explicitly as the partial derivative of invasion fitness \({f}_{i}^{^{\prime}}\) with respect to the mutant trait, and evaluated at \({x}_{i}^{^{\prime}}\to {x}_{i}\). Consequently, we have the following selection gradient of resident species \(i\) in the ecological network (see Appendix S1):
In Eq. (4), the selection gradient is imposed by two factors: the sensitivity of the intrinsic rate of increase to the trait change (first term on the right hand side), and the sensitivity of the total interaction pressure experienced by an average individual of species \(i\) to its trait change (second term on the right hand side). The canonical equation of Adaptive Dynamics states that the rate of trait evolution is proportional to the selection gradient (Dieckmann and Law 1996; Geritz et al. 1998): \({\dot{x}}_{i}={v}_{i}{s}_{i}\), where \({v}_{i}\) reflects a compound factor of trait variability of species \(i\) in the network and is chosen to be a small positive number in practice to scale the pace of trait evolutionary dynamics relative to ecological dynamics.
In addition, based on Eq. (1) we can compute the sensitivity of the demographic fitness of species \(i\) to its trait change (\(\partial {f}_{i}/\partial {x}_{i}\)), defined as the partial derivative of its per-capita population change rate with respect to its trait. Some algebraic manipulation suggests that this sensitivity of fitness to trait change is related to, but distinct from, the selection gradient, \(\partial {f}_{i}/\partial {x}_{i}\ne {s}_{i}\); see Appendix S2. They only become the same when the resident population is removed (\({n}_{i}=0\) and thus itself becomes the mutant). Our formulation of the selection gradient in Eq. (4) follows the relaxed definition in Adaptive Dynamics (Dieckmann and Law 1996; Geritz et al. 1998) and is different to the eco-evolutionary network model that does not consider the difference of the two (fitness sensitivity to trait versus selection gradient) and assumes the selection gradient as a linear function of trait changes (Guimarães Jr et al. 2017).
Propagule influx can alter such trait evolution driven by biotic interactions within an ecological network. As mentioned above, with an influx of propagules the amount of population change of species \(i\) from time \(t\) to \(t+\tau\) equals the amount of influx propagules \({\gamma }_{i}\left(t\right)\tau\) plus the amount of population change within the network, \({n}_{i}\left(t+\tau \right)-{n}_{i}\left(t\right)={\gamma }_{i}\left(t\right)\tau +{n}_{i}\left(t\right){f}_{i}\left(t\right)\tau\). Let \({z}_{i}\left(t\right)\) be the trait value of influx propagules at time \(t\), so that we can derive the following trait evolutionary dynamics with propagule influx after straightforward algebraic manipulations (see Appendix S3):
In Eq. (5), the trait evolution is driven by two components. The first term on the right hand side depicts how the difference between the trait of influx propagules and the resident trait, \(\left({z}_{i}-{x}_{i}\right)\), can steer the trait dynamics, while the evolutionary force from this component diminishes either when the influx propagules have the same trait as the resident population (\({z}_{i}={x}_{i}\)) or when the influx rate relative to the resident population size drops. This latter scenario could happen either by means of halting the influx (\({\gamma }_{i}\to 0)\) or with the natural increase of the resident population size. This first term therefore describes the effect of propagule pressure on the trait evolution. Although propagule pressure can have a lasting effect on population dynamics (Eq. 2), its role in steering trait evolution is declining with the natural increase of the resident population size (Eq. 5). The second term on the right hand side of Eq. (5) is the relaxed canonical equation of Adaptive Dynamics that captures trait evolution from directional selection along the selection gradient defined in Eq. (4) using population size at the standing point instead of population size at equilibrium. At an evolutionarily singular point (\({\dot{x}}_{i}=0\)), we could further discern more complex scenarios (e.g. disruptive selection and evolutionarily stable strategy) by exploring higher-order derivatives of the right-hand side of Eq. (5) with respect to the focal trait.
Invasiveness in and invasibility of an ecological network
An emerging view in invasion science regarding the performance of a non-native species is that it depends on both the propagule pressure and non-native species traits relative to the traits of resident species. Once introduced, a species becomes a ‘resident’ of the recipient network, although it retains its non-native label. The anticipated demographic performance, or the invasiveness, of the non-native species can be predefined as its per-capita population growth rate \({f}_{A}={\dot{n}}_{A}/{n}_{A}\); the normalisation of \({\dot{n}}_{A}\) by \({n}_{A}\) makes the demographic performance comparable among species. According to its ecological dynamics in Eq. (2), we thus have the following demographic performance:
It is clear that the demographic performance of a non-native species can be partitioned into four components (the right-hand terms in Eq. (6), respectively; schematically shown in Fig. 1): propagule pressure, intrinsic rate of increase, intraspecific density dependence, and trait-mediated interspecific interaction pressure (total per-capita interaction strength from other resident species in the network).
A detailed landscape of invasion fitness, illustrating the purpose of the concepts encompassed in the generic model proposed in this paper. Invasiveness of a non-native species (\({\dot{n}}_{A}/{n}_{A}\), vertical axis) is determined by its trait (\({x}_{A}\)) and population size (\({n}_{A}\)) as the horizontal plane, as well as propagule pressure (\({\gamma }_{A}\)). The blue surface represents invasiveness for once-off introduction (\({\gamma }_{A}=0\)), while the yellow surface represents invasiveness with a constant rate of propagule influx (\({\gamma }_{A}>0\)). Green plane represents zero fitness (for reference purpose). For once-off introduction (\({\gamma }_{A}=0\)), invasion fitness (blue surface) divides the trait axis into a number of positive (+) and negative (−) performance pockets when the initial population size \({n}_{A}\) is small. A non-native species possessing a trait value located in the positive pockets in the trait space will successfully invade, while other traits fail. For a particular non-native trait, its invasiveness also depends on the initial population size: for instance, when \({n}_{A}<{a}_{A}\) the non-native population could suffer from positive density dependence (Allee effect) and fail to establish, but can establish when \({n}_{A}>{a}_{A}\); ultimately, demographic and invasiveness will be constrained by negative density dependence (thus the blue and yellow curves eventually bend downwards when the population size \({n}_{A}\) becomes too large (e.g. exceeding the carrying capacity). Results from actual simulations for specific ecological networks are shown in Fig. 2
We could map these four components onto the unified framework for biological invasions that considers three stages along the continuum of introduction, naturalisation, and invasion (Blackburn et al. 2011). The introduction phase is represented by the first component of the demographic performance, \({\gamma }_{A}/{n}_{A}\), and emphasizes the umbrella effect of propagule pressure. If there is a constant influx of propagules (the yellow surface for \({\gamma }_{A}>0\) in Fig. 1), the non-native species can always grow from a zero initial population size (\({n}_{A}=0\)), with the initial population dynamics solely determined by propagule pressure: \({\dot{n}}_{A}={\gamma }_{A}\). However, this does not guarantee successful establishment and invasion as the roles of other components start to kick in with the increase of population size while the effect of propagule pressure recedes.
As population size (\({n}_{A}\)) increases during the post-introduction invasion stages (naturalisation and eventually invasion), the demographic performance of a non-native species gradually switches to become controlled by the ecological forces of its intrinsic growth rate and combined biotic interaction strength. As part of the post-introduction stages, the second component in Eq. (6) reflects the intrinsic rate of growth, \({r}_{A}\). Large positive values of \({r}_{A}\) contribute to faster population growth, whereas negative values indicate unsuitable abiotic environments or the presence of a critical Allee effect. For instance, if the non-native species experiences a critical Allee effect (i.e. demographic performance increases with density from negative to positive at low population sizes), without a constant propagule influx (\({\gamma }_{A}=0\)), the initial propagule size from the initial introduction needs to exceed the Allee threshold (\({n}_{A}(0)>{a}_{A}\), with \({a}_{A}\) the Allee threshold; blue surface in Fig. 1) to allow the establishment and growth of the non-native population.
The third component in the demographic performance of a non-native species is also important during the post-introduction invasion stages; this reflects the density-dependent self-regulation, with normally \(\alpha \left({x}_{A},{x}_{A}\right)<0\) forcing the demographic fitness to become negative at large population size (Fig. 1). Indeed, any introduced species will eventually become constrained by some negative density dependence when its population size reaches its carrying capacity. Being density dependent, the contribution of this component to the demographic performance and trait dynamics increases proportionally to the population size. This also means that we can ignore the role of this component when considering the initial performance and trait dynamics (when the population size is small).
The fourth component in the demographic performance of a non-native species (Eq. 6) captures the role of interspecific biotic interactions from other resident species in the ecological network. It is the sum of interaction strength from all individuals of other resident species, where the interaction strength depends on the position of the non-native trait in the trait space relative to the resident traits. Through the influence of trait-mediated demographic fitness, interspecific interactions can constrain or expand the fundamental niche of an incoming non-native species in the trait space, forming pockets of positive or negative trait-dependent invasiveness (along the trait axis in Fig. 1). These pockets of positive performance are opportunistic empty niches waiting to be invaded. Consequently, the invasibility of an ecological network is encapsulated by the empty niches in an ecosystem that remain under-exploited by resident species at a given time. Should an introduced species possess a trait within such an empty niche, it can increase its population size from rare, even without continuous influx of propagules. This means that the empty niches in an ecological network can be defined as pockets of trait space (\({x}_{A}\in E\)) with positive demographic performance (\({\dot{n}}_{A}/{n}_{A}>0\)) under zero propagule influx (\({\gamma }_{A}=0\)) and negligible initial propagule size (\({n}_{A}\to 0\)), visualised along the trait axis on the blue surface in Fig. 1. The total width of empty niches over the entire feasible range of trait values thus defines the invasibility of an ecological network, while the invasiveness of a particular invader (with its trait given) is defined by the corresponding height on the demographic performance surface (Fig. 1; Hui et al. 2016).
The evolutionary dynamics of the non-native trait, \({\dot{x}}_{A}\), can also be formulated according to Eq. (5), which gives
Note, as the trait of influx propagules is the trait of the initial non-native population (\({z}_{A}={x}_{A}\)), the first term in Eq. (7) (propagule pressure) does not contribute to the initial trait dynamics of the non-native species. It only starts to steer the trait dynamics once the trait difference emerges although its influence wanes with increasing population size. Propagule pressure (first term on the right hand side) and the selection gradient (second term on the right hand side) can either work synergistically to speed up the trait evolution when the two forces have the same sign; they can also cancel each other to slow down the trait evolution. Equations (2) together with (5), (6) and (7), capture the eco-evolutionary dynamics of all involved species in an ecological network invaded by non-native species. In the next section, we use four examples to illustrate how this generic theoretical model can be used to explore the effect of an invasive trait in mediating invasiveness in complex ecological networks.
Trait-mediated interactions and invasiveness
A crucial step in contextualising the above theoretical model is to parameterise the strengths of trait-mediated interactions between species. This is different from the theoretical framework of May (1972) and Allesina and Tang (2012) where the interaction strength (\({\alpha }_{ij}\)) is considered phenomenologically only, and is not explicitly considered as a function of the traits between involved species. As such, our model also differs from adaptive networks focusing on the evolution of interaction strength (Valdovinos et al. 2018). Instead, we consider the strength of a biotic interaction between two species to be dependent on the traits of both involved species, and it is such trait-mediated interactions that then make the invasiveness dependent on the non-native species’ trait and its position relative to those of the resident species in the network. As illustrated below, different types of interactions can be possibly synthesised into an overarching formula, with the trait-mediated per-capita interaction strength expressed as the Gaussian-type interaction kernel of the trait difference between two interacting species (Doebeli and Dieckmann 2000; Hui et al. 2018):
For a competitive interaction, the sign of the interaction strength is negative and \(\mu =0\). For mutualism, the sign is positive with normally \(\mu =0\). This implies stronger interactions between species with similar traits. For antagonistic interactions (e.g. species \(i\) is the predator, while species \(j\) the prey), the sign of \(\alpha \left({x}_{i},{x}_{j}\right)\) is positive for the predator whereas it is negative for the prey, with \(\mu >0\) the optimal trait difference between the predator and the prey (e.g. in terms of their body sizes). The coefficient \(\sigma\) represents the width of the interaction kernel and depicts the reciprocal of interaction specialisation. A small \(\sigma\) implies that interactions only effectively occur between species with similar traits, whereas a large \(\sigma\) suggests that species with large trait differences can still interact. Coevolution in ecological networks with such trait-mediated interactions has been widely discussed in the literature (e.g. Hui et al. 2018). Although there can be other forms of interaction kernels (e.g. Gallien et al. 2018), they are nevertheless infrequent and suitable only for specific systems. We therefore focus only on the above generic form of interaction kernels. In the next section we provide four examples for invasions into specific ecological networks with species engaging in trait-mediated interactions according to the above Gaussian form.
A competitive community
As we will show, our model suggests that to become invasive, under the scenario of horizontal resource competition, the trait of the non-native species needs to be distinct from those of resident species to elude intense interspecific competition. A horizontal ecological community involves only resource competition between resident species from the same functional guild and is typical in ecology (e.g. light competition between trees in forests).
Using the generic form of Eq. (8), the trait-mediated interaction strength between two competitive species can be formulated as \(\alpha \left({x}_{i},{x}_{j}\right)=-\mathrm{exp}\left(-{\left({x}_{i}-{x}_{j}\right)}^{2}/(2{\sigma }_{P}^{2})\right)\), with \({\sigma }_{P}\) the width of the competition kernel. This kernel is symmetric and the strongest for intraspecific competition (\(\alpha \left({x}_{i},{x}_{i}\right)=-1\)). According to Eqs. (2) and (5), we can thus formulate the ecological and evolutionary dynamics of all resident species before invasion. According to Eqs. (6) and (7), we can formulate a non-native species invading an ecological community with \({S}_{p}\) number of resident species. Here, we only discuss the invasiveness (i.e. the demographic performance) of the non-native species invading this community (Fig. 2a and Appendix S4):
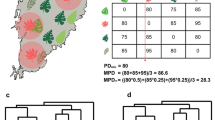
Let \({d}_{Aj}={x}_{A}-{x}_{j}\) be the trait distance between the non-native species and resident species \(j\); \({J}_{P}=\sum {n}_{j}\) the total number of individuals of all resident species (i.e. the community size), and \({w}_{j}={n}_{j}/{J}_{P}\) the relative abundance of species \(j\). Using the Taylor series to expand an exponential function, we have \(\mathrm{exp}\left(-z\right)\approx 1-z\). We define the following centrality of a non-native species with trait \({x}_{A}\) in competitive community P (Fig. 3):
Four examples of eco-evolutionary dynamics on demographic fitness landscapes (see Fig. 1). a A competitive community; b a bipartite mutualistic network between animals and plants with within-guild competition, facing the introduction of a non-native plant species; c a bipartite antagonistic network with within-guild competition, facing the introduction of a non-native resource species; d a food web with interspecific competition. White/grey dots represent trait and population size of resident species [white for plants and grey for animals in (b); white for resources and grey for consumers in (c)]. Arrows represent the joint ecological dynamics (projection along the population size vertical axis) and evolutionary dynamics (projection along the trait horizontal axis) of resident species. For (b) and (c), grey dots and arrows indicate the other functional guild relative to the non-native species. The background colour represents the demographic performance of an incoming non-native (\({\dot{n}}_{A}/{n}_{A}\)), with black contour lines representing zero invasion fitness. White bell-shaped lines represent within-guild non-native trait centrality (measured by Eq. 10), while grey bell-shaped lines represent centrality for exploiting non-native mutualism (b) and non-native resource (c). In (d) dotted line represents centrality for consuming the non-native resource, while dashed line represents the centrality for the non-native to consume resident resource species\(.\) Parameters: a: \(r=1\), \({\sigma }_{P}^{2}=0.005\), \({\gamma }_{A}=0.001\). b: \(r=1\), \({\sigma }_{P}=0.1\), \({\sigma }_{M}=0.05\), \({\gamma }_{A}=0.001\); c: \(r=1\), \({\sigma }_{R}={\sigma }_{P}=0.05\), \({\sigma }_{G}=0.1\), \({\mu }_{G}=0.2,{\gamma }_{A}=0.001\); d: \(r=1\), \({\sigma }_{P}^{2}={\sigma }_{G}^{2}=0.005\), \({\mu }_{G}=0.1\), \({\gamma }_{A}=0.001\). See Appendix S4 for Matlab code
An illustration of centrality over the two-dimensional trait space. Green circles indicate trait positions of twenty species generated randomly (green circle centres), with the corresponding abundances (proportional to the radius) randomly generated from a geometric distribution with a mean of 5. The background contour surface was computed according to Eq. (10) as the centrality
In other words, the centrality of the non-native species within the competitive community is the inverse of the sum of weighted squares of the trait distance between resident species and the non-native species, with the relative abundances of resident species as weights (Fig. 3). This definition reflects the overall pressure of biotic interactions experienced by the non-native species. Our centrality metric is related to, but distinct from, the typical harmonic centrality in network science (Marchiori and Latora 2000). A variant of the denominator has been proposed as an index of biotic novelty (Schittko et al. 2020), and therefore our centrality also captures functional familiarity of an invader to its recipient community.
For simplicity, we consider a small once-off introduction (\({\gamma }_{A}=0\) and \({n}_{A}\to 0\)) and therefore derive the following inequality to ensure the demographic performance \({\dot{n}}_{A}/{n}_{A}\) to be positive:
This suggests that for a non-native species to be able to invade a competitive community the centrality of its trait must be lower than a threshold set by the competition kernel. Invasion is more likely to succeed in communities with more specialised competitors (i.e. a narrower kernel); that is, such communities beget a high level of invasibility. Importantly, the requirement of low centrality suggests that the non-native species needs to possess a trait lying at the periphery of the traits of resident species in the community. In other words, to become invasive, the trait of the non-native species needs to be distinct from those of resident species to elude intense interspecific competition. This conforms to recent macroecological evidence derived from exploring the distributions of traits of vascular plants when divided into natives, archaeophytes and neophytes (Divíšek et al. 2018).
A bipartite mutualistic network
Bipartite mutualistic networks are ubiquitous in nature (Bronstein 2015); they include pollination networks, seed dispersal networks, and below-ground networks of plant–microbe symbiosis (e.g. Steidinger et al. 2019). Within the context of a bipartite mutualistic network, our model suggests that to become a successful invader, a species needs to possess not only traits that are more similar to the centroid of the traits of its mutualistic partners, but also traits that occur towards the periphery of the trait space of resident species from the same guild.
Using the generic form of Eq. (8), the trait-mediated strength of mutualistic interaction can be formulated as \(\alpha \left({x}_{i},{y}_{j}\right)=\mathrm{exp}\left(-{\left({x}_{i}-{y}_{j}\right)}^{2}/(2{\sigma }_{M}^{2})\right)\), where \({x}_{i}\) and \({y}_{j}\) are traits of two species from separate guilds that are engaging in assortative interactions with \({\sigma }_{M}\) the width of the interaction kernel of mutualism. To keep the model analytically tractable, we used the simplest linear functional response for the mutualistic term. A number of \({S}_{P}\) species within the same guild (e.g. flowering plants) are assumed to engage in resource competition as described in the previous section, while a number of \({S}_{M}\) species in the other guild engage in mutualistic interactions with the non-native species. Due to symmetry of interactions, we only formulate the demographic performance of a non-native species from functional guild P (Fig. 2B and Appendix S4):
Similarly, we can define the centrality of the optimal mutualistic position for the non-native species in the trait space of its mutualistic partner guild as \({C}_{A}^{\{M\}}=1/\sum_{k=1}^{{S}_{M}}{d}_{Ak}^{2}{w}_{k}\), with \({d}_{Ak}={x}_{A}-{y}_{k}\) and \({w}_{k}={n}_{k}/{J}_{M}\). Following the same line, let \({\rho }_{P|M}={J}_{P}/{J}_{M}\) be the ratio of community size between the focal guild and the mutualistic partner guild, and we have the following inequality for positive demographic performance,
To get a rough picture of this inequality, if \({\rho }_{P|M}=1\) and \({\sigma }_{P}^{2}={\sigma }_{M}^{2}\), the above inequality can be shortened to \({C}_{A}^{\{M\}}>{C}_{A}^{\{P\}}\); this implies that the invader’s trait should be closer to the centroid of its mutualistic partners than to the centroid of its within-guild competitors so to reap the benefit from mutualism and elude the harm from competition. For the simple scenario where species within the same guild do not engage in resource competition, the above inequality becomes \({C}_{A}^{\{M\}}>1/(2{\sigma }_{M}^{2})\). We can thus conclude that a successful invader in a mutualistic network needs to possess traits towards the centroid of the traits of its mutualistic partners, and in contrast, towards the periphery of the traits of those resident species from the same guild. This also implies that the degree of trait dispersion within functional guilds and the level of trait overlapping between functional guilds could signal the invasibility of a mutualistic network.
Similar to our formulation of the demographic performance, but with a nonlinear functional response for the mutualistic term, the model proposed by Bastolla et al. (2009) has shown that low pressure from interspecific competition is necessary to ensure species coexistence in a mutualistic network. This is consistent with our condition to elude competition pressure via peripheral trait positioning for invasion success. Moreover, the nested structure that characterises many mutualistic networks has been shown to play a key role in reducing competition (Bastolla et al. 2009). It is, however, debatable whether the nested structure can also affect the success of an invasion into such nested mutualistic networks (Minoarivelo and Hui 2016b; Valdovinos et al. 2018). Although the nested structure can minimise the negative pressure from competition, it does not necessarily facilitate the establishment of an invader. As shown here, the invader needs to minimise competition pressure and simultaneously maximise mutualistic gain to ensure a successful invasion.
A bipartite antagonistic network
Bipartite antagonistic networks are also ubiquitous in nature (e.g. Morris et al. 2014; Nuwagaba et al. 2015), and include predator–prey networks and host-parasite networks. Within the context of a bipartite antagonistic network, our model below suggests that to become a successful invader, when the non-native is a resource species (e.g. prey), it should exhibit traits that ensure the traits of its optimal consumer locating at the periphery of the consumer trait space (to reduce consumption rates by residents); when the non-native is consuming resident resources, the position of the non-native species’ optimal resource should be close to the centroid of resident resource trait space (to maximize consumption rates of the invader).
We use subscript C and R to denote the consumer and the resource guilds respectively. Using the generic form of Eq. (8), the trait-mediated interaction strength of resource consumption can be formulated as \(\alpha \left({y}_{C},{x}_{R}\right)=\mathrm{exp}\left(-{\left({y}_{C}-{x}_{R}-{\mu }_{G}\right)}^{2}/(2{\sigma }_{G}^{2})\right)\), where \({x}_{R}\) and \({y}_{C}\) are traits of the resource species and the consumer species from two different guilds, with \({\mu }_{G}>0\) the optimal consumer to resource trait gap and \({\sigma }_{G}\) the width of the resource consumption kernel. We further assume that there are competitive interactions between species within the same guild. Consequently, if the non-native species is part of the resource guild, we have (Fig. 2c and Appendix S4):
Following the same procedure and let \({C}_{A+{\mu }_{G}}^{\{C\}}=1/\sum_{k=1}^{{S}_{C}}{d}_{Ak}^{2}{w}_{k}\), with \({d}_{Ak}={y}_{k}-({x}_{A}+{\mu }_{G})\) and \({w}_{k}={n}_{k}/{J}_{C}\), be the centrality of the maximum consumption position in the consumer trait space to consume the non-native resource species, and \({C}_{A}^{\left\{R\right\}}\) the centrality of the non-native species within the resource guild (as defined in Eq. 10), we thus have the following inequality for a non-native resource species to have positive demographic performance:
Again, if \({\rho }_{R|C}=1\) and \({\sigma }_{R}^{2}={\sigma }_{G}^{2}\), the above inequality can be shortened to \(1/{C}_{A}^{\left\{R\right\}}+1/{C}_{A+{\mu }_{G}}^{\{C\}}>{\sigma }_{G}^{2}\); that is, \({C}_{A+{\mu }_{G}}^{\{C\}}<1/({\sigma }_{G}^{2}-1/{C}_{A}^{\left\{R\right\}})\), the non-native resource species will have a better demographic performance when its optimal consumer position (with trait \({x}_{A}+{\mu }_{G}\)) has a lower centrality. Without competition within the guild, we simply have \({C}_{A+{\mu }_{G}}^{\{C\}}<1/(2{\sigma }_{G}^{2})\). This means that if the non-native resource can only be effectively consumed by resident species at the periphery of the consumer trait space, the non-native resource will experience low levels of consumption from the resident consumers and thus have a higher chance of establishing and invading.
If the non-native species is part of the consumer guild, we have the following:
Let \({C}_{A-{\mu }_{G}}^{\{R\}}=1/\sum_{k=1}^{{S}_{R}}{d}_{Ak}^{2}{w}_{k}\), with \({d}_{Ak}=\left({y}_{A}-{\mu }_{G}\right)-{x}_{k}\) and \({w}_{k}={n}_{k}/{J}_{R}\), be the centrality of the non-native consumer’s optimal resource position in the resource trait space, and \({C}_{A}^{\left\{C\right\}}\) the centrality of the non-native species in the consumer trait space (as defined by Eq. 10). We have the following inequality to ensure the non-native consumer a positive demographic performance:
This bears similarity to the inequality of Eq. (13). Following the similar arguments, if \({\rho }_{C|R}=1\) and \({\sigma }_{C}^{2}={\sigma }_{G}^{2}\), the above inequality can be shortened to \({C}_{A-{\mu }_{G}}^{\{R\}}>{C}_{A}^{\{C\}}\); that is, the optimal resource position of the non-native species needs to be more central in the resource trait space than its position in the competitive consumer trait space. If we ignore within-guild competition, the inequality becomes \({C}_{A-{\mu }_{G}}^{\{R\}}>1/(2{\sigma }_{G}^{2})\); that is, the position of the non-native species’ optimal resource should be closer to the centroid in the resource trait space, to ensure the invasion success of the non-native consumer.
A multitrophic food web
Food webs are a mixture of consumers and resources in a multitrophic ecological network (e.g. freshwater and oceanic food webs), where body size can be the key indicator of who eats whom. Each species in a food web could have three functional roles: as a competitor, a resource, or a consumer. In the context of multitrophic food webs, our model below suggests that the invader’s optimal consumer position should lean towards the trait periphery while its optimal resource position should be central in the trait space. To ensure elevated performance, the non-native species, thus, needs to move from trait periphery to centroid with the increase of its trophic level.
A non-native species invading a food web thus has the following demographic performance (Fig. 2d and Appendix S4):
Following the similar procedure as above, we have the following inequality for a non-native species to possess positive demographic performance in a food web:
Assuming no competition, the above inequality becomes \({C}_{A+{\mu }_{G}}<1/(2{\sigma }_{G}^{2}+1/{C}_{A-{\mu }_{G}})\) or equivalently \({C}_{A-{\mu }_{G}}>1/(1/{C}_{A+{\mu }_{G}}-2{\sigma }_{G}^{2})\), in addition to \({C}_{A+{\mu }_{G}}<{C}_{A-{\mu }_{G}}\). This means that the invader’s optimal consumer position (\({x}_{A}+{\mu }_{G}\)) should be towards the trait periphery while its optimal resource position (\({x}_{A}-{\mu }_{G}\)) should be central in the trait space. That is, the ideal trait position of a non-native species drifts from periphery to centre with the increase of its trophic level.
Discussion
Our results illustrate that for a non-native species to successfully invade a network, it must possess traits positioned at the centre of the traits of its facilitators and optimal resources in order to reap the benefit from positive interactions. It must simultaneously possess traits at the periphery of the traits of its competitors and optimal consumers in order to elude the harm from negative interactions. This we call the central-to-reap, edge-to-elude trait strategy for elevated invasiveness in an ecological network. Results from the model therefore highlight the central-to-reap, edge-to-elude trait strategy for elevated invasiveness and the possibility to reveal such trait positions for any given ecological networks using the proposed trait centrality that accounts for both trait differences between the prospective non-native species and the resident species and the relative abundance of the resident species.
We have expanded the concept of invasion fitness in evolutionary game theory to accommodate unique elements associated with biological invasions. This has allowed us to theoretically explore and elucidate the invasiveness of a non-native species invading an ecological network. Previous eco-evolutionary models of ecological networks have also used the concept of an adaptive landscape to model trait evolution and its role in shaping network architectures (Minoarivelo and Hui 2016a; Guimarães Jr et al. 2017). We have added a step by proposing that the framework not only allows us to model the trait evolution of resident species but also allows us to explore the outcome of an invasion event. Our mathematical model allows us to visualise species performance associated with both the non-native traits relative to those of the resident species in the ecological network and the role of propagule pressure. This model, therefore, merges insights from invasion science, evolutionary ecology, community ecology, and ecological networks. We discuss here a list of directly related issues and future research needs for which our model can offer tentative solutions. We first discuss the deep meaning and implication of the central-to-reap, edge-to-elude trait strategies for understanding and forecasting invasiveness. We then discuss how performance can be related to the availability of empty opportunistic niches and the penetration of invasion barriers. Finally, we discuss the data format and methodologies that are needed to parameterise and implement our model.
Central to reap, edge to elude
The four examples used to demonstrate the utility of our mathematical model provide evidence of a cohesive trait strategy: to be successful, invaders need to position their traits relative to the trait distributions of resident species from different functional guilds. They must also mitigate negative interactions by occupying peripheral trait positions and increase positive interactions by seeking central trait positions. This trait strategy of central-to-reap, edge-to-elude, highlights the leverage trait position in an ecological network for a non-native species to achieve elevated invasiveness. A similar strategy in repeated games, known as win-stay, lose-shift, describes heuristic learning of a player’s opponent by sticking to the same strategy that yielded a positive payoff in the last round of play but shifting to an alternative strategy if it was a loss. This strategy has been confirmed as the most robust winning strategy in game theory (Nowak and Sigmund 1993). In the multiplayer games of an ecological network, the central-to-reap, edge-to-elude trait strategy also reflects the ‘winning’ trait position that ensures a non-native species outcompeting other resident species in a community. The emergent central-to-reap, edge-to-elude trait strategy can be tested with assemblage-level trait data from multiple functional guilds (e.g. Divíšek et al. 2018).
The central-to-reap, edge-to-elude strategy, of course, only broadly describes the trait position for elevated invasiveness, since we only used the first-order Taylor series for interaction strength approximation. Future elaborations that consider nonlinear or higher-order interactions and more accurate numerical schemes should generate additional hypotheses (e.g. a more complex landscape of invasion fitness emerged in Fig. 2 when nonlinear interaction strengths in ecological networks were not approximated by the first-order Taylor series). For instance, within the hull of traits of the resident species (Fig. 3), an introduced species with a similar trait to a resident species, may have a greater probability of successful establishment due to the presence of required niches to ensure its survival. However, this same introduced species may suffer severely from biotic resistance, ultimately limiting its invasiveness (Divíšek et al. 2018). In contrast, an introduced species with a trait sitting between the traits of two resident species faces an uncertain outcome: either there is an empty niche to allow invasion, or no niche available for invasion (Hui et al. 2016). However, it should be noted that excessive elaborations could increase the intrinsic system complexity, creating computational irreducibility and actually lowering the realised predictability (Beckage et al. 2011). Given the contextual complexity of any invasion event (Pyšek et al. 2020), a fine balance of system elaboration that can clearly contain system uncertainty should be preferred (Latombe et al. 2019a).
Empty niches and invasion barriers
Within the discussions on the role of functional traits in determining invasiveness in the literature (e.g. Catford et al. 2019; Enders et al. 2020), the prominent hypotheses in invasion science are centred around the concept of opportunistic ecological niches (e.g. Simberloff 1981; Herbold and Moyle 1986; Shea and Chesson 2002). An empty niche is defined by the specific absence of a species along particular gradients in the resource space (Hutchinson 1957; Holt 2009). Ecological niches in communities have been argued to be largely unsaturated and thus open to invasion (Simberloff 1981; Walker and Valentine 1984; Rohde 2005). The presence of unsaturated niches has been hypothesised to explain the lack of biotic resistance to some biological invasions and the lack of impact in some invaded communities (Mack et al. 2000; Sax et al. 2007). In a fitness landscape, empty niches are represented by pockets of positive fitness in the trait space that are ‘waiting’ to be filled through invasion or incremental trait evolution (Figs. 1 and 2). Non-native species with traits that match these empty niches can establish in the newly invaded environment without the necessity of intensively competing with and affecting native species. The concept of invasiveness and invasibility can thus be measured by the shape and quantity of such empty niches in an ecological network (Lonsdale 1999; Shea and Chesson 2002; Hui et al. 2016). With the concept of the landscape of invasion fitness, these two concepts—niches and traits—are therefore closely mirrored, as in our model.
Coupled with rapid environmental changes, coevolution of entangled biotic interactions can drive change in trait-mediated and density-dependent interaction strengths (Thompson 2013), creating both empty niches and invasion barriers that are dynamic at both ecological and evolutionary time scales. As highlighted in our model (Figs. 1 and 2), such empty opportunistic niches with positive invasion fitness are enclosed by valleys of negative invasion fitness in the trait space (also see, Hui et al. 2016). Similar to empty niches, ecological and evolutionary barriers are also constructed through trait-mediated biotic interactions that can drastically bend and reshape the invasion fitness landscape. Negative (antagonistic) interactions, such as those involving competitors and predators, can constrain the fundamental niche and form ecological barriers to invasion with respect to specific non-native traits. In contrast, positive (mutualistic) interactions can expand the fundamental niches through the provision of mutualistic benefits into otherwise unsuitable niche space (Rodriguez-Cabal et al. 2012; Stachowicz 2012; Afkhami et al. 2014), thereby unravelling ecological barriers to invasion.
Once a non-native species has engaged with resident species in coevolving dynamics, the network of biotic interactions can drastically adjust its fundamental niche to form the realised niche accessible to the non-native species. Coevolution in the context of invasion could impose evolutionary barriers that could eventually prohibit trait radiation among resident species of a community. Evolutionary barriers are mechanisms in place that prohibit directional evolution of resident species with certain traits. In other words, rare mutants with traits similar to those of resident species cannot establish and replace the resident trait. This would constrain the distribution of functional traits in the community and potentially create empty niches that can only be filled by the invaders with large trait differences. For instance, coevolution via facilitative interactions can generate positive reinforcing feedbacks (e.g. mutualism). Such reinforcing feedbacks can lead to a lock-in of trait evolution in a sub-optimal state in terms of functional trait distribution in resident species. This can happen when selfish mutualists benefit excessively from the interactions thereby creating evolutionary barriers that thwart further radiation of traits (Minoarivelo et al. in prep). Such empty niches set up by evolutionary barriers can also give rise to priority effects: the traits and sequence/history of invasion can greatly affect how the fitness landscape—and therefore invisibility—of an ecological network unfolds (Minoarivelo and Hui 2018). Over ecological time scales that are relevant to invasion management, coevolution can therefore have a great effect on the demographic and invasiveness of both resident and non-native species (Saul and Jeschke 2015; Le Roux et al. 2017).
Trait-mediated interaction strength
To unveil the landscape of invasion fitness associated with an ecological network, we need to elucidate the interaction strength as a function of traits and relative abundances of involved species (Catford et al. 2019), and then compute the fitness landscape of the ecological network over trait space (Figs. 1 and 2). That is, to effectively implement our model we need to choose a suite of traits and explain interaction strength as the degree of trait matching or difference. A suite of traits, rather than a single trait, are often needed to determine interaction strength (Eklöf et al. 2013). Unlike previous evolutionary models of ecological networks (Guimarães Jr et al. 2017; Valdovinos et al. 2018), our theoretical model possesses a generic feature to further accommodate a high-dimensional trait space. The choice of traits is worth considering briefly. The selected traits describe the functional role of the interacting species—more precisely, they define the strength of interaction and fitness, or demographic consequence between species pairs. Species functional traits normally obey strong allometric relationships and can be mostly expressed in a two-dimensional trait space (e.g. based on the first two axes from the principal component analyses or non-metric multidimensional scaling of the species-by-trait matrix; Díaz et al. 2016). In practice, we need to transform these allometric traits before calculating the level of trait matching or difference (e.g. consider that the trait represents the logarithm of the body size of resident species). This formulation of interaction strength using trait matching or difference (such as in Eq. 9) has reasonably strong empirical support, and has often been used to formulate many kinds of ecological networks with coevolving traits (e.g. Brännström et al. 2011; Nuismer et al. 2013; Guimarães Jr et al. 2017; Hui et al. 2018). As traits are essentially the strategies each species plays in the evolutionary game, we can also expand the dimension of functional traits to include other life-history strategies that are able to differentiate demographic performance between species, for example, preference, plasticity, phenology, and phylogeny (van Kleunen et al. 2010; Landi et al. 2018a).
Interaction strength itself can also be considered as being adaptive (e.g. Valdovinos et al. 2010, 2018; Zhang et al. 2013; Gibert and Yeakel 2019). In such cases, an interaction that is increasing or decreasing its strength due to selective forces can be considered within our framework as involved traits evolving respectively towards trait convergence or trait divergence. In this sense, the two frameworks, targeting adaptive traits of species (our model) and adaptive interaction strength between species, have reached a consistent strategy for invasion success in mutualistic networks—to position the non-native trait for better mutualistic gain (Valdovinos et al. 2018). However, the implicit role of traits in the evolutionary dynamics of interaction strength could mask the role of trait dispersion in an ecological network and the trait convergence-divergence between interacting species, for instance, in pollination networks where interaction strength can be computed as the foraging efficiency of pollinators and considered as the evolving trait (Valdovinos et al. 2018). Doing so, however, ignores that an interaction is the game played between two interacting species and that the traits of the plant species being pollinated can also be important for the interaction strength, as demonstrated in Darwin’s coevolution race (Toju 2011; Zhang et al. 2013). Nevertheless, models focusing solely on evolving interaction strength are often easier to parameterise especially for large networks, while we anticipate that the challenge facing the parameterisation of trait-mediated adaptive networks could potentially be overcome in near future. Specifically, advancing eco-informatics documenting both species traits and interaction strengths could allow us soon to parameterise the interaction kernel function (e.g. Eq. 9) for any given ecological networks.
Both the demographic performance of species and the strength of biotic interactions are scale dependent (Araújo and Rozenfeld 2014; Hui et al. 2017), and such scale dependency dictates the contribution of biotic interactions to the structure and functioning of ecological networks (Galiana et al. 2018). For instance, biotic interactions can be discerned between non-native and native plant assemblages of reserve networks hundreds of kilometres apart (Latombe et al. 2018) but not when comparing non-native versus native ant assemblages of oceanic islands thousands of kilometres apart (Latombe et al. 2019b). With the increase of spatial scales, demographic performance also exhibits distinct accumulation curves, normally with inflated growth and reduced volatility associated with successful non-native invaders (e.g. gypsy moths Lymantria dispar in the northeast US; Hui et al. 2017). When a study system is geographically too large to allow individuals a reasonable chance of encountering and interacting with each other during one life span, we should rather break down the large spatial network into meta-networks or meta-ecosystems (Gravel et al. 2016), depicting interlinked local networks via dispersal of propagules. Invasiveness in a meta-network depends not only on the non-native traits and propagule pressure, but also on the spatial entry points—whether to start an invasion from the spatial periphery or centroid; this adds an extra dimension to our model. Moreover, temporal scale can also profoundly affect the strengths of biotic interactions and thus the timing for elevated invasiveness (CaraDonna and Waser 2020). Exploring the effects of spatial and temporal scales on the invasiveness of a non-native species and the invasibility of an ecological network, will be natural extensions to our current model.
A careful integration of empirical data with theory will allow us to map the entire network of biotic interactions in a community. In addition to the potential of using functional traits (e.g. body size) to build basic network structures (Gravel et al. 2013), the strength of biotic interactions can be inferred from other interaction proxies (Morales-Castilla et al. 2015). For instance, because biotic interactions greatly affect species co-distribution in metacommunities (Cazelles et al. 2016), changes in the structure and strength of biotic interactions can be realised as changes in species co-distribution in an ecological network. Therefore, the consequence of species introduction into and removal from ecological networks can be captured by assemblage-level temporal turnover and resulted interaction rewiring (CaraDonna et al. 2017; Bosc et al. 2018; Keet et al. 2019). As the coexistence of resident species in an ecological network needs to abide by the stability criterion that prescribes the specific architecture of a stable network (Landi et al. 2018b), a non-native species needs to break the stability criterion to be successful, which can result in temporal turnover and changes in species abundances along the fastest direction (described by system eigenvectors) away from the current network and assemblage structures (Hui and Richardson 2019). Models implementing interaction rewiring and thus temporal turnover can greatly improve our ability to explain observed network topology (Zhang et al. 2011; Nuwagaba et al. 2015; Nnakenyi et al. 2019). There is, therefore, great potential to rapidly estimate the interaction strengths of ecological networks from network dissimilarity and interaction turnover (Poisot et al. 2012; McGeoch et al. 2019). Network science could contribute additional tools to tackle the challenges facing rapid parameterisation of ecological networks (Delmas et al. 2019). For instance, the proposed regression method in Sect. 2.1 could solve the challenge to some extent but still requires large amounts of time-series data.
Taken together, ecological opportunities and barriers can be formed dynamically and adaptively in response to the ecological novelty created via biological invasions. Nonetheless, for short- to mid-term invasion management, estimating and mapping the entire interaction network is crucial to both visualise fitness landscapes and project species-specific eco-evolutionary dynamics (Hui and Richardson 2019). To this end, the barrier scheme of the introduction-naturalised-invasion continuum (Richardson et al. 2000; Blackburn et al. 2011) needs to be expanded to accommodate the dynamic complexity of ecological networks (especially at the end of this continuum). A barrier or opportunity to invasion could quickly unravel or emerge, changing the trait centrality of non-native species in different functional guilds of the invaded ecosystem—therefore altering its invasiveness. This also suggests that pairwise native-non-native trait comparisons have limited value when seeking advances in invasion science. To this end, we urge researchers in the fields of biodiversity monitoring and informatics to devise methods that allow rapid mapping of the entire interaction network. In particular, with the rapid expansion of trait data, the baseline trait-mediated interaction strength (e.g. Eq. 9) could be parameterised and fitted using trait data combined with knowledge of relative abundances or co-distributions in local communities (e.g. Brousseau et al. 2018).
Code availability
Included in the Online Supplementary Material.
References
Afkhami ME, McIntyre PJ, Strauss SY (2014) Mutualist-mediated effects on species’ range limits across large geographic scales. Ecol Lett 17:1265–1273. https://doi.org/10.1111/ele.12332
Allesina S, Tang S (2012) Stability criteria for complex ecosystems. Nature 483:205–208. https://doi.org/10.1038/nature10832
Araújo MB, Rozenfeld A (2014) The geographic scaling of biotic interactions. Ecography 37:406–415. https://doi.org/10.1111/j.1600-0587.2013.00643.x
Bastolla U, Fortuna MA, Pascual-García A, Ferrera A, Luque B, Bascompte J (2009) The architecture of mutualistic networks minimizes competition and increases biodiversity. Nature 458:1018–1020. https://doi.org/10.1038/nature07950
Beckage B, Gross LJ, Kauffman S (2011) The limits to prediction in ecological systems. Ecosphere 2:1–12. https://doi.org/10.1890/ES11-00211.1
Blackburn TM, Pyšek P, Bacher S, Carlton JT, Duncan RP, Jarošík V, Wilson JRU, Richardson DM (2011) A proposed unified framework for biological invasions. Trends Ecol Evol 26:333–339. https://doi.org/10.1016/j.tree.2011.03.023
Bosc C, Roets F, Hui C, Pauw A (2018) Interactions among predators and plant specificity protect herbivores from top predators. Ecology 99:1602–1609. https://doi.org/10.1002/ecy.2377
Brännström Å, Loeuille N, Loreau M, Dieckmann U (2011) Emergence and maintenance of biodiversity in an evolutionary food-web model. Theor Ecol 4:467–478. https://doi.org/10.1007/s12080-010-0089-6
Brousseau PM, Gravel D, Handa IT (2018) On the development of a predictive functional trait approach for studying terrestrial arthropods. J Anim Ecol 87:1209–1220. https://doi.org/10.1111/1365-2656.12834
Bronstein JL (2015) Mutualism. Oxford University Press, Oxford
CaraDonna PJ, Waser NM (2020) Temporal flexibility in the structure of plant–pollinator interaction networks. Oikos 129:1369–1380. https://doi.org/10.1111/oik.07526
CaraDonna PJ, Petry WK, Brennan RM, Cunningham JL, Bronstein JL, Waser NM, Sanders NJ (2017) Interaction rewiring and the rapid turnover of plant–pollinator networks. Ecol Lett 20:385–394. https://doi.org/10.1111/ele.12740
Catford JA, Jansson R, Nilsson C (2009) Reducing redundancy in invasion ecology by integrating hypotheses into a single theoretical framework. Divers Distrib 15:22–40. https://doi.org/10.1111/j.1472-4642.2008.00521.x
Catford JA, Smith AL, Wragg PD, Clark AT, Kosmala M, Cavender-Bares J, Reich PB, Tilman D (2019) Traits linked with species invasiveness and community invasibility vary with time, stage and indicator of invasion in a long-term grassland experiment. Ecol Lett 22:593–604. https://doi.org/10.1111/ele.13220
Cazelles K, Mouquet N, Mouillot D, Gravel D (2016) On the integration of biotic interaction and environmental constraints at the biogeographical scale. Ecography 39:921–931. https://doi.org/10.1111/ecog.01714
Courchamp F, Fournier A, Bellard C, Bertelsmeier C, Bonnaud E, Jeschke JM, Russell JC (2017) Invasion biology: specific problems and possible solutions. Trends Ecol Evol 32:13–22. https://doi.org/10.1016/j.tree.2016.11.001
Darwin CR (1859) On the origin of species by means of natural selection, or the preservation of favoured races in the struggle for life. John Murray, London
Delmas E, Besson M, Brice MH, Burkle LA, Dalla Riva GV, Fortin MJ, Gravel D, Guimarães PR, Hembry DH, Newman EA, Olesen JM, Pires MM, Yeakel JD, Poisot T (2019) Analysing ecological networks of species interactions. Biol Rev 94:16–36. https://doi.org/10.1111/brv.12433
Dercole F, Della Rossa F, Landi P (2016) The transition from evolutionary stability to branching: a catastrophic evolutionary shift. Sci Rep 6:26310. https://doi.org/10.1038/srep26310
Díaz S, Kattge J, Cornelissen J et al (2016) The global spectrum of plant form and function. Nature 529:167–171. https://doi.org/10.1038/nature16489
Dieckmann U, Ferrière R (2004) Adaptive dynamics and evolving biodiversity. In: Ferrière R, Dieckmann U, Couvet D (eds) Evolutionary conservation biology. Cambridge University Press, Cambridge, pp 188–224. https://doi.org/10.1017/CBO9780511542022.015
Dieckmann U, Law R (1996) The dynamical theory of coevolution: a derivation from stochastic ecological processes. J Math Biol 34:579–612. https://doi.org/10.1007/BF02409751
Divíšek J, Chytrý M, Beckage B, Gotelli NJ, Lososová Z, Pyšek P, Richardson DM, Molofsky J (2018) Similarity of introduced plant species to native ones facilitates naturalization, but differences enhance invasion success. Nat Commun 9:4631. https://doi.org/10.1038/s41467-018-06995-4
Doebeli M, Dieckmann U (2000) Evolutionary branching and sympatric speciation caused by different types of ecological interactions. Am Nat 156:S77–S101. https://doi.org/10.1086/303417
Eklöf A, Jacob U, Kopp J et al (2013) The dimensionality of ecological networks. Ecol Lett 16:577–583. https://doi.org/10.1111/ele.12081
Enders M, Havemann F, Ruland F et al (2020) A conceptual map of invasion biology: Integrating hypotheses into a consensus network. Glob Ecol Biogeogr 29:978–991. https://doi.org/10.1111/geb.13082
Galiana N, Lurgi M, Claramunt-López B, Fortin MJ, Leroux S, Cazelles K, Gravel D, Montoya JM (2018) The spatial scaling of species interaction networks. Nat Ecol Evol 2:782–790. https://doi.org/10.1038/s41559-018-0517-3
Gallien L, Landi P, Hui C, Richardson DM (2018) Emergence of weak-intransitive competition through adaptive diversification and eco-evolutionary feedbacks. J Ecol 106:877–889. https://doi.org/10.1111/1365-2745.12961
Gavrilets S (2004) Fitness landscapes and the origin of species. Princeton University Press (Princeton/New Jersey).
Geritz SAH, Metz JAJ, Kisdi É, Meszéna G (1997) Dynamics of adaptation and evolutionary branching. Phys Rev Lett 78:2024–2027. https://doi.org/10.1103/PhysRevLett.78.2024
Geritz SAH, Kisdi É, Meszéna G, Metz JAJ (1998) Evolutionarily singular strategies and the adaptive growth and branching of the evolutionary tree. Evol Ecol 12:35–57. https://doi.org/10.1023/A:1006554906681
Gibert JP, Yeakel JD (2019) Eco-evolutionary origins of diverse abundance, biomass, and trophic structures in food webs. Front Ecol Evol 7:15. https://doi.org/10.3389/fevo.2019.00015
Guimarães PR Jr, Pires MM, Jordano P, Bascompte J, Thompson JN (2017) Indirect effects drive coevolution in mutualistic networks. Nature 550:511–514. https://doi.org/10.1038/nature24273
Gravel D, Poisot T, Albouy C, Velez L, Mouillot D (2013) Inferring food web structure from predator–prey body size relationships. Methods Ecol Evol 4:1083–1090. https://doi.org/10.1111/2041-210X.12103
Gravel D, Massol F, Leibold MA (2016) Stability and complexity in model meta-ecosystems. Nat Commun 7:12457. https://doi.org/10.1038/ncomms12457
Gyllenberg M, Parvinen K (2001) Necessary and sufficient conditions for evolutionary suicide. Bull Math Biol 63:981–993. https://doi.org/10.1006/bulm.2001.0253
Herbold B, Moyle PB (1986) Introduced species and vacant niches. Am Nat 128:751–760. https://doi.org/10.1086/284600
Hofbauer J, Sigmund K (1990) Adaptive dynamics and evolutionary stability. Appl Math Lett 3:75–79. https://doi.org/10.1016/0893-9659(90)90051-C
Holt RD (2009) Bringing the Hutchinsonian niche into the 21st century: Ecological and evolutionary perspectives. Proc Natl Acad Sci USA 106:19659–19665. https://doi.org/10.1073/pnas.0905137106
Hui C, Richardson DM (2017) Invasion dynamics. Oxford University Press, Oxford
Hui C, Richardson DM (2019) How to invade an ecological network. Trends Ecol Evol 34:121–131. https://doi.org/10.1016/j.tree.2018.11.003
Hui C, Minoarivelo HO, Landi P (2018) Modelling coevolution in ecological networks with adaptive dynamics. Math Methods Appl Sci 41:8407–8422. https://doi.org/10.1002/mma.4612
Hui C, Fox GA, Gurevitch J (2017) Scale-dependent portfolio effects explain growth inflation and volatility reduction in landscape demography. Proc Natl Acad Sci USA 114:12507–12511. https://doi.org/10.1073/pnas.1704213114
Hui C, Landi P, Latombe G (2020) The role of biotic interactions in invasion ecology: theories and hypotheses. In: Traveset A, Richardson DM (eds) Plant invasions: the role of biotic interactions. CAB International, Wallingford, pp 26–44. https://doi.org/10.1079/9781789242171.0002
Hui C, Richardson DM, Landi P, Minoarivelo HO, Garnas J, Roy HE (2016) Defining invasiveness and invasibility in ecological networks. Biol Invasions 18:971–983. https://doi.org/10.1007/s10530-016-1076-7
Hutchinson GE (1957) Concluding remarks. Cold Spring Harb Symp Quant Biol 22:415–427. https://doi.org/10.1101/SQB.1957.022.01.039
Keet JH, Ellis AG, Hui C, Le Roux JJ (2019) Strong spatial and temporal turnover of soil bacterial communities in South Africa’s hyperdiverse fynbos biome. Soil Biol Biochem 136:107541. https://doi.org/10.1016/j.soilbio.2019.107541
Kisdi É, Jacobs FJA, Geritz SAH (2002) Red Queen evolution by cycles of evolutionary branching and extinction. Selection 2:161–176. https://doi.org/10.1556/Select.2.2001.1-2.12
Landi P, Vonesh J, Hui C (2018a) Variability in life-history switch points across and within populations explained by Adaptive Dynamics. J R Soc Interface 15:20180371. https://doi.org/10.1098/rsif.2018.0371
Landi P, Minoarivelo HO, Brännström Å, Hui C, Dieckmann U (2018b) Complexity and stability of ecological networks: a review of the theory. Popul Ecol 60:319–345. https://doi.org/10.1007/s10144-018-0628-3
Latombe G, Richardson DM, Pyšek P, Kučera T, Hui C (2018) Drivers of species turnover of native and alien plants with different residence times vary with species commonness. Ecology 99:2763–2775. https://doi.org/10.1002/ecy.2528
Latombe G, Canavan S, Hirsch H, Hui C, Kumschick S, Nsikani MM, Potgieter LJ, Robinson TB, Saul WC, Turner SC, Wilson JRU, Yannelli FA, Richardson DM (2019a) A four-component classification of uncertainties in biological invasions: implications for management. Ecosphere 10:e02669. https://doi.org/10.1002/ecs2.2669
Latombe G, Roura-Pascual N, Hui C (2019b) Similar compositional turnover but distinct insular biogeographical drivers of native and exotic ants in two oceans. J Biogeogr 46:2299–2310. https://doi.org/10.1111/jbi.13671
Le Roux JJ, Hui C, Keet JH, Ellis A (2017) Co-introduction versus ecological fitting pathways to the establishment of effective mutualisms during biological invasions. New Phytol 215:1354–1360. https://doi.org/10.1111/nph.14593
Lehmann L, Mullon C, Akçay E, Van Cleve J (2016) Invasion fitness, inclusive fitness, and reproductive numbers in heterogeneous populations. Evolution 70:1689–1702. https://doi.org/10.1111/evo.12980
Leimar O, Sasaki A, Doebeli M, Dieckmann U (2013) Limiting similarity, species packing, and the shape of competition kernels. J Theor Biol 339:3–13. https://doi.org/10.1016/j.jtbi.2013.08.005
Lonsdale WM (1999) Global patterns of plant invasions and the concept of invasibility. Ecology 80:1522–1536. https://doi.org/10.1890/0012-9658(1999)080[1522:GPOPIA]2.0.CO;2
Lynch M, Walsh B (1998) Genetics and analysis of quantitative traits. Sinauer Associates, Sunderland
MacArthur R, Levins R (1967) The limiting similarity, convergence, and divergence of coexisting species. Am Nat 101:377–385. https://doi.org/10.1086/282505
Mack RN, Simberloff D, Lonsdale WM, Evans H, Clout M, Bazzaz FA (2000) Biotic invasions: Causes, epidemiology, global consequences, and control. Ecol Appl 10:689–710. https://doi.org/10.1890/1051-0761(2000)010[0689:BICEGC]2.0.CO;2
Marchiori M, Latora V (2000) Harmony in the small-world. Phys A 285:539–546. https://doi.org/10.1016/S0378-4371(00)00311-3
May RM (1972) Will a large complex system be stable? Nature 238:413–414. https://doi.org/10.1038/238413a0
Maynard Smith J, Price GR (1973) The logic of animal conflict. Nature 246:15–18. https://doi.org/10.1038/246015a0
McGeoch MA, Latombe G, Andrew NR, Nakagawa S, Nipperess DA, Roigé M, Marzinelli EM, Campbell AH, Vergés A, Thomas T, Steinberg PD, Selwood KE, Henriksen MV, Hui C (2019) Measuring continuous compositional change using decline and decay in zeta diversity. Ecology 100:e02832. https://doi.org/10.1002/ecy.2832
Meszena G, Kisdi E, Dieckmann U, Geritz SAH, Metz JAJ (2001) Evolutionary optimisation models and matrix games in the unified perspective of adaptive dynamics. Selection 2:193–210. https://doi.org/10.1556/Select.2.2001.1-2.14
Meszena G, Gyllenberg M, Jacobs FJ, Metz JAJ (2005) Link between population dynamics and dynamics of Darwinian evolution. Phys Rev Lett 95:078105. https://doi.org/10.1103/PhysRevLett.95.078105
Metz JAJ, Nisbet RM, Geritz SAH (1992) How should we define “fitness” for general ecological scenarios? Trends Ecol Evol 7:198–202. https://doi.org/10.1016/0169-5347(92)90073-K
Metz JAJ, Geritz SAH, Meszéna G, Jacobs FJA, van Heerwaarden JS (1996) Adaptive dynamics: A geometrical study of the consequences of nearly faithful reproduction. In: van Strien SJ, Verduyn Lunel SM (eds) Stochastic and spatial structures of dynamical systems, Proceedings of the Royal Dutch Academy of Science (KNAW Verhandelingen). North Holland, Dordrecht, pp 183–231
Minoarivelo HO, Hui C (2016a) Trait-mediated interaction leads to structural emergence in mutualistic networks. Evol Ecol 30:105–121. https://doi.org/10.1007/s10682-015-9798-z
Minoarivelo HO, Hui C (2016b) Invading a mutualistic network: to be or not to be similar. Ecol Evol 6:4981–4996. https://doi.org/10.1002/ece3.2263
Minoarivelo HO, Hui C (2018) Alternative assembly processes from trait-mediated co-evolution in mutualistic communities. J Theor Biol 454:146–153. https://doi.org/10.1016/j.jtbi.2018.06.004
Morales-Castilla I, Matias MG, Gravel D, Araújo MB (2015) Inferring biotic interactions from proxies. Trends Ecol Evol 30:347–356. https://doi.org/10.1016/j.tree.2015.03.014
Morris RJ, Gripenberg S, Lewis OT, Roslin T (2014) Antagonistic interaction networks are structured independently of latitude and host guild. Ecol Lett 17:340–349. https://doi.org/10.1111/ele.12235
Nnakenyi CA, Traveset A, Heleno R, Minoarivelo HO, Hui C (2019) Fine-tuning the nested structure of pollination networks by adaptive interaction switching biogeography and sampling effect in the Galápagos Islands. Oikos 128:1413–1423. https://doi.org/10.1111/oik.06053
Norman TM, Horlbeck MA, Replogle JM, Ge AY, Xu A, Jost M, Gilbert LA, Weissman JS (2019) Exploring genetic interaction manifolds constructed from rich single-cell phenotypes. Science 365:786–793. https://doi.org/10.1126/science.aax4438
Nowak M, Sigmund K (1993) A strategy of win-stay, lose-shift that outperforms tit-for-tat in the Prisoner’s Dilemma game. Nature 364:56–58. https://doi.org/10.1038/364056a0
Nowak MA, Sigmund K (2004) Evolutionary dynamics of biological games. Science 303:793–799. https://doi.org/10.1126/science.1093411
Nuismer SL, Jordano P, Bascompte J (2013) Coevolution and the architecture of mutualistic networks. Evolution 67:338–354. https://doi.org/10.1111/j.1558-5646.2012.01801.x
Nuwagaba S, Zhang F, Hui C (2015) A hybrid behavioural rule of adaptation and drift explains the emergent architecture of antagonistic networks. Proc R Soc B: Biol Sci 282:20150320. https://doi.org/10.1098/rspb.2015.0320
Poisot T, Canard E, Mouillot D, Mouquet N, Gravel D (2012) The dissimilarity of species interaction networks. Ecol Lett 15:1353–1361. https://doi.org/10.1111/ele.12002
Prentis PJ, Wilson JRU, Dormontt EE, Richardson DM, Lowe AJ (2008) Adaptive evolution in invasive species. Trends Plant Sci 13:288–294. https://doi.org/10.1016/j.tplants.2008.03.004
Pyšek P, Bacher S, Kühn I, Novoa A, Catford JA, Hulme PE, Pergl J, Richardson DM, Wilson JRU, Blackburn TM (2020) Framework for invasive aliens (MAFIA): disentangling large-scale context dependency in biological invasions. NeoBiota 62:407–461. https://doi.org/10.3897/neobiota.62.52787
Ricciardi A, Blackburn TM, Carlton JT, Dick JTA, Hulme PE, Iacarella JC, Jeschke JM, Liebhold AM, Lockwood JL, MacIsaac HJ, Pyšek P, Richardson DM, Ruiz GM, Simberloff D, Sutherland WJ, Wardle DA, Aldridge DC (2017) Invasion science: a horizon scan of emerging challenges and opportunities. Trends Ecol Evol 32:464–474. https://doi.org/10.1016/j.tree.2017.03.007
Richardson DM, Pyšek P, Rejmánek M, Barbour MG, Panetta FD, West CJ (2000) Naturalization and invasion of alien plants: concepts and definitions. Divers Distrib 6:93–107. https://doi.org/10.1046/j.1472-4642.2000.00083.x
Rodriguez-Cabal MA, Barrios-Garcia MN, Amico GC, Aizen MA, Sanders NJ (2012) Node-by-node disassembly of a mutualistic interaction web driven by species introductions. Proc Natl Acad Sci USA 110:16503–16507. https://doi.org/10.1073/pnas.1300131110
Rohde K (2005) Nonequilibrium ecology. Cambridge University Press, Cambridge
Saul WC, Jeschke JM (2015) Eco-evolutionary experience in novel species interactions. Ecol Lett 18:236–245. https://doi.org/10.1111/ele.12408
Sax DF, Stachowicz JJ, Brown JH, Bruno JF, Dawson MN, Gaines SD, Grosberg RK, Hastings A, Holt RD, Mayfield MM, O’Connor MI, Rice WR (2007) Ecological and evolutionary insights from species invasions. Trends Ecol Evol 22:465–471. https://doi.org/10.1016/j.tree.2007.06.009
Schittko C, Bernard-Verdier M, Heger T, Buchholz S, Kowarik I, von der Lippe M, Seitz B, Joshi J, Jeschke JM (2020) A multidimensional framework for measuring biotic novelty: how novel is a community? Glob Change Biol 26:4401–4417. https://doi.org/10.1111/gcb.15140
Shea K, Chesson P (2002) Community ecology theory as a framework for biological invasions. Trends Ecol Evol 17:170–176. https://doi.org/10.1016/S0169-5347(02)02495-3
Simberloff D (1981) Community effects of introduced species. In: Nitecki TH (ed) Biotic crises in ecological and evolutionary time. Academic Press, New York, pp 53–81
Simberloff D (2009) The role of propagule pressure in biological invasions. Annu Rev Ecol Evol Syst 40:81–102. https://doi.org/10.1146/annurev.ecolsys.110308.120304
Stachowicz JJ (2012) Niche expansion by positive interactions: realizing the fundamentals. A comment on Rodriguez-Cabal et al. Ideas Ecol Evol 5:42–43. https://doi.org/10.4033/iee.2012.5.10.c
Steidinger BS, Crowther TW, Liang J et al (2019) Climatic controls of decomposition drive the global biogeography of forest-tree symbioses. Nature 569:404–408. https://doi.org/10.1038/s41586-019-1128-0
Thompson JN (2013) Relentless evolution. University of Chicago Press, Chicago
Toju H (2011) Weevils and camellias in a Darwin’s race: model system for the study of eco-evolutionary interactions between species. Ecol Res 26:239–251. https://doi.org/10.1007/s11284-011-0807-6
Traveset A, Richardson DM (2020) Plant Invasions: the role of biotic interactions—an overview. In: Traveset A, Richardson DM (eds) Plant invasions: the role of biotic interactions. CAB International, Wallingford, pp 1–25. https://doi.org/10.1079/9781789242171.0001
Valdovinos FS, Ramos-Jiliberto R, Garay-Narváez L, Urbani P, Dunne JA (2010) Consequences of adaptive behaviour for the structure and dynamics of food webs. Ecol Lett 13:1546–1559. https://doi.org/10.1111/j.1461-0248.2010.01535.x
Valdovinos FS, Berlow EL, De Espanés PM, Ramos-Jiliberto R, Vázquez DP, Martinez ND (2018) Species traits and network structure predict the success and impacts of pollinator invasions. Nat Commun 9:2153. https://doi.org/10.1038/s41467-018-04593-y
van Kleunen M, Weber E, Fischer M (2010) A meta-analysis of trait differences between invasive and non-invasive plant species. Ecol Lett 13:235–245. https://doi.org/10.1111/j.1461-0248.2009.01418.x
Vaz AS, Kueffer C, Kull CA, Richardson DM, Schindler S, Muñoz-Pajares AJ, Vicente JR, Martins J, Hui C, Kühn I, Honrado JP (2017) The progress of interdisciplinarity in invasion science. Ambio 46:428–442. https://doi.org/10.1007/s13280-017-0897-7
von Humboldt A (1807) Essai sur la géographie des plantes. Fr. Schoell, Paris
von Neumann J, Morgenstern O (1944) The theory of games and economic behavior. Princeton University Press, Princeton
Walker TD, Valentine JW (1984) Equilibrium models of evolutionary species diversity and the number of empty niches. Am Nat 124:887–899. https://doi.org/10.1086/284322
Waxman D, Gavrilets S (2005) 20 questions on adaptive dynamics. J Evol Biol 18:1139–1154. https://doi.org/10.1111/j.1420-9101.2005.00948.x
Wright S (1932) The roles of mutation, inbreeding, crossbreeding and selection in evolution. Proc Sixth Int Congr Genetics 1:356–366
Wright S (1988) Surfaces of selective value revisited. Am Nat 131:115–123. https://doi.org/10.1086/284777
Zhang F, Hui C, Terblanche JS (2011) An interaction switch predicts the nested architecture of mutualistic networks. Ecol Lett 14:797–803. https://doi.org/10.1111/j.1461-0248.2011.01647.x
Zhang F, Hui C, Pauw A (2013) Adaptive divergence in Darwin’s race: how coevolution can generate trait diversity in a pollination system. Evolution 67:548–560. https://doi.org/10.1111/j.1558-5646.2012.01796.x
Acknowledgments
This paper emerged from a workshop on ‘Frameworks used in Invasion Science’ hosted by the DSI-NRF Centre of Excellence for Invasion Biology in Stellenbosch, South Africa, in November 2019, that was supported by the National Research Foundation of South Africa and Stellenbosch University. Key concepts featured in this paper were conceived during an earlier workshop on Networks in Ecology at the University of Vermont, USA, in August 2019. We thank Nicholas J. Gotelli, Laurent Hébert-Dufresne, Dina Mistry, Giovanni Petri and Nathan J. Sanders for lively discussion during the Vermont workshop, Sven Bacher, Fernanda Valdovinos, J.A.J. Metz, Feng Zhang, and Ulf Dieckmann for thoughtful comments, and the University of Vermont’s Presidents Office and the DSI-NRF Centre of Excellence for Invasion Biology for funding the two workshops.
Funding
CH was supported by the South African Research Chair Initiative (NRF Grant 89967). DMR acknowledges support from the University of Vermont (as a James Marsh Professor-at-Large between 2012 and 2019) and the Oppenheimer Memorial Trust (Grant 18576/03). HER was supported by the Natural Environment Research Council award number NE/R016429/1 as part of the UK‐SCAPE programme delivering National Capability. XJ was supported by Internal Funds of the KU Leuven.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Models were developed by CH, with model analyses and numerical simulations by CH, HOM and PL. The first draft of the manuscript was written by CH, DMR, HOM and PL; all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Consent to publication
All authors have consented to publish the revised version of the manuscript as it was submitted.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Appendix S1: Derivation of the selection gradient.
Appendix S2: Fitness sensitive to trait change versus selection gradient.
Appendix S3: Derivation of the trait dynamics with propagule influx.
Appendix S4: Matlab code for numerical simulations.
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hui, C., Richardson, D.M., Landi, P. et al. Trait positions for elevated invasiveness in adaptive ecological networks. Biol Invasions 23, 1965–1985 (2021). https://doi.org/10.1007/s10530-021-02484-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-021-02484-w