Abstract
Ponto-Caspian gobies have invaded many river systems in the northern hemisphere, but their colonization patterns often remain unrecognized at the early stages of invasion. Here, we analyzed the ongoing invasion of the round goby Neogobius melanostomus (Pallas, 1814) in the Elbe River (Germany) by combining literature research with electrofishing and environmental DNA (eDNA) sampling. Along the surveyed 517 km, abundances peaked in the upper and lower sections, while low or no goby abundances were detected in the middle section. Combining the results of the field survey and literature data, we inferred four dispersal modes, jointly appearing during the colonization. (1) Human mediated transport, which was responsible for the initial colonization of the tidal Elbe and for a jumpy expansion to the upstream sections. (2) Fast downstream colonization, which was particularly observed for juvenile individuals. (3) Slow upstream migration, which was also recorded for another goby species, the bighead goby Ponticola kessleri (Günther, 1861) and (4) Lateral invasion via a major shipping canal in the middle river section. The eDNA analyses generally supported these conclusions and showed a high sensitivity and thus increased detection strength at low goby densities. Our study showed that the location of the first occurrence, as well as the dispersal mode, strongly determines the invasion process of non-native gobies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gobies such as Neogobius melanostomus (Pallas, 1814) or Ponticola kessleri (Günther, 1861) are invasive species to rivers throughout Europe and North America, with their native range in the Ponto–Caspian basin. They are known for having a high reproduction success and for being strong food competitors, which together lead to a major threat to the invaded river ecosystem. A successful invasion is often followed by a decline in abundances of native fish and a reduction in diversity and number of invertebrates (Borcherding et al. 2011; Cerwenka et al. 2018). Hence, knowledge of dispersal mechanisms, their establishment and how this influences the invasion progress are important for the affected river and of high interest for riverine management. For large river systems, four dispersal patterns can be observed, which distinctly differ in the resulting dispersal speed. (1) A jumpy distribution to new river sections that can lead to colonization far from the original population (Brownscombe et al. 2012). This type of invasion is predominately human-mediated, especially via boats, ballast water, or intentional releases (Roche et al. 2013). (2) Downstream dispersal, which is faster than upstream migration (Janáč et al. 2013; Borcherding et al. 2016). (3) Active upstream migration, which is usually slow (Brownscombe and Fox 2012), and (4) Lateral migration via shipping canals, which connect naturally isolated river catchments. This invasion path may result in highly connected “super-catchments” (Leuven et al. 2009).
To understand the role and interplay of these dispersal modes in colonizing new river ecosystems requires the consideration of early invasion phases. Here, we studied the early invasion dynamics of N. melanostomus in the Elbe River (Germany) and analyzed the invasion front and reconstructed the dispersal processes by means of a comprehensive literature survey and a detailed field sampling of gobies along a 517 km free-flowing river stretch. In addition to electrofishing, we used environmental DNA (eDNA) as a powerful emerging tool allowing the detection and tracking of less abundant species. We therefore combined both monitoring methods in our field-survey, enabling a comparison of conventional fish sampling results by electrofishing with eDNA detection of N. melanostomus along the river course.
Materials and methods
Study area
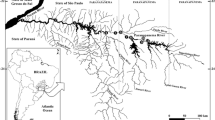
The Elbe River is an 8th order lowland river with a total length of about 1092 km and a catchment area of 148,268 km2. The spring region is in the Krkonoše Mountains (Giant Mountains), Czech Republic. After 364 km, the river passes the Czech–German border, where the German river kilometrage starts with river kilometer (rkm) “0” (Hofmann et al. 2005). We studied the free-flowing German river section from rkm 66 to rkm 582.7 (Fig. 1, Online Resource 1). A further sampling site was located in the Mulde River (1.5 Mulde km upstream of the mouth), a 314 km long non-navigable tributary that flows into the Elbe River at km 259 (Fig. 1, Online Resource 2). With no positive goby detection until now, we expected this site to be a non-invaded reference. Such reference sites may be of great interest as little is known about the small-scaled dispersal and the behavior of N. melanostomus into hitherto uninvaded water bodies (Šlapanský et al. 2020). At Elbe km 333, the Midland Canal over crosses the Elbe and joins the river via its connecting canals (Fig. 1). It is the longest artificial waterway in Germany and strongly used for navigation (Matteikat et al. 2016).
Map indicating the course of the Elbe River, the Mulde River and the Midland Canal (blue lines). Sampling sites of the present study are marked with crosses and numbers for river km. The records from the literature are marked with dots and letters. The inlet shows the Midland Canal and its connecting canals to the Elbe River: RCC (Rothenseer connecting canal, rkm 333.6), NCC (Niegripper connecting canal, rkm 343.7) and PCC (Pareyer connecting canal, rkm 371.5). Additional data on sampling sites and literature records are given in Online Resources 2 and 3
Literature study
We conducted a thorough literature review considering published documents from May 2008 (first documented goby occurrence in Elbe River) up to January 2020. The considered topic related keywords are given in Online Resource 1. Furthermore, we interviewed anglers, local fishing associations and related authorities.
Sampling
Sampling was carried out in October 2018 by means of electrofishing in the morning along 12 equidistantly (60 rkm) located sampling site along the Elbe River. An additional sampling site was in the Mulde River (Fig. 1, Online Resource 3). We considered the most common near-shore habitats present in the Elbe River, i.e. rip-rap and groynes (Brabender et al. 2016) that provide a preferred habitat of gobies (Borcherding et al. 2011). Fish sampling was carried out by point abundance electrofishing (electrofishing generator EFGI 650 at rkm 66, and electrofishing generator DEKA 7000, DEKA Gerätebau Marsberg, Germany at all other sites) along 450 m near-shore stretches at each sampling site. All caught fish were identified to species, counted and total body length was measured to the nearest mm. Gobies were weighted to the nearest mg (Sartorius GL3202i-1CEU, Germany) and removed from the water body. All other fish species were released. The age composition of N. melanostomus was estimated by using a length-frequency distribution analysis. Ages were classified according to peaks in the distribution (Duemler et al. 2016). According to the size classes, N. melanostomus was classified into four age groups, ranging from “0 + ” to “3 + ”.
eDNA sampling and analysis
In order to provide a seamless comparison of results by established methods in biomonitoring (such as electrofishing) and eDNA sampling, we collected data with both methods in the present study. Prior to electrofishing, triplicate samples of near shore water (2500 ml each) were taken from downstream to upstream from each sampling site following the protocol of Adrian-Kalchhauser and Burkhardt-Holm (2016). Samples were stored in portable fridges and filtered in the laboratory within 24 h after collection. DNA from water samples and contamination-controls was extracted using the Qiagen DNeasy PowerWater Kit (Germany). We modified the manufacturers’ guidelines and increased the buffer volume PW1 to 1.2 ml, because of the use of glass fiber filters (Whatman, GF/F grade, 07 µm) that have a higher capacity to absorb liquids compared to other filter types (e.g. polyethersulfone or cellulose). Recovery of PW1 after the lysis step was further improved by squeezing the filter with a 200 µl pipette tip while transferring the PW1 supernatant to a clean 2 ml collection tube.
PCR amplification followed the touchdown PCR protocol with bovine serum albumin (BSA) as a supplement according to Adrian-Kalchhauser and Burkhardt-Holm (2016), using primer pair SL_eDNA_NM_F1/R1 that have been specifically designed for N. melanostomus. From each filter (water samples and contamination controls), three PCR reactions were performed as technical replicates in a total volume of 10 µl each, using the GoTaq® G2 Hot Start Colorless Master Mix (Promega, Germany), and 3 µl DNA extract. Technical replicates were performed independently; positive and negative controls with N. melanostomus DNA (10 pg µl−1) or nuclease-free water as template, respectively, were included. DNA extraction and PCR reaction setup were executed by the use of a UV PCR workbench to avoid contamination risks. Pre- and post-PCR processing was performed in separate laboratory rooms. Pipettes and tube holders were regularly decontaminated under UV-treatment and all other lab equipment (forceps, scissors, etc.) and surfaces were regularly decontaminated using chlorine bleach solution (3.6% sodium hypochlorite) and ethanol before the analyses.
Results
The literature survey showed that the first record of N. melanostomus was made in May 2008 in the tidal Elbe River near the city of Hamburg and further records between 2011 and 2013 in surrounding river reaches (Fig. 2, Online Resource 2). In August 2015, a single record of N. melanostomus was documented in the upper Elbe River near the city of Ústí nad Labem, Czech Republic, 622 km upstream from the initial location. In September 2016, N. melanostomus was documented for the first time in the German Elbe reach around rkm 11. In 2017 a single individual was recorded further downstream (Figs. 1, 2, Online Resource 2). We did not find any published or unpublished goby record (based on literature documentation and pers. comm. with fishermen and related authorities) for almost the entire middle Elbe. Between the years 2012 and 2014, two subpopulations of round gobies were observed in the Midland Canal (Fig. 2, Online Resource 2), indicating the probability of N. melanostomus to enter the Elbe River through its connecting canals (Fig. 1).
Progression of N. melanostomus distribution along the Elbe River, based on a literature survey and compared with this recent field study. See Online Resources 1 and 2 for details on the literature search. The rkm refers to the official German kilometrage, which starts with rkm 0 at the Czech-German border. The asterisk at rkm -36 for Ústí nad Labem indicates the distance from the German border, which is 328 km from the source of the Elbe in the Czech Republic
In our field sampling, a total of 3211 individuals from 21 species subdivided into nine predatory- and 12 non-predatory fish species were caught (Online Resource 4). The sampling revealed the progression of N. melanostomus since the latest detection in 2017 (Fig. 2). We recorded 986 N. melanostomus individuals at the 12 sampling sites along the Elbe River, with the highest abundances found at km 66 (Fig. 3). Abundances decreased with increasing rkm until rkm 268, where no goby was detected. We found a second abundance peak at rkm 370 (Fig. 3) near the confluence of the Midland Canal (Fig. 1). Abundances decreased from there to rkm 427 and below (Fig. 3). The body lengths of N. melanostomus ranged from 30 to150 mm and differed significantly between sampling sites (Kruskal–Wallis χ2 = 453.4, P < 0.001). The smallest individuals (30–40 mm) were captured in the upstream reach, followed by individuals with body lengths between 50–110 mm in the middle section. The largest individuals (120–150 mm) were captured from rkm 420 and below (Fig. 4). The upstream sites at rkm 66 and 124.3 exhibited all age classes and showed the highest shares of smaller and thus younger individuals along the sampling sites (Fig. 4). During sampling at rkm 582.7, we recorded two specimens of P. kessleri with total lengths of 150 mm and 155 mm.
Abundance (N. melanostomus ind. 450 m−1) and biomass (total biomass in wet weight g/450 m) distribution of N. melanostomus in October 2018 in the Elbe and Mulde River. The arrow indicates the location of the junction between the Midland Canal and the Elbe at rkm 333. Data from the Mulde River, which is connected to the Elbe at rkm 259, are shown through the interruption of the x-axis
Length distribution of N. melanostomus in October 2018, based on the size classes in (cm) and related age groups from “0 + ” to “3 + ”. The arrow indicates the location of the junction between the Midland Canal and the Elbe at rkm 333. Numbers above the bars indicate the total number of captured gobies. Data from the Mulde River, which is connected to the Elbe at rkm 259, are shown through the interruption of the x-axis
In order to support the electrofishing, N. melanostomus was additionally detected through highly sensitive PCR-based eDNA sampling. Generally, PCR amplification success as a measure of goby presence was not systematically related to goby abundance as detected via electrofishing. For example, amplification success was highest at sites where N. melanostomus dominated the fish community (Fig. 5) but also at sites where native fish were dominant (rkm 472, 582.7, Fig. 5). At sites having N. melanostomus abundances of < 6 ind. 450 m−1, eDNA amplification indicated round goby presence but not throughout all replicates. The water samples from the Mulde River showed only a very weak band for one out of nine PCR replicates on agarose gels, which is indicative of low eDNA concentrations, resulting in the sampling site with the lowest amplification success (Fig. 5).
Contribution of N. melanostomus to total fish community from electrofishing and amplification success for eDNA. The left y-axis gives the electrofishing survey abundance (%) of all captured fish. The total fish abundance is given on top of each bar. The right y-axis gives the eDNA detection success of N. melanostomus (given by number of positive amplifications of the PCR triplicates per each of the three water samples analyzed per sampling site: max. 9; min. 0). The asterisk at rkm 66 indicates a complete failure of amplification from one out of three water samples, while all other PCR replicates indicate a rather high eDNA concentration of N. melanostomus. The arrow indicates the location of the junction between the Midland Canal and the Elbe at rkm 333. Data from the Mulde River, which is connected to the Elbe at rkm 259, are shown through the interruption of the x-axis
Discussion
We could show that N. melanostomus had a bimodal distribution in the studied section of the River Elbe, with no records in the intermediate section. The age structure changed longitudinally from age groups 0 + and 1 + , to downstream, with mostly adult N. melanostomus, belonging to the age groups 2 + and 3 + , indicating size-related movement patterns. Šlapanský et al. (2020), Janáč et al. (2013, 2019) or Roche et al. (2013) described similar findings of round goby movement in their studies. They observed different age classes being responsible for the invasion process, i.e., adults undertake mainly upstream movement, and younger individuals disperse rapidly downstream.
Our findings together with the invasion history deduced from the literature survey indicate at least four different dispersal modes: (1) a human-mediated dispersal via the ballast water of ships explains the initial goby records in the lower Elbe, as well as the records in the Czech Republic (Fig. 1, Online Resource 2). This outcome supports and confirms the assumption of Roche et al. (2015), Buřič et al. (2015), and Janáč et al. (2013) that consider the transport of N. melanostomus in ballast waters via shipping or human-driven goby release near Ústí nad Labem as the most probable way of N. melanostomus occurrence in the upper Elbe region. However, genetic analysis is needed to clarify whether the observed populations in the upper and lower Elbe River belong to the same or to two distinct invasion lineages. Moreover, further genetic research is needed to track the invasion route of gobies into the Elbe River (Janáč et al. 2017). (2) Downstream migration of especially early life stages, probably explains the population spread in the upper Elbe (Fig. 4). The dominance of 1 + fish at rkm 165 and 217.4 downstream of the first abundance peak suggest such a fast downstream dispersal of young fish. Our results match with previous studies that showed that downstream drift facilitated the dispersal of especially small-sized, juvenile fishes, indicating a recently colonized river section (Janáč et al. 2013, 2019; Borcherding et al. 2016; Ramler et al. 2016). It is noticeable that no young-of-the-year fish (0 +) was recorded at rkm 165 and 217.4, suggesting that live time was too short for such a long dispersal from the first abundance peak. (3) Upward migration from the first records in the tidal Elbe does hardly play any role for the lower parts of the Elbe as only very few individuals, belonging all to the age-group “2 + ” and “3 + ” were observed in this part. Therefore, an active upward migration can be excluded as a quantitative important dispersal mechanism as we identify a lack of goby occurrence along the sampling sites. (4) Lateral immigration via the Midland canal and its tributaries was probably responsible for the observed abundance and biomass peak at rkm 370 (Fig. 3). Closely upstream to this peak, a distinctly lower abundance (6 individuals) was detected, and further downstream to the peak a slightly elevated number of individuals. Both confirm a stronger downstream and slower upstream dispersal, as discussed above for the upper Elbe. Concerning P. kessleri, only two specimens were detected in the lower Elbe. This indicates a rather slow dispersal movement after the first detection in 2015 at the same lower Elbe region (Thiel et al. 2017). For the large river system Elbe, P. kessleri can not be described as an established population. Further population establishment and range expansion requires future long-term studies.
The analysis of eDNA represents a powerful tool for biomonitoring and real-time detection of aquatic macroorganisms in areas of low abundance, including invasive species. Our results indicate a tendency of density-dependent N. melanostomus detection via eDNA. While we were not able to detect N. melanostomus eDNA in all water samples or PCR replicates derived from sites with rather low abundances (< 6 ind. 450 m−1), analyzing water samples from sites with high N. melanostomus densities (> 40 ind. 450 m−1) generally resulted in an extensive amplification success. At the Mulde River, our non-invaded reference, only one PCR reaction out of nine replicates showed a very weak band on agarose gels. This is either suggestive for very low concentrations of N. melanostomus eDNA in the water sample or indicates a stochastic effect during PCR amplification that is not N. melanostomus dependable. According to Veldhoen et al. (2016), we determine that case as a negative detection. On the other hand, positive eDNA detection at rkm 268, where no N. melanostomus was detected by electrofishing, could either be due to very low abundances below the detection level for electrofishing or due to downstream drift of eDNA.
It should be noted that the data need to be interpreted cautiously as very few molecules can produce rather strong signals. Furthermore, the flow velocity in lotic systems can significantly affect the transport of eDNA particles due to variation in sedimentation, remobilization, and dilution (Jane et al. 2015). While some studies could show eDNA transport of only a few meters downstream (e.g., Pilliod et al. 2014), others reported efficient transport with no significant decrease in eDNA concentrations over several hundred meters in streams (e.g., Wacker et al. 2019) or even detection distances of more than 100 km in large rivers (e.g., Pont et al. 2018). Taking reported data on release, decay rates and depositional velocities of eDNA under natural conditions into account (reviewed in, e.g., Harrison et al. 2019 or in Evans et al. 2017 for experimental studies), the detection of far downstream transported eDNA particles seems unlikely but cannot be entirely excluded.
In summary, our eDNA data indicate a generally high sensitivity even at rather low round goby abundance levels (< 6 ind. 450 m−1) with a density-dependent variation in detection-success (see Fig. 5). Therefore, the eDNA approach seems to be ideal as an early detection method of a recent invasion of N. melanostomus already at low abundances. Nevertheless, further studies that compare the detection efficiency of electrofishing and eDNA for populations with low goby abundances are needed.
Traditional field survey techniques such as electrofishing are still required to differentiate abundance levels and biomass (but see Jane et al. 2015 and Yates et al. 2019 for positive correlations of eDNA concentrations and population abundance) and especially the age structure of invasive gobies under natural conditions.
The presented study captures the early invasion process of a large river ecosystem before continuous establishment along the river stretch is completed. The spatially explicit sampling of both density and age structure along the river stretch plus early reports of occurrence allows a reconstruction of the major dispersal patterns and their effects on the population composition. Particularly the location of the initial occurrence strongly determines the colonization progress of the total river system due to distinctly different upstream versus downstream migration speeds.
Change history
20 October 2021
A Correction to this paper has been published: https://doi.org/10.1007/s10530-021-02643-z
References
Adrian-Kalchhauser I, Burkhardt-Holm P (2016) An eDNA assay to monitor a globally invasive fish species from flowing freshwater. PLoS ONE 11:e0147558. https://doi.org/10.1371/journal.pone.0147558
Borcherding J, Staas S, Krüger S, Ondračková M, Šlapanský L, Jurajda P (2011) Non-native gobiid species in the lower river Rhine (Germany): recent range extensions and densities. J Appl Ichthyol 27:153–155. https://doi.org/10.1111/j.1439-0426.2010.01662.x
Borcherding J, Arndt H, Breiden S, Brenner K, Heermann L, Höfer S, Leistenschneider C, Lindner J, Staas S, Gertzen S (2016) Drift of fish larvae and juveniles in the lower Rhine before and after the goby invasion. Limnologica 59:53–62. https://doi.org/10.1016/j.limno.2016.05.001
Brabender M, Weitere M, Anlanger C, Brauns M (2016) Secondary production and richness of native and non-native macroinvertebrates are driven by human-altered shoreline morphology in a large river. Hydrobiologia 776:51–65. https://doi.org/10.1007/s10750-016-2734-6
Brownscombe JW, Fox MG (2012) Range expansion dynamics of the invasive round goby (Neogobius melanostomus) in a river system. Aquat Ecol 46:175–189. https://doi.org/10.1007/s10452-012-9390-3
Brownscombe JW, Masson L, Beresford DV, Fox MG (2012) Modeling round goby Neogobius melanostomus range expansion in a Canadian river system. Aquat Invasions 7:537–545. https://doi.org/10.3391/ai.2012.7.4.010
Buřič M, Bláha M, Kouba A, Drozd B (2015) Upstream expansion of round goby (Neogobius melanostomus)-first record in the upper reaches of the Elbe river. Knowl Manag Aquat Ecosyst 416:1–5. https://doi.org/10.1051/kmae/2015029
Cerwenka AF, Brandner J, Schliewen UK, Geist J (2018) Population trends of invasive alien gobies in the upper Danube River: 10 years after first detection of the globally invasive round goby (Neogobius melanostomus). Aquat Invasions 13:525–535. https://doi.org/10.3391/ai.2018.13.4.10
Duemler JJ, Kozelichki JM, Simon TP (2016) Relationships between length-weight, age, and body condition of the round goby Neogobius melanostomus (Pallas) in a tributary and harbor embayment in Southern Lake Michigan, USA. Turk J Fish Aquat Sci 16:205–212. https://doi.org/10.4194/1303-2712-v16_1_21
Evans NT, Shirey PD, Wieringa JG, Mahon AR, Lamberti GA (2017) Comparative cost and effort of fish distribution detection via environmental dna analysis and electrofishing. Fisheries 42:90–99. https://doi.org/10.1080/03632415.2017.1276329
Harrison JB, Sunday JM, Rogers SM (2019) Predicting the fate of eDNA in the environment and implications for studying biodiversity. Proc R Soc B 286:20191409. https://doi.org/10.1098/rspb.2019.1409
Hempel M, Thiel R (2013) First records of the round goby Neogobius melanostomus (Pallas, 1814) in the Elbe River, Germany. Bioinvasions Rec 2:291–295. https://doi.org/10.3391/bir.2013.2.4.05
Hofmann J, Behrendt H, Gilbert A, Janssen R, Kannen A, Kappenberg J (2005) Catchment-coastal zone interaction based upon scenario and model analyses: Elbe and the German Bight case study. Reg Environ Change 5:54–81. https://doi.org/10.1007/s10113-004-0082-y
Janáč M, Šlapanský L, Valová Z, Jurajda P (2013) Downstream drift of round goby (Neogobius melanostomus) and tubenose goby (Proterorhinus semilunaris) in their non-native area. Ecol Freshw Fish 22:430–438. https://doi.org/10.1111/eff.12037
Janáč M, Bryja J, Ondračková M, Mendel J, Jurajda P (2017) Genetic structure of three invasive gobiid species along the Danube–Rhine invasion corridor: similar distributions, different histories. Aquat Invasions 12:551–564. https://doi.org/10.3391/ai.2017.12.4.11
Janáč M, Jurajdová Z, Roche K, Šlapanský L, Jurajda P (2019) An isolated round goby population in the upper Elbe: population characteristics and short-term impacts on the native fish assemblage. Aquat Invasions 14:738–757. https://doi.org/10.3391/ai.2019.14.4.12
Jane S, Wilcox TM, MCKelvey KS, Young MK, Schwartz MK, Lowe WH, Letchers BH, Whiteley AR (2015) Distance, flow and PCR inhibition: eDNA dynamics in two headwater streams. Mol Ecol Res 15:216–227. https://doi.org/10.1111/1755-0998.12285
Leuven RSEW, van der Velde G, Baijens I, Snijders J, van der Zwart C, Lenders HJR, Bij de Vaate A (2009) The river Rhine: a global highway for dispersal of aquatic invasive species. Biol Invasions 11:1989–2008. https://doi.org/10.1007/s10530-009-9491-7
Matteikat W, Klefoth T, Emmrich M (2016) Charakteristika zweier populationen der schwarzmundgrundel Neogobius melanostomus, Pallas 1814 im Mittellandkanal. Lauterbornia 81:163–174
Pfeifer M, Völker F, Gause S (2016) Neue Fischart in Sachsen–Nachweis der Schwarzmundgrundel (Neogobius melanostomus, Pallas 1814). Fischer Angler Sachsen 4:166
Pilliod DS, Goldberg CS, Arkle RS, Waits LP (2014) Factors influencing detection of eDNA from a stream-dwelling amphibian. Mol Ecol Res 14:109–116. https://doi.org/10.1111/1755-0998.12159
Pont D, Rocle M, Valentini A, Civade R, Jean P, Maire A, Roset N, Schabuss M, Zornig H, Dejean T (2018) Environmental DNA reveals quantitative patterns of fish biodiversity in large rivers despite its downstream transportation. Sci Rep 8:10361. https://doi.org/10.1038/s41598-018-28424-8
Ramler D, Ahnelt H, Nemeschkal HL, Keckeis H (2016) The drift of early life stages of Percidae and Gobiidae (Pisces: Teleostei) in a free-flowing section of the Austrian Danube. Hydrobiologia 781:199–216. https://doi.org/10.1007/s10750-016-2845-0
Roche KF, Janáč M, Jurajda P (2013) A review of Gobiid expansion along the Danube–Rhine corridor-geopolitical changes as a driver for invasion. Knowl Manag Aquat Ecosyst 411:01–23. https://doi.org/10.1051/kmae/2013066
Roche K, Janáč M, Šlapanský L, Mikl L, Kopeček L, Jurajda P (2015) A newly established round goby (Neogobius melanostomus) population in the upper stretch of the river Elbe. Knowl Manag Aquat Ecosyst 416:33–44. https://doi.org/10.1051/kmae/2015030
Šlapanský L, Janáč M, Roche K, Jurajda P (2020) Round goby movement patterns in a non-navigable river. Can J Fish Aquat Sci 77:475–483. https://doi.org/10.1139/cjfas-2018-0488
Thiel R, Schulze M, Hempel M, Husemann M (2017) Most northerly record of the bighead goby Ponticola kessleri (Günther, 1861) in the Elbe River, Germany. Bioinvasions Rec 6:73–78. https://doi.org/10.3391/bir.2017.6.1.12
Veldhoen N, Hobbs J, Ikonomou G, Hii M, Lesperance M, Helbing CC (2016) Implementation of novel design features for qPCR-based eDNA assessment. PLoS ONE 11:e0164907. https://doi.org/10.1371/journal.pone.0164907
Völker F, Gause S (2017) Gewässerzustandsbewertung nach EU-WRRL-Teil Fische. Sächsisches Landesamt für Umwelt, Landwirtschaft und Geologie 1:9–10
Wacker S, Fossøy F, Larsen BM, Brandsegg H, Sivertsgård R, Karlsson S (2019) Downstream transport and seasonal variation in freshwater pearl mussel (Margaritifera margaritifera) eDNA concentration. Environ DNA 1:64–67. https://doi.org/10.1002/edn3.10
Yates MC, Fraser DJ, Derry AM (2019) Meta-analysis supports further refinement of eDNA for monitoring aquatic species-specific abundance in nature. Environ DNA 1:5–13. https://doi.org/10.1002/edn3.7
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised due to retrospective open access order.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nogueira Tavares, C., Brauns, M., Hille, S. et al. Tracing the colonization process of non-native gobies into a large river: the relevance of different dispersal modes. Biol Invasions 22, 2421–2429 (2020). https://doi.org/10.1007/s10530-020-02281-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-020-02281-x