Abstract
The Asian shore crab (Hemigrapsus sanguineus) and the brush-clawed shore crab (H. takanoi) both originate from east Asia but have in recent years established viable populations throughout coastal European waters. The two species are highly competitive and can occur in very high densities. Both species have been linked to the decrease of otherwise abundant native species such as the European shore crab (Carcinus maenas) and the common periwinkle (Littorina littorea). As both crabs are relatively new in European coastal waters, little is known about the full distribution range that may be achieved by these crabs and the ecological impact that may follow with an invasion of coastal habitats. In this study we investigated the potential distribution of Hemigrapsus in coastal waters of Northern Europe and the potential impact on the native competitor C. maenas. To this end we collected crabs on the Swedish west coast and on Helgoland and used these for behavioural experiments. We also collected new observation records of both species in Sweden and used these occurrences for building ecological niche models. We report that the potential distribution range of both H. sanguineus and H. takanoi extends from western Sweden to eastern Ireland (east–west) and southern Norway to western France (north–south). We also found evidence that H. takanoi can establish viable populations in nutrient-rich inshore areas of the western and southern Baltic Sea and that competition by H. sanguineus is likely to affect abundances of C. maenas in the areas where these species co-occur.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Non-indigenous species (NIS) are a global issue and the number of NIS has increased during the last decades (Cohen and Carlton 1998; European Environment Agency 2015; Zenetos et al. 2012). European waters are especially susceptible for invasive species from Asia and North America and the main invasion vectors are ballast water and transportation across the Suez Canal (Gollasch and Nehring 2006).
The Asian shore crab (Hemigrapsus sanguineus) was first discovered in Europe in 1999, found in Le Havre, France. The individuals found were already mature and breeding, thus it is believed that they arrived in Le Havre in 1998 at the latest (Breton et al. 2002). Since then, H. sanguineus has spread along the continental coast of North western Europe, from Northern France to the German North Sea coast and has recently been found in England, Wales and Scotland (Marine Biological Association 2017; Berggren and Karlsson 2017; Dauvin 2009a; Seeley et al. 2015). The species have also been found in the Black Sea by the coast of Romania and in the Mediterranean Sea off the coasts of Croatia and Tunisia (Amor et al. 2017; Micu et al. 2010; Schubart 2003), theses were however only a few or a single individual.
The brush-clawed shore crab (Hemigrapsus takanoi) was first found in La Rochelle, France, in 1994. This species was then known as Hemigrapsus penicillatus since H. takanoi was first described as a species in 2005 by Asakura and Watanabe (2005). H. penicillatus is currently not present in Europe. The current distribution of H. takanoi in Europe reaches from north-eastern Spain to the Swedish west coast as well as south-eastern England, and western and eastern Scotland. In addition, there is one observation from the German coast facing the Baltic Sea (Marine Biological Association 2017; Berggren and Karlsson 2017; Geburzi et al. 2015; Noél et al. 1997; Wood et al. 2015).
Both crabs originate from east Asia and are considered invasive alien species (IAS) as they are highly competitive in many of the locations where they have become established. Outside of Long Island, USA, the presence of H. sanguineus has reduced the number of the common periwinkle (Littorina littorea) by 80% and the native flatback mud crab (Eurypanopeus depressus) by 95% (Kraemer et al. 2007). At this location, H. sanguineus was observed in an average abundance of 120 crabs/m2, and at one instance more than 300 crabs/m2. Additionally, during both spring and fall of 2016, mean densities above 300 crabs/m2 where recorded in Rhode Island, USA, (O’Connor 2018), further indicating the potential density of the species.
As both crabs are relatively new in European coastal waters, little is known about the full distribution range that may be achieved by these crabs and the ecological impact that may follow with an invasion of coastal habitats. The goal of this study was to investigate aspects of potential distribution and impact for both invasive Hemigrapsus species in Northern Europe. To this end we designed competition experiments with a dominant native competitor, the European green crab Carcinus maenas, identified important environmental factors that influence the distributions of H. sanguineus and H. takanoi, and modelled potential distribution areas of both crabs in Northern Europe.
Materials and methods
Field sampling
Hemigrapsus sanguineus were collected by hand and with traps on the island of Helgoland, Germany, between the 2nd and 4th of August 2017. Crabs were collected during the day at low and incoming tide. The traps were placed in the subtidal zone at approx. 0.3–0.5 m depth for 1 h, meanwhile crabs were collected by turning rocks in the intertidal zone. The traps were constructed out of PVC pipe (approx. 500 mm long and 100 mm Ø) with a funnel (25 mm Ø opening) in one end and a 1 mm steel mesh in the other end. In the middle of the PVC pipe a drill out cylindrical tube was fitted, containing a crushed Mytilus edulis. The traps were weighted down by two heavy chains, attached to each side of the trap (total weight of approx. 5 kg).
The crabs were then brought to the Sven Lovén Centre for Marine Infrastructure—Kristineberg, Sweden by transport in an electrical cooling bag, set to 10 °C to minimize activity and potential cannibalism, filled up with Fucus spp. The transportation took approx. 48 h, no crabs died during transit.
Individuals of C. maenas were collected by hand and with the use of traps, in Smalsund, outside of Kristineberg on the 9th of August 2017. The traps were of the same design as those used in Helgoland. Traps were deployed in the subtidal zone at a depth of 0.2–0.5 m and left them for approx. 1 h, during which crabs were collected by turning rocks in the intertidal.
The crabs were kept in separate holding tanks before and during the experiments. The water in the tanks was filtrated sea water with a salinity of 25 ± 1 PSU and a temperature of 19 ± 1 °C. At the time of collection, the waters in Helgoland had a salinity of 33 PSU and a temperature of 19 °C.
Competition experiments with Carcinus maenas
Experiments started on the 15th of August, 11 days after the last sampling day for H. sanguineus and 6 days after the sampling of C. maenas. The experiments comprised three stages (early invasion, mid invasion, late invasion) always containing 10 crabs per aquaria but with different proportions of species and sex depending on the stage (Table 1). These stages are to reflect on the different species ratios during a potential invasion: invader species in minority, in equal number to the native, and eventually in majority. The sex ratio for both is based on the sex ratio of C. maenas found in the field along the Swedish west coast (Jungblut and Karlsson unpubl. data).
Crabs used in the experiments had both chelae intact and at least 2 legs on each side of the body, no newly moulded crabs were used. The carapace width (CW) of crabs used in the experiments had the same dimensions in both species: 20–35 mm CW for males, and 15–25 mm CW for females. To distinguish males and females apart during the experiment, red nail-polish was applied to the carapace of the females. Crabs were randomly collected from the holding tanks. Due to limited number of H. sanguineus, crabs were returned to the holding tank after the experiment. Hence these crabs may have been used multiple times. However, after each round of experiments, all crabs were feed M. edulis and then rested for at least 48 h until the next experiment started.
Three experimental aquaria were recorded simultaneously using a Logitech C920 webcam and the recording software OBS 20.0.1. The aquaria used had a surface area of 0.1 m2 (210 × 475 mm), resulting in a density of ~ 100 crabs/m2, a density common in the field. In each aquarium a thin layer of filtered shell-sand (1–5 mm grain size) without organic matter was used as a substrate.
The first experiment assessed if H. sanguineus can actively dislodge C. maenas and/or restrain its food sources at any of the three invasive stages. To this end, all crabs were kept without food in holding tanks with filtered seawater for 48 h. Thereafter, the animals were transferred to the experimental aquaria and a M. edulis (length 80 ± 10 mm with one side crushed) was placed in each aquarium. The aquaria were then recorded for 30 min. At 1, 5, 10, 15, 20, 25 and 30 min into the recording, presence or absence of feeding on the mussel was noted for each crab species and each of its sexes.
The mussels were then removed, and a man-made shelter was placed in each aquarium. The shelter was constructed by a PVC pipe (150 mm in length and 67 mm in diameter) which was attached to a plastic slab (200 × 125 mm) to keep the pipe in place. The shelter was then covered in a layer of sand and the crabs were then left for 30 min to settle down after being disturbed and then recorded for 30 min. At 1, 5, 10, 15, 20, 25 and 30 min into the recording presence or absence of crabs in the shelter was noted for each species and each sex. A minimum of half of the crab needed to be covered by the shelter to be considered “sheltered”. After the end of the experiment, the crabs were relocated to their holding tanks.
For each group (e.g. C. maenas males), a mean feeding ratio/hiding ratio was calculated based on the presence (1) or absence (0) at the food location/inside the man-made shelter at each time point (1, 5, 10 min etc.). This was done separately for each of the 6 replicates (aquaria). These 6 replicates were used to calculate the 95% confidence interval as well as a pairwise t test assuming unequal variance (Table S1).
Ecological modelling
Species and environmental data
The selected geographical area for this study include the coastal waters of Europe from the French west coast in the south to the Faroe Islands in the north, and also the Baltic Sea. Occurrence records (presence only) were obtained by sampling the Swedish West coast, through the literature (Dauvin et al. 2009; Dauvin 2009b; Van den Brink et al. 2012) as well as from the Global Biodiversity Information Facility (GBIF 2017a, b). Records obtained by the authors and collaborators were submitted to GBIF through the Swedish Species Gateway (http://www.Artportalen.se). All occurrence records were carefully revised and verified before being used in the models, while unverified records were discarded.
We used gridded environmental data available as global marine layers through Bio-Oracle (http://www.bio-oracle.ugent.be/) with a resolution of 5 arc-min (Tyberghein et al. 2012). These data layers are generated from monthly satellite data (Aqua-MODIS and SeaWiFS; https://oceancolor.gsfc.nasa.gov/) as well as in situ measured oceanographic data from the World Ocean Database 2009 (Boyer et al. 2009). We also used marine layers from AquaMaps (http://www.aquamaps.org/download/main.php) with a resolution of 30 arc-min (Kaschner et al. 2008). These layers were built from long-term averages of temporally varying environmental variables (Ready et al. 2010).
Environmental data was obtained using version 2 of the BioClim workflow (http://purl.ox.ac.uk/workflow/myexp-3725.2) available at the BioVeL portal (Hardisty et al. 2016). The workflow was used to retrieve environmentally unique points (Nix 1986; Vestbo et al. 2018) from the species occurrence files for a set of 14 environmental data sets (abbreviations used throughout the article are given in parentheses), including the following. Bio-Oracle (5 arc-min): Mean dissolved oxygen in ml/l (Oxy), Mean nitrate [NO3] [NO3 + NO2] in µmol/l (NO3), Mean phosphate in µmol/l (PO4), Maximum sea surface temperature in °C (Max SST), Minimum sea surface temperature in °C (Min SST), Sea surface temperature range in °C (Range SST), Mean calcite concentration in mol/m3 (CaCO3), Maximum chlorophyll A concentration in mg/m3 (Max ChlA), Minimum chlorophyll A concentration in mg/m3 (Min ChlA), Range of chlorophyll A concentration in mg/m3 (Range ChlA). AquaMaps (30 arc-min): Mean ice concentration in % (Ice), Mean sea surface salinity in PSU (SSS), Mean sea surface temperature °C (SST), Mean chlorophyll A concentration in mg/m3 (Mean ChlA). Chlorophyll A data sets were included as chlorophyll acts as a proxy for phytoplankton, and thus reflects the amount of nutrition in the water.
Statistical analysis
Initially, we conducted a correlation analysis (data not shown) as well as a principal component analysis (PCA) on log-transformed data using the R statistical environment 3.0.2 (R Core Team 2013). Both analyses were performed to identify the variables explaining the variation in the data set, and which can be used as predictor variables in the modelling. The PCA was also used to compare the environmental space occupied by the two species and estimated as a 9 and 10-dimensional hypervolume (Fig. S1). We used all non-correlated variables that were represented (with > 5%) in the first two components of the PCA to build the niche models.
Ecological niche modelling
We used version 20 of the ecological niche modelling (ENM) workflow (http://purl.ox.ac.uk/workflow/myexp-3355.20) to describe and compare the geographical and environmental space occupied by the two species and to estimate the potential distribution maps (PD) with favourable biotic and environmental conditions in the study region. We did not consider demographic, dispersal, or substrate properties that may also be used in species distribution modelling (Peterson et al. 2011; Reiss et al. 2014). For more information on the ENM workflows see De Giovanni et al. (2015), Holl et al. (2013), Leidenberger et al. (2015) and Vestbo et al. (2018). We executed parallel analyses with two ENM algorithms, Maximum Entropy v. 1.0 (Phillips et al. 2006; Phillips and Dudík 2008), and Support vector machine v. 0.5 (Schölkopf et al. 2001).
Models were created using each species’ maximum distribution range within the study region and the set of predictor variables identified in the PCA. Maxent models were set to run with 10.000 background points (including input points) drawn from the mask. Feature selection was automated, allowing the algorithm to combine feature types when fitting a model, and perform 500 iterations. Tolerance for detecting model convergence was set to 0.00001, while sample threshold was set to 80 (product), 10 (quadratic), and 15 (hinge). SVM models were set to execute the C-SVC algorithm with radial basis kernels, gamma values 1/k (where k is the number of layers), and a cost value of 1. All models were set to produce a probabilistic, instead of binary output.
For each species, we ran both algorithms across the above-specified environmental layers. Models were tested using fivefold cross-validation based on the area under the curve (AUC) value and omission error rate (false negative rate, OE), and subsequently projected using the same layers (native projections). The results of the ENMs were visualized as potential distribution maps (PD) maps, showing the suitable environment of a species in the region of interest. Overall, we executed 4 niche models (2 algorithms, 2 species).
Raster maps created by the niche modelling algorithms were processed using the qGIS software package v. 2.6 Brighton (Quantum GIS Development Team 2014). We produce PD maps as consensus from the raster values of both algorithms, where red indicates agreement between both algorithms according to lowest presence threshold in each cell (Pearson et al. 2007).
Results
Behavioural experiments
Food competition
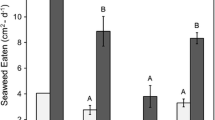
Male C. maenas showed a significant decrease in feeding ratio between the mid- and late invasion stage. The late invasion stage feeding ratio was also significantly different to the early invasion stage’s feeding ratio (Fig. 1). Likewise, female C. maenas had a significant increase in feeding ratio between the mid- and late invasion stage (Fig. 1), while no significant changes in feeding ratio were observed with either H. sanguineus groups (Fig. 1).
Male H. sanguineus had a significantly higher feeding ratio compared to male C. maenas at the late invasion stage (Fig. 1). Female H. sanguineus had significantly lower feeding ratio compared to female C. maenas at the early- and late invasion stage (Fig. 1). All crabs survived the experiments.
Shelter competition
Female H. sanguineus had a significant increase in hiding ratio between the mid- and late invasion stage. The late invasion stage hiding ratio was also significantly different to the early invasion stage’s hiding ratio (Fig. 1). Female C. maenas had a significant decrease in hiding ratio when comparing early- and late invasion stage. No significant changes in hiding ratio were observed with either male group (Fig. 1).
Female H. sanguineus had a significantly higher hiding ratio compared to female C. maenas at the late invasion stage (Fig. 1). No significant differences in hiding ratio were observed between the male groups (Fig. 1). All crabs survived the experiments.
Ecological niche models
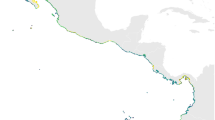
The predictor variables selected by the PCA were the same for both species: mean nitrate concentration (NO3), minimum SST (Min SST), mean calcite concentration (CaCO3) and minimum chlorophyll A concentration (Min ChlA). We also added mean surface salinity (SSS) to both models to increase the accuracy of the predictions in the Baltic Sea, which has a strong salinity gradient. The ENM models performed well in the cross-validation returning high AUC values and low omission errors (Table 2). The resulting PD maps were very similar for the two species with suitable environment throughout the Northern European continental coast between Northern France and eastern Denmark, as well as along the south coast of Norway and west coast of Sweden (Fig. 2). In the south, the potential distribution extends to the Atlantic coastline of France. In the British Isles, large areas of suitable environment are found in much of eastern and western England, Wales, southern Scotland and eastern Ireland. In the Baltic Sea, no suitable environment was found for H. sanguineus. However, for H. takanoi, suitable environment was found in the large estuaries of the Southern Baltic Sea, e.g. in the Szczecin Lagoon, the Vistula Lagoon, and the Curonian lagoon.
Discussion
Behaviour and potential ecological impact of Hemigrapsus
The behavioural experiments show that male H. sanguineus competes well with male C. maenas, even when outnumbered 2 to 5 (Fig. 1). This will benefit the specie’s establishment, as then even few individuals may be able to compete for food in areas with large abundances of C. maenas. In areas along the American east coast, where H. sanguineus occurs together with the likewise invasive green crab (C. maenas) it has been observed that H. sanguineus outcompetes juvenile C. maenas in the competition for shelter. When H. sanguineus was not present, 97% of all C. maenas were found under rocks and when H. sanguineus was present, 22% of C. maenas were found under rocks (Jensen et al. 2002). Furthermore, H. sanguineus has been found to prey on juvenile C. maenas (Lohrer and Whitlatch 2002a). In Europe, in the Dutch delta, H. takanoi occurs in high numbers and is believed to contribute to the decrease of C. maenas in the area (Van den Brink et al. 2012). Previous experimental studies have also shown that the presence of H. sanguineus alters the food preference of C. maenas, changing it from mainly mussels to red algae. This could have a negative impact on C. maenas, as a reduction in mussel consumption can result in lower growth rates (Ponat and Adelung 1980).
Female H. sanguineus had a low feeding ratio throughout all invasion stages, and if they do not feed, the females cannot produce offspring to sustain the population. It could be that females have a different food preference compared to the males and do not prey on bivalves. Previous studies have suggested that H. sanguineus mainly is herbivorous (Griffen 2011; Griffen et al. 2012; Tyrrell and Harris 2000), which could explain the low feeding ratio in this study.
If H. sanguineus establish itself along in the eastern extensions of the North Sea (Skagerrak and Kattegat), it may reduce the numbers of Littorina snails in some areas, as it has done in areas of the US (Kraemer et al. 2007), as well as M. edulis (Lohrer and Whitlatch 2002b). When comparing a single crab, previous studies have been inconclusive about whether H. sanguineus has an equal or higher feeding rate on M. edulis compared to C. maenas, supported by DeGraaf and Tyrrell (2004) but opposed by MacDonald et al. (2007) as well as Lohrer and Whitlatch (2002b). However, as H. sanguineus can occur in higher abundances compared to C. maenas, the impact on prey species such as M. edulis can potentially become higher. For native species feeding on algae (e.g. Littorina spp.), H. sanguineus might become, not only a predator but a competitor as it also feeds on algae, especially during its juvenile stage (Griffen 2011; Griffen et al. 2012; Tyrrell and Harris 2000). Further, H. sanguineus over all seem to feed more on animal tissue in laboratory experiment than in field experiments (Griffen 2011).
During the food competition experiment, differences in behaviour could be observed between the male C. maenas and H. sanguineus. C. maenas showed a more aggressive behaviour, not only towards H. sanguineus but also intraspecific aggression, and often used its chelae to pinch any crab which came close. However, H. sanguineus did not display that type of aggressive behaviour. H. sanguineus instead shoved other crabs out of its way and wedged itself in between the other crabs and the food. Once it started feeding, it used its big chelae to push away other crabs. This behaviour was however rarely seen towards its own species. Similar behaviour has been documented by Jensen et al. (2002), who wrote that H. sanguineus dislodged C. maenas from the food and then used its legs to fend off the competitor. Previous studies have also shown that H. sanguineus exhibits a higher conspecific tolerance than C. maenas (Hobbs et al. 2017), which is highly beneficial for H. sanguineus as it facilitates the species to exist in higher densities compared to C. maenas.
The effectiveness of male H. sanguineus can be seen when comparing the early and late food experiment (Table S1), where male C. maenas had a higher feeding ratio (not significantly) despite being 5 to 2. When male C. maenas were in minority (despite feeding at 1 min in all replicates), male H. sanguineus had a significantly higher feeding ratio (P = 0.0007).
There was a distinct difference in behaviour between the sexes of H. sanguineus, where males were active and competed well, while females were passive and competed poorly (Fig. 1). This is something which, to our knowledge, has not been reported in the literature before. It suggests that a species’ behaviour may vary strongly between the sexes, and future studies may further investigate the behavioural differences between males and females to improve our knowledge of H. sanguineus’ foraging behaviour.
The potential stress on the collected H. sanguineus due to a change in salinity, cannot be completely disregarded and could have affected the results of the experiments. However, this is not deemed probable as the crabs had 9 days to acclimatize to the new salinity (25 ± 1 PSU). Additionally, H. sanguineus is found in waters of similar salinity and have shown to tolerate lower salinities in experiments (Gerard et al. 1999; Jungblut et al. 2015; Ledesma and O’Connor 2001; McDermott 1999).
Potential distribution range of Hemigrapsus
Both Hemigrapsus species prefer areas with high productivity and with high levels of nitrate as indicated by the selected predictor variables. The areas are often found in near coast environments like estuaries, archipelagos, and lagoons (Fig. 2). The presence of large cities and ports in these areas as well as the traffic between them will likely facilitate further spreading of both species and may allow these crabs to establish new populations in areas that could not be reached by natural dispersal. Besides high productivity and nitrate, both species prefer areas with high amounts of calcite. This may be a factor limiting the distribution in the north, along the south-western coast of Norway, as calcite levels are low in these areas (source, Bio-Oracle).
Our ENMs suggest that both species will be able to expand their distribution ranges northwards to the southern coast of Norway, as well as southward into the Öresund and Belt regions between Denmark and Sweden (Fig. 2). Noticeably, H. takanoi finds suitable environment even in semi-enclosed lagoons along the southern coast of the Baltic Sea. Here, the alien crab could have a greater chance of establishment and impact due to the low species diversity of the area (Paavola et al. 2005). H. takanoi has already been found in the Baltic Sea, in Kiel (Geburzi et al. 2015), and it is likely that the species has arrived there via the Kiel Canal, which connects the Wadden Sea with the Baltic Sea. If this is the case, the species could rapidly increase in numbers in the bay of Kiel. In order to establish populations further east, e.g. in the Szczecin Lagoon, the Vistula Lagoon, or the Curonian lagoon, larvae or juveniles will need to travel with vessels, as the predominant surface current flows from east to west. However, many ferry lines commute between these hotspots, e.g. between Kiel and Kleipeda and may support further spread if ballast water is not treated.
In the British Isles, both species have been recorded in southern England and southern Scotland (Marine Biological Association 2017; Seeley et al. 2015). Most of the coast of England and the complete coastline of the Irish Sea were deemed suitable according to the models and the many ports in the area could facilitate dispersion, which could lead to a rapid colonization of the region. Along the French west coast both species find suitable environment according to our models, but only H. takanoi have so far been recorded in the region (Dauvin et al. 2009; Noél et al. 1997). It is possible that the Brittany peninsula acts as a barrier, restricting natural southward dispersal of H. sanguineus from the English Channel (Fig. 2). If so, there is a risk that also in this region maritime traffic may facilitate the transfer of larva to the Atlantic coast.
Future studies
Future studies should investigate the invasion process more in-depth, incl. ecological relations to other species than C. maenas. Doi et al. (2009) reported that H. takanoi could crush larger mussels (Xenostrobus securis) than that of similar sized Carcinus aestuarii. If both Hemigrapsus species have a wider range in sizes of M. edulis and Littorina spp. on which they can prey compared to C. maenas (> 35 mm in carapace width) then they could increase the predation pressure on molluscs in the intertidal and upper subtidal zones. Experiments examining size preference of these prey species are therefore needed. In addition, evolutionary studies may help to understand the potential of the species to fully adapt to Baltic conditions. Investigating H. sanguineus’ and H. takanoi’s abilities to adapt to new environments, similar to the studies done on C. maenas by Best et al. (2017) and Tepolt and Somero (2014). Finally, population genetic studies similar to that of Reusch et al. (2010) may prove helpful to better understand the invasive pathways of Hemigrapsus, which is likely to be caused by a combination of regional currents and ship traffic.
References
Amor KO-B, Amor MMB, Rifi M, Souissi JB (2017) Diversity of crustacean species from Tunis Southern Lagoon (Central Mediterranean) after an ecological restoration. Cah Biol Mar 58:49–57
Asakura A, Watanabe S (2005) Hemigrapsus takanoi, new species, a sibling species of the common Japanese intertidal crab H. penicillatus (Decapoda: Brachyura: Grapsoidea). J Crustac Biol 25:279–292
Berggren M, Karlsson R (2017) Invasiva, asiatiska krabbor längs Västkusten. Fauna & Flora 112:22–26
Best K, McKenzie CH, Couturier C (2017) Reproductive biology of an invasive population of European green crab, Carcinus maenas, in Placentia Bay. Nfld Manag 8:247–255
Boyer TP et al (2009) World ocean database 2009. In: Levitus S (ed) NOAA Atlas NESDIS 66. U.S. Government Printing Office, Washingtion, DC, 216 pp
Breton G, Faasse M, Noël P, Vincent T (2002) A new alien crab in Europe: Hemigrapsus sanguineus (Decapoda: Brachyura: Grapsidae). J Crustac Biol 22:184–189
Cohen AN, Carlton JT (1998) Accelerating invasion rate in a highly invaded estuary. Science 279:555–558
Dauvin J-C (2009a) Asian shore crabs Hemigrapsus spp. (Crustacea: Brachyura: Grapsoidea) continue their invasion around the Cotentin Peninsula, Normandy, France: status of the Hemigrapsus population in 2009. Aquat Invasions 4:605–611
Dauvin J-C (2009b) Establishment of the invasive Asian shore crab Hemigrapsus sanguineus (De Haan, 1835) (Crustacea: Brachyura: Grapsoidea) from the Cotentin Peninsular, Normandy, France. Aquat Invasions 4:467–472
Dauvin J-C, Tous Rius A, Ruellet T (2009) Recent expansion of two invasive crabs species Hemigrapsus sanguineus (de Haan, 1835) and H. takanoi Asakura and Watanabe 2005 along the Opal Coast, France. Aquat Invasions 4:451–465
De Giovanni R, Williams AR, Ernst VH, Kulawik R, Fernandez FQ, Hardisty AR (2015) ENM components: a new set of web service-based workflow components for ecological niche modelling. Ecography 39:376–383
DeGraaf JD, Tyrrell MC (2004) Comparison of the feeding rates of two introduced crab species, Carcinus maenas and Hemigrapsus sanguineus, on the blue mussel, Mytilus edulis. Northeast Nat 11:163–167
Doi W, Iinuma Y, Yokota M, Watanabe S (2009) Comparative feeding behavior of invasive (Carcinus aestuarii) and native crabs (Hemigrapsus takanoi). Crustac Res 38:1–11
European Environment Agency (2015) Trends in marine non-indigenous species. Accessed 4 Apr 2019
GBIF (2017a) Hemigrapsus sanguineus (De Haan, 1835) GBIF backbone taxonomy checklist dataset. https://doi.org/10.15468/39omei. Accessed via GBIF.org on 3 Oct 2017
GBIF (2017b) Hemigrapsus takanoi Asakura & Watanabe GBIF backbone taxonomy checklist dataset. https://doi.org/10.15468/39omei. Accessed via GBIF.org on 3 Oct 2017
Geburzi JC, Graumann G, Köhnk S, Brandis D (2015) First record of the Asian crab Hemigrapsus takanoi Asakura & Watanabe, 2005 (Decapoda, Brachyura, Varunidae) in the Baltic Sea. BioInvasions Rec 4:103–107
Gerard V, Cerrato R, Larson A (1999) Potential impacts of a western Pacific grapsid crab on intertidal communities of the northwestern Atlantic Ocean. Biol Invasions 1:353–361
Gollasch S, Nehring S (2006) National checklist for aquatic alien species in Germany. Aquat Invasions 1:245–269
Griffen BD (2011) Ecological impacts of replacing one invasive species with another in rocky intertidal areas. In: Galil BS, Clark PF, Carlton JT (eds) In the wrong place-alien marine crustaceans: distribution, biology and impacts. Springer, Berlin, pp 687–701
Griffen BD, Altman I, Bess BM, Hurley J, Penfield A (2012) The role of foraging in the success of invasive Asian shore crabs in New England. Biol Invasions 14:2545–2558
Hardisty AR et al (2016) BioVeL: a virtual laboratory for data analysis and modelling in biodiversity science and ecology. BMC Ecol 16:49
Hobbs N-VS, Cobb JS, Thornber CS (2017) Conspecific tolerance and heterospecific competition as mechanisms for overcoming resistance to invasion by an intertidal crab. Biol Invasions 19:765–772
Holl S, Garijo D, Belhajjame K, Zimmermann O, De Giovanni R, Obst M, Goble C (2013) On specifying and sharing scientific workflow optimization results using research objects. In: Proceedings of the 8th workshop on workflows in support of large-scale science. ACM, pp 28–37
Jensen GC, McDonald PS, Armstrong DA (2002) East meets west: competitive interactions between green crab Carcinus maenas, and native and introduced shore crab Hemigrapsus spp. Mar Ecol Prog Ser 225:251–262
Jungblut S, Boos K, Saborowski R, Hagen W (2015) Temperature and salinity tolerance of the Asian shore crab Hemigrapsus sanguineus. Crustaceologen-Tagung, Bremerhaven, Germany, 26–29 March 2015
Kaschner K et al (2008) AquaMaps environmental dataset: half-degree cells authority file (HCAF). World Wide Web electronic publication, www.aquamaps.org/data. Version 07/2010
Kraemer GP, Sellberg M, Gordon A, Main J (2007) Eight-year record of Hemigrapsus sanguineus (Asian shore crab) invasion in western Long Island Sound estuary. Northeast Nat 14:207–224
Ledesma ME, O’Connor NJ (2001) Habitat and diet of the non-native crab Hemigrapsus sanguineus in southeastern New England. Northeast Nat 8:63–79
Leidenberger S, De Giovanni R, Kulawik R, Williams AR, Bourlat SJ (2015) Mapping present and future potential distribution patterns for a meso-grazer guild in the Baltic Sea. J Biogeogr 42:241–254
Lohrer AM, Whitlatch RB (2002a) Interactions among aliens: apparent replacement of one exotic species by another. Ecology 83:719–732
Lohrer AM, Whitlatch RB (2002b) Relative impacts of two exotic brachyuran species on blue mussel populations in Long Island Sound. Mar Ecol Prog Ser 227:135–144
MacDonald JA, Roudez R, Glover T, Weis JS (2007) The invasive green crab and Japanese shore crab: behavioral interactions with a native crab species, the blue crab. Biol Invasions 9:837–848
Marine Biological Association (2017) Ongoing United Kingdom MarLIN Shore GBIF.org
McDermott JJ (1999) Natural history and biology of the Asian shore crab Hemigrapsus sanguineus in the western Atlantic: a review, with new information. In: Marine bioinvasions: proceedings of a conference January
Micu D, Nita V, Todorova V (2010) First record of the Japanese shore crab Hemigrapsus sanguineus (de Haan, 1835) (Brachyura: Grapsoidea: Varunidae) from the Black Sea. Aquat Invasions 5:1–4
Nix HA (1986) A biogeographic analysis of Australian elapid snakes. In: Atlas of Australian elapid snakes. Bureau of Flora and Fauna, Canberra, pp 4–15
Noél PY, Tardy E, d’Acoz CdU (1997) Will the crab Hemigrapsus penicillatus invade the coasts of Europe? C R Acad Sci-Ser III 320:741–745
O’Connor NJ (2018) Changes in Population Sizes of Hemigrapsus sanguineus (Asian Shore Crab) and Resident Crab Species in Southeastern New England (2010–2016). Northeast Nat 25:197–202
Paavola M, Olenin S, Leppäkoski E (2005) Are invasive species most successful in habitats of low native species richness across European brackish water seas? Estuar Coast Shelf Sci 64:738–750
Pearson RG, Raxworthy CJ, Nakamura M, Townsend Peterson A (2007) Predicting species distributions from small numbers of occurrence records: a test case using cryptic geckos in Madagascar. J Biogeogr 34:102–117
Peterson AT, Sobersón J, Pearson RG, Anderson RP, Martínez-Meyer E, Nakamura M, Araújo MB (2011) Ecological niches and geographic distributions, vol 49. Princeton University Press, Princeton
Phillips SJ, Dudík M (2008) Modeling of species distributions with Maxent: new extensions and a comprehensive evaluation. Ecography 31:161–175
Phillips SJ, Anderson RP, Schapire RE (2006) Maximum entropy modeling of species geographic distributions. Ecol Model 190:231–259
Ponat A, Adelung D (1980) Studies to establish an optimal diet for Carcinus maenas II. Protein and lipid requirements Marine Biology 60:115–122
R Core Team (2013) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Ready JS et al (2010) Predicting the distributions of marine organisms at the global scale. Ecol Model 221:467–478
Reiss H et al (2014) Benthos distribution modelling and its relevance for marine ecosystem management. ICES J Mar Sci 72(2):297–315
Reusch TB, Bolte S, Sparwel M, Moss AG, Javidpour JJME (2010) Microsatellites reveal origin and genetic diversity of Eurasian invasions by one of the world’s most notorious marine invader, Mnemiopsis leidyi (Ctenophora). Mol Ecol 19:2690–2699
Schölkopf B, Platt JC, Shawe-Taylor J, Smola AJ, Williamson RC (2001) Estimating the support of a high-dimensional distribution. Neural Comput 13:1443–1471
Schubart CD (2003) The East Asian shore crab Hemigrapsus sanguineus (Brachyura: Varunidae) in the Mediterranean Sea: an independent human-mediated introduction. Sci Mar 67:195–200
Seeley B, Sewell J, Clark PF (2015) First GB records of the invasive Asian shore crab, Hemigrapsus sanguineus from Glamorgan, Wales and Kent, England. Mar Biodivers Rec 8:1–4
Quantum GIS Development Team (2014) Quantum GIS geographic information system. Open Source Geospatial Foundation Project. http://qgis.osgeo.org
Tepolt CK, Somero GN (2014) Master of all trades: thermal acclimation and adaptation of cardiac function in a broadly distributed marine invasive species, the European green crab, Carcinus maenas. J Exp Biol 217:1129–1138
Tyberghein L, Verbruggen H, Pauly K, Troupin C, Mineur F, De Clerck O (2012) Bio-ORACLE: a global environmental dataset for marine species distribution modelling. Glob Ecol Biogeogr 21:272–281
Tyrrell M, Harris L (2000) Potential impact of the introduced Asian shore crab, Hemigrapsus sanguineus, in Northern New England: diet, feeding preferences, and overlap with the green crab, Carcinus maenas. In: Marine bioinvasions: proceedings of the first national conference, Cambridge, MA, 24–27 Jan 1999. MIT Sea Grant College Program, pp 208–220
Van den Brink A, Wijnhoven S, McLay C (2012) Competition and niche segregation following the arrival of Hemigrapsus takanoi in the formerly Carcinus maenas dominated Dutch delta. J Sea Res 73:126–136
Vestbo S, Obst M, Quevedo Fernandez FJ, Intanai I, Funch PJ (2018) Present and potential future distributions of Asian horseshoe crabs determine areas for conservation. Front Mar Sci 5:164
Wood CA, Bishop JDD, Davies CJ, Delduca EL, Hatton JC, Herbert RJH, Clark PF (2015) Hemigrapsus takanoi Asakura and Watanabe, 2005 (Crustacea: Decapoda: Brachyura: Grapsoidea): first records of the brush-clawed shore crab from Great Britain. Bioinvasions Rec 4:109–113. https://doi.org/10.3391/bir.2015.4.2.07
Zenetos Α et al (2012) Alien species in the Mediterranean Sea by 2012. A contribution to the application of European Union’s Marine Strategy Framework Directive (MSFD). Part 2. Introduction trends and pathways. Mediterr Mar Sci 13:328–352
Acknowledgements
We would like to acknowledge and thank Simon Jungblut for making it possible for the first author to visit Helgoland and collect H. sanguineus as well as for the help when collecting the crabs. This study is supported by the EU’s Seventh Framework Program Project BioVeL (www.biovel.eu, Grant No. 283359), the Swedish LifeWatch program funded by the Swedish Research Council (Grant No. 829-2009-6278).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Karlsson, R., Obst, M. & Berggren, M. Analysis of potential distribution and impacts for two species of alien crabs in Northern Europe. Biol Invasions 21, 3109–3119 (2019). https://doi.org/10.1007/s10530-019-02044-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-019-02044-3