Abstract
Competitive dynamics between native and exotic species can influence both the success of exotics in the novel environment as well as diversity and abundance of native species. Invasive species are often characterized by multiple introductions in the novel range, which can lead to population differentiation for invasion characteristics. Here we use two invasive lineages of the exotic grass, Aegilops triuncialis L., to determine if these lineages differ in their response to competitors and in their persistence in the invaded range. We find that one lineage is negatively affected by competitor presence in both flowering phenology and reproductive output, while the other lineage shows no response in either trait. Furthermore, we find that the two lineages were introduced at different times and are each capable of replacing the other as the most abundant lineage of a given county. Our results demonstrate that genetic lineage is a more important determinant of competitive response than neighbor identity, and that the two lineages may employ alternate invasion mechanisms. Because management techniques are decided upon based on traits that confer invasiveness, our results highlight the importance of considering intraspecific variation in the invaded range.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The success of an invader in a novel environment is dependent on key traits of that invader (Prinzing et al. 2002; Smith and Knapp 2001) as well as traits of the resident community (Levine et al. 2004). Invasive species can surpass comparable native species in fitness, size and growth rates (van Kleunen et al. 2010), and thus competitive superiority is employed as a mechanism of invasion for some species (Levine et al. 2003; Vila and Weiner 2004). Competition for resources between individuals of different species plays an important role in plant community composition (Booth and Grime 2003; Gibson et al. 2012), so if invasive species successfully outcompete natives, this could result in reduced native species abundance and diversity. Consequently, understanding the outcomes of competitive interactions is critical to understanding ecosystem assembly and invasive species impacts, and is of interest to a wide variety of scientists and resource managers.
The competitive abilities of an individual can be broken down into two traits: their competitive effect, which is the impact an individual has on their neighbor’s growth; and their competitive response, which is the impact on the individual’s growth due to the lower resource levels resulting from neighbor presence (Goldberg and Landa 1991). Recent work tends to focus on the competitive effects of invasive species to answer important questions regarding how they impact the communities they invade (Fried et al. 2014). However, competitive response allows one to investigate the ability of exotic species to invade a novel environment with native plant competitors. Competitive outcomes between native and exotic species are driven by competitive effects and competitive responses, and these dynamics are further mediated by intraspecific variation (Turkington 1996).
Intraspecific variation in invasive species is determined by the amount of genetic variation that accompanies colonization of the novel environment. Invasion events are frequently characterized by multiple introductions that tend to be genetically distinct from one another (Durka et al. 2005). The introduced populations face novel environmental conditions that result in unique selection pressures compared to the native range (Mooney and Cleland 2001). If multiple introductions of invaders result in genetic variation in the introduced range, multiple genotype by environment interactions can lead to selection for different suites of traits and thus different competitive abilities among invasive populations. In fact, invasive lineages of exotic species can differ in important reproductive and phenotypic traits (Meimberg et al. 2010). However, most work to date has been based on the assumption that competitive effect or response is invariant within a species (Bennett et al. 2013; Mariotte et al. 2012) or that differences are between native and introduced populations of an invasive species (Bossdorf et al. 2005; Kumschick et al. 2013). A better approach is to consider the role of intraspecific variation within the invaded range to account for potential trait differences resulting from dynamic relationships between introduction events and subsequent selection.
To test if invasive lineages differ in their competitive ability we used two genetic lineages of the invasive grass, Aegilops triuncialis L.. Aegilops triuncialis is native to Eurasia and currently invading grasslands throughout California and southern Oregon where it has reduced native plant numbers and extent (Batten et al. 2006). Meimberg et al. (2010) showed that all of twenty sampled invasive populations identified with one of three lineages that differ in their range sizes as well as in reproductive traits. Populations within a lineage were found to be inbred, exhibiting no genetic variation within a lineage, and therefore act as replicates for a lineage. We investigated whether established populations of the two most abundant lineages of A. triuncialis differed in their competitive response to three common grassland species that are known to persist after A. triuncialis invasion. Additionally, we used herbarium records to study the historical distribution of lineages, and to test the hypothesis that the most competitive lineage introduced to a county is the lineage currently found at this location.
Methods
Species selection and seed source
Aegilops triuncialis is a selfing, annual grass native to Eurasia, which invades arid and semi-arid grasslands throughout northern California. Meimberg et al. (2010) sampled invasive populations throughout California and identified three lineages (East, West and South) with the East and West lineages occupying much larger areas than the South (Meimberg et al. 2010). To minimize maternal effects, we used offspring from a common garden experiment (Espeland and Rice 2012), which included 11 populations (8 East and 3 West, defined as populations 102, 103, 105, 106, 107, 115, 117, 119, 122, 124 and 125 in Meimberg et al. 2010). Populations sampled in Meimberg et al. (2010) lacked within population variation, and nearby populations were usually of the same lineage. “West” populations were found in the western part of northern California, specifically Mendocino, Colusa, Napa, Solano, and Yolo Counties. East populations were in the eastern section of the invaded range, and included Lake, Sierra, Sacramento, Yuba, El Dorado, Amador, and Butte Counties. The South lineage was found in the (relatively) southern counties of Tuolumne and Calaveras. Populations therefore acted as replicates within a lineage.
In order to test the competitive response of these 11 populations of A. triuncialis, we chose two native competitor species that persist in California grasslands following A. triuncialis invasion (Aigner and Woerly 2011), as well as an exotic grass that is found throughout California (Cal-IPC 2006): (1) Poa secunda J. Presl, a native perennial bunchgrass found throughout California and the western United States; (2) Festuca microstachys Nutt., an annual grass of short stature native to California, and (3) Festuca perennis (L.) Columbus & J.P. Sm., an exotic annual/biennial grass that was introduced throughout California as a forage species (Cal-IPC 2006) and is larger than P. secunda and F. microstachys. Taxonomic nomenclature throughout this manuscript follows Baldwin et al. (2012). Seeds for P. secunda and F. microstachys were obtained from Hedgerow Farms (Winters, CA USA), a commercial entity that produces native seeds for large-scale restoration projects throughout California. Field seeds of P. secunda and F. microstachys were originally collected by Hedgerow Farm from stands in Yolo County, CA in 1997 and 2002, respectively. These seeds were grown in common gardens in Yolo County, and the seeds used here were produced within one to three years of experimental planting. Seeds for F. perennis were collected from University of California-Davis McLaughlin Natural Reserve (Lake County, CA USA), and collected within one to three years of the date of the experiment. Consequently all seeds were produced in a similar location and then evaluated in a greenhouse in Colorado. Note that A. triuncialis is currently invasive in both Yolo and Lake Counties.
Perennial competition
We used the native grass P. secunda to test the competitive response of A. triuncialis to perennial grass competition. Aegilops triuncialis is more likely to encounter P. secunda as a plant than as a seed, and thus we established plants of P. secunda prior to planting A. triuncialis. Two seeds of P. secunda were planted into 120 pots (five cm in diameter) in a greenhouse in March 2012 using 4-P Mix, Professional Formula potting soil (Fafard, Agawam, MA, USA). Pots were placed on a greenhouse bench and thinned to a single (larger) individual after germination. Plants were kept well watered and experienced the equivalent of a 1 year life cycle after which they went dormant. In December 2012 dormant plants were removed from pots and split down the middle. Both halves of each plant were then repotted into the center of a 12.7 cm diameter pot, leaving 3 cm space between each half. We used this approach to ensure A. triuncialis, when planted, interacted with P. secunda in the pot. Pots were then watered for two weeks allowing all P. secunda individuals to come out of dormancy. Two A. triuncialis seeds from the same seed family (i.e. full-sibling individuals) within a population of a particular lineage [40 seeds from Population (pop) 102, 6 from pop 103, 40 from pop 105, 16 from pop 106, 12 from pop 107, 8 from pop 115, 32 from pop 117, 18 from pop 119, 40 from pop 122, 18 from pop 124, and 10 from pop 125] were then sown into each P. secunda pot, 60 pots per lineage (Table 1). Aegilops triuncialis was thinned to a single (larger) individual after germination. Three weeks after planting, we applied a nitrogen addition treatment to half the pots (n = 60 pots, n = 30 per lineage) at the application rate of 10 g/m2 weekly over the growing season.
Annual competition
In January 2013 two A. triuncialis seeds (equivalent in family/population origins per competitor to Perennial Competition above) were sown into the center of 12.7 cm diameter pots filled with 4-P Mix, Professional Formula potting soil (Fafard, Agawam, MA, USA). Eight F. microstachys or F. perennis seeds were then planted around the A. triuncialis seed (2–3 cm radius from seed) to create maximum competition for A. triuncialis. This resulted in 120 pots per competitor (F. microstachys or F. perennis) with 60 pots sown with East lineage seeds [3 from population (pop) 103, 8 from pop 106, 6 from pop 107, 4 from pop 115, 16 from pop 117, 9 from pop 119, 9 from pop 125 and 5 from pop 125] and 60 with West lineage seeds [20 from population (pop) 102, 20 from pop 105, and 20 from pop 122] per competitor (Table 1). Plants were thinned to a single A. triuncialis individual in the center of the pot surrounded by four competitors (the smaller of the two individuals was removed). Plants were allowed to grow for three weeks prior to N addition, after which half of the pots received weekly N additions at the same application rate as the P. secunda pots (see Perennial Competition).
Data and biomass collection
Dates of germination, first flower, and death were recorded for all individuals (A. triuncialis and competitors). Flowering stalks that senesced were collected weekly to avoid loss of biomass or release of seeds. The remaining aboveground biomass of all individuals was harvested on the date of final A. triuncialis senescence. All individuals that had not senesced 210 days after planting were destructively harvested. Biomass was oven dried at 60 °C for 48 h and weighed.
Statistical analysis
For A. triuncialis the response variables days to flower, total biomass and total spikelets were analyzed as a function of lineage, competitor species and their interaction to test for the effect of competitor species on phenotypic traits of each lineage. Total spikelets and total biomass were square-root transformed to meet assumptions of normality. Total competitor biomass was retained as a random covariate but never accounted for greater than 3% of variation. At the end of the experiment both control and N addition pots indicated non-limiting levels of N in the soil, thus, we averaged our results over N treatments.
Herbaria samples
We investigated differences between current and historical distributions of lineages with herbaria samples to test if competitive response explains current patterns of lineage abundance within counties. Meimberg et al. (2010) sampled populations in 14 counties in 2007 and identified populations in each county as belonging to a single lineage (East, West or South). Of the 14 counties sampled, we were able to obtain plant material for the first recorded introduction into 10 of those counties (n = 11 samples: one from each county except for Sacramento where there were two) from herbaria throughout northern California (See Online Resource 1). We were not able to amplify DNA from herbaria samples representing initial introductions into Colusa, Sierra, Tuolumne or Yolo counties. As a control, we used freeze-dried tissue sampled from previously genotyped individuals belonging to each of the three invasive lineages to be run alongside herbaria samples. Leaf tissue from these samples was collected and freeze-dried for 48 h immediately after collection. Dried leaf tissue from herbaria samples and controls was placed individually into 2 mL micro-centrifuge tubes with three 1.99 mm ball bearings and ground to a fine powder in a modified paint shaker. DNA was then extracted from each ground sample using the CTAB technique as described by Doyle and Doyle (1987). Measurements for microsatellite polymorphisms in each sample were performed according to the protocol Meimberg et al. (2010) used to originally identify the three invasive lineages, and run on the same sequence analyzer as the original study. We ran each herbarium sample two times with reference samples for each run, and merged the results for simplicity. Any alleles inconsistent between technical replicates were deleted from the analysis and the genotype at that marker was considered missing data. Additionally, if we had data for less than five marker genotypes for a single sample we did not use the sample. Microsatellite data from herbaria samples were compared to samples of current invasive populations in order to determine whether they identified with previously classified lineages or represented a unique lineage/introduction. We then compared initial introduction identity (lineage) as identified in the herbaria samples with the lineage occupying the same county in 2007 (as per Meimberg et al. 2010).
Results
Flowering time and reproductive output
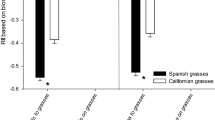
Lineages usually differed in flowering phenology and, in the case of the West lineage, days to flower depended on competitor species, while the East lineage consistently flowered at the same time across all treatments. The West lineage flowered significantly earlier than the East lineage when grown alone (p < 0.02) as well as when grown in competition with F. microstachys and F. perennis (p < 0.0001 for both, Fig. 1). When grown in competition with P. secunda, lineages did not differ in days to flower (p > 0.5, Fig. 1). Within the West lineage, days to flower when grown in competition with F. microstachys or F. perennis did not differ from the control (p = 0.7985 and p = 0.8745, respectively, Fig. 1; Online Resource 2), but the West lineage flowered significantly later when grown in competition with P. secunda (p < 0.03, Fig. 1). For the East lineage, days to flower did not differ from the control for any competitor treatments (p = 0.9771, p = 0.9579, and p = 0.9005 for F. microstachys, F. perennis, and P. secunda, respectively, Fig. 1; Online Resource 2).
Mean days to flower per lineage in all treatments, the East lineage with filled circles and West lineage with open circles (error bars are one standard error of the mean). The West always flowered significantly earlier than the East in all treatments except P. secunda. Mean days to flower for the West was no different when in competition with F. microstachys or F. perennis compared to the control, but was later when in competition with P. secunda. Mean days to flower for the East was no different in any treatment compared to the control
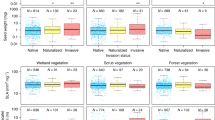
Lineages generally differed in reproductive output, and, in the case of the West lineage, spikelet production depended on competitor species, while the East lineage generally produced the same number of spikelets across all treatments. The West lineage produced significantly more spikelets than the East lineage when grown alone, as well as when grown in competition with F. microstachys and F. perennis (p < 0.0001 for all, Fig. 2; Online Resource 2). When grown in competition with P. secunda, lineages did not differ from each other in spikelet production (p > 0.2, Fig. 2; Online Resource 2). When the West lineage was grown in competition with F. microstachys or F. perennis, spikelet production did not differ from the control (p = 0.9362 and p = 0.3255, respectively, Fig. 2; Online Resource 2), but the West lineage produced significantly fewer spikelets when grown in competition with P. secunda compared to the control (p < 0.0001, Fig. 2, Online Resource 2). The East lineage produced significantly more spikelets when grown with F. microstachys compared to the control (p < 0.05, Fig. 2; Online Resource 2). Spikelet production did not differ for the East when grown with F. perennis (p = 0.5136) or P. secunda (p = 0.6013) compared to the control (Fig. 2; Online Resource 2).
Mean number of spikelets produced per lineage in all treatments, the East lineage with filled circles and West lineage with open circles (error bars are one standard error of the mean). Spikelet production was square-root transformed for analysis, but data presented represent raw values. Number of spikelets produced by the West was significantly higher than the East in all treatments except P. secunda competition. Number of spikelets produced by the West was no different when in competition with F. microstachys or F. perennis compared to the control, but was lower when in competition with P. secunda. Number of spikelets produced by the East was higher when grown in competition with F. microstachys compared to the control, and did not differ when grown with F. perennis or P. secunda compared to the control
Biomass
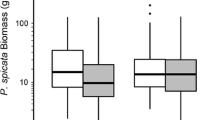
Aegilops triuncialis lineages did not significantly differ in biomass production across treatments (p = 0.2379, Fig. 3; Online Resource 2), thus we analyzed competitive response across lineages. Biomass production across lineages did not differ when grown in competition with F. microstachys compared to the control (p = 0.9691, Fig. 3; Online Resource 2). Biomass production was lower when A. triuncialis was grown in competition with F. perennis and P. secunda compared to when grown alone (p = 0.0591 and p < 0.0001, respectively, Fig. 3; Online Resource 2).
Mean aboveground biomass production produced per lineage in all treatments, the East lineage with filled circles and West lineage with open circles (error bars are one standard error of the mean). Aboveground biomass production was analyzed across lineages, and was not significantly different when grown with F. microstachys or F. perennis compared to the control, and was significantly lower when grown with P. secunda compared to the control
Herbaria results
Although we find evidence for the introduction of unique lineages during the invasion of A. triuncialis, in general we see that both the East and West lineages can persist after introduction. Of 11 herbaria samples, six had the same multi-locus genotype (i.e. lineage) as is currently found in the county (Table 2). Introductions in three counties represented unique multi-locus genotypes; these were Amador, Calaveras, and Sacramento Counties (Table 2). The Amador sample was most similar to the South lineage, but contained one private allele, as well as two alleles that are found in the East and West but not the South. The Calaveras sample contained five private alleles and was consequently the sample that was most divergent from current lineages. The Sacramento sample was most similar to the East lineage, and contained two private alleles although they only differed by a single base pair.
Discussion
Our findings demonstrate that genetic lineage of the invasive grass A. triuncialis is a better predictor of competitive response than identity of the competitor species. In other words, genetic variation within A. triuncialis contributed more to competitive outcomes than interspecific variation among competitors. Furthermore, we found an interaction between lineage and competition such that the East lineage exhibited no significant negative response to competition in flowering phenology or spikelet production, while the West lineage was affected by competition in both metrics (Figs. 1, 2). This result supports those of previous work demonstrating genetic lineages respond differently to competition (Cahill et al. 2005; Latta et al. 2004; Muller and Bartelheimer 2013). We add to this body of work by showing that genetic lineages within the invaded range respond differently to competition. In addition, we find the degree of competitive response is dependent on the interaction between lineage and competitor species.
Lineages responded similarly to competition treatments in biomass production, but their reproductive trait responses differed. For aboveground biomass production, P. secunda exerted the strongest competitive effect on A. triuncialis (Fig. 3). Although both lineages exhibited a reduction in biomass, the West lineage flowered later and produced fewer spikelets when in competition with P. secunda, while the East lineage acted no differently than when grown alone for both of these traits (Figs. 1, 2). The West lineage showed similar trends of later flowering and fewer spikelets when in competition with F. perennis. However, neither impact was shown when in competition with the annual grass F. microstachys. Conversely, the East lineage was consistent in its lack of competitive response by reproductive traits across all treatments: flowering phenology did not change and spikelet production never decreased. Differences in competitive response between the East lineage and West lineage are likely due to different resource uptake and allocation strategies that may in turn reflect differences in invasion mechanisms between the two lineages (Tilman 1995).
Our results demonstrate two different competitive strategies among lineages, with the East showing no response to competition while reproductive traits of the West lineage were impacted by the presence of a neighbor. When the West lineage produced less biomass under competition, it also produced fewer spikelets (Figs. 1, 3), indicating that reproductive output may be biomass dependent. These results are consistent with well-established research showing positive relationships between vegetative and reproductive output (Weiner et al. 2009). The East, however, exhibited a contrasting pattern of consistently flowering at the same time and producing similar numbers of spikelets, regardless of biomass accumulation. The consistency in seed output by the East despite competition may serve as a successful mechanism of invasion for this lineage. However, as these results are based on each lineage experiencing a single year competition, these findings should be considered as preliminary research. Factors that vary and/or accumulate among years (such as precipitation patterns, perennial grass sizes, plant-soil feedbacks, and transgenerational plasticity) have the potential to influence competitive outcomes. How lineages of A. triuncialis respond to these factors should be tested in future experiments.
Our microsatellite DNA analysis of herbarium specimens indicates that both lineages successfully persisted after being introduced 55% of the time, but there is also evidence that each lineage is capable of replacing the other (Table 2). The East was found in more sampling locations in 2007 than the West (Meimberg et al. 2010), but the East also appeared in our herbaria comparisons 20 years earlier than the West (Table 2). Thus, the greater range size of the East may be a result of its earlier introduction, its lack of response to competition, or a combination of the two. We identified the East as the most invasive lineage based on its larger range, and this determination was further substantiated by the results of our competition study. The success of the West lineage may be due to colonization of sites where there is minimal competition, such as recently disturbed areas or serpentine grasslands (Batten et al. 2006).
However, the West lineage also occupies one county where the East was originally introduced (Table 2), indicating the potential for the West to outcompete or replace the East. Other than in Lake County, the East and West occupy discrete ranges (Meimberg et al. 2010).
Our results highlight the potential use and importance of herbaria studies for future research in invasion ecology (Marsico et al. 2010). Previous works have utilized herbaria specimen to track shifts in plant phenology over time (Primack et al. 2004), human-mediated movement of crop species around the globe (Roullier et al. 2013), and to reconstruct the spread of invasive species in the novel range (Delisle et al. 2003). Here we show the potential to expand herbarium studies to investigate the movement of invasive lineages throughout the course of invasion. The potential to identify lineages that have more rapid range expansion or greater persistence could allow for identification of genetic factors and phenotypic traits that influence success of lineages in the novel range.
Understanding intraspecific variation in invasive species may inform more effective management. Both lineages are able to sustain reproductive output regardless of the competitor, but the West is able to produce greater numbers of spikelets if it is able to acquire more biomass. Therefore, management strategies such as increasing native species competition may be more successful with West lineages. Efforts to reduce inflorescence development such as mowing may be more useful in populations of the East as reproductive output will likely be more consistent in this lineage regardless of competition.
Our study underlines the need to examine intraspecific variation in invasive species response to common competitors in the invaded range, as the two lineages used in our study showed important distinctions in their responses to the same competitor. Furthermore, we demonstrate that genetic identity of the target individual is more important to competitive response than presence of a neighbor. Although intraspecific variation in competition has been recognized as an important factor in competitive response and interactions (Cahill et al. 2005; Gustafson et al. 2004; Latta et al. 2004), invasive traits tend to be thought of as uniform within species (but see Bossdorf et al. 2008; Kinter and Mack 2004). Here we show evidence that lineages can differ in their response to competition, and that one lineage may use its resilience to competition as a mechanism of invasion, while the other appears more vulnerable to competition and may depend on disturbance or other factors that limit direct plant–plant interactions for its success. Additionally, our results show that genetic variation within a species can determine competitive outcomes more than interspecific variation among competitors. This has profound implications for community assembly rules as well for mechanisms of invasion and effective control of invasive species.
References
Aigner PA, Woerly RJ (2011) Herbicides and Mowing to control barb goatgrass (Aegilops triuncialis) and restore native plants in serpentine grasslands. Invasive Plant Sci Manag 4:448–457. doi:10.1614/ipsm-d-11-00027.1
Baldwin BG, Goldman DH, Keil DJ, Patterson R, Rosatti TJ, Wilken DH (2012) The jepson manual: vascular plants of California, 2nd edn. University of California Press, Berkeley
Batten KM, Scow KM, Davies KF, Harrison SP (2006) Two invasive plants alter soil microbial community composition in serpentine grasslands. Biol Invasions 8:217–230. doi:10.1007/s10530-004-3856-8
Bennett JA, Lamb EG, Hall JC, Cardinal-McTeague WM, Cahill JF (2013) Increased competition does not lead to increased phylogenetic overdispersion in a native grassland. Ecol Lett 16:1168–1176. doi:10.1111/ele.12153
Booth RE, Grime JP (2003) Effects of genetic impoverishment on plant community diversity. J Ecol 91:721–730. doi:10.1046/j.1365-2745.2003.00804.x
Bossdorf O, Auge H, Lafuma L, Rogers WE, Siemann E, Prati D (2005) Phenotypic and genetic differentiation between native and introduced plant populations. Oecologia 144:1–11. doi:10.1007/s00442-005-0070-z
Bossdorf O, Lipowsky A, Prati D (2008) Selection of preadapted populations allowed Senecio inaequidens to invade Central Europe. Divers Distrib 14:676–685. doi:10.1111/j.1472-4642.2008.00471.x
Cahill JF, Kembel SW, Gustafson DJ (2005) Differential genetic influences on competitive effect and response in Arabidopsis thaliana. J Ecol 93:958–967. doi:10.1111/j.1365-2745.2005.01013.x
Cal-IPC (2006) California invasive plant inventory Publ. No. 2006-02. California Invasive Plant Council. http://www.cal-ipc.org/ip/. Accessed 21 Jan 2016
Delisle F, Lavoie C, Jean M, Lachance D (2003) Reconstructing the spread of invasive plants: taking into account biases associated with herbarium specimens. J Biogeogr 30:1033–1042
Doyle J, Doyle JL (1987) Genomic plant DNA preparation from fresh tissue-CTAB method. Phytochem Bull 19:11–15
Durka W, Bossdorf O, Prati D, Auge H (2005) Molecular evidence for multiple introductions of garlic mustard (Alliaria petiolata, Brassicaceae) to North America. Mol Ecol 14:1697–1706. doi:10.1111/j.1365-294X.2005.02521.x
Espeland EK, Rice KJ (2012) Within- and trans-generational plasticity affects the opportunity for selection in barbed goatgrass (Aegilops triuncialis). Am J Bot 99:2058–2062. doi:10.3732/ajb.1200372
Fried G, Laitung B, Pierre C, Chagué N, Panetta F (2014) Impact of invasive plants in Mediterranean habitats: disentangling the effects of characteristics of invaders and recipient communities. Biol Invasions 16:1–20. doi:10.1007/s10530-013-0597-6
Gibson DJ, Allstadt AJ, Baer SG, Geisler M (2012) Effects of foundation species genotypic diversity on subordinate species richness in an assembling community. Oikos 121:496–507. doi:10.1111/j.1600-0706.2011.19447.x
Goldberg DE, Landa K (1991) Competitive effect and response—hierarchies and correlated traits in the early stages of competition. J Ecol 79:1013–1030. doi:10.2307/2261095
Gustafson DJ, Gibson DJ, Nickrent DL (2004) Competitive relationships of Andropogon gerardii (Big Bluestem) from remnant and restored native populations and select cultivated varieties. Funct Ecol 18:7. doi:10.1111/j.0269-8463.2004.00850.x
Kinter CL, Mack RN (2004) Comparing phenotype and fitness of native, naturalized and invasive populations of downy brome (cheatgrass, Bromus tectorm). In: Hild AL, Shaw NL, Meyer SE, Booth DT, McArthur ED (comp.) Seed and soil dynamics in shrubland ecosystems: Proceedings, Laramie, vol 8, pp 12–16
Kumschick S, Hufbauer RA, Alba C, Blumenthal DM (2013) Evolution of fast-growing and more resistant phenotypes in introduced common mullein (Verbascum thapsus). J Ecol 101:378–387. doi:10.1111/1365-2745.12044
Latta RG, MacKenzie JL, Vats A, Schoen DJ (2004) Divergence and variation of quantitative traits between allozyme genotypes of Avena barbata from contrasting habitats. J Ecol 92:57–71. doi:10.1046/j.0022-0477.2004.00852.x
Levine JM, Vila M, D’Antonio CM, Dukes JS, Grigulis K, Lavorel S (2003) Mechanisms underlying the impacts of exotic plant invasions. Proc R Soc B Biol Sci 270:775–781. doi:10.1098/rspb.2003.2327
Levine JM, Adler PB, Yelenik SG (2004) A meta-analysis of biotic resistance to exotic plant invasions. Ecol Lett 7:15. doi:10.1111/j.1461-0248.2004.00657.x
Mariotte P, Buttler A, Johnson D, Thebault A, Vandenberghe C (2012) Exclusion of root competition increases competitive abilities of subordinate plant species through root–shoot interactions. J Veg Sci 23:1148–1158. doi:10.1111/j.1654-1103.2012.01432.x
Marsico TD et al (2010) PERSPECTIVE: underutilized resources for studying the evolution of invasive species during their introduction, establishment, and lag phases. Evol Appl 3:203–219. doi:10.1111/j.1752-4571.2009.00101.x
Meimberg H, Milan NF, Karatassiou M, Espeland EK, McKay JK, Rice KJ (2010) Patterns of introduction and adaptation during the invasion of Aegilops triuncialis (Poaceae) into Californian serpentine soils. Mol Ecol 19:5308–5319. doi:10.1111/j.1365-294X.2010.04875.x
Mooney HA, Cleland EE (2001) The evolutionary impact of invasive species. Proc Natl Acad Sci USA 98:5446–5451. doi:10.1073/pnas.091093398
Muller B, Bartelheimer M (2013) Interspecific competition in Arabidopsis thaliana: root hairs are important for competitive effect, but not for competitive response. Plant Soil 371:167–177. doi:10.1007/s11104-013-1675-3
Primack D, Imbres C, Primack RB, Miller-Rushing AJ, Del Tredici P (2004) Herbarium specimens demonstrate earlier flowering times in response to warming in Boston. Am J Bot 91:1260–1264. doi:10.3732/ajb.91.8.1260
Prinzing A, Durka W, Klotz S, Brandl R (2002) Which species become aliens? Evol Ecol Res 4:385–405
Roullier C, Benoit L, McKey DB, Lebot V (2013) Historical collections reveal patterns of diffusion of sweet potato in Oceania obscured by modern plant movements and recombination. Proc Natl Acad Sci USA 110:2205–2210. doi:10.1073/pnas.1211049110
Smith MD, Knapp AK (2001) Physiological and morphological traits of exotic, invasive exotic, and native plant species in tallgrass prairie. Int J Plant Sci 162:785–792. doi:10.1086/320774
Tilman D (1995) The resource-ratio hypothesis of plant succession. Am Nat 125(6):827–852
Turkington R (1996) Intergenotypic interactions in plant mixtures. Euphytica 92:15
van Kleunen M, Weber E, Fischer M (2010) A meta-analysis of trait differences between invasive and non-invasive plant species. Ecol Lett 13:235–245. doi:10.1111/j.1461-0248.2009.01418.x
Vila M, Weiner J (2004) Are invasive plant species better competitors than native plant species? Evidence from pair-wise experiments. Oikos 105:229–238. doi:10.1111/j.0030-1299.2004.12682.x
Weiner J, Campbell L, Pino J, Echarte L (2009) The allometry of reproduction within plant populations. J Ecol 97:1220–1233. doi:10.1111/j.1365-2745.2009.01559.x
Acknowledgements
The authors would like to thank Kevin Rice for providing the seeds used in the experiment, as well as Shea Roberts and Sam Williams for their help in data collection and maintenance of the experiment. The authors would also like to thank staff at the Univeristy of California- Berkeley, University of California- Davis, California Department of Food and Agriculture, and the California Academy of Sciences for assistance and access to herbarium samples for use in this study, as well as to the three anonymous reviewers for their comments.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Gomola, C.E., Espeland, E.K. & McKay, J.K. Genetic lineages of the invasive Aegilops triuncialis differ in competitive response to neighboring grassland species. Biol Invasions 19, 469–478 (2017). https://doi.org/10.1007/s10530-016-1366-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-016-1366-0