Abstract
The ability of a biocatalyst to tolerate furan inhibitors present in hemicellulose hydrolysates is important for the production of renewable chemicals. This study shows EMFR9, a furfural-tolerant mutant of ethanologenic E. coli LY180, has also acquired tolerance to 5-hydroxymethyl furfural (5-HMF). The mechanism of action of 5-HMF and furfural appear similar. Furan tolerance results primarily from lower expression of yqhD and dkgA, two furan reductases with a low Km for NADPH. Furan tolerance was also increased by adding plasmids encoding a NADPH/NADH transhydrogenase (pntAB). Together, these results support the hypothesis that the NADPH-dependent reduction of furans by YqhD and DkgA inhibits growth by competing with biosynthesis for this limiting cofactor.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lignocellulosic biomass represents a potential feedstock for microbial conversion to renewable fuels and chemicals. Prior to fermentation, carbohydrate components (cellulose and hemicellulose) must be converted to soluble sugars using acids, enzymes, or a combination (Cheng et al. 2008; Wyman et al. 2005; Um et al. 2003). During steam pretreatment with mineral acids, 5-hydroxymethyl furfural (5-HMF) and furfural are produced as minor but toxic side products from the dehydration of hexose and pentose sugars, respectively (Martinez et al. 2000a; Palmqvist and Hahn-Hagerdal 2000b). 5-HMF has been shown to retard growth and fermentation of ethanologenic E. coli (Zaldivar et al. 1999) and Saccharomyces cerevisiae (Almeida et al. 2008; Palmqvist and Hahn-Hagerdal 2000a; Taherzadeh et al. 2000).
Furans can be removed from hemicellulose hydrolysates by over-liming to pH 10 at elevated temperatures (Martinez et al. 2000a). This process requires the efficient separation of hydrolysate syrups from cellulosic fibers, specialized equipment for lime mixing, separation of syrups from insoluble calcium salts, and creates a solid waste for disposal. The development of furan-resistant biocatalysts could eliminate much of this process complexity. Several enteric bacterial genera (Klebsiella, Enterobacter, Escherichia, Citrobacter, Edwardsiella, Proteus) as well as yeasts convert 5-HMF into 5-hydroxymethyl furfuryl alcohol, a less toxic compound (Boopathy et al. 1993; Palmqvist and Hahn-Hagerdal 2000a; Zaldivar et al. 1999). S. cerevisiae produces multiple oxidoreductases (YGL157W, ADH6, and a mutated ADH1) that can reduce both 5-HMF and furfural to less toxic products (Almeida et al. 2008, 2009; Heer et al. 2009; Liu and Moon 2009). Increased expression of these genes was shown to be beneficial for some aspects of 5-HMF tolerance although none have been shown to increase the minimum inhibitory concentration of furfural. Gorsich et al. (2006) identified many gene inactivations in S. cerevisiae that increased sensitivity to furfural and 5-HMF. Over-expression of one gene, ZWF1 (glucose 6-phosphate dehydrogenase), increased tolerance to furfural.
A furfural-resistant mutant (strain EMFR9) of an ethanologenic E. coli was recently isolated (Miller et al. 2009b). In contrast to studies with yeast, resistance in EMFR9 was found to result from decreased expression of two NADPH-dependent furfural oxidoreductases (yqhD and dkgA) which encode half of the cellular furfural reductase activity. Like biosynthetic enzymes, both have low Km values for NADPH.
In this study, we demonstrate that EMFR9 is also resistant to 5-HMF. Both furans appear to inhibit the growth of E. coli in xylose minimal media by a similar mechanism. Furan reduction by YqhD and DkgA limits biosynthesis by depleting NADPH pools.
Materials and methods
Strains, media, and growth conditions
Strains and plasmids used in this study have been previously described (Miller et al. 2009a, b). These include LY180 (an ethanologenic derivative of E. coli), EMFR9 (furfural-tolerant derivative of LY180), LY180ΔyqhD, LY180ΔdkgA, LY180ΔyqhD ΔdkgA, pLOI4301 containing yqhD. Plasmids pLOI4303 containing dkgA (Miller et al. 2009b), and pLOI4316 containing pntAB (Miller et al. 2009a) were also used. Cultures were grown at 37°C in AM1 minimal media (Martinez et al. 2007) containing 20 g l−1 xylose (solid medium), 50 g l−1 xylose (Bioscreen C growth analyzer and tube cultures), or 100 g l−1 (pH-controlled fermentations).
Tolerance to HMF was tested using 13 × 100 mm closed tubes containing 4 ml AM1 and 5-HMF as indicated. When appropriate, antibiotics were included for plasmid maintenance. Tubes were inoculated to an initial density of 0.05 OD550nm. Growth was measured as the OD550 value after incubation (60 rpm) for 48 h. To examine the effects of pntAB on furan tolerance, a multiwall plate containing 400 μl AM1 (and 5-HMF or furfural) per well was inoculated as above. OD(420–580nm bandwidth) was measured for 72 h using a Bioscreen C growth analyzer (Oy Growth Curves, Helsinki, Finland).
For fermentation experiments, seed cultures of LY180 and EMFR9 were grown overnight in small fermentors (37°C, 200 rpm) containing 350 ml of AM1 medium. Broth was maintained at pH 6.5 by the automatic addition of 2 N KOH. Upon reaching mid-log phase, experimental fermenters were inoculated to an initial cell density of 0.1 OD550 (33 mg dry cell weight l−1). Cell mass (OD550) and furan levels were monitored at 12-h intervals as described previously (Martinez et al. 2000b).
Furan reduction in vivo was measured using pH-controlled fermenters. Furans were added when the cultures reached approximately 1 OD550 using a 10% w/v stock solution. Cell mass and 5-HMF were measured after 0, 15, 30, and 60 min.
In vitro furan reduction
Culture tubes (13 × 100 mm) containing AM1 and 0.1 mM IPTG were inoculated to 0.05 OD550 and incubated at 37°C. These were harvested at a density of 1–2 OD550. Cell pellets were washed once with 100 mM potassium phosphate buffer (pH 7.0), and resuspended in buffer at a density of 10 OD550. Samples (1 ml) were added to 2 ml tubes containing Lysing Matrix B and disrupted (20 s) using a FastPrep-24 (MP Biomedicals, Solon, OH). Furan-dependent oxidation of NADPH was measured at 340 nm using a DU 800 spectrophotometer (Beckman Coulter, Fullerton, CA). Reactions (200 μl total volume; 37°C) contained 50 μl crude extract, 0.2 mM NADPH, and 20 mM 5-HMF. Protein was measured using the BCA assay (Thermo Scientific, Rockford, IL).
Statistical analysis
Data are presented as an average ± SD (n ≥ 3). Statistical comparisons (2-tailed student t test) were made using Graphpad Prism software (La Jolla, CA).
Results
Strain EMFR9 exhibits increased tolerance to 5-HMF
Mutations present in the furfural-resistant mutant, EMFR9, also increased resistance to 5-HMF (Fig. 1). At 1 g 5-HMF l−1, growth and ethanol production by EMFR9 were equal to that of LY180 (parent) in the absence of 5-HMF (Fig. 1a, b). 5-HMF was rapidly metabolized by EMFR9 during the initial 24 h of fermentation with no detrimental effect on cell yield or ethanol yield. The growth of LY180 was completely inhibited by 1.0 g 5-HMF l−1, although 5-HMF levels declined slowly during incubation (Fig. 1a, b, c). No decline was observed without inoculation (data not shown) confirming that this is the result of metabolic activity.
Effect of 5-HMF on anaerobic growth and fermentation. Cells were grown in AM1 mineral salts media with xylose (100 g xylose l−1). a Cell mass during growth with 1 g 5-HMF l−1; b ethanol production during fermentation with 1 g 5-HMF l−1; c reduction of 5-HMF (1.0 g l−1) during fermentation; d cell mass during growth with 2.5 g 5-HMF l−1; e ethanol production during fermentation with 2.5 g 5-HMF l−1; f reduction of 5-HMF (2.5 g 5-HMF l−1) during fermentation. Parallel fermentations without 5-HMF are included (dashed lines) in panels a and b for comparison. All data are plotted as a mean with standard deviation (n = 3). Symbols for all: open square, LY180; and filled circle, EMFR9
With EMFR9, ethanol production and growth were slowed by inclusion of 2.5 g 5-HMF l−1 but proceeded to completion after 96 h (Fig. 1d, e, f). Cell and ethanol yields with this higher level of 5-HMF were comparable to LY180 without 5-HMF. The level of 5-HMF declined rapidly and completely with EMFR9. With LY180, metabolism of 5-HMF was slow and incomplete (Fig. 1f).
Effects of YqhD and DkgA on 5-HMF tolerance
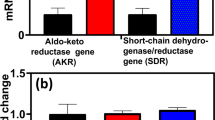
Furfural tolerance in EMFR9 was previously demonstrated to result from the silencing of two NADPH-dependent oxidoreductases, YqhD and DkgA (Miller et al. 2009b). Genes encoding these activities were cloned into pCR2.1 TOPO, transformed into EMFR9, and induced with 0.1 mM IPTG. Cells were harvested, disrupted, and tested for 5-HMF reductase activity (Fig. 2a). Expression of yqhD and dkgA individually from plasmids resulted in a 5-fold increase in the rate of 5-HMF-dependent oxidation of NADPH, confirming that YqhD and DkgA use 5-HMF as a substrate.
Effect of YqhD and DkgA on the in vitro reduction of 5-HMF and on 5-HMF tolerance. a Specific activity for 5-HMF reduction in vitro. Activity was measured in lysed cell extracts (2 mM NADPH, 20 mM 5-HMF). b Effect of yqhD and dkgA expression from plasmids on the cell yield of EMFR9 (resistant mutant). Experiments were performed in tube cultures with AM1 medium containing 50 g xylose l−1 and 1.0 g 5-HMF l−1 (48 h incubation). Note that inclusion of kanamycin for plasmid maintenance lowers 5-HMF tolerance. Induced (Ind.) were grown with 0.1 mM IPTG. c Effect of yqhD and dkgA deletions on the cell yield of LY180 (parent). Experiments were performed in tube cultures with AM1 medium containing 50 g xylose l−1 and 2.5 g 5-HMF l−1 (48 h incubation). All data are plotted as a mean with standard deviation (n = 4)
The individual expression of yqhD and dkgA from plasmids decreased the tolerance of EMFR9 to 5-HMF (Fig. 2b). Addition of kanamycin (12.5 mg l−1) for plasmid maintenance decreased 5-HMF tolerance in all strains, requiring the use of a lower concentration of 5-HMF (1 g l−1) in this experiment. Plasmid pCR2.1 is leaky for the expression of cloned genes in the absence of IPTG (Purvis et al. 2005). Even uninduced expression of yqhD was sufficient to restore the sensitivity of EMFR9 to 5-HMF. Growth inhibition by 5-HMF was further increased by yqhD induction. Expression of dkgA was less effective and required induction to restore 5-HMF sensitivity in EMFR9. Differences in effectiveness between these two oxidoreductases are consistent with the lower apparent Km of YqhD (8 μM) for NADPH compared to 23 μM for DkgA (Miller et al. 2009b).
Deletion of yqhD from LY180 increased tolerance to 2.5 g 5-HMF l−1 (Fig. 2c). Deletions in which markers remained in the chromosome were less effective but confirmed that the inactivation of yqhD was beneficial for 5-HMF tolerance in all cases.
Increasing the availability of NADPH increases 5-HMF tolerance
The proton-translocating transhydrogenase pntAB (Keseler et al. 2009) was over-expressed in LY180 (Fig. 3) to increase the availability of NADPH. In the absence of inhibitor (Fig. 3a), both LY180 with the vector (control) and LY180 (pTrc99a pntAB) grew at the same rate. Induction of LY180 (pTrc99a-pntA) with IPTG (0.01 mM) was detrimental in the absence of 5-HMF. Uninduced LY180 (pTcr99a pntAB), however, grew more rapidly than the vector control (Fig. 3b, c) in the presence of 5-HMF (0.9 and 1.8 g l−1). A similar benefit of pntAB was observed previously with furfural (Miller et al. 2009a). Thus the inhibition of growth by both furans appears to result from furan reduction, depleting the pool of NADPH required for biosynthesis. In addition, over-expression of pntAB led to an increase in overall growth after 72 h, even in the absence of furfural, indicating that biosynthesis may be limited by NADPH under these conditions.
Effect of pntAB expression from plasmids on 5-HMF tolerance. Experiments were conducted using the Bioscreen C growth curve analyzer with AM1 medium containing 50 g xylose l−1 and 5-HMF as indicated. All data are plotted as a mean with standard deviation (n = 10). Connecting points have been omitted for clarity. a No supplement; b supplemented with 0.9 g 5-HMF l−1; and c supplemented with 1.8 g 5-HMF l−1. Symbols for all: open triangle, LY180 (pTrc99a-control); open circle, LY180 (pTrc99a-pntAB) uninduced; filled circle, LY180 (pTrc99a-pntAB) induced with 0.01 mM IPTG
Sulfur assimilation and cysteine biosynthesis have a particularly high requirement for NADPH. Supplementing with cysteine was previously shown to increase furfural tolerance in E. coli LY180 (Miller et al. 2009a) but was of less benefit for 5-HMF tolerance (Fig. 4). Growth of LY180 was partially inhibited by 1 g 5-HMF l−1 and completely restored by supplementing with 100 μM cysteine. Growth in the presence of 2.5 g 5-HMF l−1 was not restored by 100 or 1,000 μM cysteine (Fig. 4b). Unlike furfural, cysteine supplements did not increase the MIC for 5-HMF.
Effect of l-cysteine on 5-HMF tolerance of LY180. Experiments were performed in tube cultures with AM1 medium containing 50 g xylose l−1 and 5-HMF (24 h incubation). Cultures were supplemented with filter-sterilized l-cysteine as indicated. All data are plotted as a mean with standard deviation (n = 4). a 1 g 5-HMF l−1; b 2 g 5-HMF l−1
Conclusions
Selection of an E. coli mutant (EMFR9) for resistance to growth inhibition by furfural (Miller et al. 2009b) was accompanied by resistance to 5-HMF. Both of these furans are important toxins in acid hydrolysates of hemicellulose (Almeida et al. 2009; Gorsich et al. 2006; Zaldivar et al. 1999). At low concentration, both furans were less toxic to cellular functions in E. coli than the consequences of their metabolism. During catabolism, low Km NADPH-dependent oxidoreductases (YqhD and DkgA) compete with biosynthesis for this key cofactor and inhibit growth (Fig. 5). Genes encoding these enzymes (yqhD and dkgA) were silenced in the furan-resistant mutant. Sensitivity was restored by plasmids encoding the production of either enzyme. Resistance was acquired in the parent by deletion of yqhD.
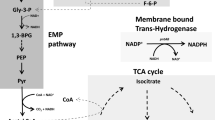
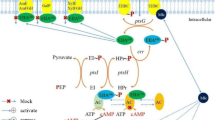
Proposed mechanism for growth inhibition by furans during ethanol fermentation. Xylose or other sugar is initially catabolized to CO2, acetaldehyde, and low levels of other intermediates. Energy is conserved in the form of ATP. NADH serves as the dominant electron carrier, and is regenerated during fermentation by the reduction of acetaldehyde to ethanol. Small amounts of NADPH are produced for essential biosynthetic reactions by genes such as those listed. NADP+ and ADP are regenerated during biosynthesis (and maintenance). Two oxidoreductases (YqhD and DkgA) with low apparent Km values for NADPH are proposed to inhibit biosynthesis and growth by depleting the supply of NADPH
Cells are quite limited in their ability to produce NADPH during fermentative growth on xylose. The most likely source of NADPH under these conditions would be the transhydrogenases (sthA and pntAB). Plasmid expression of pntAB was found to increase resistance to both furfural (Miller et al. 2009a, b) and 5-HMF. A small amount of NADPH would be produced by isocitrate dehydrogenase (icd) during 2-ketoglutarate biosynthesis. Additional NADPH could be produced during gluconeogenesis using malic enzyme (maeB) and by the pentose phosphate pathway (glucose dehydrogenase, gnd; glucose 6-phosphate dehydrogenase, zwf).
The effects of 5-HMF and furfural may not be identical, however. Growth inhibition by these compounds differed in the presence of supplemental cysteine. Sulfur assimilation and cysteine have a high requirement for NADPH. Addition of low levels of cysteine increased the MIC of furfural, presumably by sparing NADPH for other biosynthetic functions (Miller et al. 2009a). Unlike furfural, cysteine addition provided a very limited benefit for 5-HMF tolerance. Thus it is likely that both 5-HMF and furfural have additional inhibitory actions that are not shared.
References
Almeida JRM, Roder A, Modig T, Laadan B, Liden G, Gorwa-Grauslund MF (2008) NADH- vs NADPH-coupled reduction of 5-hydroxymethyl furfural (HMF) and its implications on product distribution in Saccharomyces cerevisiae. Appl Microbiol Biotechnol 78:939–945
Almeida JRM, Bertilsson M, Gorwa-Grauslund MF, Gorsich S, Liden G (2009) Metabolic effects of furaldehydes and impacts on biotechnological processes. Appl Microbiol Biotechnol 82:625–638
Boopathy R, Bokang H, Daniels L (1993) Biotransformation of furfural and 5-hydroxymethyl furfural by enteric bacteria. J Ind Microbiol 11:147–150
Cheng KK, Cai BY, Zhang JA, Ling HZ, Zhou YH, Ge JP, Xu JM (2008) Sugarcane bagasse hemicellulose hydrolysate for ethanol production by acid recovery process. Biochem Eng J 38:105–109
Gorsich SW, Dien BS, Nichols NN, Slininger PJ, Liu ZL, Skory CD (2006) Tolerance to furfural-induced stress is associated with pentose phosphate pathway genes ZWF1, GND1, RPE1, and TKL1 in Saccharomyces cerevisiae. Appl Microbiol Biotechnol 71:339–349
Heer D, Heine D, Sauer U (2009) Resistance of Saccharomyces cerevisiae to high furfural concentration is based on NADPH-dependent reduction by at least two oxidoreductases. Appl Environ Microbiol. doi:10.1128/AEM.01649-9
Keseler IM, Bonavides-Martinez C, Collado-Vides J, Gama-Castro S, Gunsalus RP, Johnson DA, Krummenacker M, Nolan LM, Paley S, Paulsen IT, Peralta-Gil M, Santos-Zavaleta A, Shearer AG, Karp PD (2009) EcoCyc: a comprehensive view of Escherichia coli biology. Nucleic Acids Res 37:D464–D470
Liu ZL, Moon J (2009) A novel NADPH-dependent aldehyde reductase gene from Saccharomyces cerevisiae NRRL Y-12632 involved in the detoxification of aldehyde inhibitors derived from lignocellulosic biomass conversion. Gene 446:1–10
Martinez A, Rodriguez ME, York SW, Preston JF, Ingram LO (2000a) Effects of Ca(OH)2 treatments (“overliming”) on the composition and toxicity of bagasse hemicellulose hydrolysates. Biotechnol Bioeng 69:526–536
Martinez A, Rodriguez ME, York SW, Preston JF, Ingram LO (2000b) Use of UV absorbance to monitor furans in dilute acid hydrolysates of biomass. Biotechnol Prog 16:637–641
Martinez A, Grabar TB, Shanmugam KT, Yomano LP, York SW, Ingram LO (2007) Low salt medium for lactate and ethanol production by recombinant Escherichia coli B. Biotechnol Lett 29:397–404
Miller EN, Jarboe LR, Turner PC, Pharkya P, Yomano LP, York SW, Nunn D, Shanmugam KT, Ingram LO (2009a) Furfural inhibits growth by limiting sulfur assimilation in ethanologenic Escherichia coli strain LY180. Appl Environ Microbiol 75:6132–6141
Miller EN, Jarboe LR, Yomano LP, York SW, Shanmugam KT, Ingram LO (2009b) Silencing of NADPH-dependent oxidoreductase genes (yqhD and dkgA) in furfural-resistant ethanologenic Escherichia coli. Appl Environ Microbiol 75:4315–4323
Palmqvist E, Hahn-Hagerdal B (2000a) Fermentation of lignocellulosic hydrolysates. I: inhibition and detoxification. Bioresour Technol 74:17–24
Palmqvist E, Hahn-Hagerdal B (2000b) Fermentation of lignocellulosic hydrolysates. II: inhibitors and mechanisms of inhibition. Bioresour Technol 74:25–33
Purvis JE, Yomano LP, Ingram LO (2005) Enhanced trehalose production improves growth of Escherichia coli under osmotic stress. Appl Environ Microbiol 71:3761–3769
Taherzadeh MJ, Gustafsson L, Niklasson C, Liden G (2000) Physiological effects of 5-hydroxymethylfurfural on Saccharomyces cerevisiae. Appl Microbiol Biotechnol 53:701–708
Um BH, Karim MN, Henk LL (2003) Effect of sulfuric and phosphoric acid pretreatments on enzymatic hydrolysis of corn stover. Appl Biochem Biotechnol 105:115–125
Wyman CE, Dale BE, Elander RT, Holtzapple M, Ladisch MR, Lee YY (2005) Comparative sugar recovery data from laboratory scale application of leading pretreatment technologies to corn stover. Bioresour Technol 96:2026–2032
Zaldivar J, Martinez A, Ingram LO (1999) Effect of selected aldehydes on the growth and fermentation of ethanologenic Escherichia coli. Biotechnol Bioeng 65:24–33
Acknowledgments
The authors gratefully acknowledge research support by grants from the U.S. Department of Energy (DE-FG36-08GO88142, DE-FC36-GO17058). This work was facilitated by the EcoCyc database (Keseler et al. 2009).
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Miller, E.N., Turner, P.C., Jarboe, L.R. et al. Genetic changes that increase 5-hydroxymethyl furfural resistance in ethanol-producing Escherichia coli LY180. Biotechnol Lett 32, 661–667 (2010). https://doi.org/10.1007/s10529-010-0209-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-010-0209-9